Myosin V functions as a vesicle tether at the plasma membrane to control neurotransmitter release in central synapses
Figures
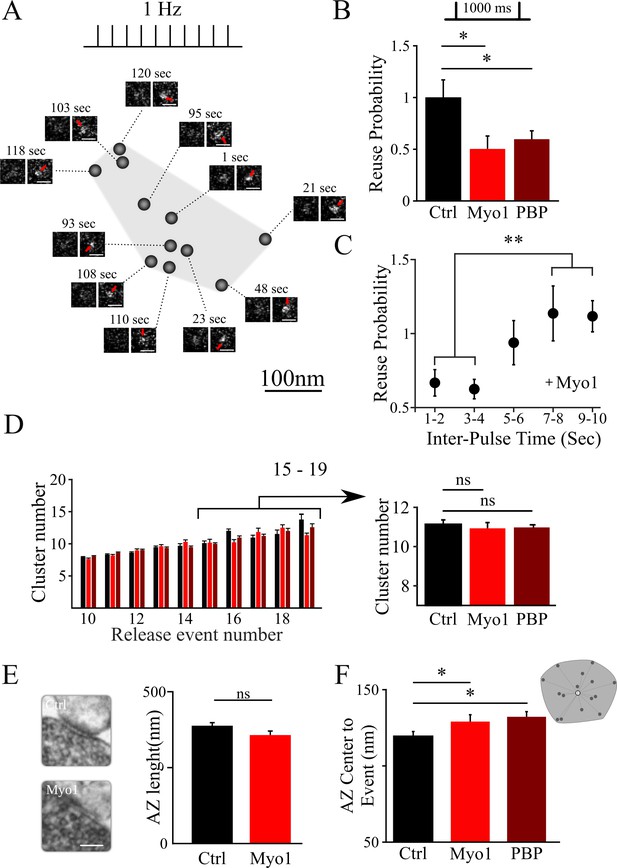
Inhibition of myosin V impairs the reuse of release sites in hippocampal synapses.
(A) Sample spatial distribution of release events within a single hippocampal bouton evoked by 1 Hz stimulation. Scale bar: 1 µm. (B) Effect of myosin V inhibition with Myo1 or PBP on release site reuse was evaluated using a paired-pulse protocol as a probability that the same site is reused for two sequential stimuli 1000 ms apart, normalized to the same measurement in control condition. (C) Effect of myosin V inhibition with Myo1 on the probability that the same site is reused for two stimuli at different inter-stimulus intervals (in the range of 1 – 10 s) normalized to the same measurement in control conditions and binned for two sequential intervals. (D) Effect of myosin V inhibition on the average number of clusters/release sites detected in individual boutons plotted as a function of the number of release events observed (left). Pooled average number of clusters for boutons with 15 to 19 detected events in different conditions indicated (right). (E) Large-Area Scanning Electron Microscopy (LaSEM) of hippocampal boutons in culture showing that no significant changes in the AZ size were detected as a result of myosin V inhibition with Myo1. Examples of control and Myo1 treatment are shown (left). Scale bar: 200 nm. (F) Effect of myosin V inhibition on the distribution of distances of release events to AZ center evoked at 1 Hz. **=P < 0.01, *=P < 0.05, two-sample t-test; ns – not significant.
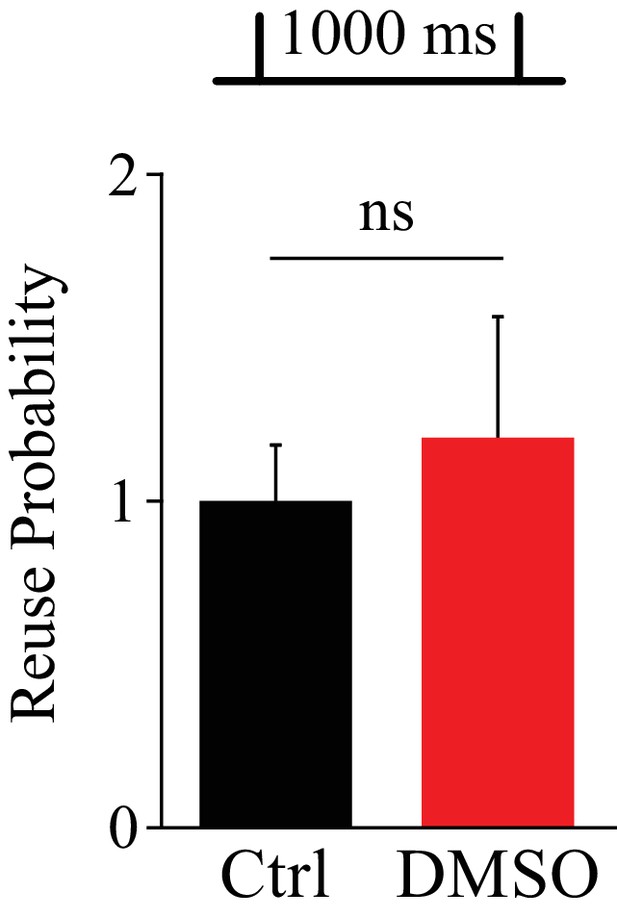
DMSO alone has no effect on release site reuse.
Proportion of release sites that are reused at an inter-stimulus interval of 1000 ms (experiment in Figure 1B) in the presence of 0.5% DMSO normalized to the control value.
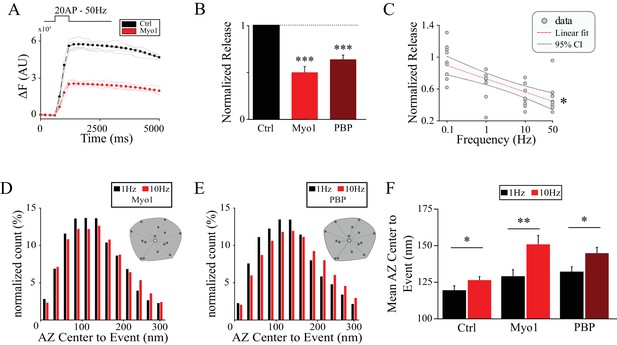
Myosin V controls release site refilling during high-frequency stimulation.
(A) Examples of vGlut1-pHluorin responses at single hippocampal boutons to 20 stimuli trains at 50 Hz for Ctrl (black) and Myo1 (red). (B) Ratio of vGlut1-pHluorin responses in (A) at individual boutons for Myo1 or PBP normalized to the control. (C) Effect of myosin inhibition with Myo1 on the amplitude of vGlut1-pHluorin responses to 20 stimuli trains at individual boutons as a function of the train frequency, averaged across all boutons in the movie and normalized to a control at a corresponding frequency. (D–E) Effect of myosin inhibition with Myo1 (D) or PBP (E) on the distribution of distances of release events to AZ center evoked at 1 Hz vs 10 Hz. (F) Analysis of data in (D,E) showing a mean distance of release events to the AZ center per bouton at 1 Hz or 10 Hz for Ctrl, Myo1 and PBP, respectively. ***=P < 0.001, **=P < 0.01, *=P < 0.05, two-sample t-test; ns – not significant.
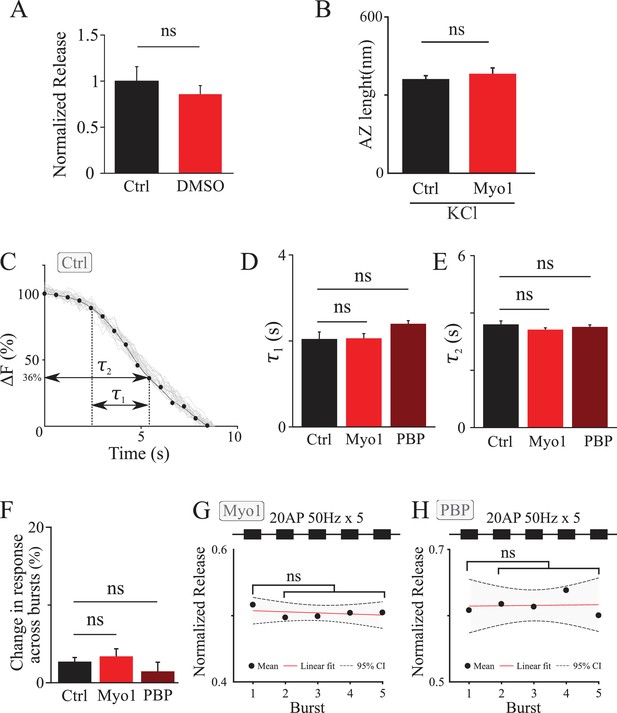
Controls for vesicle release measurements during high-frequency stimulation.
(A) vGlut1-pHluorin responses evoked by a high-frequency train (experiment in Figure 2A,B) in the presence of 0.5% DMSO normalized to the control value. (B) LaSEM of hippocampal boutons in culture showing that no significant changes in the AZ size were detected as a result of myosin V inhibition with Myo1 (20 min) during a sustained depolarization with 55 mM KCl (10 min). (C,D,E) Myosin V inhibition does not affect endocytosis dynamics. (C) Sample (grey) and average (black) bouton fluorescence decay after 20 stimuli at 50 Hz in control conditions. Quantification of the data in (C) shows no significant changes in the exponential component of fluorescence decay time (τ1) between Ctrl, Myo and PBP (D), or in the total fluorescence decay time including the initial plateau (τ2) (E). (F, G, H) Analysis of synaptic responses to five consecutive bursts of 20 stimuli at 50 Hz separate by 20 s. Change in synaptic response in two consecutive bursts, measured for each two consecutive bursts and averaged. No significant differences were observed between Ctrl, Myo1, or PBP (F). (A) mplitude of synaptic responses to each of five consecutive bursts in the presence of Myo1 (G) or PBP (H) normalized to control.
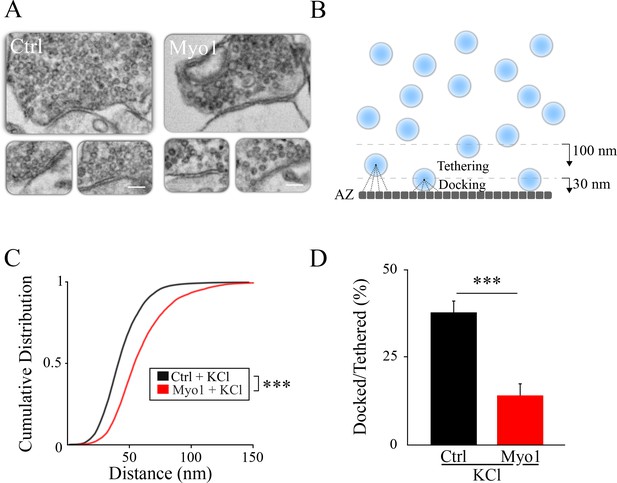
Inhibition of myosin V causes a vesicle docking defect during sustained activity.
(A) LaSEM of individual hippocampal boutons in cultures depolarized by KCl application (55 mM) for 10 min in the presence or absence of Myo1 (20 min), immediately followed by fixation. (B–D) Membrane opposite to the PSD was divided into 0.5 nm sections (B) and the distance from each section to the closest vesicle was determined and plotted as a cumulative histogram (C). This subset of closest vesicles was subsequently used to estimate the relation between docked and tethered vesicle populations (D). We considered vesicle as ‘docked’ when the distance from the AZ section to the vesicle center was under 30 nm and ‘tethered’ when the distance was under 100 nm. Scale bar: 200 nm. ***=P < 0.001; **=P < 0.01; two-sample KS-test (C) or two-sample t-test (D). ns = not significant.
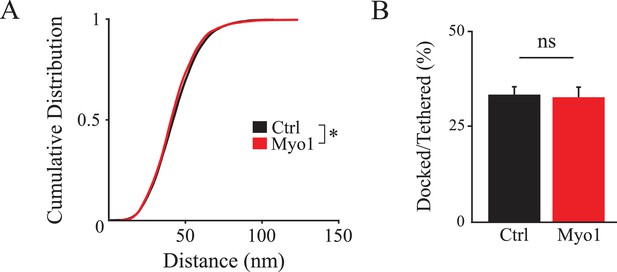
Ultrastructural analysis of vesicle docking under basal conditions in the absence of KCl.
(A) Distribution of distances from the AZ to the closest vesicles is similar but slightly left-shifted in the presence of Myo1 under basal conditions in the absence of KCl. (B) The ratio of docked/tethered vesicles is not affected by inhibition of myosin V with Myo1 under basal conditions in the absence of KCl. *=P < 0.05; ns - not significant; two-sample KS-test (A); two-sample t-test (B).
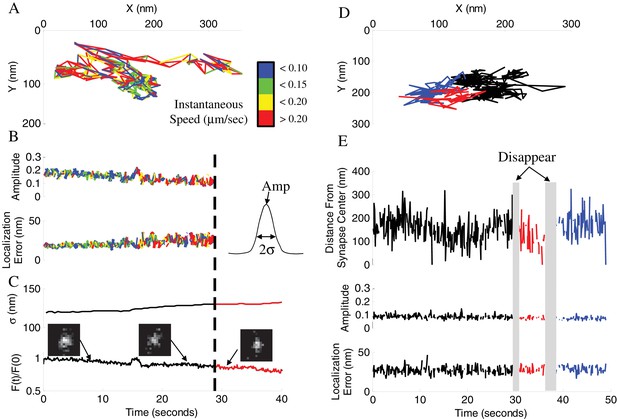
Vesicle disappearance/reappearance events.
(A) Sample track of a vesicle that disappeared during observation. Track is color-coded by instantaneous vesicle speed. (B) Parameters of vesicle image from (A) before and after vesicle disappearance indicated by a dotted line. Amplitude of the vesicle image (PSF) normalized to image bit-depth (214-1) (Top), and localization error of vesicle position as a function of time (Bottom). (C) Half-width of a Gaussian fit to the whole-synapse image (σ, Top) and whole-synapse integrated intensity (Bottom) before (Black) and after (Red) vesicle detection was lost, for the same vesicle as in A. Raw images (insets) of vesicle image at different time points show that vesicle is still present in the synapse after detection is lost. (D) Sample track of a vesicle that undergoes multiple disappearance/reappearance events. Track is color-coded with initial track shown in black, the first reappearance in red, the second reappearance in blue. (E) Parameters of vesicle track from (D). Vesicle position over time plotted as a distance from center of the bouton (Top). Amplitude of the vesicle image (Middle) and localization error of vesicle position (Bottom) as a function of time. Periods of vesicle disappearance are highlighted in grey. The color coding is the same as in (D).
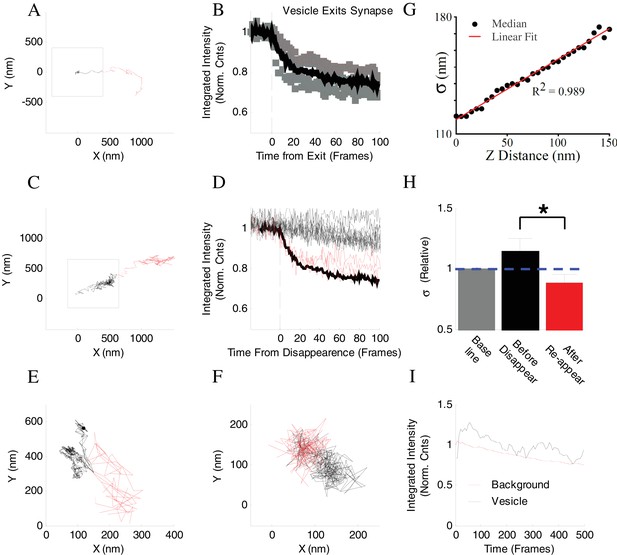
Controls for vesicle tracking analyses.
(A) Example of a vesicle track that was observed to leave a synapse (Square) to the axon under normal detection criteria. The part of the track that was observed within the synapse is shown in black, while outside the synapse in red. (B) Integrated intensity of four different boutons in which a vesicle exiting the bouton was observed under normal detection criteria, similar to an example in (A). Intensity is normalized by first frame and plotted as a function of time, with T = 0 defined as the time the vesicle is known to leave the synapse (dashed line). The mean integrated intensity is determined from all four examples (Solid line). (C) Example of a vesicle track that exits a synapse (Square) at the point when detection is lost. Vesicle was tracked within the synapse using standard PSF width restriction of 1.5 pixels (Black). The same vesicle can be tracked well beyond the initial point of disappearance and is observed exiting the synapse under less stringent detection criteria (PSF width restriction of 3 pixels, Red). (D) Representative integrated bouton intensity plotted as a function of time for 30 different boutons in which vesicle detection was lost. Intensity is normalized to the first frame and plotted as a function of time, where T = 0 is defined as the time vesicle detection was lost. The vast majority of boutons do not show a significant change in intensity after vesicle detection is lost (black). Only three boutons show a large decrease after vesicle detection is lost (Red) with reduction in intensity similar to changes observed for the known vesicle exit events (solid black line from (B). (E, F). Representative examples of vesicle tracks (left example is the same as in Figure 4A, but rotated 90°) that could be tracked well beyond the initial point of disappearance under less stringent detection criteria (portion of the track before disappearance is shown in black, after disappearance in red). (G) The PSF σ as a function of the distance along the Z-axis for 40 nm green fluorescent beads. (H) Relative changes in σ values for a representative subset of 16 disappearing vesicles. σ–values were determined for the last frame just before disappearance, and the first frame just after re-appearance and normalized to baseline determined as an average σ–value for the first 5 frames of vesicle tracks. (I) Comparison of normalized intensities of the vesicle image amplitude and nearby background to compare the rates of photo bleaching (N = 6 representative samples). The background (Red) was from a process near each vesicle, but outside the synapse, and normalized to its value in the first frame. The vesicle intensity (Black) has been background subtracted and normalized to its value in the first frame. *=P < 0.05; two-sample t-Test.
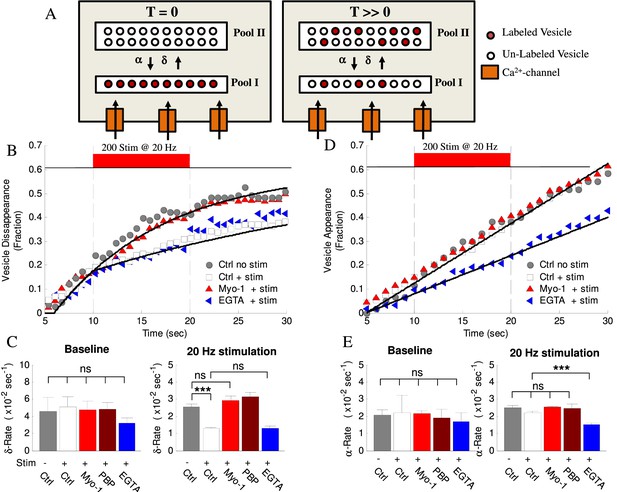
Activity- and myosin V- dependent vesicle shuttling between a membrane and an inner pools.
(A) Model hypothesis of vesicle exchange between two pools resulting in observed disappearance and re-appearance. At T = 0 all labeled vesicles are assumed to be in Pool I; vesicles undergo transition toward Pool II at rate (δ) and re-appear with rate (α). With time (T >> 0) the experimentally observed δ–rate is lower because un-labeled vesicles replace the labeled ones and disappear at the same rate but without being counted. (B) Vesicle disappearance as a function of time plotted as a fraction of total vesicles observed. Ctrl-No Stim represents vesicle disappearance rate in the absence of activity at any time during observation; other data show effects of stimulation (+Stim, 20 Hz applied at 10 – 20 s period) without (white) or with (red) myosin-V inhibition (Myo-1), or EGTA-AM (blue). Computational model is shown as solid lines. Note that the X-axis starts at T = 5 s because all tracks were required to be observed for at least 5 s to be included in analysis. (C) Mean disappearance rate (δ) from exponential-recovery fits to data in (B) at baseline (5–10 s) and during stimulation (10 – 20 s). Error-bars are mean-residual of fits to data. (D) Same as (B) but for vesicle appearances in different conditions indicated plotted as a fraction of total vesicles observed. Linear fits to data show constant rates (solid lines). (E) Same as (C) for the mean appearance rate (α) from linear fits to data in (D). Error-bars are mean-residual of linear fits to data. ***=P < 0.001; two-sample KS-test. ns = not significant.
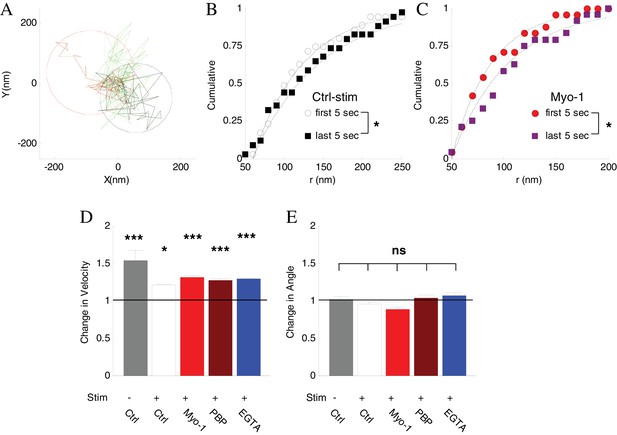
Inhibtion of myosin V does not affect vesicle mobility before disappearance.
(A) Spatial analysis of vesicle displacement based on the radius of a circle encompassing 95% of vesicle trajectory for the first 5 s (50 frames, black circle) and for the last 5 s (red circle) before disappearance. (B) Cumulative distributions for the spatial analysis in (A) in control with stimulation conditions compared for all disappearing vesicles and fit to exponential recovery function (solid line). (C) Same as (B) in the presence of Myo-1 and stimulation. (D) The ratio of instantaneous speed during the last 2 s and the first 2 s of observation. Error-bars are determined from residuals of cumulative fits. (E) The ratio of angular displacement during the last 2 s and the first 2 s of observation. Error-bars are determined from SEM. ***=P < 0.001; *=P < 0.05; two-sample t-test (D) or two-sample KS-test (E).
Tables
Table of all data values and statistical analyses.
Data table columns are formatted as (i) corresponding figure location; (ii) conditions being statistically compared and separated by ‘/’; (iii) number of samples (synapses, dishes, cultures) used for each test; (iv) mean values and errors for each condition separated by ‘/’ and corresponding to conditions in column (i); (v) statistical test used for comparison; (vi) P-value resulting from the statistical comparison.
| Figure number | Conditions | NSyn | NDishes | Ncultures | mean ± sem | Stastical test | P val |
|---|---|---|---|---|---|---|---|
| Basal Pr (related to 1) | Ctrl/Myo-1 | 367/259 | 15/15 | 4/3 | 0.064 ± 0.001/0.056 ± 0.002 | Two-sample t-test | 0.004 |
| Ctrl/PBP | 367/839 | 15/11 | 4/3 | 0.064 ± 0.001/0.067 ± 0.001 | Two-sample t-test | 0.15 | |
| 1(B) | Ctrl/Myo-1 | 367/259 | 15/15 | 4/3 | 1.000 ± 0.006/0.503 ± 0.004 | Two-sample t-test | 0.02 |
| Ctrl/PBP | 367/839 | 15/11 | 4/3 | 1.000 ± 0.006/0.5971 ± 0.003 | Two-sample t-test | 0.02 | |
| 1(C) | 1–4/7–10 | 367/259 | 15/15 | 4/3 | 0.63 ± 0.06/1.036 ± 0.1028 | Two-sample t-test | 0.003 |
| 1(D) | Ctrl/Myo-1 | 367/259 | 15/15 | 4/3 | 11.1 ± 0.2/10.9 ± 0.3 | Two-sample t-test | 0.58 |
| Ctrl/PBP | 367/839 | 15/11 | 4/3 | 11.1 ± 0.2/11.0 ± 0.1 | Two-sample t-test | 0.56 | |
| 1(E) | Ctrl/Myo-1 | 137/81 | 3/3 | 3/3 | 378 ± 10/364 ± 13 | Two-sample t-test | 0.38 |
| 1(F) | Ctrl/Myo-1 | 367/259 | 15/15 | 4/3 | 120 ± 3/129 ± 4 | Two-sample t-test | 0.04 |
| Ctrl/PBP | 367/839 | 15/11 | 4/3 | 120 ± 3/132 ± 3 | Two-sample t-test | 0.01 | |
| 2(B) | Ctrl/Myo-1 | - | 9/10 | 3/3 | 0.50 ± 0.06 | Two-sample t-test | <0.001 |
| Ctrl/PBP | - | 9/7 | 3/3 | 0.64 ± 0.04 | Two-sample t-test | <0.001 | |
| Slope (related to 2B) | Ctrl/Myo-1 | - | 9/10 | 3/3 | 0.52 ± 0.05 | Two-sample t-test | <0.001 |
| Ctrl/PBP | - | 9/7 | 3/3 | 0.62 ± 0.04 | Two-sample t-test | <0.001 | |
| 2(C) | Ctrl/Myo-1 | - | 36/34 | 11/11 | y = 0.75269–0.17806 x (Linear Fit) | Linear Fit | <0.001 |
| 2(F) | 1 Hz/10 Hz, Ctrl | 367/254 | 15/15 | 4/6 | 120 ± 3/126 ± 2 | Two-sample t-test | 0.04 |
| 1 Hz/10 Hz, Myo-1 | 259/862 | 15/10 | 3/3 | 129 ± 4/151 ± 6 | Two-sample t-test | 0.004 | |
| 1 Hz/10 Hz, PBP | 839/988 | 11/11 | 3/3 | 132 ± 3/145 ± 4 | Two-sample t-test | 0.01 | |
| 3(C) | Ctrl/Myo-1, KCl | 86/67 | 3/3 | 3/3 | 43.19 ± 0.02/59.32 ± 0.03 | Two-sample KS-test | <0.001 |
| 3(D) | Ctrl/Myo-1, KCl | 86/67 | 3/3 | 3/3 | 37 ± 3/13 ± 3 | Two-sample t-test | <0.001 |
| 5(C) Baseline | Ctrl(-)/Ctrl(+) | 29/122 | 27/64 | 5/12 | 0.048 ± 0.017/0.05 ± 0.0012 | Two-sample KS-test | 0.42 |
| Ctrl(-)/Myo-1(+) | 29/69 | 27/40 | 5/8 | 0.048 ± 0.017/0.047 ± 0.01 | Two-sample KS-test | 0.42 | |
| Ctrl(-)/PBP(+) | 29/21 | 27/47 | 5/8 | 0.048 ± 0.017/0.048 ± 0.008 | Two-sample KS-test | 0.42 | |
| Ctrl(-)/EGTA(+) | 29/51 | 27/30 | 5/6 | 0.048 ± 0.017/0.03 ± 0.006 | Two-sample KS-test | 0.88 | |
| Ctrl(-)/DMSO(+) | 29/44 | 27/34 | 5/5 | 0.048 ± 0.017/0.046 ± 0.01 | Two-sample KS-test | 0.43 | |
| Ctrl(+)/Myo-1(+) | 122/69 | 64/40 | 12/8 | 0.05 ± 0.0012/0.047 ± 0.01 | Two-sample KS-test | 0.43 | |
| Ctrl(+)/PBP(+) | 122/21 | 64/47 | 12/8 | 0.05 ± 0.0012/0.048 ± 0.008 | Two-sample KS-test | 0.43 | |
| Ctrl(+)/EGTA(+) | 122/51 | 64/30 | 12/6 | 0.05 ± 0.0012/0.03 ± 0.006 | Two-sample KS-test | 0.88 | |
| Ctrl(+)/DMSO(+) | 122/44 | 64/34 | 12/5 | 0.05 ± 0.0012/0.04608 ± 0.01053 | Two-sample KS-test | 0.13 | |
| Myo-1(+)/PBP(+) | 69/21 | 40/47 | 8/8 | 0.047 ± 0.01/0.04831 ± 0.00749 | Two-sample KS-test | 0.88 | |
| Myo-1(+)/EGTA(+) | 69/51 | 40/30 | 8/6 | 0.047 ± 0.01/0.03201±0.00639 | Two-sample KS-test | 0.43 | |
| PBP(+)/EGTA(+) | 21/51 | 47/30 | 8/6 | 0.048 ± 0.008/0.03 ± 0.006 | Two-sample KS-test | 0.43 | |
| 5(C) 20 Hz stimulation | Ctrl(-)/Ctrl(+) | 29/122 | 27/64 | 5/12 | 0.026 ± 0.002/0.013 ± 0.0003 | Two-sample KS-test | <0.001 |
| Ctrl(-)/Myo-1(+) | 29/69 | 27/40 | 5/8 | 0.026 ± 0.002/0.029 ± 0.003 | Two-sample KS-test | 0.16 | |
| Ctrl(-)/PBP(+) | 29/21 | 27/47 | 5/8 | 0.026 ± 0.002/0.03 ± 0.002 | Two-sample KS-test | 0.63 | |
| Ctrl(-)/EGTA(+) | 29/51 | 27/30 | 5/6 | 0.026 ± 0.002/0.013 ± 0.002 | Two-sample KS-test | <0.001 | |
| Ctrl(+)/Myo-1(+) | 122/69 | 64/40 | 12/8 | 0.013 ± 0.0003/0.02907 ± 0.00264 | Two-sample KS-test | 0.02 | |
| Ctrl(+)/PBP(+) | 122/21 | 64/47 | 12/8 | 0.013 ± 0.0003/0.03 ± 0.002 | Two-sample KS-test | 0.02 | |
| Ctrl(+)/EGTA(+) | 122/51 | 64/30 | 12/6 | 0.013 ± 0.0003/0.013 ± 0.002 | Two-sample KS-test | 0.63 | |
| Ctrl(+)/DMSO(+) | 122/44 | 64/34 | 12/5 | 0.013 ± 0.0003/0.012 ± 0.0006 | Two-sample KS-test | 0.66 | |
| Myo-1(+)/PBP(+) | 69/21 | 40/47 | 8/8 | 0.029 ± 0.003/0.03 ± 0.002 | Two-sample KS-test | 0.16 | |
| Myo-1(+)/EGTA(+) | 69/51 | 40/30 | 8/6 | 0.029 ± 0.003/0.013 ± 0.002 | Two-sample KS-test | 0.02 | |
| PBP(+)/EGTA(+) | 21/51 | 47/30 | 8/6 | 0.03 ± 0.002/0.013 ± 0.002 | Two-sample KS-test | 0.007 | |
| 5(B) | MC-Model (-) | 100 | 1000 | 8.50E-02 | |||
| MC-Model (+) | 100 | 1000 | 3.00E-02 | ||||
| 5(E) Baseline | Ctrl(-)/Ctrl(+) | 60/343 | 27/64 | 5/12 | 0.02 ± 0.003/0.02 ± 0.01 | Two-sample KS-test | 0.14 |
| Ctrl(-)/Myo-1(+) | 60/174 | 27/49 | 5/8 | 0.02 ± 0.003/0.02 ± 0.002 | Two-sample KS-test | 0.59 | |
| Ctrl(-)/PBP(+) | 60/55 | 27/34 | 5/8 | 0.02 ± 0.003/0.02 ± 0.005 | Two-sample KS-test | 0.14 | |
| Ctrl(-)/EGTA(+) | 60/63 | 27/30 | 5/6 | 0.02 ± 0.003/0.016 ± 0.005 | Two-sample KS-test | 0.14 | |
| Ctrl(+)/Myo-1(+) | 343/174 | 64/40 | 12/8 | 0.02 ± 0.01/0.02 ± 0.002 | Two-sample KS-test | 1 | |
| Ctrl(+)/PBP(+) | 343/55 | 64/34 | 12/8 | 0.02 ± 0.01/0.02 ± 0.005 | Two-sample KS-test | 0.59 | |
| Ctrl(+)/EGTA(+) | 343/63 | 64/30 | 12/6 | 0.02 ± 0.01/0.016 ± 0.005 | Two-sample KS-test | 0.31 | |
| Myo-1(+)/PBP(+) | 174/55 | 40/34 | 8/8 | 0.02 ± 0.002/0.02 ± 0.005 | Two-sample KS-test | 0.89 | |
| Myo-1(+)/EGTA(+) | 174/63 | 40/30 | 8/6 | 0.02 ± 0.002/0.016 ± 0.005 | Two-sample KS-test | 0.14 | |
| 5(E) 20 Hz stimulation | Ctrl(-)/Ctrl(+) | 60/343 | 27/64 | 5/12 | 0.025 ± 0.001/0.02 ± 0.0006 | Two-sample KS-test | 0.59 |
| Ctrl(-)/Myo-1(+) | 60/174 | 27/49 | 5/8 | 0.025 ± 0.001/0.025 ± 0.0005 | Two-sample KS-test | 1 | |
| Ctrl(-)/PBP(+) | 60/55 | 27/34 | 5/8 | 0.025 ± 0.001/0.024 ± 0.003 | Two-sample KS-test | 0.31 | |
| Ctrl(-)/EGTA(+) | 60/63 | 27/30 | 5/6 | 0.025 ± 0.001/0.015 ± 0.001 | Two-sample KS-test | 0.001 | |
| Ctrl(+)/Myo-1(+) | 343/174 | 64/40 | 12/8 | 0.02 ± 0.0006/0.025 ± 0.0005 | Two-sample KS-test | 0.89 | |
| Ctrl(+)/PBP(+) | 343/55 | 64/34 | 12/8 | 0.02 ± 0.0006/0.024 ± 0.003 | Two-sample KS-test | 0.31 | |
| Ctrl(+)/EGTA(+) | 343/63 | 64/30 | 12/6 | 0.02 ± 0.0006/0.015 ± 0.001 | Two-sample KS-test | 0.001 | |
| Myo-1(+)/PBP(+) | 174/55 | 40/34 | 8/8 | 0.025 ± 0.0005/0.024 ± 0.003 | Two-sample KS-test | 0.14 | |
| Myo-1(+)/EGTA(+) | 174/63 | 40/30 | 8/6 | 0.025 ± 0.0005/0.015 ± 0.001 | Two-sample KS-test | 0.001 | |
| Displacement (related to 6B) | Ctrl(-) Last 5/First 5 s | 12 | 27 | 5 | 1.17 ± 0.12 | Two-sample t-test | 0.001 |
| Ctrl(+) Last 5/First 5 s | 35 | 64 | 12 | 1.08 ± 0.05 | Two-sample t-test | 0.02 | |
| Myo-1(+) Last 5/First 5 s | 24 | 40 | 8 | 1.18 ± 0.12 | Two-sample t-test | <0.001 | |
| 6(D) Change in Velocity | Ctrl(-)/Ctrl(-) | 12 | 27 | 5 | 1.54 ± 0.13 | Two-sample t-test | <0.001 |
| Ctrl(+)/Ctrl(+) | 35 | 64 | 12 | 1.2 ± 0.016 | Two-sample t-test | 0.022 | |
| Myo-1(+)/Myo-1(+) | 24 | 40 | 8 | 1.31 ± 0.02 | Two-sample t-test | <0.001 | |
| PBP(+)/PBP(+) | 17 | 47 | 8 | 1.27 ± 0.03 | Two-sample t-test | 0.002 | |
| EGTA(+)/EGTA(+) | 20 | 30 | 6 | 1.29 ± 0.02 | Two-sample t-test | 0.002 | |
| 6(E) Change in Angle | Ctrl(-)/Ctrl(-) | 12 | 27 | 5 | 1.02 ± 0.04 | Two-sample KS-test | 1 |
| Ctrl(+)/Ctrl(+) | 35 | 64 | 12 | 0.95 ± 0.04 | Two-sample KS-test | 1 | |
| Myo-1(+)/Myo-1(+) | 24 | 40 | 8 | 0.88 ± 0.03 | Two-sample KS-test | 0.65 | |
| PBP(+)/PBP(+) | 17 | 47 | 8 | 1.03 ± 0.04 | Two-sample KS-test | 1 | |
| EGTA(+)/EGTA(+) | 20 | 30 | 6 | 1.06 ± 0.04 | Two-sample KS-test | 1 | |
| Figure 1—figure supplement 1(A) | Ctrl/DMSO | - | 15/8 | 4/3 | 1.00 ± 0.17/0.97 ± 0.16 | Chi-squared test | 0.89 |
| Figure 2—figure supplement 1(A) | Ctrl/DMSO | - | 7/9 | 3/3 | 1.0 ± 0.1/0.86 ± 0.09 | Two-sample t-test | 0.43 |
| Figure 2—figure supplement 1(B) | Ctrl/Myo-1 | - | 9/10 | 3/3 | 2.74 ± 0.5/3.42 ± 0.98 | Two-sample t-test | 0.56 |
| Ctrl/PBP | - | 9/7 | 3/3 | 2.74 ± 0.5/1.47 ± 1.18 | Two-sample t-test | 0.3 | |
| Figure 2—figure supplement 1(C) | Ctrl/Myo-1, KCl | 86/67 | 3/3 | 3/3 | 38 ± 3/14±3 | Two-sample t-test | 0.40 |
| Figure 2—figure supplement 1(E) | Ctrl/Myo-1 | - | 9/10 | 3/3 | 2.05 ± 0.16/2.06 ± 0.11 | Two-sample t-test | 0.93 |
| Ctrl/PBP | - | 9/7 | 3/3 | 2.05 ± 0.16/2.39 ± 0.07 | Two-sample t-test | 0.09 | |
| Figure 2—figure supplement 1(F) | Ctrl/Myo-1 | - | 9/10 | 3/3 | 3.6 ± 0.1/3.42 ± 0.06 | Two-sample t-test | 0.2 |
| Ctrl/PBP | - | 9/7 | 3/3 | 3.6 ± 0.1/3.51 ± 0.07 | Two-sample t-test | 0.57 | |
| Figure 2—figure supplement 1(G) | Ctrl/Myo-1 | - | 9/10 | 3/3 | y = 0.50915–0.0016552 x (Linear Fit) | Linear Fit | 0.57 |
| Ctrl/Myo-1; 1 st vs 2–5 | - | 9/10 | 3/3 | 0.52 ± 0.06/0.50 ± 0.02 | Two-sample t-test | 0.81 | |
| Figure 2—figure supplement 1(H) | Ctrl/PBP | - | 9/7 | 3/3 | y = 0.61391 + 0.00052562 x (Linear Fit) | Linear Fit | 0.93 |
| Ctrl/PBP; 1 st vs 2–5 | - | 9/7 | 3/3 | 0.61 ± 0.04/0.62 ± 0.02 | Two-sample t-test | 0.85 | |
| Figure 3—figure supplement 1(A) | Ctrl/Myo-1, KCl | 86/67 | 3/3 | 3/3 | 43.23 ± 0.02/41.15 ± 0.02 | Two-sample KS-test | <0.001 |
| Figure 3—figure supplement 1(B) | Ctrl/Myo-1 | 137/81 | 3/3 | 3/3 | 34 ± 2/33 ± 3 | Two-sample t-test | 0.84 |
| Figure 4—figure supplement 1(H) | Ctrl Disappear/Re-appear | 16/16 | 64/64 | 12/12 | 1.14 ± 0.105/0.88 ± 0.062 | Two-sample t-test | 0.04 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Chemical compound, drug | MyoVin-1 (Myo-1) | EMD Millipore | Cat. No. 475984 | |
| Chemical compound, drug | Pentabrom opseudilin (PBP) | FISHER SCIENTIFIC | Cat. No. 501015859 | |
| Chemical compound, drug | EGTA-AM | FISHER SCIENTIFIC | Cat. No. E1219 | |
| Chemical compound, drug | SGC5 | VWR | Cat. No. 89410–772 | |
| Chemical compound, drug | DL-AP5 Sodium salt | Tocris | Cat. No. 3693 | |
| Chemical compound, drug | CNQX disodium salt | Tocris | Cat. No. 1045 | |
| Chemical compound, drug | HEPES | Sigma | Cat. No. H4034 | |
| Chemical compound, drug | D-(+)-Glucose | Sigma | Cat. No. G 7021 | |
| Chemical compound, drug | Calcium chloride dihydrate | Sigma | Cat. No. 223506 | |
| Chemical compound, drug | Magnesium chloride hexahydrate | Sigma | Cat. No. M9272 | |
| Chemical compound, drug | Minimum Essential Media (MEM) - No Phenol Red | thermofisher | Cat. No. 51200–038 | |
| Chemical compound, drug | Characterized Fetal Bovine Serum | hyclone | Cat. No. SH30071.03 | |
| Chemical compound, drug | Penicillin- Streptomycin (5,000 U/mL) | thermofisher | Cat. No. 15070063 | |
| Chemical compound, drug | N-2 Supplement (100X) | thermofisher | Cat. No. 17502048 | |
| Chemical compound, drug | Donor Equine Serum | hyclone | Cat. No. SH30074.03 | |
| Chemical compound, drug | Sodium Pyruvate (100 mM) | thermofisher | Cat. No. 11360–070 | |
| Chemical compound, drug | Neurobasal- A Medium | thermofisher | Cat. No. 10888–022 | |
| Chemical compound, drug | B-27 Supplement (50X), serum free | thermofisher | Cat. No. 17504–044 | |
| Chemical compound, drug | GlutaMAX Supplement | thermofisher | Cat. No. 35050061 | |
| Chemical compound, drug | Earle’s Balanced Salts | sigmaaldrich | Cat. No. E3024 | |
| Chemical compound, drug | Corning Collagen I, Rat | Fisher Scientific | Cat. No. 354236 | |
| Chemical compound, drug | Cover Glasses | Fisher Scientific | Cat. No. 12-545-80 | |
| Chemical compound, drug | Papain | Worthington Biochemical | Cat. No. LS003126 | |
| Chemical compound, drug | PDL (poly-D-lysine) | BD Bios ciences | Cat. No. 40210 | |
| Chemical compound, drug | Trypsin-EDTA (0.05%), phenol red | therm ofisher | Cat. No. 25300–054 | |
| Biological sample (Rattus norvegicus, Female) | Sprague-Dawley Timed-Pregnant rat, E15 pups | Charles River | Pups of both genders | |
| Software, algorithm | MATLAB | MathWorks | RRID:SCR_001622 | |
| Software, algorithm | u-track2.0 | Jaqaman, K., et al. (2008), Nat.Meth. 5, 695–702 | Gaudenz Danuser Lab | |
| Software, algorithm | ImageJ | https://imagej.nih.gov/ij/ | RRID:SCR_003070 | |
| Software, algorithm | Fiji | http://fiji.sc | RRID:SCR_002285 | |
| Software, algorithm | inkscape | RRID:SCR_014479 | ||
| Other | VGluT1- pHluorin | Drs. Robert Edwards and Susan Voglmaier (UCSF) | Genetically encoded optical indicator of vesicle release and recycling |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.39440.013





