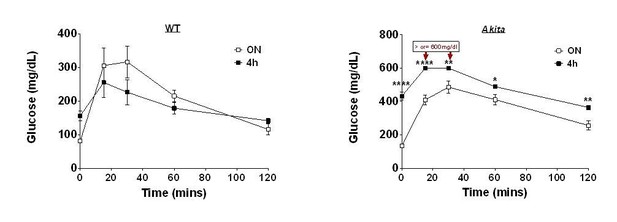
Inhibition of mTORC1 by ER stress impairs neonatal β-cell expansion and predisposes to diabetes in the Akita mouse
Figures
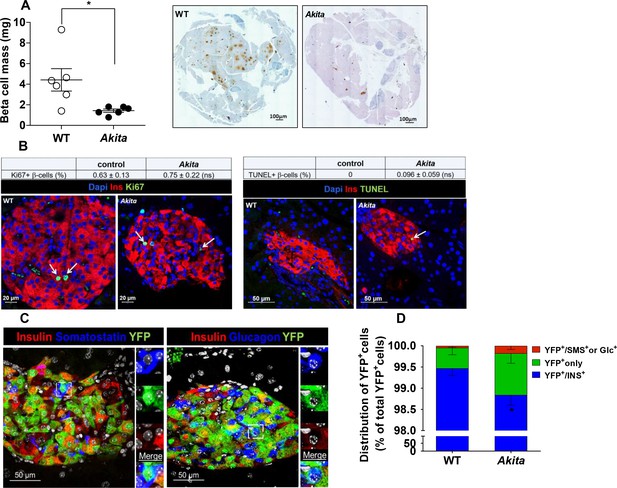
β-Cell mass, turnover and differentiation in adult Akita mice.
Analyses were performed on 2- to 3-month-old Akita mice and age-matched controls. (a) β-cell mass (n = 6 in each group); (b) β-cell proliferation and apoptosis assessed by staining for insulin and Ki67 (n = 6–7 mice in each group; a total of 4909 wild type (WT) and 2523 Akita β-cells were quantified) or TUNEL (n = 4–5 mice in each group; 2592 WT and 1754 Akita β-cells). The percentage of Ki67+ and TUNEL+β-cells is shown in the table above; (c–d) β-cell differentiation was assessed by lineage tracing. Wild-type and Akita mice were crossed with RIP-Cre:Rosa26-Yfp reporter mice; (c) pancreatic sections of Akita mice were immunostained for insulin and somatostatin or glucagon. Lineage-traced β-cells (YFP+) expressing somatostatin or glucagon is shown in squares and zoomed in; (d) quantification of insulin-expressing β-cells (percentage of insulin+/YFP+ cells), insulin-degranulated β-cells (percentage of insulin-/YFP+ cells) and of cells with misexpression of somatostatin or glucagon (percentage of somatostatin+ or glucagon+/YFP+ cells) in WT and Akita mice is shown; *p<0.05.
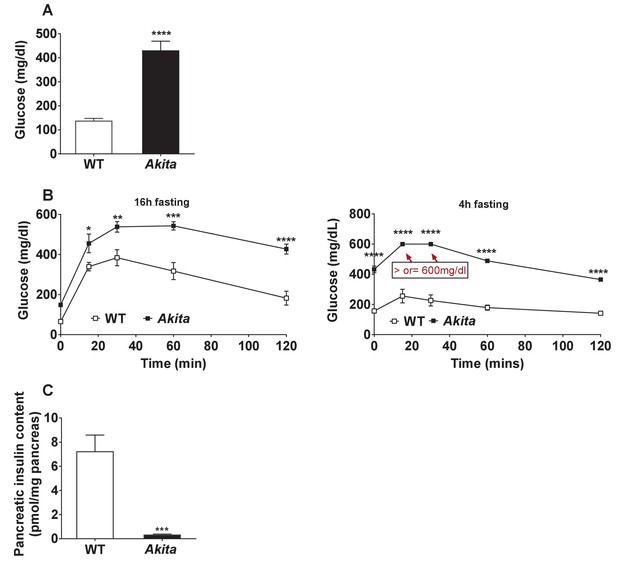
Glycemia and β-cell function in adult Akita and control mice.
(a) Fed blood glucose (n = 11–15 in each group); (b) IPGTT- glucose (1.5 g/kg) was injected intraperitoneally after an overnight or 4 hr fast (n = 4–5 in each group); (c) pancreatic insulin content analyzed by ELISA on whole pancreas extracts (n = 4–5 in each group). *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001.
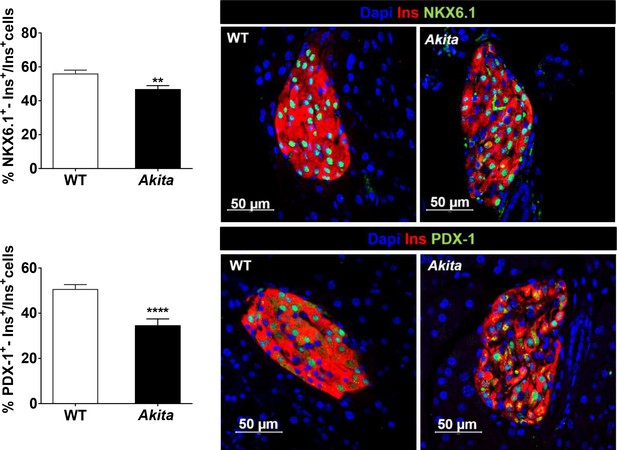
NKX6.1 and PDX-1 expression in adult Akitaβ-cells.
Pancreatic sections of 2- to 3-month-old Akita and control mice were stained for NKX6.1 (n = 5 mice in each group; 3348 WT and 2370 Akita β-cells) or PDX-1 and insulin (n = 5–7 mice in each group; 4580 WT and 2320 Akita β-cells). Values are mean ± SE. **p<0.01, ****p<0.0001.
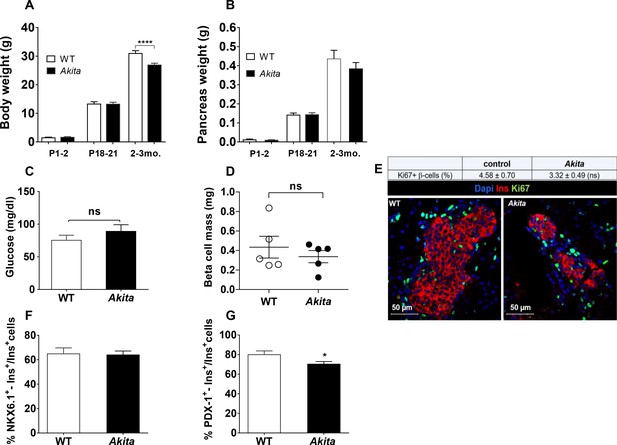
Dynamic changes of body and pancreas growth and glycemia, β-cell mass, proliferation and differentiation in Akita and control mice at P1-2.
(a) body weight, (b) pancreas weight of wild-type and Akita mice at P1-2, P19-21 and at the age of 2–3 months. (a) P1-2: WT (n = 8); Akita mice (n = 4), P19-21: WT (n = 21); Akita mice (n = 23), 2–3 months: WT (n = 33); Akita mice (n = 39); (b) P1-2: WT (n = 8); Akita mice (n = 4), P19-21: n = 14 in each group, 2–3 months: n = 17 mice in each group). (c) fed blood glucose (n = 7–8 mice in each group); (d) β-cell mass (n = 5 mice in each group); (e) β-cell proliferation assessed by immunostaining for insulin and Ki67 (n = 4 mice in each group; 1886 WT and 1483 Akita β-cells). The percentage of Ki67+ β-cells is shown in the table above; (f–g) quantification of β-cells (insulin+) expressing NKX6.1 (n = 3–4 mice in each group; 1148 WT and 1808 Akita β-cells) and PDX-1 (n = 3–5 mice in each group; 1364 WT and 1507 Akita β-cells). *p<0.05, ****p<0.0001.
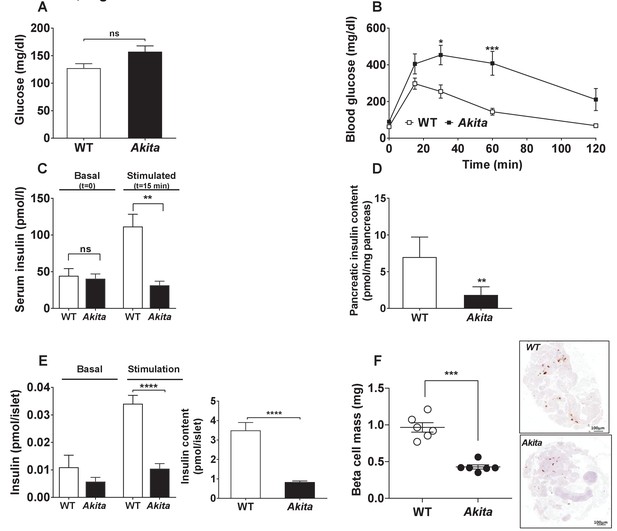
Metabolic state, β-cell function and mass in pre-weaning (P19-21) Akita mice and age-matched controls.
(a) fed blood glucose (n = 7 in each group); (b) IPGTT- glucose (1.5 g/kg) was injected intraperitoneally after an overnight fast (n = 5 in each group); (c) glucose-stimulated insulin secretion in vivo. Insulin was measured before and 15 min following IP glucose injection (1.5 g/kg); (d) pancreatic insulin content (n = 4–5 in each group); (e) basal (3.3 mmol/l glucose) and stimulated (16.7 mmol/l glucose) insulin secretion and insulin content of Akita and control islets analyzed by static incubations. Islets were divided into 4 batches of 25 islets per group (n = 3); (f) β-cell mass (n = 6 mice in each group). *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001.
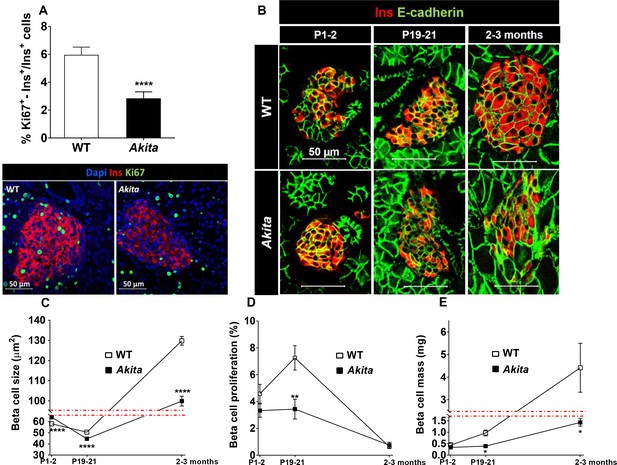
Dynamic changes in β-cell expansion in Akita and control mice.
(a) β-cell proliferation assessed by immunostaining for insulin and Ki67 (n = 6 mice in each group; 2541 WT and 3391 Akita β−cells); (b) β-cell size at P1-2 (newborn, n = 4–5 mice in each group; 334 WT and 435 Akita β-cells), P19-21 (pre-weaning, n = 3 mice in each group; 330 WT and 364 Akita β−cells) and in adult mice (2–3 month-old, n = 3 mice in each group; 266 WT and 417 Akita β−cells) assessed by immunostaining for E-cadherin and insulin. Quantifications of β-cell size (c), proliferation (d), and mass (e) are shown. *p<0.05, **p<0.01, ****p<0.0001.
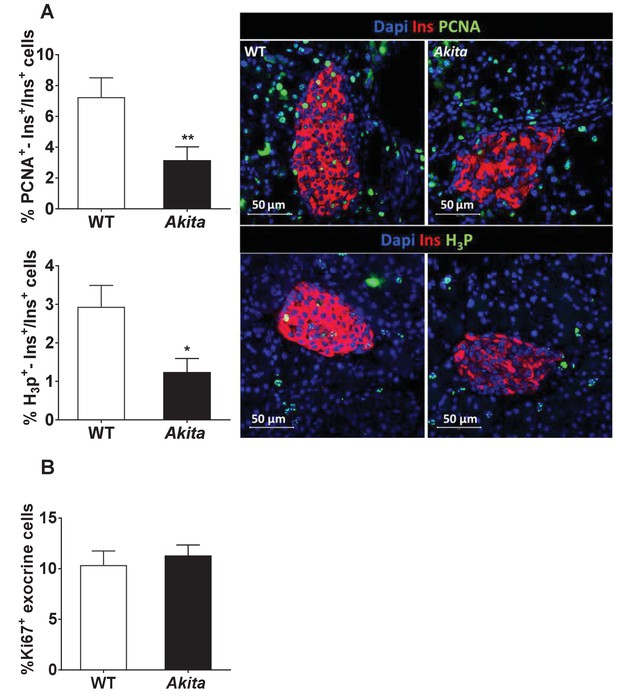
Proliferation of β-cells and exocrine cells in pre-weaning Akita and control mice.
Analyses were performed on pancreatic sections of Akita mice and age-matched controls at 19–21 days stained for proliferation markers. (a) β-Cell proliferation was assessed by staining for insulin and PCNA (n = 3 mice in each group; 1241 WT and 1944 Akita β cells), or phospho-Histone H3 (H3P n = 3 mice in each group; 1176 WT and 1982 Akita β−cells). (b) Proliferation of pancreatic exocrine cells in pre-weaning Akita and control mice. Pancreatic sections of pre-weaning Akita and control mice were stained for insulin and Ki67 and exocrine cells surrounding the islets were used for quantification. The percentage of proliferating exocrine cells (Ki67+/INS-) is shown (n = 6 mice in each group; 2486 WT and 2670 Akita cells). *p<0.05, **p<0.01.
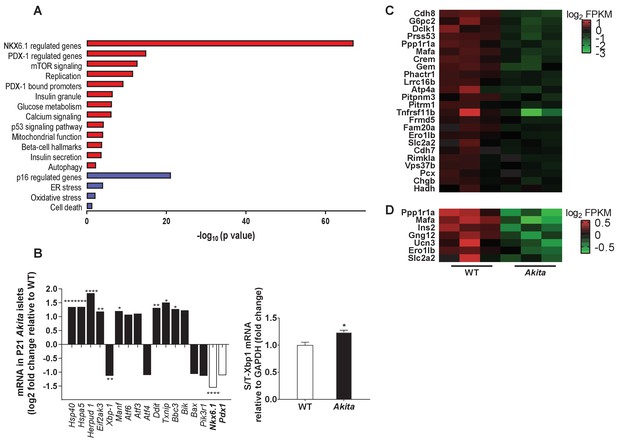
Transcriptomic analysis of ER stress markers and β-cell gene signature in neonate Akita islets.
(a) RNA-seq comparing the transcriptome of islets from P19-21 Akita and age-matched control mice (n = 3 samples in each group, each sample is a pool of islets from three mice). Columns represent pathways that are differentially regulated in Akita mice; (b) expression of UPR and apoptosis genes and of Nkx6.1 and Pdx1 in islets of Akita compared to control mice at P19-21. Spliced and total Xbp1 were also quantified by qPCR. The spliced/total Xbp1 ratio is shown beside (n = 3); (c–d) heat map of genes regulated by NKX6.1 (c) and PDX-1 (d) in Akita islets and controls. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001.
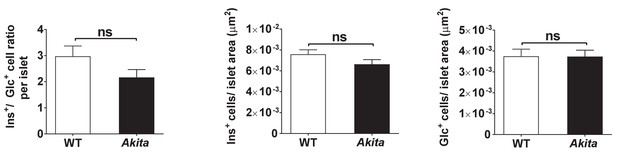
Islet composition of wild-type and Akita mice.
β/α cell ratio and number of β and α cells per islet area in P19-21 wild-type and Akita mice (n = 44 and 35 islets isolated from three to four mice in each group).
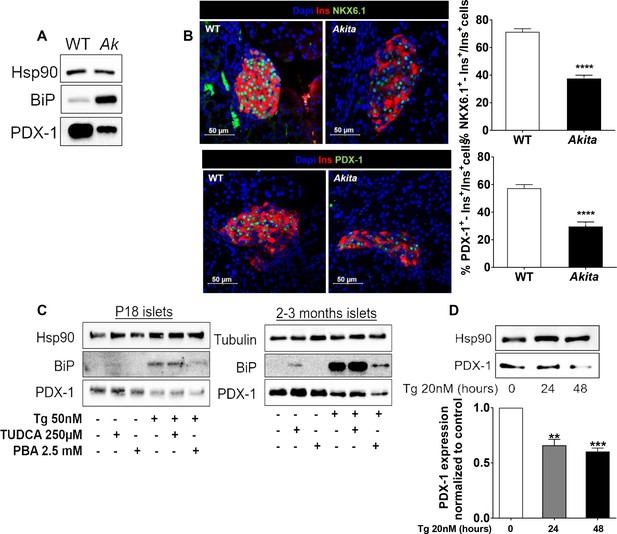
Effects of ER stress on the expression of β-cell transcription factors in neonate Akita islets (P19-21) and islets treated with thapsigargin.
(a) PDX-1 and BiP protein level analyzed by Western blotting (n = 3, each sample is a pool of islets from four to six mice); (b) quantification of NKX6.1 (n = 3 mice in each group; 1646 WT and 728 Akita β−cells), and PDX-1 (n = 3 mice in each group; 1534 WT and 844 Akita β−cells) expressing β-cells. Pancreatic sections were immunostained for NKX6.1 or PDX-1 and insulin. The percentage of NKX6.1- and PDX-1-positive β-cells is shown. (c) Islets from young (P19-21) and adult wild-type mice were treated with low-dose thapsigargin (50 nmol/l) and TUDCA (250 µmol/l) or PBA (2.5 mmol/l) for 48 hr with daily media changes and further analyzed by western blotting for PDX-1 and BiP (n = 3, each sample is a pool of islets from six to nine mice); (d) INS-1E cells were treated with 20 nmol/l thapsigargin for 24 and 48 hr followed by western blotting for PDX-1. **p<0.01, ***p<0.001, ****p<0.0001.
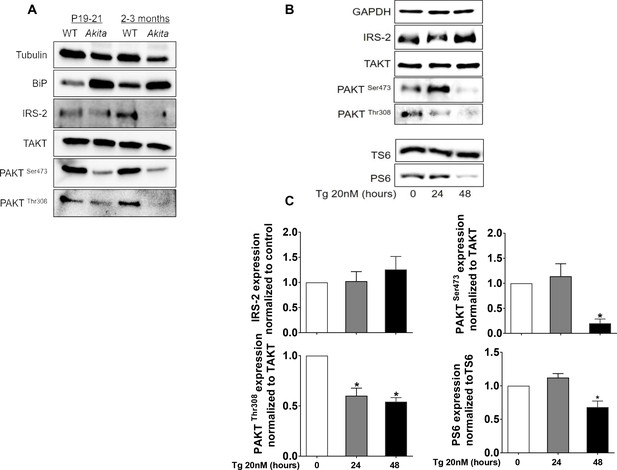
Effects of ER stress on IRS2/Akt signaling in Akita islets and in INS-1E treated with low-dose thapsigargin.
(a) IRS2/Akt signaling in islets from neonate (P19-21) and adult wild-type and Akita mice. Each sample is a pool of islets from 4 to 15 mice (n = 4 for neonate islets and n = 2 for adult islets). (b–c) INS-1E cells were treated with 20 nmol/l thapsigargin for 24 and 48 hr followed by western blotting for IRS2, total and phosphorylated Akt (Ser473 and Thr308) and S6 (Ser240/244). A representative experiment (b) and quantification (c) are shown (n = 4–6). *p<0.05.
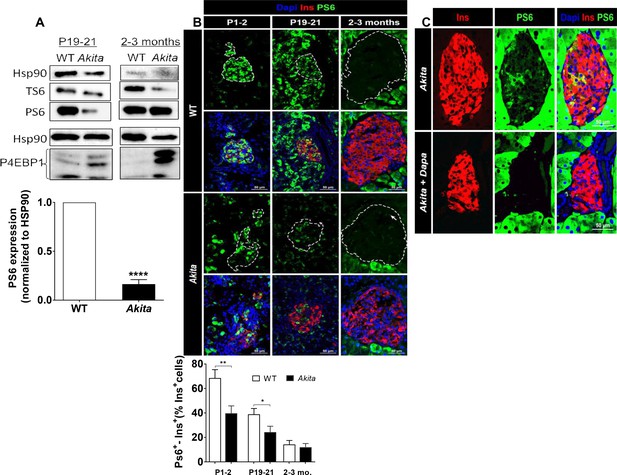
mTORC1 signaling in neonate and adult Akita islets.
(a) Western blot analysis of S6 and 4EBP1 phosphorylation in islets of neonate (P19-21) and adult wild-type and Akita mice. Quantification of phosphorylated S6 in neonate Akita compared to control islets is shown (n = 3, each sample is a pool of islets from 4 to 7 mice); (b) immunostaining for phospho-S6 on pancreatic sections of P1-2, P19-21 and adult Akita mice and age-matched controls and quantifications of the percentage of S6+ β-cells (P1-2: n = 4 mice in each group; 1159 WT and 1655 Akita β−cells; P19-21: n = 6 mice in each group; 2259 WT and 1567 Akita β−cells; adult: n = 4–5 mice in each group; 2391 WT and 1383 Akita β-cells). Islet boundaries are marked by dotted line; (c) adult Akita mice were treated with 25 mg/kg dapagliflozin in drinking water for 72 hr. Blood glucose in dapagliflozin-treated Akita mice was ~ 200 mg/dl compared to ~ 500 mg/dl in control Akita mice. Pancreatic sections were immunostained for insulin and phospho-S6 (n = 3 mice in each group). *p<0.05, **p<0.01, ****p<0.0001.
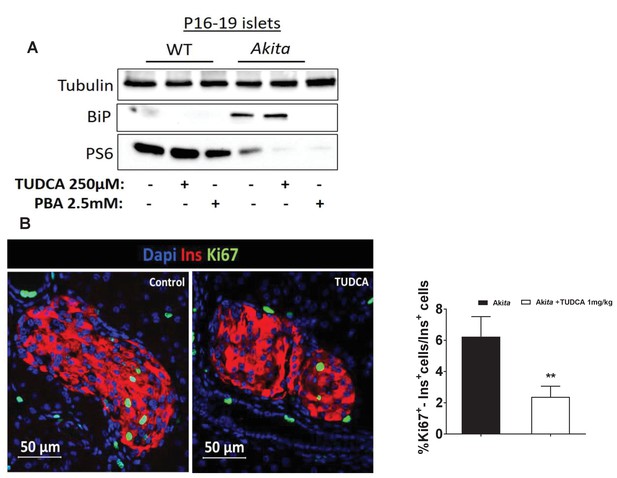
Effects of chemical chaperones on mTORC1 activity in neonate Akita islets and controls.
(a) islets of P16-19 Akita and wild-type (WT) mice were treated with 250 μmol/l TUDCA or 2.5 mmol/l PBA for 48 hr followed by western blotting for BiP and phosphorylated S6 (n = 3, each sample is a pool of islets from 4 to 9 mice); (b) effects of TUDCA on β-cell proliferation in neonate Akita mice (P18-20). TUDCA (1 mg/kg) was injected IP twice daily for 48 hr followed by immunostaining of pancreatic sections for Ki67 and insulin (n = 4–6 mice in each group; 1403 control Akita and 2183 TUDCA-treated Akitaβ-cells). **p<0.01.
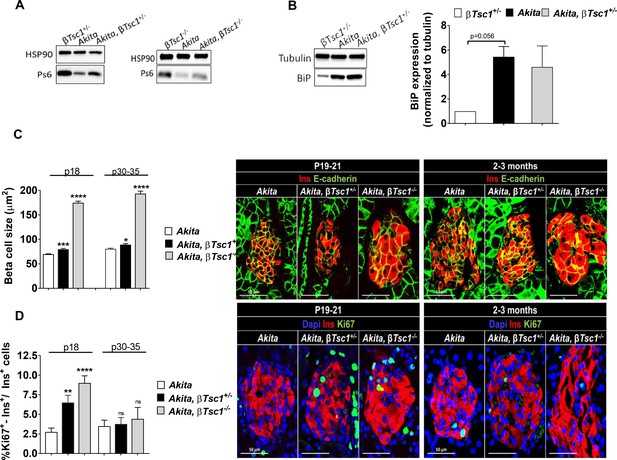
Effects of mTORC1 activation in neonate Akita β-cells on β-cell size and proliferation.
Studies were performed on heterozygous and homozygous βTsc1 knockout Akita mice (RIP-Cre:Tsc1flox/+:Akita (Akita, βTsc1+/-) and RIP-Cre:Tsc1flox/flox:Akita (Akita, βTsc1-/-). Tsc1flox/+:Akita and Tsc1flox/flox:Akita were used as Akita controls. RIP-Cre:Tsc1flox/+ mice (βTsc1+/-) and RIP-Cre:Tsc1flox/flox mice (βTsc1-/-) were used as WT controls (a, b). (a) Western blotting for phospho-S6 on islets from homozygous and heterozygous knockout mice and matched controls (n = 4, each sample is a pool of islets from two to four mice); (b) Western blotting and quantification of BiP expression in wild-type, Akita and Akita, βTsc1 +/- mice (n = 4, each sample is a pool of islets from two to four mice); (c) β-cell size was assessed by immunostaining for insulin and E-cadherin (n = 400–500 β-cells per group), (d) β-cell proliferation was assessed by immunostaining for insulin and Ki67 (n = 1200–1400 β−cells per group). Quantifications and representative images are shown. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001.
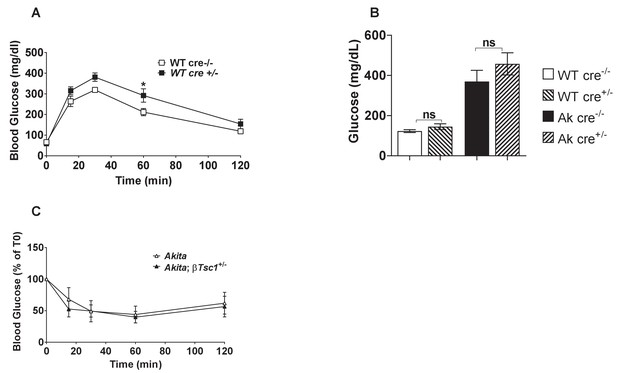
Metabolic characterization of RIP-Cre mouse.
(a) IPGTT- glucose (1.5 gr/kg) was injected after an overnight fast to adult RIP-Cre and non-transgenic control mice (n = 3); (b) fed blood glucose of adult RIP-Cre mice and RIP-Cre:Akita mice compared to non-transgenic controls (n = 6–8); (c) insulin tolerance test on Akita and RIP-Cre:Akita mice (n = 3). *p<0.05.
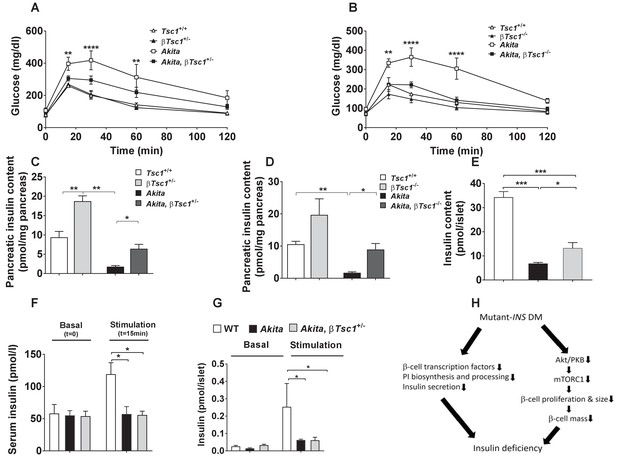
Effects of mTORC1 activation in neonate Akita β-cells on diabetes.
(a–b) IPGTT at P30-35: glucose (1 g/kg) was injected IP after an overnight fast; (a) heterozygous Tsc1 knockout Akita mice (RIP-Cre:Tsc1flox/+:Akita (Akita, βTsc1+/-) and matched controls: Tsc1flox/+ mice (Tsc1+/+), RIP-Cre:Tsc1flox/+ mice (βTsc1+/-), and Tsc1flox/+:Akita (Akita) (n = 3–5 mice in each group); (b) homozygous Tsc1 knockout Akita mice (RIP-Cre:Tsc1flox/flox:Akita (Akita, βTsc1-/-) and matched controls: Tsc1flox/flox mice (Tsc+/+), RIP-Cre:Tsc1flox/flox mice (βTsc1-/-), and Tsc1flox/flox:Akita (Akita) (n = 3–5 in each group); (c–d) pancreatic insulin content of heterozygous and homozygous Tsc1 knockout Akita mice and matched controls at P30-35 (WT (n = 7), Akita (n = 11), Akita, βTsc1+/- (n = 3) and Akita, βTsc1-/- (n = 4); (e) islet insulin content. (f–g) Effects of mTORC1 activation in neonate Akita β-cells on insulin secretion in vivo and ex vivo. (f) insulin secretion in response to IP glucose injection (n = 6 mice in each group); (g) islets were isolated from Tsc1flox/+ WT mice (WT), Tsc1flox/+:Akita (Akita) and RIP-Cre:Tsc1flox/+:Akita (Akita, βTsc1+/-) mice and insulin secretion assessed following static incubations at basal (3.3 mmol/l) and stimulated (16.7) mmol/l glucose. (h) a model of the pathophysiology of permanent neonatal diabetes. *p<0.05 **, p<0.01, ***, p<0.001****, p<0.0001.
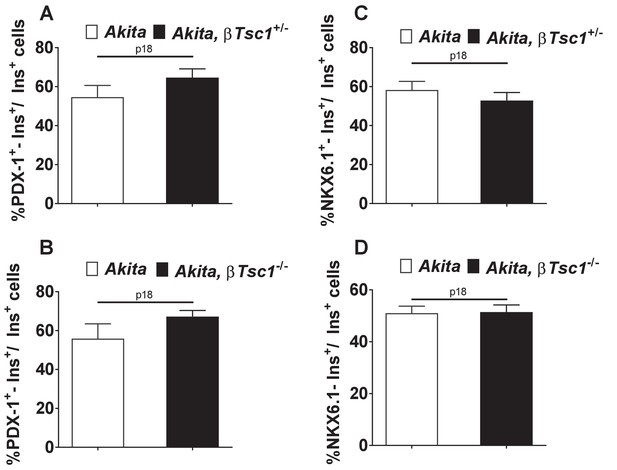
Effects of mTORC1 activation in Akitaβ-cells on PDX-1 (a, b) and NKX6.1 expression (c, d).
Heterozygous (a, c) and homozygous (b, d) βTsc1 knockout Akita and age-matched controls were sacrificed at P18. Pancreatic sections were stained for insulin and PDX-1 (Akita (n = 1760 β-cells); Akita, βTsc1+/- (n = 814 β-cells); Akita, βTsc1-/- (n = 1458 β−cells) or NKX6.1 (Akita (n = 1500 β-cells); Akita, βTsc1+/- (n = 1438 β-cells); Akita, βTsc1-/- (n = 1584 β-cells).
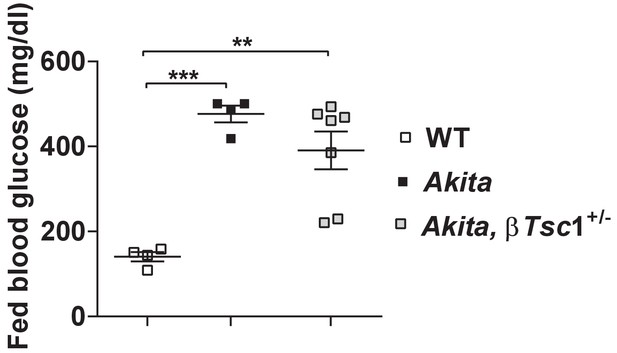
Fed blood glucose of Tsc1flox/+ mice (WT), Tsc1flox/+:Akita (Akita) and heterozygous Tsc1 knockout RIP-Cre:Tsc1flox/+:Akita (Akita,βTsc1+/-) mice at the age of 2–3 months.
Blood glucose levels are the mean of the last three consecutive glucose measurements.
**p<0.01, ***p<0.001.
Tables
Transcriptome changes in P19-21 Akita islets compared to age-matched controls (n = 3 per each group).
https://doi.org/10.7554/eLife.38472.013| Gene symbol | log2 Fold change | p value | Gene symbol | log2 Fold change | p value | |
|---|---|---|---|---|---|---|
| β cell signature | Growth factors and mTOR signaling | |||||
| Pcsk1 | −2.1101 | 1.1341E-13 | Dapp1 | −2.0554 | 7.7092E-09 | |
| Mafa | −1.9801 | 2.1124E-12 | Egfr | −2.0185 | 2.0824E-08 | |
| Igsf11 | −1.7632 | 2.8185E-05 | Cth | −1.9445 | 4.9302E-05 | |
| Insulin II | −1.7070 | 5.9006E-14 | Igf2 | −1.5846 | 4.3954E-03 | |
| Ucn3 | −1.5979 | 2.8193E-06 | Tubg1 | −1.5134 | 3.6446E-06 | |
| Nkx6-1 | −1.5542 | 5.0005E-11 | Sqle | −1.4758 | 5.5074E-07 | |
| Vdr | −1.4299 | 3.0451E-08 | IGF1r | −1.4735 | 1.5160E-05 | |
| Slc2a2 | −1.4352 | 5.3528E-04 | Tpi1 | −1.4516 | 1.4280E-15 | |
| Insulin I | −1.3988 | 1.7436E-11 | Btg2 | −1.4350 | 1.5855E-03 | |
| Nkx2-2 | −1.2408 | 8.1578E-04 | Elovl6 | −1.3484 | 3.1052E-03 | |
| Elovl5 | −1.3484 | 6.7970E-08 | ||||
| Insulin secretion, Insulin granules | Mllt11 | −1.3283 | 9.1320E-04 | |||
| Sytl4 | −2.1903 | 6.6331E-18 | Ppa1 | −1.3162 | 4.9816E-04 | |
| Pcsk1 | −2.1101 | 1.1341E-13 | Uchl5 | −1.3085 | 8.1939E-03 | |
| Vgf | −2.1003 | 2.1515E-18 | IGF2r | −1.2440 | 1.9131E-05 | |
| Gng12 | −1.6254 | 3.0041E-07 | ||||
| Syt5 | −1.5143 | 1.3141E-04 | Cell cycle, replication | |||
| Iqgap1 | −1.4997 | 2.8284E-08 | Pak6 | −2.1602 | 1.0026E-12 | |
| Chrm3 | −1.4571 | 1.2136E-05 | Tmem71 | −2.0697 | 3.8133E-12 | |
| Gng4 | −1.4365 | 2.7040E-06 | S100a10 | −2.0519 | 9.5186E-07 | |
| Chgb | −1.3739 | 1.4523E-03 | Spc25 | −1.8739 | 1.1632E-05 | |
| Gpr119 | −1.3712 | 2.6136E-03 | Mpp6 | −1.8457 | 3.7249E-06 | |
| Ptprn | −1.3342 | 1.7018E-04 | Plagl1 | −1.8253 | 1.5119E-12 | |
| Nup93 | −1.7850 | 1.1110E-06 | ||||
| Calcium signaling | Orc6 | −1.6260 | 1.5634E-04 | |||
| Npy | −3.2026 | 3.9755E-13 | Tmem144 | −1.6091 | 2.9040E-03 | |
| Crem | −1.9204 | 2.4035E-09 | Vrk1 | −1.5578 | 1.0175E-08 | |
| Gem | −1.8187 | 2.6277E-05 | Shmt1 | −1.5410 | 1.6394E-03 | |
| Dusp1 | −1.5182 | 9.3142E-10 | Mcm3 | −1.5330 | 4.7341E-04 | |
| Plat | −1.5137 | 8.2853E-03 | Plch1 | −1.5305 | 6.9677E-11 | |
| Tpcn2 | −1.4906 | 3.5030E-03 | Hells | −1.5287 | 2.7324E-03 | |
| Mif | −1.4597 | 9.0441E-04 | Mns1 | −1.5270 | 5.1788E-06 | |
| Vcl | −1.3870 | 3.6093E-03 | Plat | −1.5137 | 8.2853E-03 | |
| Serca2 | −1.2529 | 5.7379E-03 | Tubg1 | −1.5134 | 3.6446E-06 | |
| Dnmt1 | −1.4820 | 3.7757E-06 | ||||
| ER sress | Junb | −1.4651 | 5.6951E-04 | |||
| Herpud1 | 1.8461 | 4.6638E-24 | Pcna | −1.4557 | 7.8393E-04 | |
| Nucb1 | 1.4045 | 3.0361E-08 | Cast | −1.4562 | 9.4477E-04 | |
| Hspa5 | 1.3517 | 4.1005E-04 | Net1 | −1.4507 | 1.6503E-03 | |
| Dnajc3 | 1.3468 | 3.2649E-06 | Myo5a | −1.4252 | 1.2734E-03 | |
| Ddit3 | 1.3153 | 7.5196E-03 | Alms1 | −1.4229 | 1.1049E-03 | |
| Manf | 1.2009 | 3.2893E-02 | Chaf1a | −1.4137 | 6.1889E-03 | |
| Lig1 | −1.4101 | 9.1110E-04 | ||||
| Oxidative stress | Ramp2 | −1.3860 | 4.0237E-03 | |||
| Gstp1 | 1.6177 | 6.5804E-06 | Nphp4 | −1.3854 | 8.2551E-03 | |
| Txnip | 1.5075 | 1.0152E-02 | Mcm6 | −1.3587 | 5.1079E-03 | |
| Gstz1 | 1.4388 | 4.9681E-04 | Ywhah | −1.3561 | 7.3805E-05 | |
| Tubb4b | −1.3543 | 1.8881E-04 | ||||
| Cell death | Rgs3 | −1.3424 | 3.2137E-04 | |||
| Card14 | 4.3450 | 8.5040E-32 | Bex2 | −1.3389 | 5.1152E-04 | |
| Gdf15 | 3.1410 | 1.6785E-27 | Clic1 | −1.3291 | 3.9543E-04 | |
| Bmp3 | 2.5051 | 5.7201E-18 | Polh | −1.3169 | 9.9122E-03 | |
| Proc | 2.0628 | 1.5909E-07 | Tpm4 | −1.3125 | 3.3364E-04 | |
| Rorc | 1.9612 | 7.4298E-07 | Uchl5 | −1.3085 | 8.1939E-03 | |
| Bdnf | 1.8487 | 8.2780E-05 | Kpnb1 | −1.3081 | 3.7374E-05 | |
| Herpud1 | 1.8461 | 4.6638E-24 | Phf6 | −1.3048 | 5.1520E-05 | |
| Creb3l1 | 1.7775 | 5.6240E-11 | Pitpnm1 | −1.3032 | 5.6223E-04 | |
| Eph7 | 1.7461 | 1.7641E-04 | Aim1 | −1.3023 | 9.1456E-03 | |
| Pde3a | 1.7236 | 2.3400E-04 | Cdk5rap2 | −1.3022 | 1.7257E-03 | |
| Ascl1 | 1.7219 | 6.1969E-04 | ||||
| Mpz | 1.7049 | 1.2768E-04 | Mitochondria and electron transport chain | |||
| Relt | 1.6992 | 3.9164E-07 | Ndufs2 | −1.6249 | 6.6049E-16 | |
| Cnr1 | 1.6906 | 1.3432E-03 | Sdhc | −1.2008 | 9.8858E-04 | |
| Osgin1 | 1.6732 | 1.0137E-06 | Cox6a2 | −5.1088 | 1.1287E-24 | |
| Vip | 1.6721 | 1.5422E-03 | Cox6c | −1.2150 | 2.0212E-02 | |
| Gstp1 | 1.6177 | 6.5804E-06 | Pdk1 | 2.0052 | 4.3578E-11 | |
| Klf11 | 1.6138 | 6.0024E-05 | Pdk2 | 1.5897 | 1.6104E-04 | |
| Rgn | 1.5784 | 4.3567E-03 | Pdk4 | 1.3010 | 6.7512E-02 | |
| Dlc1 | 1.5682 | 9.9181E-05 | Pcx | −1.3978 | 4.8333E-09 | |
| Rass2f | 1.5651 | 5.6005E-03 | Fh1 | −1.8589 | 1.7194E-07 | |
| Wnt4 | 1.5573 | 4.4795E-08 | ||||
| Tle1 | 1.5488 | 1.0352E-13 | Non-beta cell hormones | |||
| Fgb | 1.5356 | 7.8501E-03 | Glucagon | 1.3727 | 7.0468E-03 | |
| Bmpr1b | 1.5335 | 3.1844E-03 | Somatostatin | 1.1510 | 6.5084E-02 | |
| Pycr1 | 1.5308 | 4.0752E-04 | Pancreatic polypeptide | 1.3546 | 1.9982E-03 | |
| Cd44 | 1.5265 | 9.8606E-03 | Ghrelin | 1.2195 | 1.3036E-01 | |
| Nod1 | 1.5259 | 5.4041E-05 | ||||
| Rasgrf2 | 1.5068 | 2.4891E-04 | ||||
| Dapk1 | 1.5036 | 7.5147E-06 | ||||
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.38472.022