Consolidation alters motor sequence-specific distributed representations
Figures
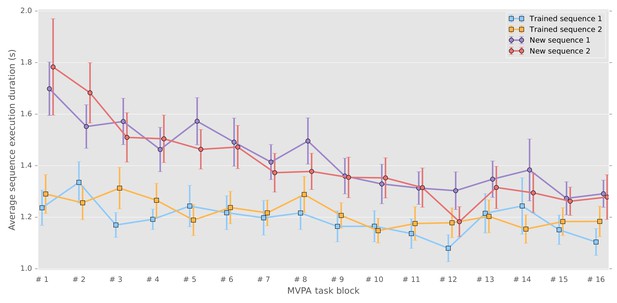
Correct sequence durations (average and standard error of the mean across participants) across the MVPA task blocks.
https://doi.org/10.7554/eLife.39324.002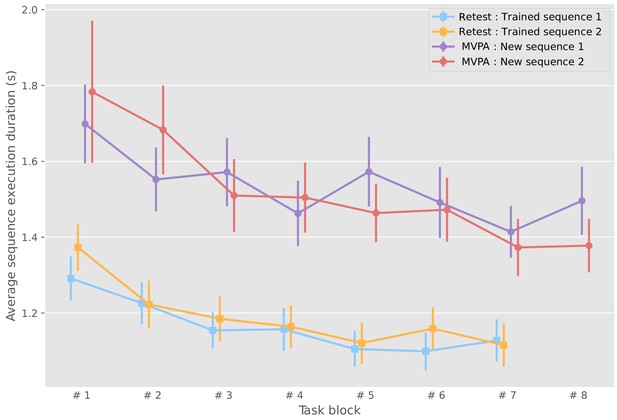
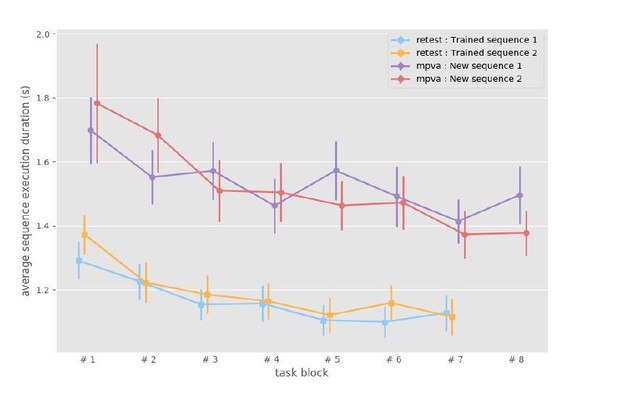
Sequence duration of trained sequences in the first blocks for each sequence on the last experimental day (mean and standard error across participants).
https://doi.org/10.7554/eLife.39324.003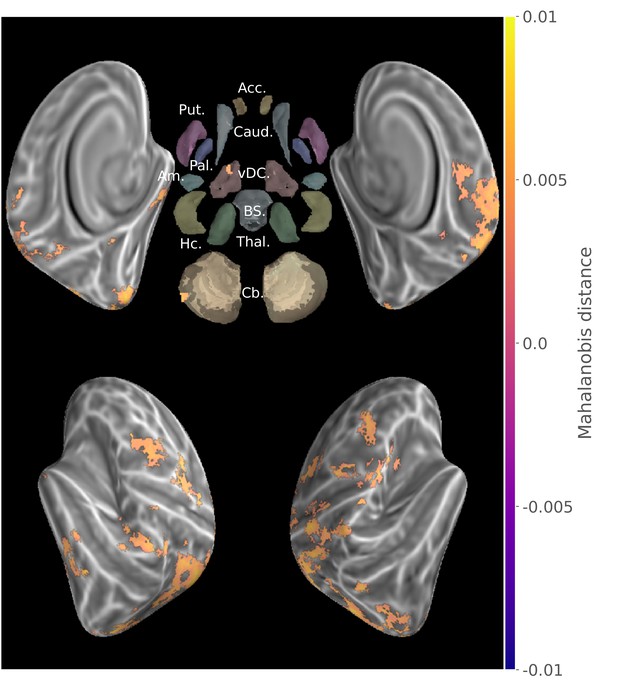
Group searchlight conjunction of new and consolidated sequences discriminability maps (z-score thresholded at p < .05 TFCE-cluster-corrected) showing a large distributed set of cortical regions showing sequence disciminative patterns at both learning stages.
Regions of interest with Freesurfer colors: Acc.:Accumbens; Put.:Putamen; Caud.:Caudate; Pal.:Pallidum; vDC:ventral diencephalon; Am.:Amygdala; Hc.:Hippocampus; Thal.:Thalamus; Cb.:Cerebellum; BS:brain-stem.
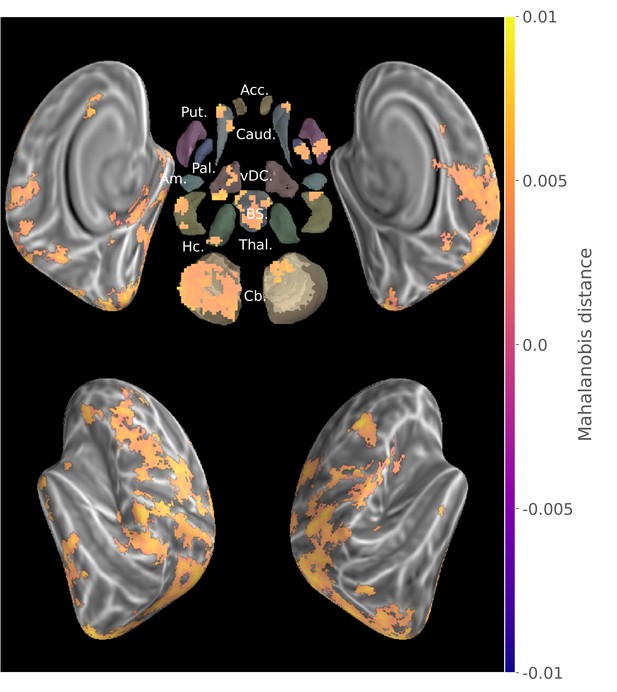
Group searchlight map of cross-validated Mahalanobis distance between the two new sequences (z-score thresholded at p < .05 TFCE-cluster-corrected).
https://doi.org/10.7554/eLife.39324.005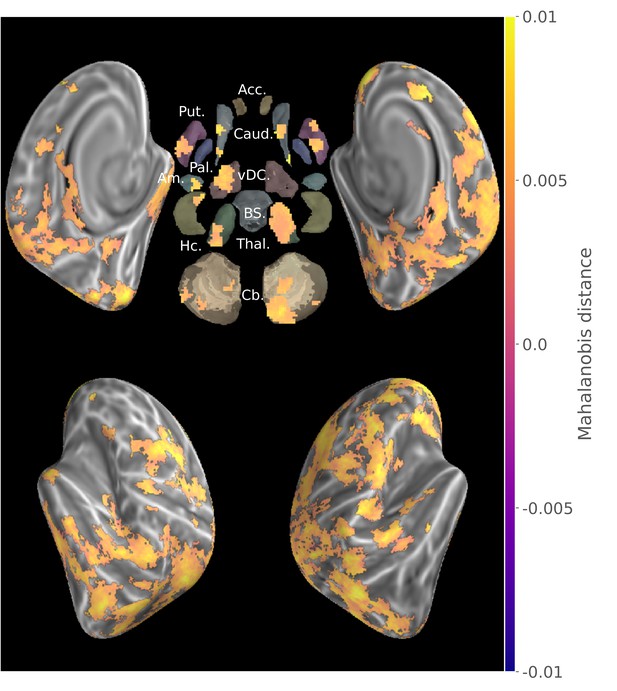
Group searchlight map of cross-validated Mahalanobis distance between the two consolidated sequences (z-score thresholded at p < .05 TFCE-cluster-corrected).
https://doi.org/10.7554/eLife.39324.006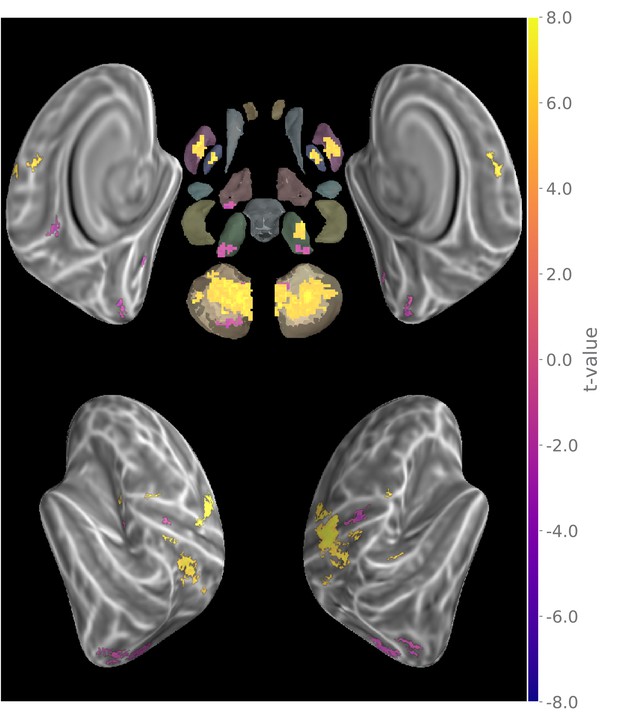
Main effect of motor sequence execution during MVPA task.
(t-value thresholded at p < 05 FDR-corrected).

Illustration of the method: for each neighborhood centered on a gray-matter coordinate, we computed cross-validated Mahalanobis distance matrices.
The two maps of interest, respectively representing the distances between new sequences and between consolidated sequences, were then analyzed at the group level.
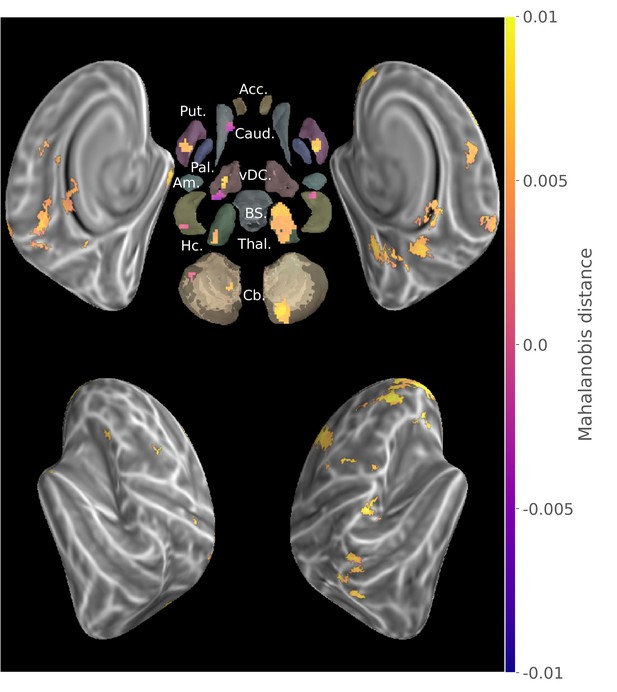
Conjunction of group searchlight contrast (paired t-test) between consolidated and new sequences discriminability maps and separate group discriminability maps for new and consolidated sequences (z-score thresholded at p < .05 TFCE-cluster-corrected) showing a reorganization of the distributed memory trace between these two stages.
Acc.:Accumbens; Put.:Putamen; Caud.:Caudate; Pal.:Pallidum; vDC:ventral diencephalon; Am.:Amygdala; Hc.:Hippocampus; Thal.:Thalamus; Cb.:Cerebellum; BS:brain-stem.
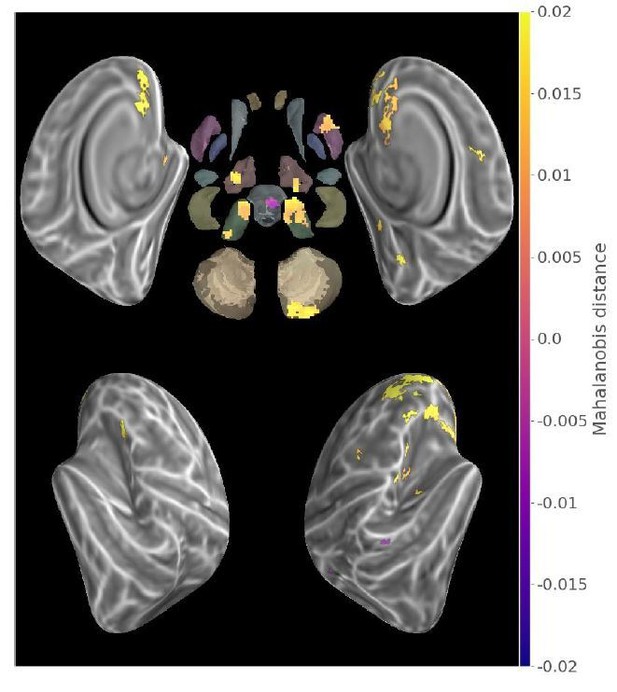
Contrast between within-trained and within-consolidated sequence mahalanobis distances in the second acquisition run (p<.05 Monte-Carlo TFCE corrected).
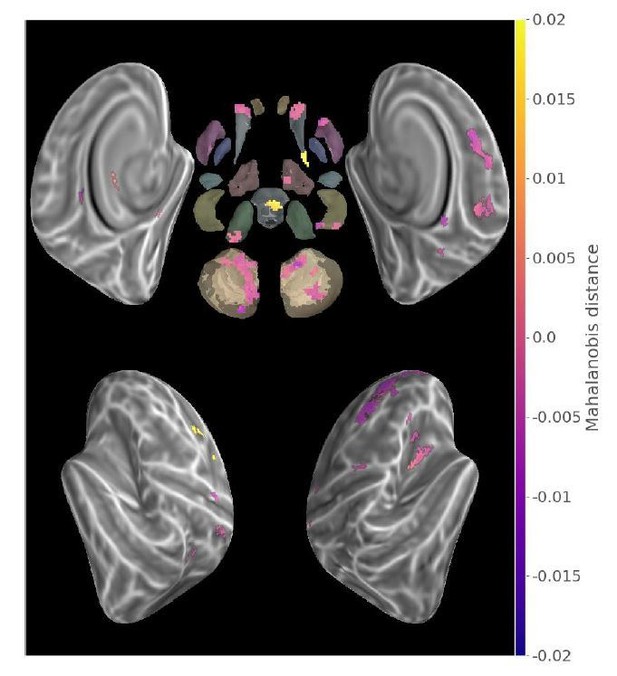
Contrast (run2-run1) between new sequences Mahalanobis distances (p<.05 Monte-Carlo TFCE corrected).

Main effect of motor sequence execution during MVPA task.
(t-value thresholded at p<.05 FDR-corrected).
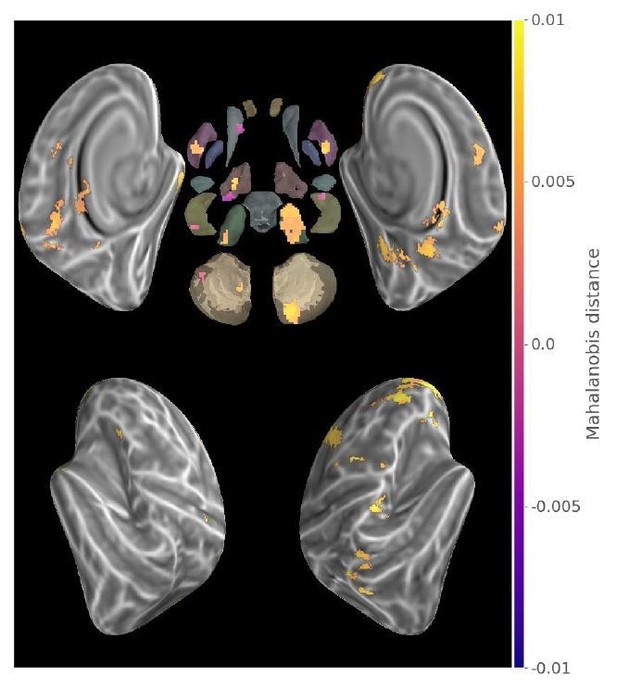
Conjunction of Consolidated>New contrast map and exclusive disjunction of Consolidate>0 and New>0 maps (p<.05 Monte-Carlo TFCE corrected).
Additional files
-
Supplementary file 1
Test for differences in speed (mean duration to perform a correct sequence) per block between trained and new sequences.
- https://doi.org/10.7554/eLife.39324.012
-
Supplementary file 2
Test for differences in accuracy (number of correct sequences over the five repetitions in a block) between trained and new sequences.
- https://doi.org/10.7554/eLife.39324.013
-
Supplementary file 3
Test for differences in speed and accuracy between the new sequences.
- https://doi.org/10.7554/eLife.39324.014
-
Supplementary file 4
Test for differences in speed and accuracy between the consolidated sequences.
- https://doi.org/10.7554/eLife.39324.015
-
Source code 1
MNI coordinates of main-effect clusters center-of-mass.
- https://doi.org/10.7554/eLife.39324.016
-
Source code 2
MNI coordinates of contrast clusters center-of-mass.
- https://doi.org/10.7554/eLife.39324.017
-
Transparent reporting form
- https://doi.org/10.7554/eLife.39324.018