Spatiotemporal mosaic self-patterning of pluripotent stem cells using CRISPR interference
Figures
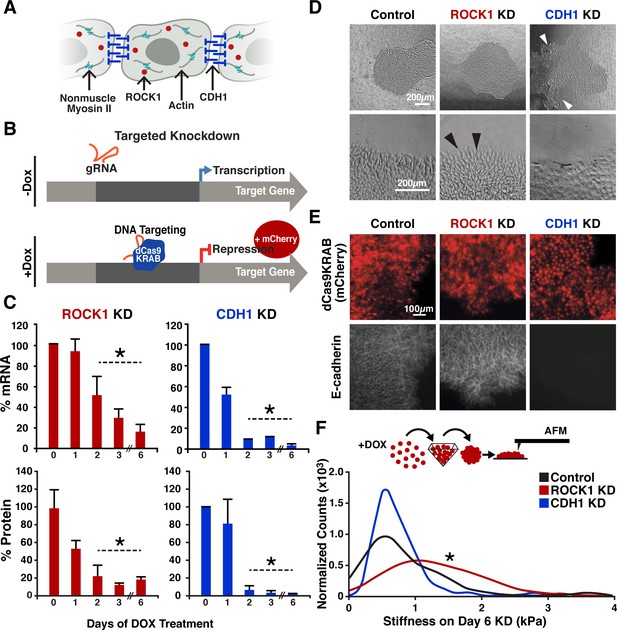
CRISPRi of ROCK1 and CDH1 modulate physical properties of the cell.
(A) Schematic of ROCK1 and CDH1 within a cell. CDH1 is a trans-membrane adhesion molecule that locates to the borders of cells and ROCK1 is a cytoplasmic kinase that acts upon non-muscle myosin II. (B) Schematic of the CRISPRi system. Doxycycline addition to the hiPSC culture media leads to the expression of mCherry and dCas9-KRAB to induce knockdown of target gene. (C) qPCR and western blot quantification of knockdown timing; knockdown of both mRNA and protein were achieved by day three of DOX treatment when compared to untreated hiPSCs (p<0.05, n = 3, data represent mean ± SD). (D) Brightfield imaging of knockdown hiPSCs indicated morphological differences in colony shape (white arrows) and cell extensions (black arrows) at colony borders. (E) Live reporter fluorescence for dCas9-KRAB expression (red) and immunostaining for CDH1 (gray) demonstrated loss of CDH1 in induced CDH1 CRISPRi hiPSCs, but maintenance of CDH1 contacts in the off-target control and ROCK1 KD hiPSCs. (F) Atomic force microscopy (AFM) of knockdown populations exhibited a twofold increase in Young’s elastic modulus of ROCK1 knockdown cells compared to control and CDH1 knockdown cells (p<0.05, n = 36, 65, 72 force points for Control, ROCK1 KD, and CDH1 KD, respectively, area under curve = 1).
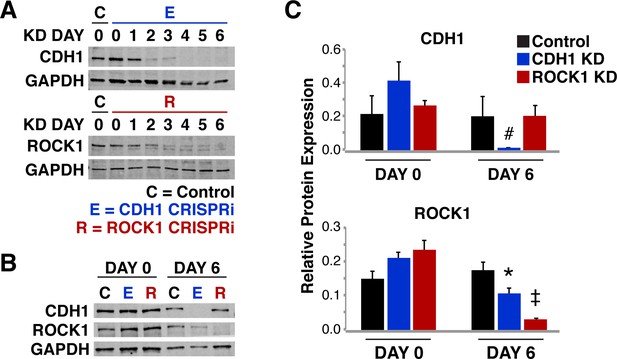
Protein KD time course for ROCK1 and CDH1.
(A) Western blot reflecting KD time course of ROCK1 and CDH1 over 6 days. (B) Western blot of ROCK1 and CDH1 protein levels in KD cells. (C) Densitometry quantification of CDH1 and ROCK1 protein levels in both the ROCK KD cells and the CDH1 KD cells. CDH1 was only significantly different in cells where CDH1 was KD. ROCK1 protein levels exhibited a significant decrease in both CDH1 and ROCK1 (p<0.01).
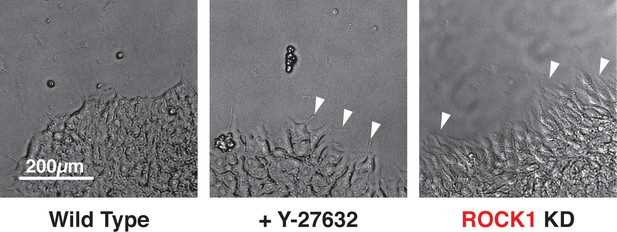
Morphology of ROCK1 KD hiPSCs.
Morphology of ROCK1 KD cells compared to WT cells treated with small molecule inhibitor of ROCK1, Y-27632. The ROCK1 KD and the Y-27632 treated cells displayed cell protrusions (white arrowheads) at the borders of colonies when compared to uninhibited WT cells.
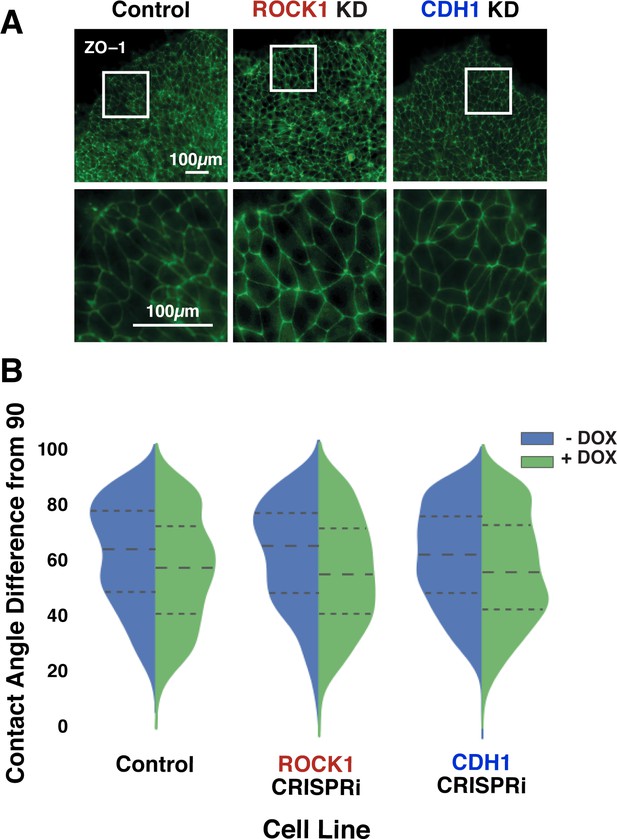
Maintenance of tight junctions in KD hiPSCs.
(A) Immunostaining of zona occludens 1 (ZO-1) in pure populations of hiPSCs after 6 days of induced KD of ROCK1 or CDH1. (B) Population distribution of contact angles before and after DOX treatment measured at ZO-1 junctions. The contact angles of KD cells did not differ significantly compared to the off-target CRISPRi guide control. Dotted lines delineate quartiles. (p<0.05; n = 15 colonies, 50 angles per colony).
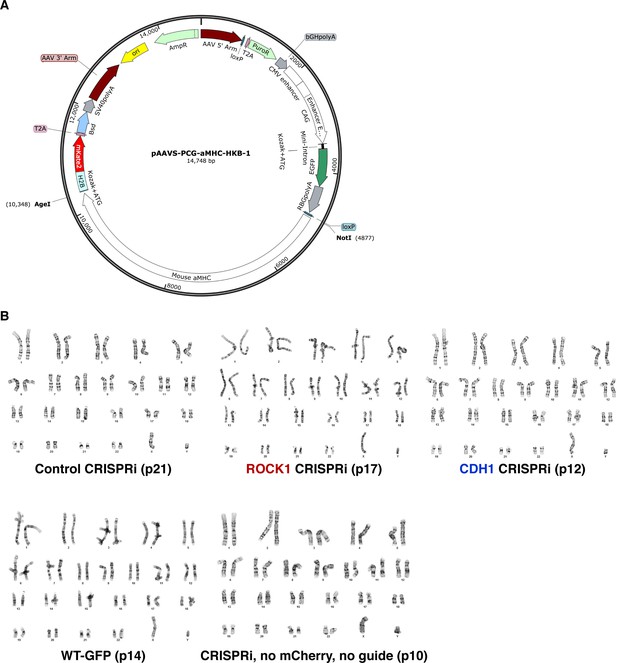
Maintenance of normal hiPSC karyotypes.
(A) Vector map of constitutive GFP cloned into WT hiPSCs to create the WT-GFP line. (B) Karyotypes of all cell lines used in experiments displayed no chromosomal defects. Passage numbers are indicated by p#.
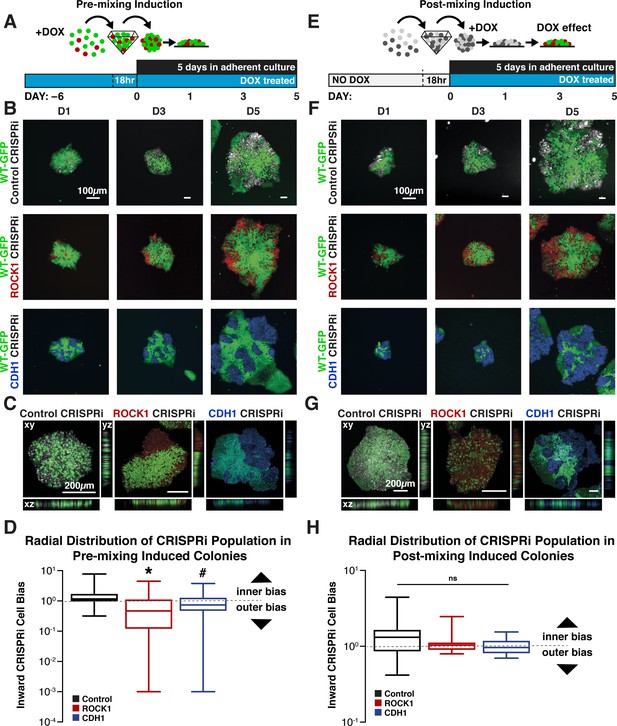
Cell-autonomous pattern emergence in mixed population colonies.
(A) Schematic of experimental timeline. WT-GFP and ROCK1- or CDH1- CRISPRi hiPSCs were pretreated with doxycycline for 6 days before aggregation in pyramidal microwells and re-plating as mixed colonies. (B) Live cell imaging of pattern emergence over time from mixing colonies. Control populations remain mixed, ROCK1 KD hiPSCs cluster radially at borders of colonies, and CDH1 KD populations sort themselves from WT-GFP hiPSCs regardless of location within colony. (C) Confocal microscopy of patterned colonies of hiPSCs with KD induction prior to mixing. (D) Quantification of the radial distribution of KD cells in pre-induced mixed colonies. The ratio of inner cell area to outer cell area normalized to total cell area is displayed (n = 25,* and # indicate significance, p<0.05). (E) Schematic of experimental timeline for WT-GFP and ROCK1- or CDH1-CRISPRi hiPSCs treated with doxycycline upon re-plating as mixed colonies. (F) Live cell imaging of pattern emergence in post-mixing induction colonies, where CRISPRi KD is induced after cell population mixing. (G) Confocal microscopy of patterned hiPSC colonies with KD induction upon mixing populations, where ROCK KD cells stack vertically with WT-GFP hiPSCs. (H) Quantification of the radial distribution of KD cells in post-induced mixed colonies. The ratio of inner cell area to outer cell area normalized to total cell area is displayed (n = 20).
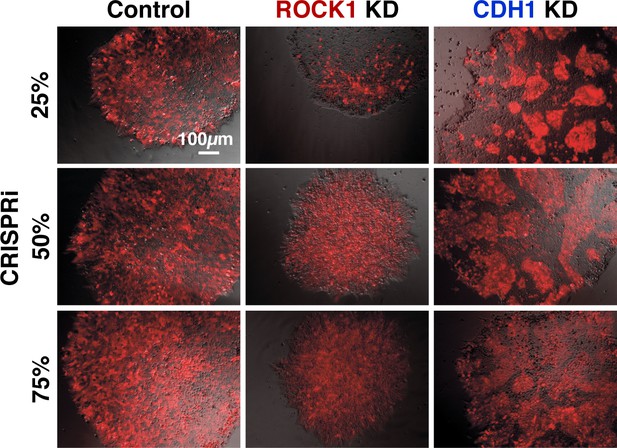
Pattern emergence of variable ratios of mixed populations.
Live imaging on mixed colonies of CRISPRi KD hiPSCs with WT hiPSCs, where a picture was taken every 30 min for 12 hr, 4 days after the start of KD and mixing of two populations (no color = no guide, red = KD population). The displayed images were taken at the start of day four. The ratio of CRISPRi KD cells was varied at 25%, 50%, and 75% of the overall population.
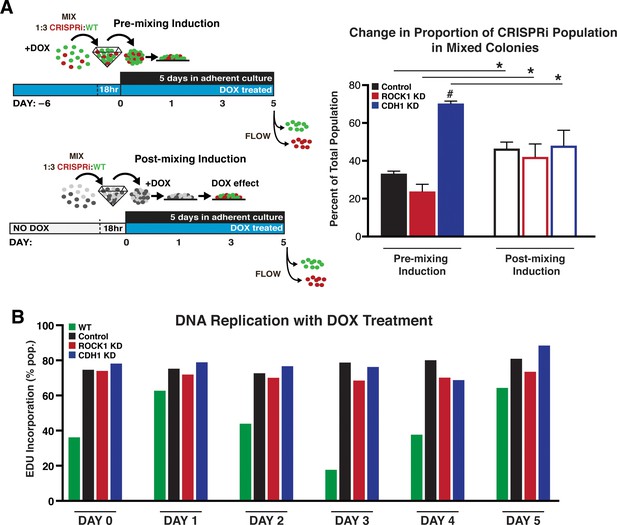
Cell population change over time.
(A) Percent of CRISPRi KD cells after 5 days in mixed culture measured by flow cytometry. (n = 10,000 events, three biological reps; * indicated significance between pre and post mixed and # indicates significance from other KDs; p<0.05) (B) EdU incorporation overtime in pure populations as KD is induced by DOX treatment (n = 10,000 events).
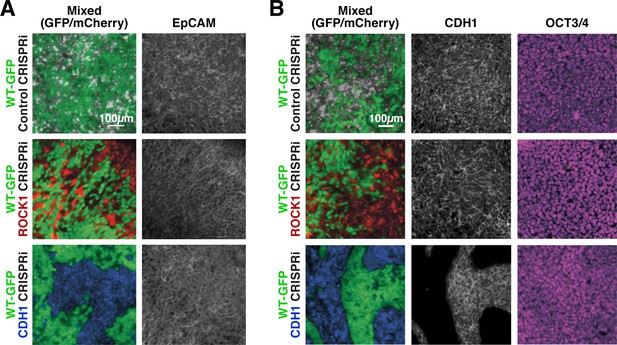
Maintenance of nuclear pluripotency markers and epithelial phenotype.
(A) Immunostaining of EpCAM for mixed colonies displayed relatively uniform expression regardless of KD. (B) Immunostaining for E-cadherin (CDH1) and OCT3/4 in patterned hiPSC colonies demonstrating nuclear localized OCT3/4 throughout the mixed populations.
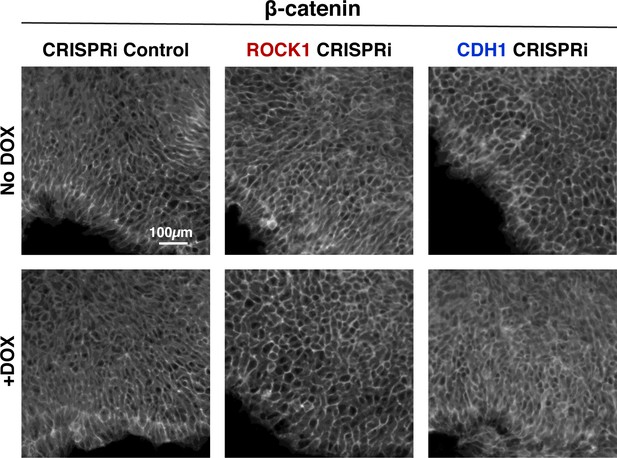
Maintenance of cell junction localized β-catenin in KD cells.
Immunostaining of β-catenin pure hiPSC colonies on day six of KD. β-catenin remains localized to the cell junctions in hiPSCs before and after DOX treatment to induce KD of ROCK1 and CDH1, respectively.
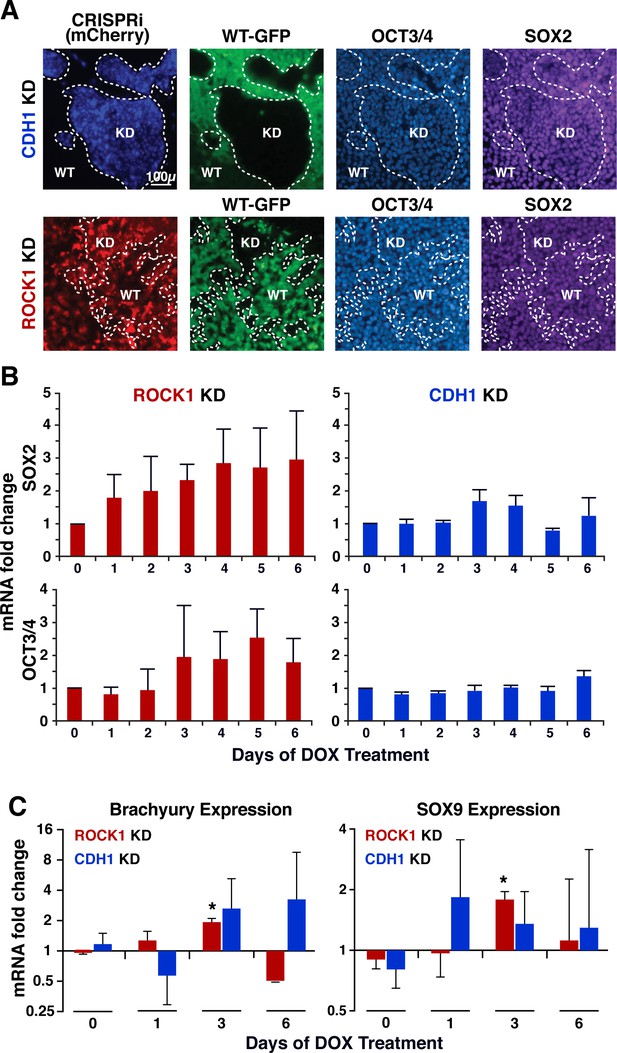
Pluripotent and early germ layer gene expression in KD cells.
(A) Immunostaining of OCT3/4 and SOX2 in mixed colonies on day six of KD, where borders between cell populations are denoted by dotted lines. (B) Gene expression of OCT3/4 and SOX2 in ROCK1 KD and CDH1 KD hiPSCs quantified by mRNA fold change (n = 3). (C) Gene expression of markers of the primitive streak (Brachyury) and neural crest (SOX9) in pure populations of ROCK1 KD or CDH1 KD hiPSCs quantified by mRNA fold change (n = 3, * indicates significance; p<0.05).
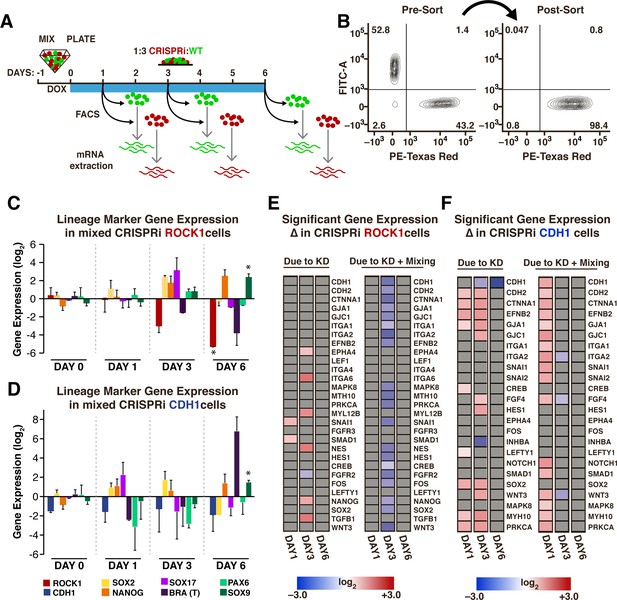
Transient gene expression changes in mixed populations.
(A) Schematic of experimental timeline; WT-GFP and ROCK1- or CDH1-CRISPRi hiPSCs were mixed and re-plated prior to KD induction. Different cell populations were isolated by FACS for mRNA extraction on days 1, 3, and 6 after KD induction. (n = 3 per condition). (B) Representative scatter plot of a FACS-sorted population of mCherry +cells (indicating KD induction) with >98% purity. (C,D) Plots of specific mRNA expression changes at days 1, 3, and 6 in KD cell populations that have been mixed with WT. (* and # indicate significance, p<0.05). (E,F) Heat maps display fold change expression of genes found to display significant changes in ROCK1 or CDH1 KD cells mixed with WT-GFP hiPSCs when compared to time-matched, off-target control hiPSCs. Grey color indicates non-significance. Significance (p<0.05, n = 3) was determined using a one-way analysis of variance (ANOVA) followed by post-hoc pairwise comparisons by Tukey’s tests to determine the effect of mixing populations, the effect of solely KD, and the effect of KD within a mixed population.
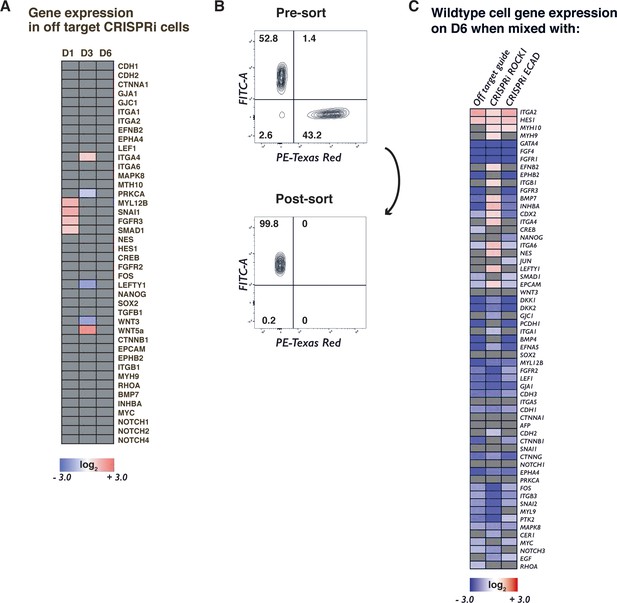
Gene expression changes in WT-GFP cells mixed with CRISPRi cells.
(A) Heat map showing gene expression changes due to mixing two populations together without induction of CRISPRi in sub population where grey indicates non-significance. (B) Example panel of FACS-sorted population of GFP+ (i.e. WT) cells with >99% purity. (C) Heat map of significant gene expression fold changes in WT-GFP cells mixed for 6 days with OTG control, ROCK1 KD, or CDH1 KD hiPSCs when compared to time-matched, pure WT-GFP populations. Grey color indicates non-significance. Significance (p<0.05, n = 3) was determined using an ANOVA taking into account the independent effects of mixing populations, KD alone, and KD within a mixed population.
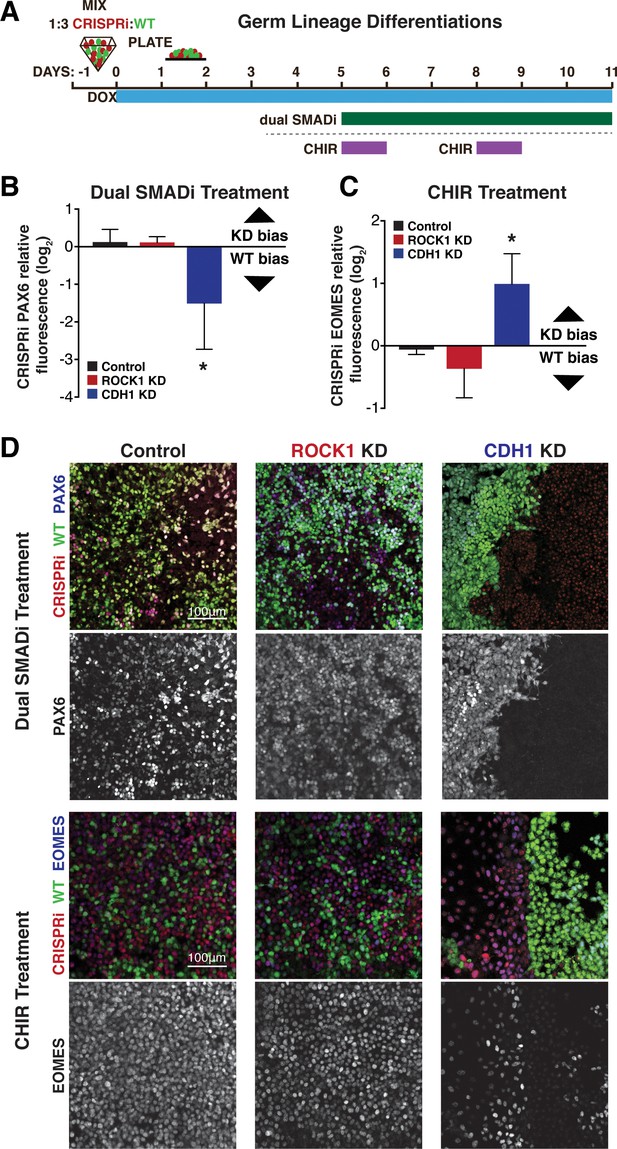
Germ lineage differentiations of mixed colonies.
(A) Schematic of a 6 day ectoderm directed differentiation using dual SMAD inhibition or a complementary mesendoderm directed differentiation using CHIR treatment. (B) Quantification of PAX6 protein presence by immunofluorescence in WT and KD cells of mixed colonies after dual SMAD inhibition treatment. (n = 4 biological replicates with 25 images per biological replicate; * indicates significance; p<0.05) (C) Quantification of EOMES protein presence by immunofluorescence in WT and KD cells of mixed colonies after CHIR treatment. (D) Representative images of mixed colony differentiations with either dual SMAD inhibition or CHIR treatment stained for PAX6 and EOMES, respectively.
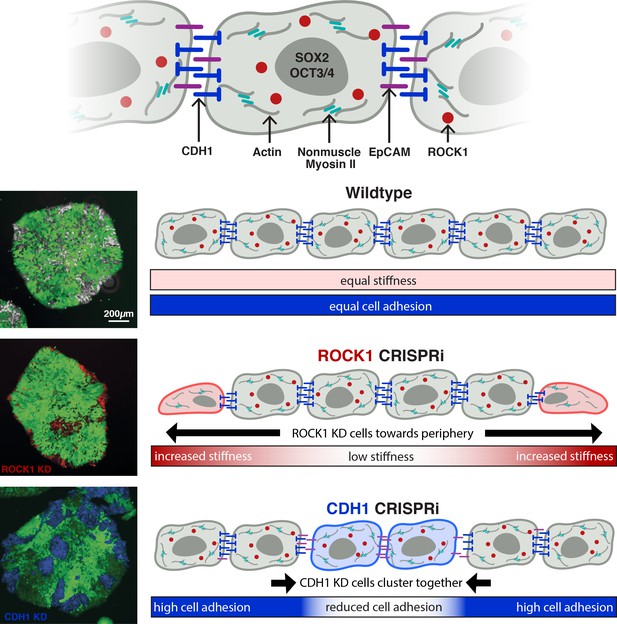
Inducible pattern emergence through the KD of molecules that affect hiPSC physical properties.
(A) Schematic of working model of sub-population manipulation where controlled changes in cellular stiffness or cellular adhesion result in specific colony pattern formation. With mosaic KD, ROCK1 produces continuous radial separation of KD cells from WT, whereas CDH1 displays discrete islands of KD cells within the WT population.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (H. sapien, male) | WT-GFP | this paper | hiPSC line containing constitutative GFP in AAVS1 locus | |
| Cell line (H. sapien, male) | CRISPRi no guide | Mandegar et al., 2016; DOI 10.1016/j.stem.2016.01.022 | hiPSC line containing DOX inducible dCas9KRAB | |
| Cell line (H. sapien, male) | CRISPRi control | Mandegar et al., 2016; DOI 10.1016/j.stem.2016.01.022 | hiPSC line containing DOX inducible dCas9KRAB and gRNA to KCNH2 | |
| Cell line (H. sapien, male) | CRISPRi ROCK1 | Mandegar et al., 2016; DOI 10.1016/j.stem.2016.01.022 | hiPSC line containing DOX inducible dCas9KRAB and gRNA to ROCK1 | |
| Cell line (H. sapien, male) | CRISPRi CDH1 | this paper | hiPSC line containing DOX inducible dCas9KRAB and gRNA to CDH1 | |
| Antibody | mouse anti-ROCK1 | AbCAM | ab58305 | (1:200) |
| Antibody | mouse anti-CDH1 | AbCAM | ab1416 | (1:200) |
| Antibody | goat anti-GAPDH | Invitrogen | PA1-9046 | (1:10000) |
| Antibody | goat anti-OCT3/4 | SantaCruz | sc8629 | (1:400) |
| Antibody | mouse anti-SOX2 | AbCAM | ab7935 | (1:400) |
| Antibody | mouse anti-Zo1 | LifeTechnologies | lifetech 339100 | (1:400) |
| Antibody | rabbit anti -NANOG | AbCAM | ab21624 | (1:300) |
| Antibody | mouse anti-Bcatenin | BD Biosciences | BD610154 | (1:200) |
| Antibody | mous anti-EpCAM | Millipore | MAB4444 | (1:200) |
| Antibody | Alexa 488- or 647- secondaries | Life Technologies | (1:500) | |
| Other | Hoescht stain | LifeTechnologies | (1:10000) | |
| Recombinant DNA reagent | gRNA-CKB (plasmid) | Mandegar et al., 2016; DOI 10.1016/j.stem.2016.01.022 | vector containing gRNA and selection markers (blasticidin resistance, mKate fluorescence) | |
| Software, algorithm | GraphPad Prism | GraphPad Prism (https://graphpad.com) | RRID:SCR_015807 | |
| Software, algorithm | ImageJ | ImageJ (http://imagej.nih.gov/ij/) | RRID:SCR_003070 | |
| Software, algorithm | Python, scikit image | scikit-image contributors et al., 2014: DOI 10.7717/peerj.453 | ||
| Chemical compound, drug | mTeSR1 medium | STEMCELL Technologies | STEMCELL Technologies: 85850 | |
| Chemical compound, drug | Matrigel | Corning Life Sciences | Corning Life Sciences: 356231 | |
| Chemical compound, drug | Accutase | STEMCELL Technologies | STEMCELL Technologies: 7920 | |
| Chemical compound, drug | Blasticidin | ThermoFisher Scientific | ThermoFisher Scientific: R21001 | |
| Chemical compound, drug | Doxycycline | Sigma Aldrich | Sigma Aldrich: D9891 | |
| Chemical compound, drug | Y-27632 ROCK inhibitor | Selleckchem | Selleckchem: S1049 | |
| Chemical compound, drug | Puromycin | Sigma Aldrich | Sigma Aldrich: P8833 | |
| Chemical compound, drug | SB 435142 | Stemgent | Stemgent: 04-0010-05 | |
| Chemical compound, drug | LDN 193189 | Selleckchem | Selleckchem: S2618 | |
| Chemical compound, drug | CHIR 99021 | Selleckchem | Selleckchem: CT99021 | |
| Chemical compound, drug | TRIzol LS Reagent | ThermoFisher Scientific | ThermoFisher Scientific: 10296028 | |
| Sequenced-based reagent | gRNAs | This paper | See Supplementary file 1 | |
| Sequenced-based reagent | RT-qPCR primers | This paper | See Supplementary file 1 | |
| Commercial assay or kit | MycoAlert Mycoplasma Detection Kit | Lonza | Lonza: LT07218 | |
| Commercial assay or kit | Pierce BCA Protein Assay kit | ThermoFisher Scientific | ThermoFisher Scientific: 23250 | |
| Commercial assay or kit | RNeasy Mini Kit | QIAGEN | QIAGEN: 74106 | |
| Commercial assay or kit | iScript cDNA Synthesis kit | BIORAD | BIORAD: 1708891 | |
| Commercial assay or kit | Fast SYBR Green Master Mix | ThermoFisher Scientific | ThermoFisher Scientific: 4385612 | |
| Commercial assay or kit | Click-iT EdU Alexa 647 Imaging Kit | ThermoFisher Scientific | ThermoFisher Scientific: C10340 | |
| Commercial assay or kit | Direct-zol RNA MiniPrep Plus kit | ZYMO Research | ZYMO: R2061 |
Additional files
-
Supplementary file 1
Supplementary tables.
(1) Table 1 displays guide RNA sequences used for CRISPRi knockdown. (2) Table 2 outlines primer sequences used for gene expression analysis by quantitative PCR. (3) Table 3 shows gene targets and primer sequences used for the Fluidigm 96.96 array gene expression analysis.
- https://doi.org/10.7554/eLife.36045.018
-
Transparent reporting form
- https://doi.org/10.7554/eLife.36045.019





