Cytokinin transfer by a free-living mirid to Nicotiana attenuata recapitulates a strategy of endophytic insects
Figures
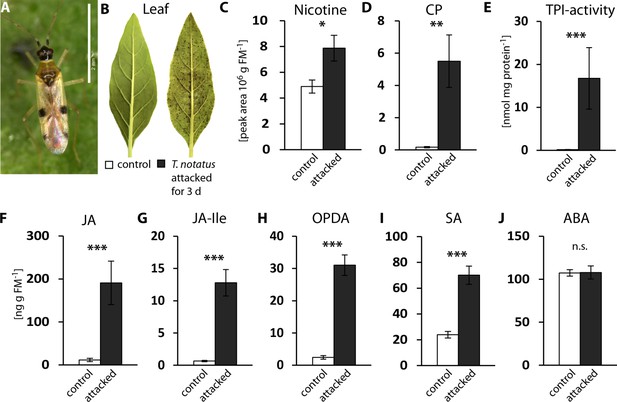
Tupiocoris notatus feeding induces JA-dependent defense responses in Nicotiana attenuata.
(A) T. notatus adult. (B) Representative pictures of a control leaf of N. attenuata and a leaf after 3 d of continuous T. notatus feeding. (C–J) Defense metabolites and stress-related phytohormone levels induced by 3 d of T. notatus attack (filled columns) compared with control leaves (open) from unattacked plants: (C) nicotine, (D) caffeoylputrescine (CP), (E) trypsin proteinase inhibitor (TPI) activity, (F) jasmonic acid (JA), (G) jasmonic acid-isoleucine conjugate (JA-Ile), (H) cis-(+)−12-oxophytodienoic acid (OPDA), (I) salicylic acid (SA) and (J) abscisic acid (ABA). Wilcoxon-Mann-Whitney test was used to identify statistically significant differences between control and attacked leaves. (C) nicotine: N = 6, W = 5, p=0.022; (D) CP: N = 6, W = 0, p=0.001, (E) TPI: N = 7, W = 0, p<0.001, (F) JA: N = 7, W = 0, p<0.001, (G) JA-Ile: N = 7, W = 0, p<0.001; (H) OPDA: N = 7, W = 0, p<0.001; (I) SA: N = 7, W = 0, p<0.001: (J) ABA: N = 7, W = 20, p=0.620. *p<0.05, **p<0.01, ***p<0.001, n.s.: not significant. Error bars depict standard errors. FM: fresh mass. For raw data see Raw_data_FIGURE_1 (Dryad: Brütting et al., 2018).
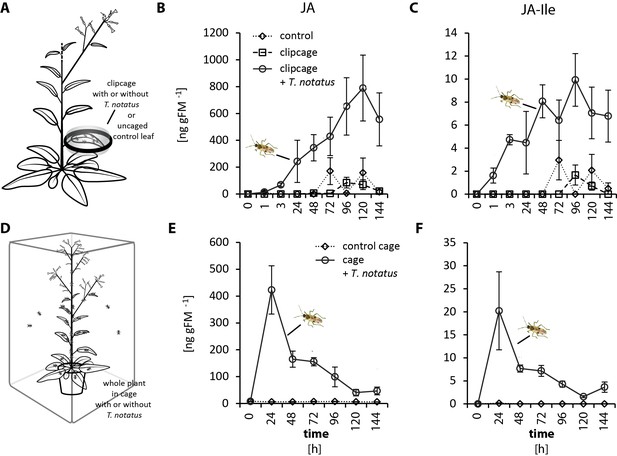
Tupiocoris notatus feeding increases levels of JA and JA-Ile.
(A) Experimental setup corresponding to (B) and (C). On each plant, we enclosed one leaf in a plastic clipcage with (clipcage+ T. notatus) or without (clipcage) 20 T. notatus. Additionally, we collected uncaged control leaves (control, dotted line). (B) Jasmonic acid (JA) and (C) jasmonic acid-isoleucine conjugate (JA-Ile) were monitored over 144 hr. Data were analyzed with ANCOVA on square root-transformed data with mirid as factor and time as continuous explanatory variable. (B) JA: time F1,51 = 38.42, p<0.001; mirid F1,51 = 93.82, p<0.001; time*mirid F1,51 = 10.89, p=0.002. (C) JA-Ile: time F1,51 = 10.17, p=0.002; mirid F1,51 = 110.10, p<0.001; time*mirid F1,51 = 1.292, p=0.261). Error bars depict standard errors (N ≥ 3). (D) Experimental setup corresponding to (E) and (F). A whole plant was caged in an insect cage with (cage+ T. notatus) or without (control cage) T. notatus adults. (E) JA and (F) JA-Ile kinetics were monitored over 144 hr after the start of herbivore exposure. Data were analyzed with generalized least squares (GLS) model with mirid as factor and time as continuous explanatory variable. (E) JA: time F1,44 = 14.878, p<0.001; mirid F1,44 = 30.218, p<0.001; time*mirid F1,44 = 14.471, p<0.001. (F) JA-Ile: time F1,44 = 4.123, p=0.048; mirid F1,44 = 33.920, p<0.001; time*mirid F1,44 = 3.978, p=0.052. Error bars depict standard errors (N = 4). FM: fresh mass. For raw data see Raw_data_FIGURE_1_S1 (Dryad: Brütting et al., 2018).
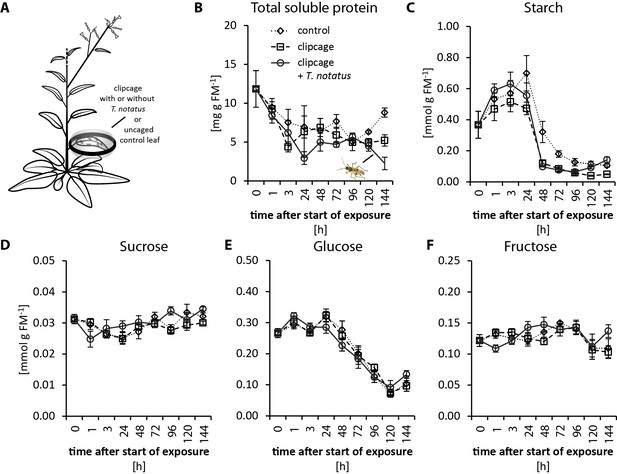
Tupiocoris notatus feeding on single leaves does not significantly change nutrient levels.
(A) Experimental setup: On each plant we enclosed one leaf in a plastic clipcage with (clipcage +T. notatus; solid line) or without (clipcage, dashed line) 20 T. notatus. Additionally, we collected uncaged control leaves (control, dotted line). (B) Total soluble proteins (TSP), (C) starch, (D) sucrose, (E) glucose and (F) fructose were analyzed in a time-kinetic from 1 to 144 hr. Statistically significant differences were identified with ANCOVA with mirid as factor and time as continuous explanatory variable. (B) TSP: log transformed time F1,51 = 14.317, p<0.001; mirid F1,51 = 2.438, p=0.125; time*mirid F1,51 = 0.479, p=0.492. (C) log transformed starch: time F1,51 = 137.376, p<0.001; mirid F1,51 = 3.749, p=0.058; time*mirid F1,51 = 2.651, p=0.110. (D) sucrose: time F1,51 = 13.847, p<0.001; mirid F1,51 = 3.883, p=0.054; time*mirid F1,51 = 5.894, p=0.019. (E) glucose: time F1,51 = 173.06, p<0.001; mirid F1,51 = 0.050, p=0.823; time*mirid F1,51 = 0.107, p=0.745. (F) fructose: log transformed time F1,51 = 0.505, p=0.480; mirid F1,51 = 0.433, p=0.513; time*mirid F1,51 = 5.798, p=0.020. Error bars depict standard errors (N ≥ 3). FM: fresh mass. For raw data see Raw_data_FIGURE_2 (Dryad: Brütting et al., 2018).
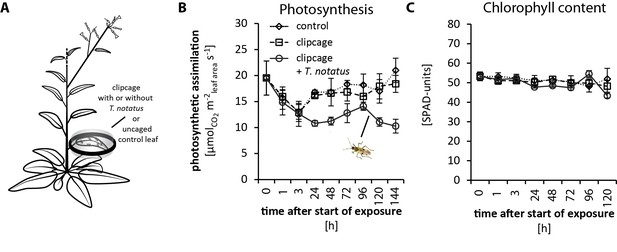
Tupiocoris notatus feeding on single leaves decreases photosynthetic rates while not influencing chlorophyll contents.
(A) Experimental setup: On each plant we enclosed one leaf in a plastic clipcage with (clipcage +T. notatus; solid line) or without (clipcage, dashed line) 20 T. notatus. Additionally, we collected untreated control leaves (control, dotted line). (B) photosynthetic rates and (C) chlorophyll contents in a time-kinetic from 1 to 144 hr (120 hr). Statistically significant differences were identified with ANCOVA with mirid as factor and time as continuous explanatory variable. (B) photosynthetic rates: time F1,51 = 0.846, p=0.362; mirid F1,51 = 41.466, p<0.001; time*mirid F1,51 = 10.802, p=0.002; (C) chlorophyll content: time F1,45 = 2.423, p=0.127; mirid F1,45 = 1.721, p=0.196; time*mirid F1,45 = 1.670, p=0.203; Error bars depict standard errors (N ≥ 3). For raw data see Raw_data_FIGURE_2_S1 (Dryad: Brütting et al., 2018).
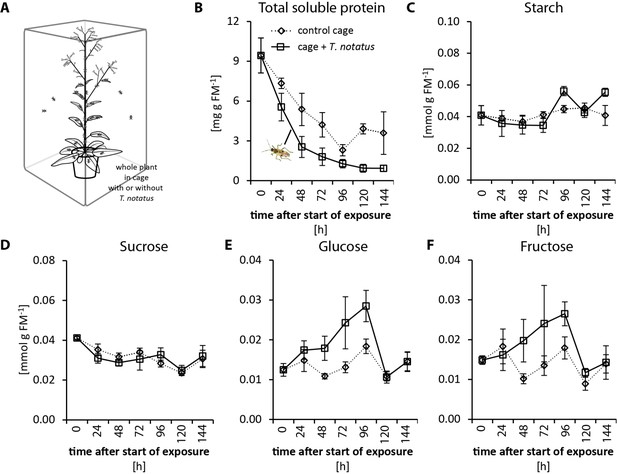
Tupiocoris notatus feeding on whole plants only slightly alters nutrient levels in attacked leaves of Nicotiana attenuata, mainly decreasing protein contents.
(A) Experimental setup: whole plants were caged with (cage+ T. notatus attacked; solid line) or without (control cage, dotted line) T. notatus. B) Protein, (C) starch, (D) sucrose, (E) glucose and (F) fructose were monitored in a time-kinetic from 24 to 144 hr. Statistically significant differences were identified with ANCOVA with mirid as factor and time as continuous explanatory variable. (B) TSP: time F1,43 = 25.145, p<0.001; mirid F1,43 = 19.672, p<0.001; time*mirid F1,43 = 0.102, p=0.75. (C) starch: time F1,44 = 13.949, p<0.001; mirid F1,44 = 0.342, p=0.561; time*mirid F1,44 = 4.932, p=0.031. (D) log transformed sucrose: time F1,43 = 0.111, p=0.740; mirid F1,43 = 3.834, p=0.057; time*mirid F1,43 0.721, p=0.401. (E) log transformed glucose: time F1,44 = 0.672, p=0.417; mirid F1,44 = 0.066, p=0.798; time*mirid F1,44 4.760, p=0.035. (F) fructose: time F1,44 = 0.890, p=0.351; mirid F1,44 = 0.229, p=0.634; time*mirid F1,44 1.83, p=0.183. Error bars depict standard errors. (N ≥ 3). FM: fresh mass. For raw data see Raw_data_FIGURE_2_S2 (Dryad: Brütting et al., 2018).
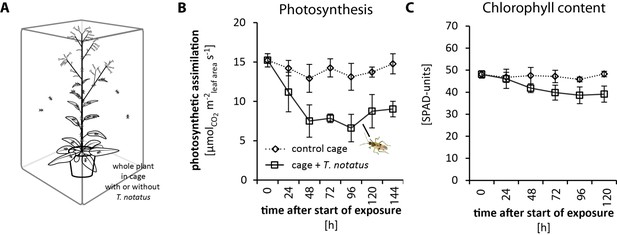
Tupiocoris notatus feeding on whole plants decreases photosynthetic rates and chlorophyll contents in attacked leaves of Nicotiana attenuata.
(A) Experimental setup: whole plants were caged with (cage+ T. notatus; solid line) or without (control cage, dotted line) T. notatus. (B) Photosynthetic assimilation rates and (C) chlorophyll contents were monitored in a time-kinetic from 24 to 144 hr (120 hr) after the start of herbivore exposure. Statistically significant differences were identified with ANCOVA with mirid as factor and time as continuous explanatory variable. (B) photosynthetic rates: time F1,44 = 0.196, p=0.660; mirid F1,44 = 102.063, p<0.001; time*mirid F1,44 1.234, p=0.273. (C) chlorophyll content: time F1,36 = 5.244, p=0.028; mirid F1,36 = 40.128, p<0.001; time*mirid F1,36 7.215, p=0.011. Error bars depict standard errors (N = 4). For raw data see Raw_data_FIGURE_2_S3 (Dryad: Brütting et al., 2018).
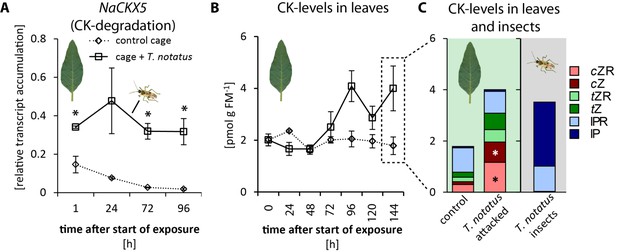
Tupiocoris notatus contain large amounts of CKs in their bodies and their feeding alters Nicotiana attenuata’s cytokinin (CK) metabolism.
(A) Transcript accumulations of NaCKX5: cytokinin oxidase/dehydrogenase 5 (which inactivates CKs by oxidation) and (B) CK levels in leaves: sum of cis-zeatin (cZ), trans-zeatin (tZ), N6-isopentenyladenine (IP) and their ribosides (cZR, tZR, IPR) in leaves exposed to T. notatus feeding (cage +T. notatus, solid line) and control leaves (control cage, dotted line) at different times after herbivore exposure. (C) Single CK types in leaves after 144 hr of exposure to T. notatus and in the insect bodies. Two-way ANOVA on log2-transformed data followed by Welch t-test with Bonferroni corrections between control and T. notatus cage for each harvest time were used to analyze A) (mirid F1,13 = 158.2, p<0.001; time F3,13 = 12.52, p<0.001; time*mirid F3,13 7.015, p=0.005). ANCOVA with mirid as factor and time as continuous explanatory variable was used to analyze B) (log2 transformed data: time F1,44 = 7.335, p=0.010; mirid F1,44 = 8.609, p=0.005; time*mirid F1,44 = 15.243 p<0.001). (C) was analyzed with Wilcoxon-Mann-Whitney test between control and attacked leaves for each CK type. • p<0.1, *p<0.05. Error bars depict standard errors (A): N ≥ 2; (B, C): N = 4). FM: fresh mass. For raw data see Raw_data_FIGURE_3 (Dryad: Brütting et al., 2018).
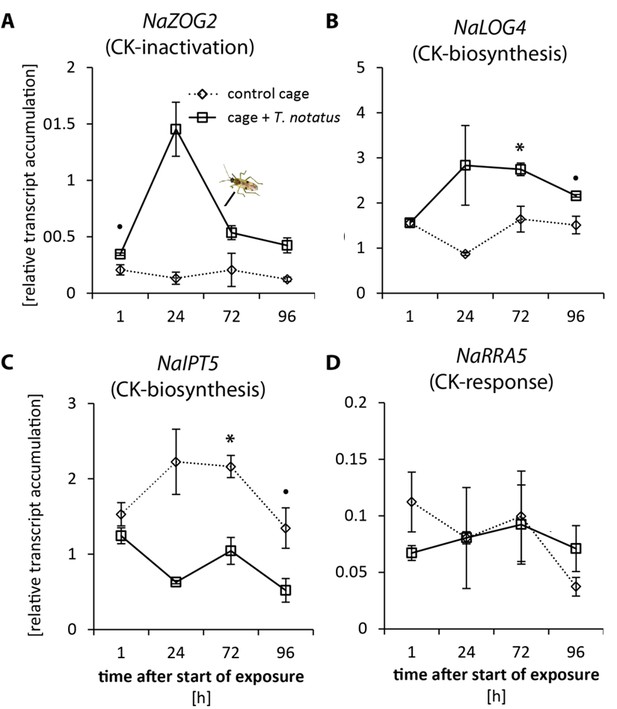
Tupiocoris notatus feeding alters transcript levels of cytokinin inactivation and biosynthetic genes in attacked leaves of Nicotiana attenuata.
Relative transcript accumulations (NaActin as reference gene) in leaves infested with T. notatus (cage+ T. notatus, solid line) and control leaves (control cage, dotted line) at different harvest times after the start of herbivore exposure. (A) NaZOG2: zeatin-O-glucosyltransferase 2. (B) log2 transformed NaLOG4: cytokinin riboside 5'-monophosphate phosphoribohydrolase LOG (LONELY GUY) 4. (C) NaIPT5: isopentenyltransferase 5. (D) NaRRA5: CK response regulator 5. Statistically significant differences were identified by two-way ANOVAs with factors mirid and time followed by Welch t-test with Bonferroni corrections between control and T. notatus cage for each harvest time. (A) NaZOG2: time F3,13 = 10.45, p<0.001; mirid F1,13 = 49.36, p<0.001; time*mirid F3,13 12.83, p<0.001; B) NaLOG4: time F3,13 = 2.092, p=0.151; mirid F1,13 = 28.28, p<0.001; time*mirid F3,13 5.372, p=0.013; (C) NaIPT5: time F3,13 = 4.274, p=0.026; mirid F1,13 = 41.21, p<0.001; time*mirid F3,13 3.030, p=0.068; (D) NaRRA5: time F3,13 = 0.996, p=0.426; mirid F1,13 = 0.039, p=0.847; time*mirid F3,13 0.643, p=0.601. • p<0.1, *p<0.05. Error bars depict standard errors (N ≥ 2). For raw data see Raw_data_FIGURE_3_S1 (Dryad: Brütting et al., 2018).
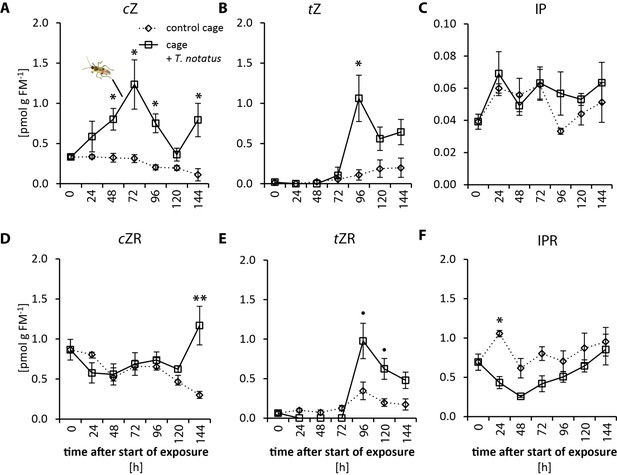
Tupiocoris notatus feeding on whole plants alters cytokinin (CKs) levels in attacked Nicotiana attenuata leaves.
Whole plants were caged with (cage+ T. notatus, solid line) or without (control cage, dotted line) T. notatus and CK concentrations in attacked leaves were quantified in a time kinetic from 24 to 144 hr after herbivore exposure. (A) cis-Zeatin (cZ), (B) trans-zeatin (tZ), (C) N6-isopentenyladenine (IP) and their ribosides (D) cZR, (E) tZR, (F) IPR. Statistically significant differences were identified by two-way ANOVAs (TWA) followed by TukeyHSD post hoc test (C), (D and F) or by generalized least squares (GLS) model followed by Wilcoxon–Mann–Whitney tests with Bonferroni corrections between control and T. notatus cage for each harvest time (A, B and E). (A) cZ: time F5,36 = 3.046, p=0.021; mirid F1,36 = 44.561, p<0.001; time*mirid F5,36 = 2.688, p=0.037. (B) tZ: time F5,36 = 4.347, p=0.003; mirid F1,36 = 18.392, p<0.001; time*mirid F5,36 = 4.898, p=0.002. (C) IP: time F5,36 = 1.351, p=0.266; mirid F1,36 = 2.249, p=0.142; time*mirid F5,36 = 0.579, p=0.716. (D) cZR: time F5,36 = 3.097, p=0.020; mirid F1,36 = 9.234, p=0.004; time*mirid F5,36 = 3.985, p=0.006. (E) tZR: time F5,36 = 9.421, p<0.001; mirid F1,36 = 11.277, p=0.002; time*mirid F5,36 = 6.177, p<0.001. (F) IPR: time F5,36 = 3.490, p=0.011; mirid F1,36 = 20.016, p<0.001; time*mirid F5,36 = 1.141, p=0.357. • p<0.1, *p<0.05, **p<0.01. Error bars depict standard errors (N = 4). FM: fresh mass. For raw data see Raw_data_FIGURE_3_S2 (Dryad: Brütting et al., 2018).
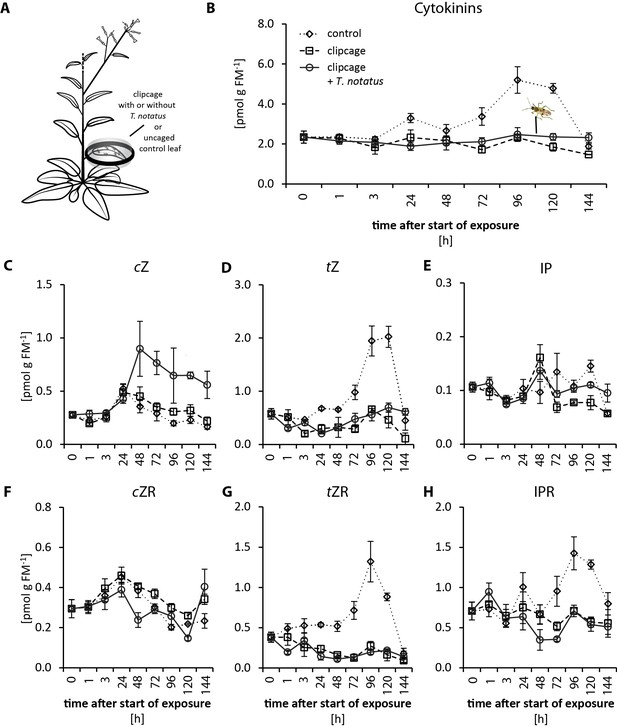
Influence of Tupiocoris notatus single-leaf feeding on cytokinin levels of attacked and unattacked leaves.
(A) Experimental setup: a single leaf was enclosed in a plastic clipcage with (clipcage+ T. notatus; solid line) or without 20 T. notatus (clipcage, dashed line). Additionally, unenclosed control leaves (control, dotted line) were also collected. (B–H) CK values in leaves at different time-points after the start of herbivore exposure: (B) sum of cis-zeatin (cZ), trans-zeatin (tZ), N6-isopentenyladenine (IP) and their ribosides (cZR, tZR, IPR). (C) cZ, (D) tZ, (E) IP, (F) cZR, (G) tZR and H) IPR. Statistically significant differences were identified by two-way ANOVAs (C, E) or by ANCOVA with mirid as factor and time as continuous explanatory variable (B, D, F, G, H). (B) sum of CKs: time F1,51 = 0.041, p=0.841; mirid F1,51 = 1.934, p=0.170; time*mirid F1,51 = 5.270, p=0.026. (C) cZ: time F7,39 = 5.855, p<0.001; mirid F1,39 = 31.43, p<0.001; time*mirid F7,39 = 1.596, p=0.166. (D) tZ: log transformed time F1,51 = 3.024, p=0.088; mirid F1,51 = 0.986, p=0.325; time*mirid F1,51 = 2.572, p=0.115. (E) IP: time F7,39 = 11.538, p<0.001; mirid F1,39 = 7.134, p=0.011; time*mirid F7,39 = 2.279, p=0.048. (F) cZR: log transformed time F1,51 = 1.083, p=0.303; mirid F1,51 = 5.066, p=0.03; time*mirid F1,51 = 0.256, p=0.615. (G) square root transformed tZR: log transformed time F1,51 = 9.910, p=0.003; mirid F1,51 = 1.164, p=0.286; time*mirid F1,51 = 0.688, p=0.411 hr IPR: log transformed time F1,51 = 11.909, p=0.001; mirid F1,51 = 1.484, p=0.229; time*mirid F1,51 = 1.813, p=0.184. Error bars depict standard error (N ≥ 3). FM: fresh mass. For raw data see Raw_data_FIGURE_3_S3 (Dryad: Brütting et al., 2018).
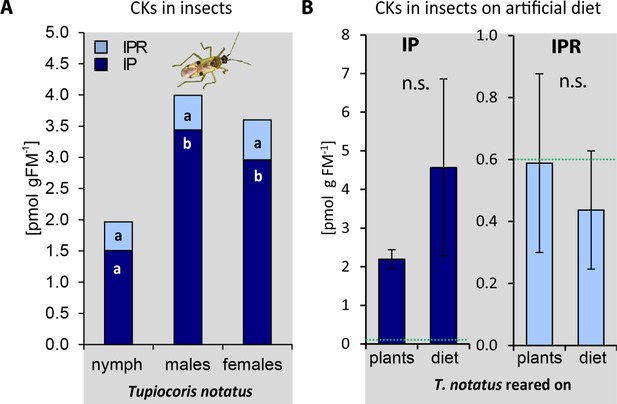
Tupiocoris notatus contains large amounts of N6-isopentenyladenine (IP) in their bodies independently of stage, sex or food source.
(A) IP and IPR (N6-isopentenyladenosine) in T. notatus nymphs and adult males and females. One-way ANOVAs with Tukey HSD post hoc test (N ≥ 3). IP: F2,10 = 17.92, p<0.001; IPR, F2,10 = 0.89, p=0.441. (B) IP and IPR in T. notatus adults reared on plants or on artificial diet. Green dotted lines present representative levels of IP and IPR in leaf tissues on which the insects had fed. Wilcoxon-Mann-Whitney test. IP: N = 7; W = 38, p=0.281; IPR N = 7; W = 35, p=0.463 (Figure 3—figure supplement 4b). n.s, not significant. Error bars depict standard errors (N = 7). FM: fresh mass. For raw data see Raw_data_FIGURE_3_S4 (Dryad: Brütting et al., 2018).
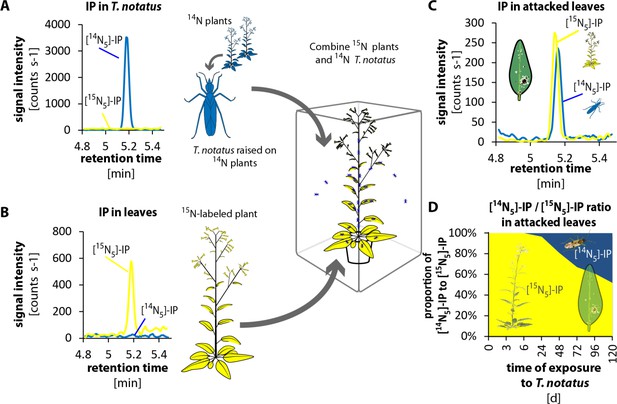
Tupiocoris notatus transfers IP to leaves of its host plant.
(A) and (B) Experimental setup and chromatograms of IP: (A) T. notatus raised on 14N-grown hydroponic plants grown contain only [14N5]-IP in their bodies. (B) Plants raised on a hydroponic medium containing only a 15N containing N-source have only [15N5]-IP in leaves. 15N labeled plants and 14N labeled insects were placed in the same cage for 5 days. Ratio of [14N5]-IP (originating from insects, blue) and [15N5]-IP (from host plant, yellow) were determined in attacked leaves. (C) Chromatograms of [14N5]-IP and [15N5]-IP in the leaves of 5d attacked plants. (D) Ratio of [14N5]-IP and [15N5]-IP at different harvest times after the start of exposure to T. notatus (N = 5). For raw data see Raw_data_FIGURE_4 (Dryad: Brütting et al., 2018).
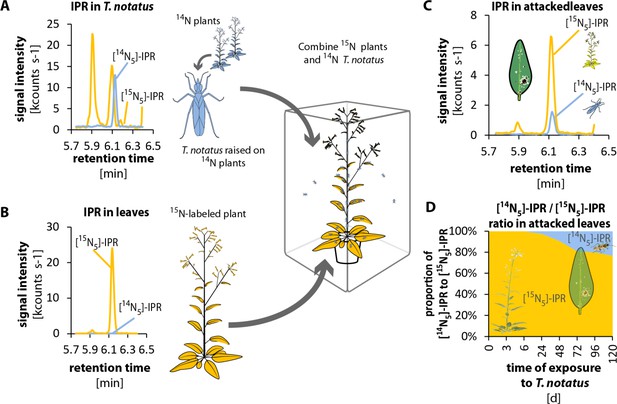
Tupiocoris notatus transfers IPR to leaves of its host plant.
(A) and B) Experimental setup and chromatograms of IPR: (A) T. notatus raised on 14N-grown hydroponic plants contain only [14N5]-IPR in their bodies. (B) Plants raised on a hydroponic medium containing only a 15N containing N-source contain only [15N5]-IPR in their leaves. 15N labeled plants and 14N labeled insects were placed in the same cage for 5 days. Ratio of [14N5]-IPR (originating from insects, blue) and [15N5]-IPR (from host plant, orange) were determined in attacked leaves. (C) Chromatograms of [14N5]-IPR and [15N5]-IPR in the leaves of 5d attacked plants. (D) Ratio of [14N5]-IPR and [15N5]-IPR at different harvest times after start of the exposure to T. notatus (N = 5). For raw data see Raw_data_FIGURE_4_S1 (Dryad: Brütting et al., 2018).
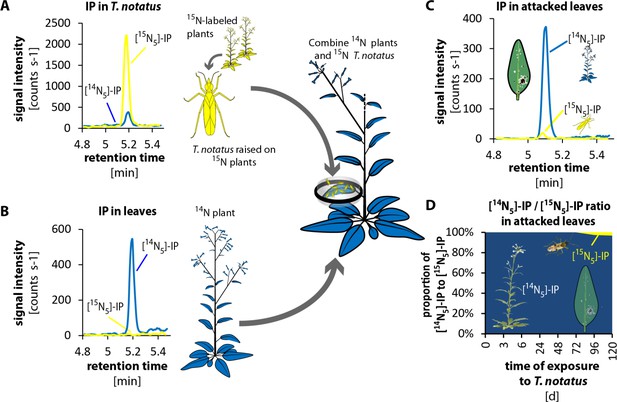
Twenty Tupiocoris notatus individuals transfer detectable amounts of IP to leaves.
(A) and (B) Experimental setup and chromatograms of IP: (A) T. notatus raised on 15N-grown hydroponic plants contain only [15N5]-IP in their bodies. (B) Plants raised on a hydroponic medium containing only 14N containing N-source contain only [14N5]-IP in leaves. 20 15N-labeled insects were placed in small cages on one leaf of 14N-grown plants for 5 days. Ratio of [15N5]-IP (originating from insects, yellow) and [14N5]-IP (from host plant, blue) were determined in attacked leaves. (C) Chromatograms of [15N5]-IP and [14N5]-IP in the leaves that were attacked for 5d. (D) Ratio of [15N5]-IP and [14N5]-IP at different harvest times after the start of exposure to T. notatus (N ≥ 3). For raw data see Raw_data_FIGURE_4_S2 (Dryad: Brütting et al., 2018).
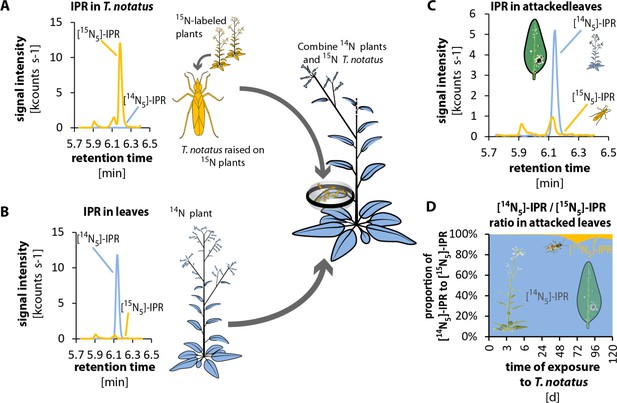
Twenty Tupiocoris notatus individuals transfer detectable amounts of IPR to leaves.
(A) and (B) Experimental setup and chromatograms of IP: (A) T. notatus raised on 15N-grown hydroponic plants contain only [15N5]-IPR in their body. (B) Plants raised on a hydroponic medium containing only 14N containing N-source contain only [14N5]-IPR in leaves. 20 15N labeled insects were placed in small cages on one leaf of 14N-grown plants for 5 days. Ratio of [15N5]-IPR (originating from insects, orange) and [14N5]-IPR (from host plant, blue) were determined in attacked leaves. (C) Chromatograms of [15N5]-IPR and [14N5]-IPR in the leaves that were attacked for 5d. (D) Ratio of [15N5]-IPR and [14N5]-IPR at different harvest times after the start of exposure to T. notatus (N ≥ 3). For raw data see Raw_data_FIGURE_4_S3 (Dryad: Brütting et al., 2018).
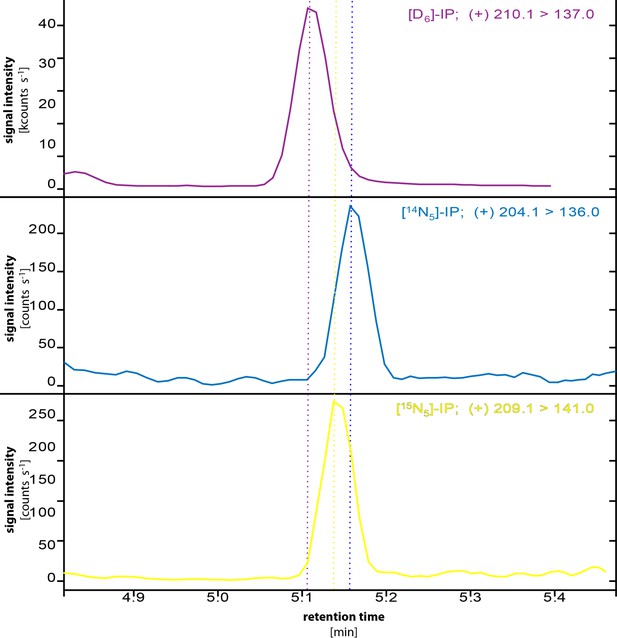
Chromatograms of IP, [D6]-IP, [15N5]-IP.
Dashed lines show the retention-time shifts between unlabeled [14N5]-IP, [D6]-IP (internal standard) and [15N5]-IP. Color coding follows that of the chromatograms. The monitored parental → daughter ion transitions are given in the top right of each chromatogram.
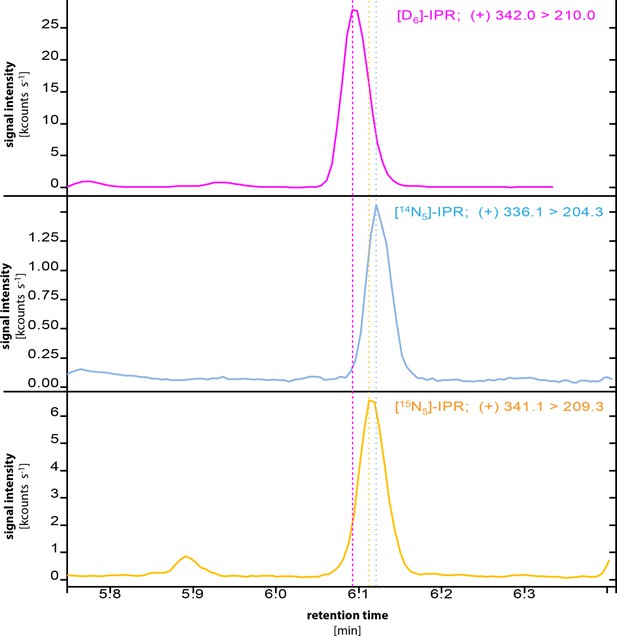
Chromatograms of IPR, [D6]-IPR and [15N5]-IPR.
Dashed lines show the retention-time shifts between unlabeled [14N5]-IP, [D6]-IP (internal standard) and [15N5]-IP. Color coding follows that of the chromatograms. The monitored parental → daughter ion transitions are given in the top right of each chromatogram.
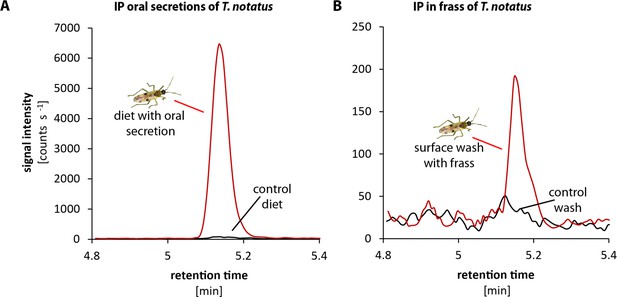
Tupiocoris notatus contain large quantities of IP in their saliva and small amounts in their frass.
Chromatograms showing the signal intensity of an MS/MS- trace for IP (204.1 → 136.0). (A) IP signal of pure sugar solution (black line) or sugar solution fed upon by T. notatus for 5 days (red line). The sugar solution was covered with a thin layer of parafilm that allowed piercing and feeding on the solution and prevented contamination by T. notatus frass. (B) Chromatograms of the surface wash of the parafilm covering the sugar solution after T. notatus feeding (red line, covered with visible frass spots) or without (control wash, black line). Chromatograms shown represent one out of six replicates.
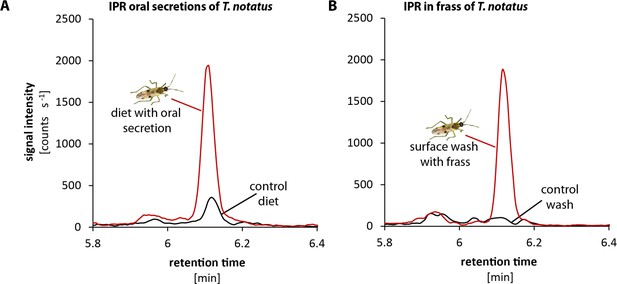
Tupiocoris notatus contain IPR in their saliva and frass.
Chromatograms showing the signal intensity of a MS/MS- trace for IPR (336.1 → 204.1). (A) IPR signal of pure sugar solution (black line) or sugar solution that has been used as diet for T. notatus for 5 days (red line). The sugar solution was covered with a thin layer of parafilm that allowed piercing and feeding on the solution and prevented contamination with T. notatus frass. (B) Chromatograms of the surface wash of the parafilm covering the sugar solution after T. notatus feeding (red line, covered with visible frass spots) or without (control wash, black line). Chromatograms shown represent one out of six replicates.
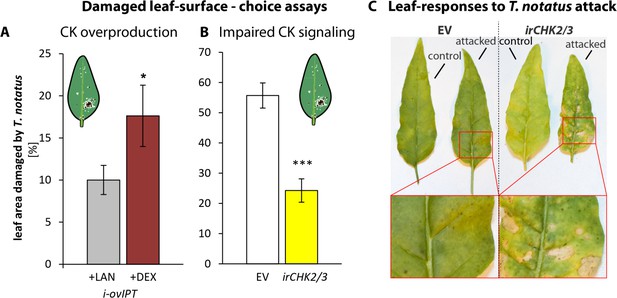
Cytokinin-regulated traits mediate Tupiocoris notatus feeding preferences and alter leaf responses to feeding.
(A) and (B): Surface damage on N. attenuata plants after 10 d of T. notatus feeding. (A) T. notatus could choose between dexamethasone-inducible isopentenyltransferase-overexpressing plants (i-ovipt) treated with dexamethasone-containing lanolin paste (+DEX) or lanolin paste without dexamethasone as control (+LAN; figure based on data from Schäfer et al., 2013). Statistically significant differences were identified with pairwise t-test: N = 7, p=0.032. (B) Choice between empty vector (EV) and irchk2/3 plants silenced in the two cytokinin receptor genes NaCHK2 and NaCHK3 (irchk2/3). Pairwise t-test: N = 6, p<0.001. Error bars depict standard errors. *p<0.05, ***p<0.001. (C) Representative pictures of leaves of EV or irchk2/3 plants with or without T. notatus damage. Magnifications show necrotic lesions occurring only in irchk2/3 plants after several days of mirid feeding. For raw data see Raw_data_FIGURE_6 (Dryad: Brütting et al., 2018).
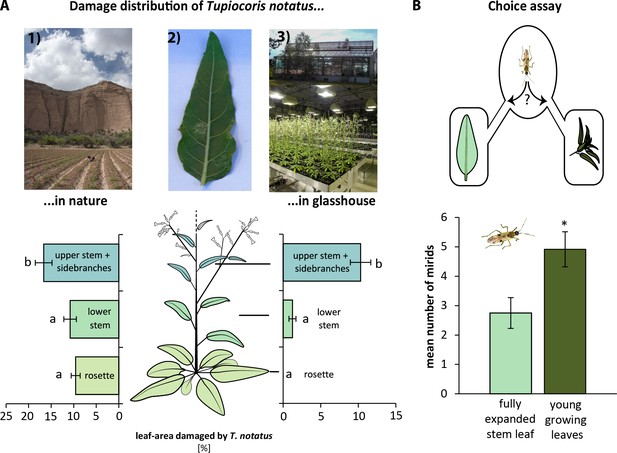
Tupiocoris notatus prefers to feed on young leaves.
(A) Distribution of T. notatus damage in flowering Nicotiana attenuata plants grown in the field and glasshouse. Upper panel: picture 1) field plot at Lytle Preserve, Utah, 2) a typically damaged leaf, 3) glasshouse and plants in the glasshouse. Lower panel: Damaged leaf area in % in field and in glasshouse. Leaves were classified as rosette leaves, lower to mid stem leaves and upper stem leaves and side branches as indicated in the schematic drawing (left). One-way ANOVAs followed by Tukey HSD post hoc tests were used to identify statistically significant differences. Field plants: N = 21, F2,33 = 5.729, p=0.007; glasshouse plants: N = 4, F2,9 = 45.5, p<0.001. Different letters indicate significant differences (p<0.01), error bars depict standard errors. (B) Choice assay: 10 mirids were placed in an arena with two tubes connected to either a fully-grown leaf or young growing leaves with an apical stem. Number of mirids on each side was counted after 12 hr. Pairwise t-test: N = 12, p=0.026. Error bars depict standard errors. For raw data see Raw_data_FIGURE_6_S1 (Dryad: Brütting et al., 2018).
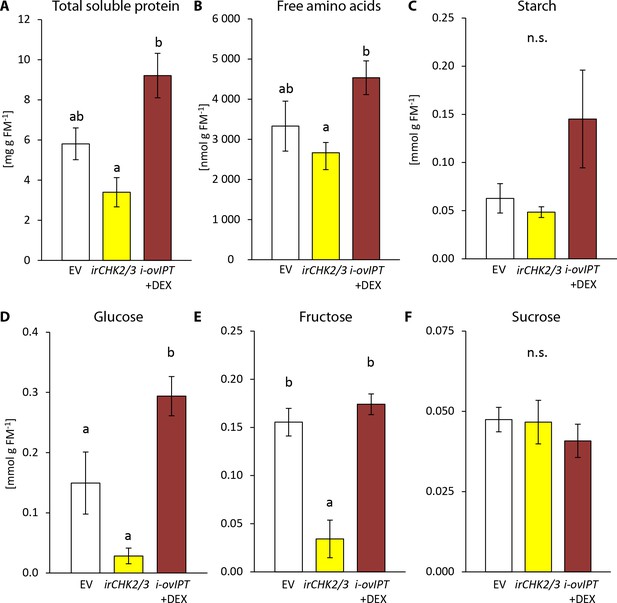
Transgenic Nicotiana attenuata plants altered in their cytokinin metabolism are also altered in their nutrient contents.
We compared nutrient contents of empty vector (EV) plants, plants silenced in the two cytokinin receptor genes NaCHK2 and NaCHK3 (irchk2/3) and dexamethasone-inducible isopentenyltransferase-overexpressing plants (i-ovipt) treated with dexamethasone-containing lanolin paste (DEX) leading to spatially-regulated increased CK levels. Concentrations were determined in untreated rosette leaves of N. attenuata: (A) protein, (B) free amino acids, (C) starch, (D) glucose, (E) fructose and (F) sucrose. Significant differences were identified with one-way ANOVAs followed by Tukey HSD post hoc tests. (A) protein: F2,9 = 10.74, p=0.004; (B) free amino acids: F2,9 = 4.27, p=0.050; (C) log2 transformed starch: F2,9 = 2.208, p=0.166; D) glucose: F2,9 = 18.89, p<0.001; (E) fructose: F2,9 = 15.43, p=0.001; (F) sucrose: F2,9 = 0.375, p=0.698. Error bars depict standard errors (N = 4). FM: fresh mass. For raw data see Raw_data_FIGURE_7 (Dryad: Brütting et al., 2018).
Additional files
-
Supplementary file 1
Calculations of the minimum amount of IP transferred by a single mirid in clip-cage experiment and estimation of the number of feeding mirids required to transfer the measured amount of IP in the whole-plant experiment.
- https://doi.org/10.7554/eLife.36268.025
-
Supplementary file 2
Sequences of primers used for real-time qPCR.
- https://doi.org/10.7554/eLife.36268.026
-
Supplementary file 3
Multi-reaction monitoring settings for the quantification of [14N5]-, [15N5]- and deuterated cytokinins in positive ionization mode.
- https://doi.org/10.7554/eLife.36268.027
-
Transparent reporting form
- https://doi.org/10.7554/eLife.36268.028





