Lysosomal cholesterol export reconstituted from fragments of Niemann-Pick C1
Figures
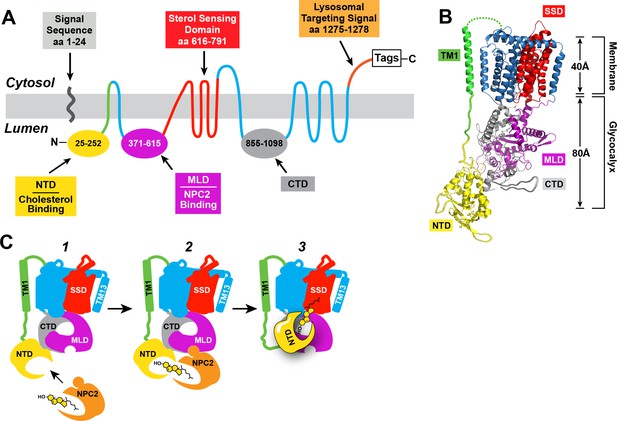
Human NPC1: topology, structure, and function.
(A) Predicted topology of human NPC1 based on the data of Davies and Ioannou (2000). Functional domains of the protein are shown in different colors. NTD, N-terminal domain; MLD, middle lumenal domain; CTD, C-terminal domain. (B) Structure of full-length NPC1 as determined by cryo-electron microscopy (Gong et al., 2016) was accessed from the Protein Data Bank (PDB: 3JD8) and color-matched to the topology map in (A). Image was generated using the PyMOL Molecular Graphics System, Version 2.0 Schrödinger, LLC. TM1, transmembrane helix 1; SSD, sterol-sensing domain. (C) Model for cholesterol transfer. (1) NPC2 brings cholesterol to NPC1, which is embedded in the lysosomal membrane. (2) NPC2 binds to the MLD of NPC1 and transfers its cholesterol to the NTD of NPC1 in a hydrophobic handoff (Kwon et al., 2009; Deffieu and Pfeffer, 2011). (3) Cholesterol is transferred from the NTD to the membrane – embedded SSD.
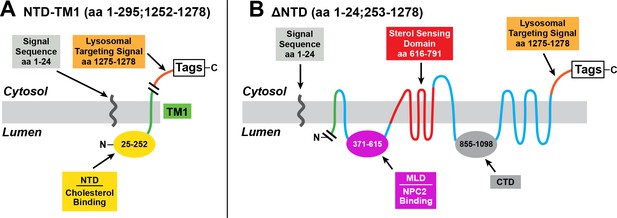
Mutant versions of human NPC1 used in cholesterol esterification assays.
(A) pNTD-TM1 encodes the signal sequence of human NPC1, followed sequentially by the NTD, TM1, a lysosomal targeting signal, and epitope tags. (B) p∆NTD encodes NPC1 with a deletion of the NTD (amino acids 25–252). The cleaved signal sequence is shown.
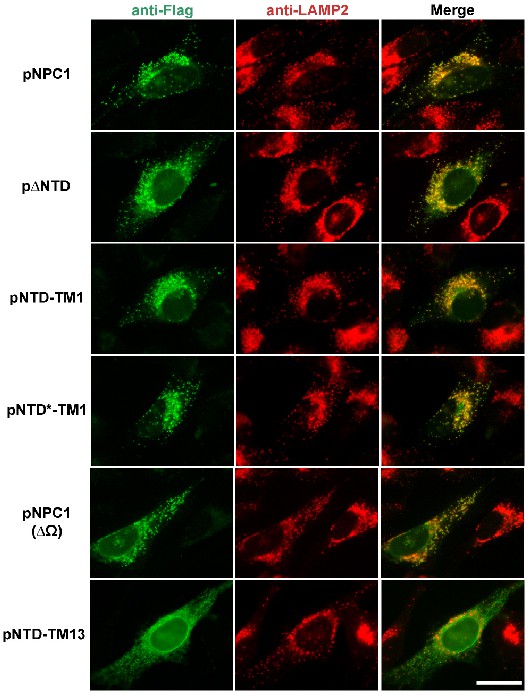
Localization of mutant NPC1 proteins to lysosomes.
SV589 cells were transfected with the indicated plasmids encoding Flag-tagged fragments of NPC1 as described in Materials and methods. Cells were fixed and double stained with 0.8 µg/ml of rabbit monoclonal anti-Flag IgG (green) together with 1 µg/ml of mouse monoclonal anti-LAMP-2 IgG (red), and images were merged (yellow). LAMP-2 is a marker for lysosomes. Immunofluorescence microscopy was performed as described in Materials and methods. Scale bar, 20 µm.
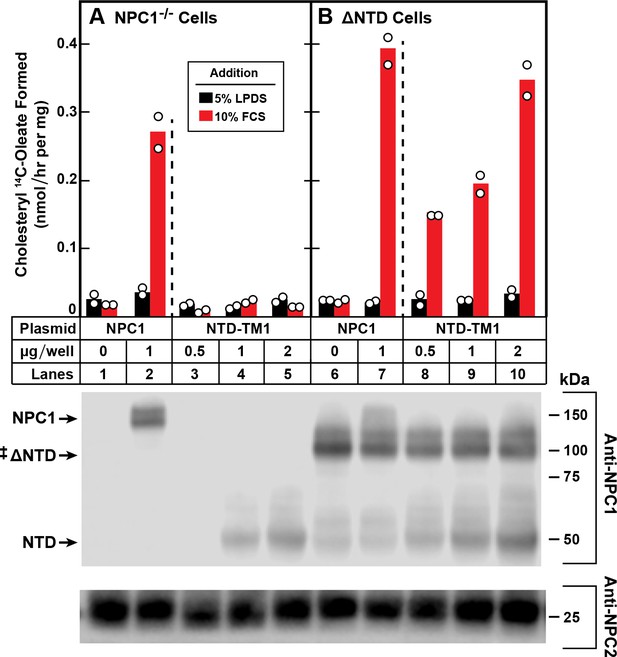
Trans-complementation between NTD and ∆NTD fragments of NPC1.
On day 0, NPC1-/- cells (A) and NPC1-/- cells stably expressing ∆NTD (B) were set up in medium A with 5% FCS at 2.5 × 105 cells/60 mm dish. On day 1, monolayers were switched to fresh medium A (without antibiotics) with 5% LPDS and then transfected with the indicated plasmids encoding either full-length NPC1 or NTD-TM1. All dishes received FuGENE HD and a total of 2 µg DNA/dish adjusted with pcDNA3.1. After incubation for 24 hr, cells were washed once with PBS and switched to medium A with 5% LPDS containing 50 µM sodium compactin and 50 µM sodium mevalonate. On day 3, the cells received fresh medium B containing compactin and mevalonate in the presence of either 5% LPDS or 10% FCS containing lipoproteins. After incubation for 4 hr at 37°C, each cell monolayer was pulse-labeled for 2 hr with 0.1 mM sodium [14C]oleate (8568 dpm/nmol). The cells were then harvested for measurement of their content of cholesteryl [14C]oleate and [14C]triglycerides as described in Materials and methods. Each bar indicates the mean of duplicate incubations with individual values shown. The mean cellular content of [14C]triglycerides in the presence of FCS was not significantly different in NPC1-/- and ∆NTD cells (11.0 and 11.8 nmol/hr per mg protein, respectively). The bottom panel shows immunoblots of whole cell extracts (40 μg) using 0.36 μg/ml of rabbit monoclonal anti-NPC1 and 1.8 μg/ml of mouse monoclonal anti-NPC2. ‡ denotes the endogenous, stably transfected ∆NTD.
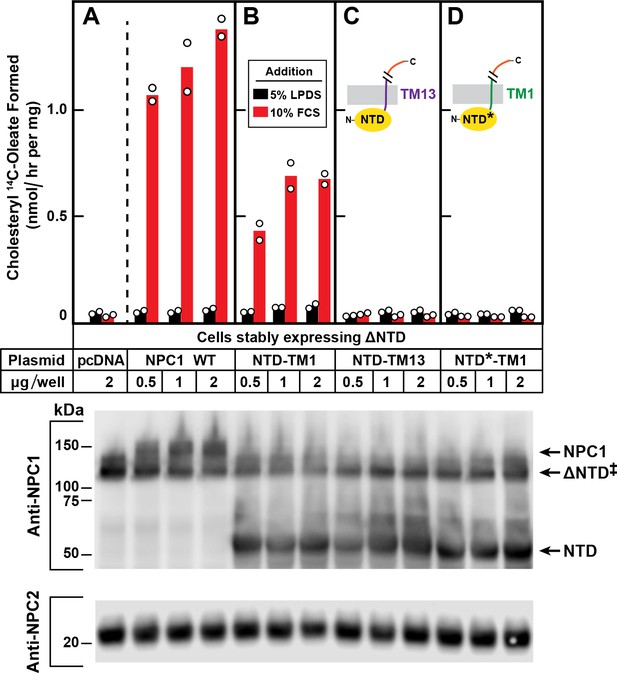
Restoration of cholesterol transport to ∆NTD sequences requires that the NTD of NPC1 localize to lysosomes and bind cholesterol.
On day 0, ∆NTD cells were set up and transfected on day one as described in Figure 4 with the indicated amount of one of the following plasmids: pcDNA3.1 (control) NPC1 (A), pNTD-TM1 (B), pNTD-TM13 (C), or pNTD*-TM1 (D). After incubation for 24 hr, cells were switched to medium A with 5% LPDS containing 50 µM sodium compactin and 50 µM sodium mevalonate. On day 3, the cells received fresh medium B containing compactin and mevalonate in the presence of either 5% LPDS or 10% FCS. After incubation for 4 hr at 37°C, each cell monolayer was pulse-labeled for 2 hr with 0.1 mM sodium [14C]oleate (9019 dpm/nmol). The cells were then harvested for measurement of their content of cholesteryl [14C]oleate and [14C]triglycerides. Each bar indicates the mean of duplicate incubations with individual values shown. The mean cellular content of [14C]triglycerides in the presence of FCS was not significantly different in cells transfected with pNPC1, pNTD-TM1, pNTD-T13, and pNTD*-TM1 (13.3, 12.7, 12.3, and 13.3 nmol per hr/mg protein, respectively). The bottom panel shows immunoblots of whole cell extracts (40 µg/lane) using 0.36 µg/ml of rabbit monoclonal anti-NPC1 and 1.8 µg/ml of mouse monoclonal anti-NPC2. ‡ denotes the endogenous, stably transfected ∆NTD.
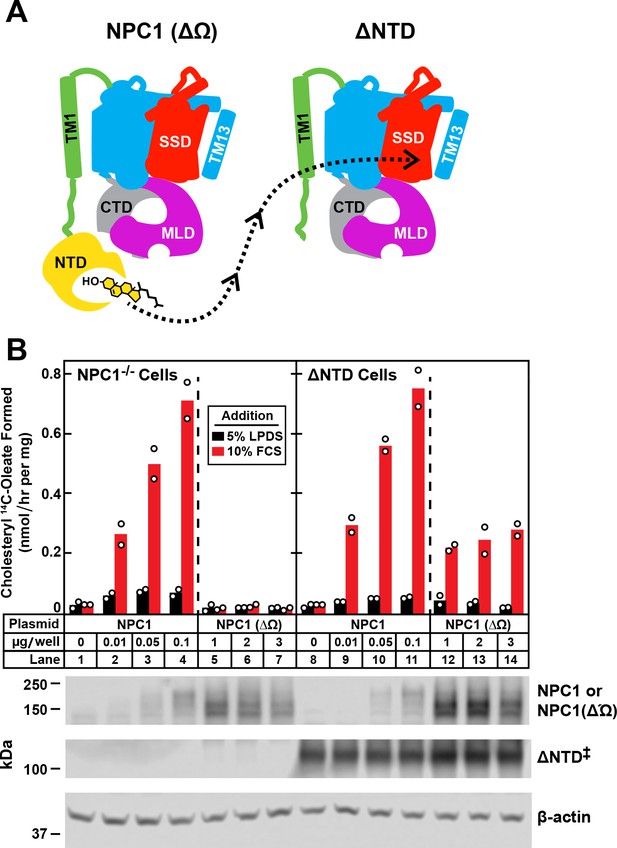
Transfer of cholesterol from NPC1(∆Ω) to ∆NTD as determined by cholesterol esterification assay.
(A) Model showing how the NTD of NPC1(∆Ω) might transfer its cholesterol to the SSD of ∆NTD. (B) Cholesterol esterification assay. On day 0, NPC1-/- cells and ∆NTD cells were set up for experiments as described in Figure 4 and transfected on day one with the indicated plasmids. All dishes contained a total of 3 µg of DNA adjusted with pcDNA3.1. After incubation for 24 hr, cells were washed once with PBS and switched to medium A with 5% LPDS containing 50 µM sodium compactin and 50 µM sodium mevalonate. On day 3, the cells received fresh medium B containing compactin and mevalonate in the presence of either 5% LPDS or 10% FCS. After incubation for 4 hr at 37°C, each cell monolayer was pulse-labeled for 2 hr with 0.1 mM sodium [14C]oleate (9452 dpm/nmol). The cells were then harvested for measurement of their content of cholesteryl [14C]oleate and [14C]triglycerides. Each bar indicates the mean of duplicate incubations with individual values shown. The mean cellular content of [14C]triglycerides in the presence of FCS was not significantly different in NPC1-/- and ∆NTD cells (11.8 and 14.8 nmol/hr per mg protein, respectively). The bottom panel shows immunoblots of whole cell extracts (40 μg) using 3.6 μg/ml rabbit polyclonal anti-NPC1(NTD) that detects both NPC1 ((lanes 2–4, 9–11) and NPC1 (∆Ω) (lanes 5–7, 12–14), 0.36 μg/ml rabbit monoclonal anti-NPC1 that detects ∆NTD* (lanes 8–14), and 0.2 μg/ml of mouse monoclonal anti-β-actin. ‡ denotes the endogenous, stably transfected ∆NTD.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Chemical compound | Sodium dodecyl sulfate (SDS) | Sigma-Aldrich | 71736 | |
| Chemical compound | Benzonase nuclease | Sigma-Aldrich | E1014 | |
| Chemical compound | Bovine serum albumin | Sigma-Aldrich | A7284 | |
| Chemical compound | [1-14C]Oleic acid (50 mCi/mmol) | PerkinElmer, Waltham, MA | NEC317050UC | |
| Chemical compound | SuperSignal West Pico Chemiluminescent Substrate | Thermo Fisher Scientific | 34580 | |
| Chemical compound | Zeocin | Life Technologies, Grand Island, NY | R25005 | |
| Chemical compound | FuGENE HD | Promega Corporation, Madison, WI | E2311 | |
| Chemical compound | Formaldehyde | Sigma-Aldrich | F8775 | |
| Chemical compound | Penicillin-Streptomycin Solution | Corning | 30–002 Cl | |
| Chemical compound | Methanol | Fisher Scientific, Hampton, NH | A412 | |
| Chemical compound | Hexane | Fisher Scientific | H292 | |
| Chemical compound | Isopropanol | Fisher Scientific | A416 | |
| Chemical compound | Heptane | Fisher Scientific | H350 | |
| Chemical compound | Ethyl ether | Fisher Scientific | E138 | |
| Chemical compound | Acetic acid | Fisher Scientific | A38C | |
| Chemical compound | Sodium compactin | Brown et al. (1978) | NA | |
| Chemical compound | Sodium mevalonate | Brown et al. (1978) | NA | |
| Other | L-Glutamine-free DMEM | Sigma-Aldrich | D5546 | culture medium |
| Other | DMEM-low glucose (1000 mg/l) | Sigma-Aldrich | D6046 | culture medium |
| Other | Ham’s F-12 medium and Dulbecco’s modified Eagle’s medium containing 2.5 mM L- glutamine (DMEM) | Corning, Manassas, VA | 10–090-CV | culture medium |
| Other | Newborn calf lipoprotein-deficient serum (LPDS, d < 1.215 g/mL) | Goldstein et al. (1983) | NA | culture serum |
| Commercial assay or kit | Bolt 4–12% Bis-Tris Plus gradient gels | Thermo Fisher Scientific, Waltham, MA | NW04125BOX | |
| Commercial assay or kit | QuikChange II XL Site-Directed Mutagenesis Kit | Agilent Technologies, Santa Clara, CA | 200522 | |
| Antibody | Rabbit monoclonal IgG against Flag | Sigma-Aldrich, St. Louis, MO | F7425, RRID: AB_439687 | |
| Antibody | Mouse monoclonal IgG against LAMP-2 | BD Biosciences, Franklin Lakes, NJ | 555803, RRID: AB_396137 | |
| Antibody | Rabbit monoclonal IgG against amino acids 1261–1278 of human NPC1 | Abcam, Cambridge, UK | ab134113 | |
| Antibody | Rabbit polyclonal IgG against NPC1(NTD)-His8-FLAG | Infante et al. (2008) | Clone 491B | |
| Antibody | Goat anti-rabbit IgG conjugated to AlexaFluor 488 | Invitrogen, Carlsbad, CA | A-11008, RRID: AB_143165 | |
| Antibody | Goat anti-mouse IgG conjugated to AlexaFluor 594 | Invitrogen | A-11005, RRID: AB_141372 | |
| Antibody | Mouse monoclonal HRP-conjugated IgG against β-actin | Cell Signaling Technology, Danvers, MA | 12262, RRID: AB_2566811 | |
| Antibody | Horse anti-mouse IgG conjugated to HRP | Cell Signaling Technology | 7076, RRID: AB_330924 | |
| Antibody | Goat anti-rabbit IgG conjugated to HRP | Cell Signaling Technology | 7074, RRID: AB_2099233 | |
| Antibody | Mouse monoclonal IgG against human NPC2 | Wang et al. (2010) | Clone 13G4 |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.38564.008





