Heterogeneous absorption of antimicrobial peptide LL37 in Escherichia coli cells enhances population survivability
Figures
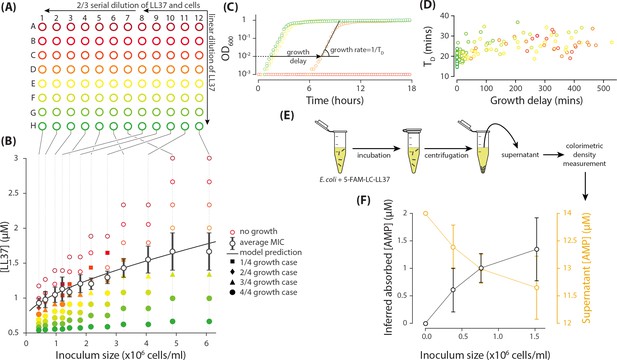
Measurement of the inoculum effect and peptide absorption by E. coli cells.
(A) A two-dimensional dilution scheme, which includes a linear dilution of LL37 peptides in columns 7 and 12 followed by two separate 2/3 dilution series of the cells and LL37 peptides on columns 12 to 8 and 7 to 1. (B) Each well represents a different combination of densities of LL37 peptides and E. coli cells from which we can extract the MIC as a function of inoculum size by monitoring growth of the culture in individual wells. The solid data points refer to the wells with growing culture and different marker symbols refer to the number of repeated trial outcomes that resulted in growing cultures. The empty data points refer to wells with no visible growth. A theoretical model developed later in this work nicely fits the average MIC. Data represent four biological repeats where the average and standard deviations of MIC are depicted with black symbols and lines. (C) The growth of the cultures were monitored by an automated plate reader in terms of OD600. Growing cultures reach a yield comparable to each other while non-growing cultures do not exhibit consistent increase in OD600. Data are examples from column 11 of Figure 1AB and they follow the same color coding. (D) Analysis of the growth in sub-MIC cultures reveal that growth is delayed depending on the LL37 concentration, but the doubling time of the cells shows no considerable change. Data are from the same experiments as in panel B and follow the same color coding as panels A and B. (E) Through colorimetric measurement of the concentration of a fluorescently tagged analogue of LL37 peptide (5-FAM-LC-LL37), we can quantify the amount of peptides remaining in the supernatant after incubation with E. coli cells. (F) The amount of 5-FAM-LC-LL37 peptides remaining in the supernatant decreases with inoculum size (the initial AMP concentration is 14 µM). The amount of absorbed AMPs by the cells are inferred by subtracting the final (supernatant) from the initial concentration of AMPs. The results are the average of 4 biological repeats. Average and standard deviations are depicted in the figure.
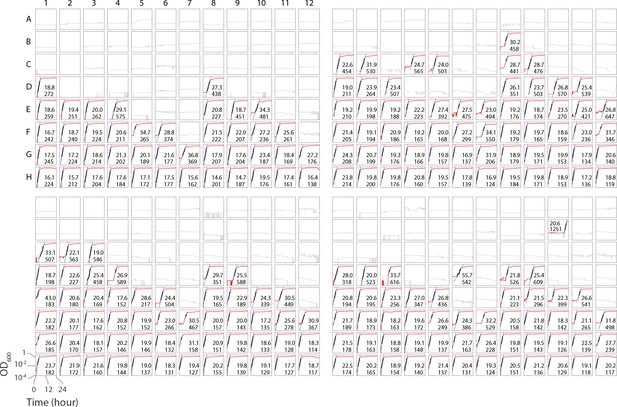
The full experimental data obtained from the microplate reader consisting of the four trials performed as reported in Figure 1 of the main text.
Each box refers to one well, where the red curve shows the growth in terms of OD600, the black overlap shows the detected exponential growth region, and the blue line depicts the exponential fit to the data. The two numbers in each box represent the calculated doubling time (in the top) and the T0.1 (in the bottom) both in terms of minutes. The T0.1 refers to the time point where the OD600 first hits 0.1. The boxes with no number and gray curve refer to the wells that did not show any growth.
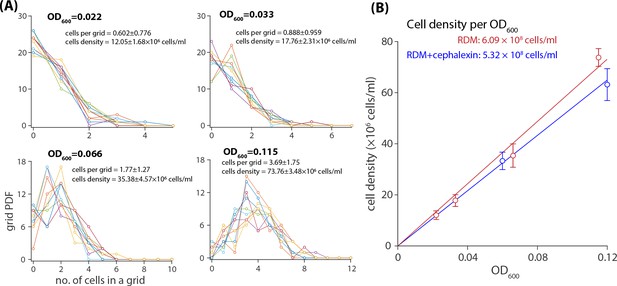
(A) A histogram of E.coli population grown in RDM to various OD600 in hemocytometer grids with a measured volume of . (B) The linear regression of the cell density as a function of OD600 provides a conversion factor for the calculation of respective cell densities beginning with OD600. Red refer to the cells grown in RDM and blue refer to a separate experiment where the same strain was grown in RDM in the presence of μg∕mL cephalexin.
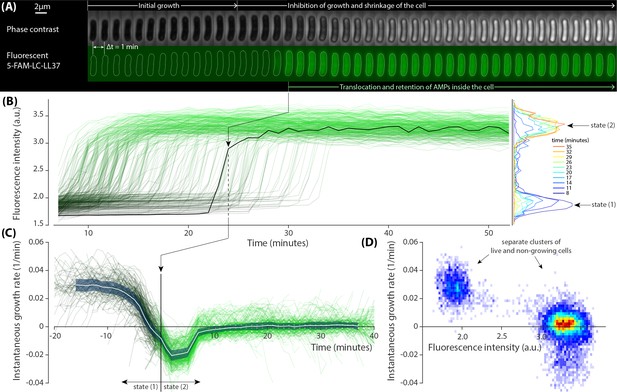
Growth inhibition of E. coli cells by dye-tagged LL37 peptides.
(A) Sample phase contrast and fluorescent time-lapse images of an E. coli cell treated with a lethal dosage of the dye-tagged LL37 peptides (5-FAM-LC-LL37). Phase contrast images show inhibition of growth and slight shrinkage of the cell after a brief growth period. The fluorescent channel shows the distribution, translocation and retention of the peptides. (B) Abrupt transition in the fluorescent signal of 383 cells that occur over a span of more than 30 min. The black line corresponds to the sample shown in panel A. The fluorescence intensity histogram is depicted for different time points on the right. (C) The instantaneous growth rate of individual cells collapse on each other when plotted in referenced to the peptide translocation point. The average behavior (white line) of the collapse shows that the translocation happens shortly after the inhibition of growth and shrinkage of the cell. The shaded area denotes the standard deviation. (D) The two dimensional distribution of instantaneous growth rate and fluorescent signal from all time points depict well-separated clusters referring to a binary response of the cells to the peptides.
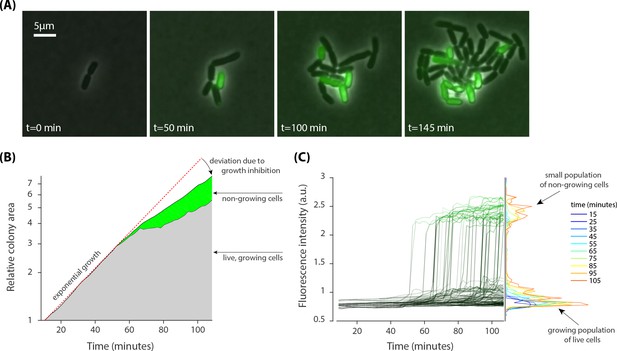
Heterogenous growth inhibition and peptide absorption at a sub-MIC concentration of peptides.
(A) Sample time-lapse images (overlay of phase contrast and fluorescent image) of dividing E. coli cells show that growth of only some cells is inhibited in a growing colony. Phase contrast images show growth of a micro-colony and the fluorescent channel (green) shows translocation of the dye-tagged LL37 peptides (5-FAM-LC-LL37) in the cells whose growth is inhibited. (B) Relative growth of the total area of 13 separate colonies, consisting of 280 cells, depicts initial exponential expansion (grey area) until the appearance of non-growing subpopulation (green area). (C) An abrupt transition in fluorescent signal is observed when growth is inhibited in cells. The transition for individual cells is similar to that in above MIC cultures (see Figure 2).
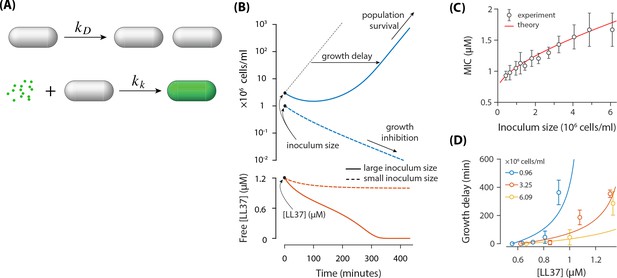
A theoretical model based on the absorption of peptides in E. coil cells.
(A) E. coli cells replicate with a rate of and they get killed with a rate of Each dead cell quickly absorbs AMPs. (B) Demonstration of the inoculum effect for the initial AMP concentration [LL37]=1.2 μM. A culture with high inoculum size ( cells/ml) survives as peptides deplete (solid lines) whereas growth in small inoculum size ( cells/ml) is inhibited with excess peptides remaining in the solution (dashed lines). Despite the survival of culture with high inoculum size, the growth is delayed. (C) Comparison of the MIC between the theoretical model and the experimental data. (D) Growth delay as a function of [LL37]. Solid lines are theoretical results (not a fit) and circles represent experimental data from microplate experiments (Figure 1). The delay is calculated with respect to the lowest AMP concentration (row H) of Figure 1AB.
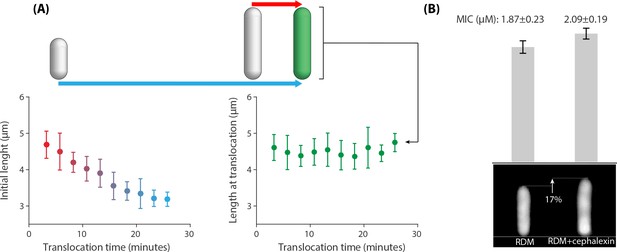
Correlations between the peptide translocation time and cell size for 383 cells that were treated with an above-MIC concentration of 5-FAM-LC-LL37 peptides reported in Figure 2.
(A Left panel) A negative correlation between initial cell length and AMP translocation time indicates resistance of small cells to peptides until a later time points in their life cycle. (A Right panel) Cell length at translocation time and the time of translocation are not correlated. Average and standard deviation are depicted for the data binned based on the x axis. (B) Average cell size affects the MIC values. Addition of μM∕mL of cephalexin to the growth media (RDM) increases cell size by 17% (sample newborn cells shown in the bottom panel), which resulted in an increase in the MIC. The inoculum size of cells/ml are used for this experiment. Twelve replicates with three biological repeats were used for this data.
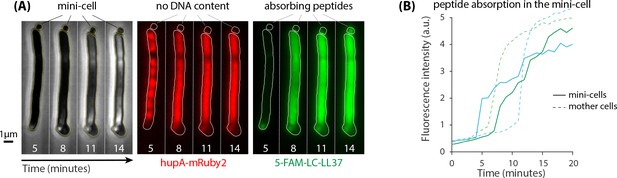
minCDE strain of E.coli was used to test the significance of interactions between peptides and DNA as well as other intracellular content.
(A) Lack of septum positioning system produces enucleated mini-cells (left panel) that do not contain DNA as indicated by the localization of hupA-mRuby2 protein (middle panel). Mini-cells absorb a large amount of 5-FAM-LC-LL37 peptides similar to wild-type cells (right panel) (B) Fluorescence intensity per area of two mini-cells suggests a significant peptide interactions with enucleated mini-cells. The rate of absorption of AMPs in mini-cells is slower than in the neighboring mother cells with DNA content.
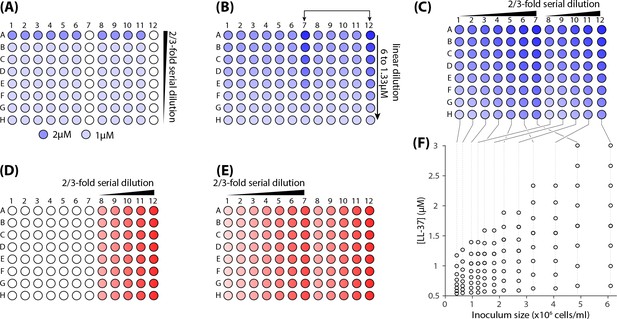
Steps for preparing a two-dimensional dilution series of LL37 and cells.
(A) 50 l/well of a 1 M solution of LL37 in the growth medium (RDM) was transferred to all of the wells excluding row A, columns 7 and 12. Next, 150 l/well of a 2 M solution was transferred to the wells of row A, leaving columns 7 and 12 blank. A 2/3-fold serial dilution was then carried out vertically on all rows, A through H. (B) 150 l/well of a linear dilution series of LL37 was was transferred to columns 7 and 12, with highest concentrations located in row A, while the lowest in row H. The concentrations chosen were 6, 5.34, 4.67, 4, 3.33, 2.67, 2 and 1.33 M. (C) A 2/3-fold serial dilution was carried out horizontally from Columns 12 through 8 and Columns 7 through 1. The final volume of the solution in each well was 100 l. (D) A cell culture was diluted to a final OD600 of 0.02 and 50 l/well was transferred into the wells located in column 12. A 2/3-fold dilution series was performed in a separate reservoir and 50 l/well was transferred to column 11. The process was repeated for columns 10, 9, and 8. (E) The cell culture diluted to the OD600 = 0.016 (80% of the culture used for column 12). 50 l/well was transferred into the wells located in column 7. For columns 6 through 1, the same volume was transferred to each well after repeating 2/3-fold dilutions of this culture. (F) The final concentration of LL37 and cell densities in the microplate.
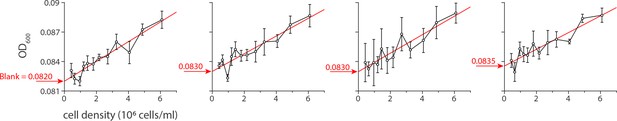
The value of the blank OD600 was determined by extrapolating the relationship between the initial readings of the OD600 generated by the plate reader against the known cell densities of the culture used in the microplate.
https://doi.org/10.7554/eLife.38174.014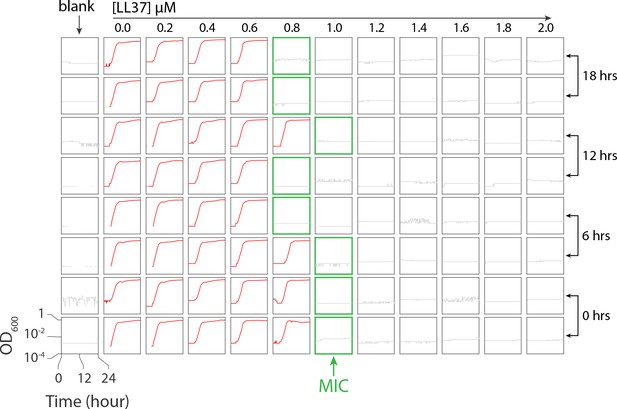
The microplate scheme for testing stability of LL37 in 37°C.
The ‘blank’ column contains growth media to ensure the absence of contaminants in the media. Columns 2 through 12 contain a linear concentration gradient of LL37 from 0 to 2 M. Four different pre-incubation times of LL37 solution in 37°C were tested. The red curve depicts bacterial growth, whereas the grey curve corresponds to wells for which growth was inhibited. Green borders refer to the MIC consisting of the lowest LL37 concentration responsible for inhibiting growth.
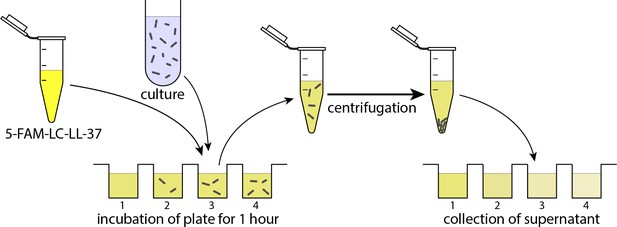
The schematic illustration of the experimental procedure for measuring the remaining peptides in the solution after absorption by bacteria.
First, the solution of 5-FAM-LC-LL37 in RDM and the cell culture is transferred to and incubated at 37°C in a microplate. The culture is then collected and centrifuged. The supernatant is transferred to another plate for fluorescence reading. The fluorescent signal is correlated with the peptide density. The final concentration of peptides (wells on the right) is dependent on the cell density.
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.38174.011





