Biosynthesis of histone messenger RNA employs a specific 3' end endonuclease
Figures
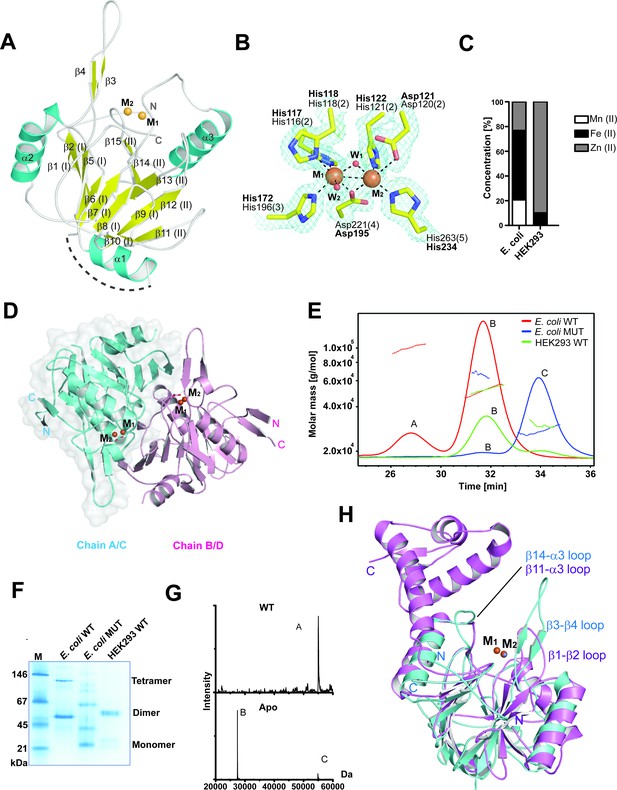
MBLAC1 is a dimeric metallo β-lactamase fold protein related in structure to LACTB2.
(A) View from an MBLAC1 crystal structure (PDB ID: 4V0H) showing secondary structure elements and the di-metal containing active site. Helices are in cyan, β-strands in yellow, and metal ions are orange spheres. Dashed line (in gray) indicates unrefined residues (aa 51–66). (B) View of metal binding and active site residues with a representative electron density map (3.0 σ mFo-DFc OMIT; cyan mesh) for the sidechains of His116 (Nε2 to M1: 2.33 Å), His118 (Nε1 to M1: 2.35 Å), Asp120 (Oδ2 to M2: 2.9 Å), His121 (Nε2 to M2: 2.37 Å), His196 (Nε2 to M1: 2.27 Å), Asp221 (Oδ2 to M1: 2.14 Å; Oδ2 to M2: 2.12 Å), His263 (Nε2 to M2: 2.06 Å) and the two water molecules (red spheres) which coordinate (black dashed lines) to the metals (orange spheres). The BBL MBL numbering system is used (Galleni et al., 2001) (active site motif numbers in parentheses) (Pettinati et al., 2016). Numbering as used in our PDB deposited structure is in bold. (C) Inductively coupled plasma mass spectrometry (ICP-MS) experiments reveal that in human cells MBLAC1 binds zinc. (D) MBLAC1 dimer model calculated by PISA (Krissinel and Henrick, 2007). A surface representation is shown for protomer A/C. (E) Multi-angle laser light scattering (MALS) analyses of E. coli produced wt and active site variant MBLAC1, and of HEK293 produced wtMBLAC1. Peak A likely represents a trimer (~80000 Da); peak B (~54000 Da) a dimer; and peak C (~27000 Da) a monomer. wt MBLAC1 is predominantly dimeric, the active site variant is mainly monomeric. The latter observation correlates with loss of activity of active site variant MBLAC1 (main text Figure 4a), the proposal that the dimer is catalytically active, and the observation that the active site is close to the dimer interface. MALS experiments were carried out at the Biophysical Services of the Biochemistry Dept., Oxford University. (F) Non-denaturing PAGE analyses indicates wt MBLAC1 produced in either E. coli or HEK293 cells is predominantly dimeric; a higher oligomeric state (tetramer) is also observed for the bacterial produced wt protein. The active site substituted enzyme produced in E. coli shows loss of oligomerization. wtMBLAC1 produced in either, E. coli or HEK293 cells shows the same oligomerization behavior. (G) Non-denaturing electrospray ionization mass spectrometry deconvoluted spectra of recombinant MBLAC1 produced in E. coli indicates that MBLAC1 is dimeric, binding two metal ions. Peak A (54880 Da) represents the dimer with two divalent transition metal ions (zinc or iron) bound to each monomer (+224 Da); peaks B and C (27440 and 54880 Da, respectively) correspond to monomer and dimer without bound metal, following metal removal using EDTA. (H) Superimposition of the MBLAC1 (cyan; metals in orange) and LACTB2 folds (PDB ID: 4AD9) (Levy et al., 2016) (pink; metals in grey) reveals a high degree of overall similarity (RMSD 2.2 Å over 153 Cα atoms).
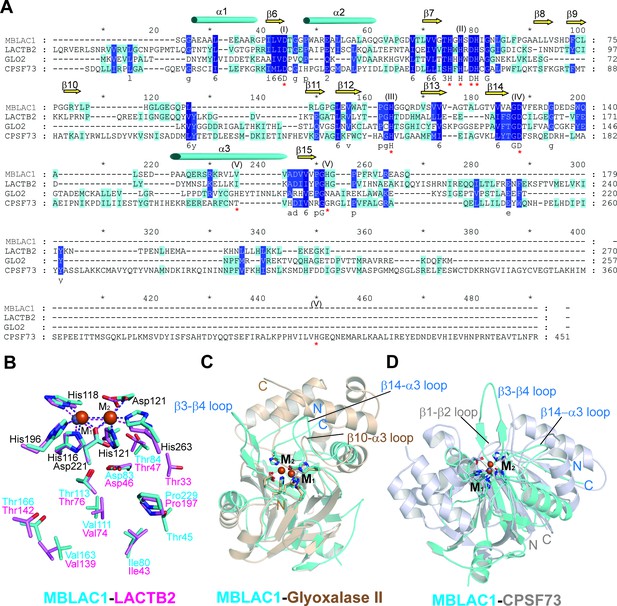
Structural comparison of MBLAC1 with ribonucleases from the MBL superfamily and with glyoxalase II.
(A) Clustal Omega (Sievers et al., 2011) alignment of sequences for the MBL domains of human MBLAC1, LACTB2, CPSF73, and glyoxalase II. Secondary structure elements are derived from the MBLAC1 structure (PDB ID 4V0H). β-Sheets are yellow arrows and α-helices are blue cylinders. Residues are colored based on conservation: dark blue represents highest conservation, blue the second highest, light blue the third highest, and no color the least conserved. The conserved metal binding residues are indicated by a red asterisk (*) with the corresponding MBL motif in parentheses. (B) Superimposed active site residues from MBLAC1 (cyan) and LACTB2 (pink). Apparently identical residues in the proximity of the active site are shown. Zinc and iron ions are gray and orange spheres, respectively. The BBL MBL numbering system is used (Galleni et al., 2001). (C) Superposition of MBLAC1 (cyan), with glyoxalase II (PDB ID: 1QH5) (brown) structures. Loops of interest, as discussed in the main text, are indicated. Detailed comparison, in particular of the loops surrounding the active site, supports the biochemical evidence that MBLAC1 is not a glyoxalase II; only one of these loops (β10-α3 loop/glyoxalase II; β14-α3 loop/MBLAC1), is in glyoxalase II where it is present in a shorter form. Zinc and iron ions are gray and orange spheres, respectively. (D) Superimposition of MBLAC1 (cyan) with CPSF73 (PDB ID: 2I7T) (grey). Note that only the MBL domains were used in the superimposition. Zinc and iron ions are gray and orange spheres, respectively.
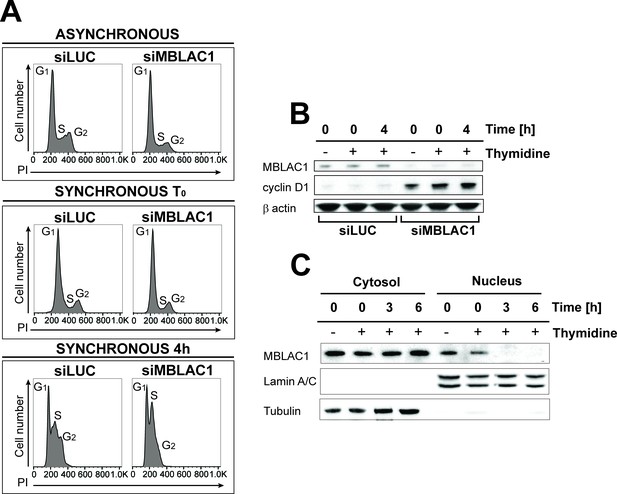
Loss of MBLAC1 impairs cell cycle progression.
(A) Cell cycle analyses of MBLAC1-depleted cells. Analyses were on unsynchronised cells, or after cell synchronization (using a double thymidine block) as indicated. Flow cytometry profiles were obtained by PI staining. Control siRNA (siLUC) transfected cells were used for reference. (B) Western blots evaluating knockdown efficiency of MBLAC1 after siRNA treatment, in unsynchronised and synchronised cells. Cyclin D1 (G1 marker) levels were analysed. β-Actin was used as a control. Control siRNA (siLUC) transfected cells were used for reference. (C) Western blot analysis showing cellular distribution of endogenous MBLAC1 after subcellular fractionation in unsynchronised (-) and early S-phase synchronised HeLa cells (+) at different time points after release from a double thymidine block. Note the presence of MBLAC1 in the nucleus in early S-phase. α-Tubulin and lamin A/C were used as cytosolic and nuclear markers, respectively.
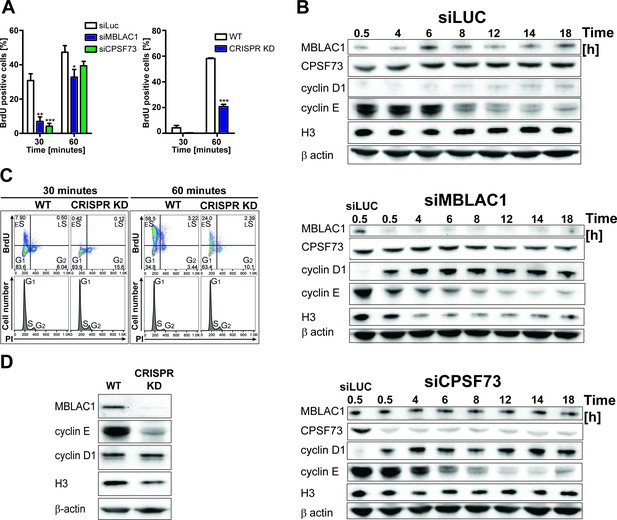
Loss of MBLAC1 impairs normal entrance to S-phase and S-phase progression.
(A) Comparative quantification of BrdU-positive cells corresponding to early-S cells in MBLAC1 and CPSF73-depleted cells and MBLAC1 CRISPR KD (CRISPR/Cas9 mediated KD) cells after synchronization. Control siRNA (siLUC) transfected cells and wt HeLa cells were used for comparison. Error bars represent SEM from three independent biological replicates. Asterisks indicate the statistical significance using the two way ANOVA tool followed by the Bonferroni multiple comparison tool in Graphpad Prism five software: *, p<0.05; **, p<0.001; ***, p<0.0001. Unlabeled variations were considered not statistically relevant. (B) Western blot analyses evaluating MBLAC1 and CPSF73 knockdown efficiency and effects on cyclin D1, cyclin E and histone H3 levels. (C) Cell cycle analysis of MBLAC1 CRISPR/Cas9 mediated stable knockdown (CRISPR KD) cells after synchronization in early S-phase. Flow cytometry profiles obtained by double-staining with an anti-BrdU antibody (upper panels) and PI (lower panels). wt HeLa cells were used for reference. Abbreviations: ES, early S-phase; LS, late S-phase. (D) Western blot analyses evaluating MBLAC1 CRISPR/Cas9-mediated stable knockdown efficiency and effects on cyclin D1, E and histone H3 levels (with synchronised cells treated with BrdU, 30-min incubation).
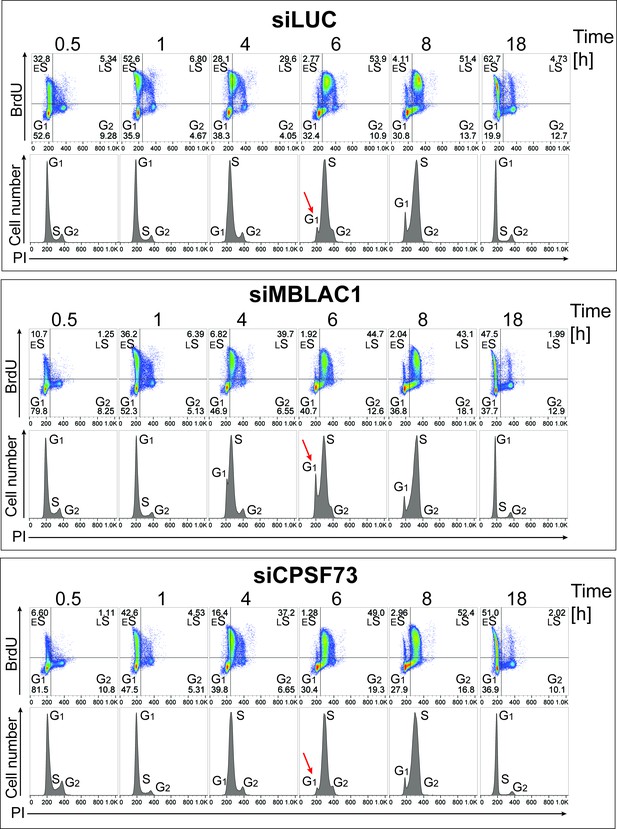
Loss of MBLAC1 impairs normal entering of S-phase and S-phase progression.
Cell cycle analyses of MBLAC1 and CPSF73 depleted cells after synchronization in early S-phase over a complete cell cycle (18 hr). Flow cytometry profiles obtained by a double-staining with anti-BrdU antibody (upper panels) and PI (lower panels). Control siRNA (siLUC) transfected cells were used for reference. Abbreviations: ES, early S-phase; LS, late S-phase. Red arrows indicate G1/early S cells 6 hr post-release from the double thymidine block.
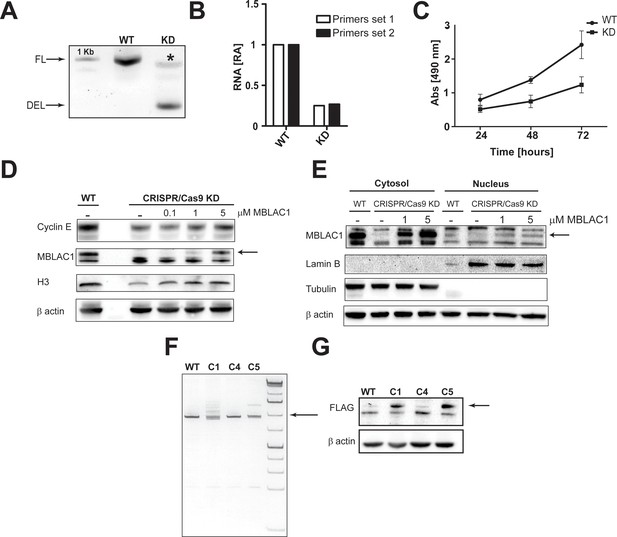
MBLAC1 CRISPR/Cas9 stable knockdown and FLAG knock in validation, wild-type-like cell phenotype restoration and MBLAC1 interactors.
(A) PCR analyses (genomic DNA) evaluating the MBLAC1 gene deletion efficiency using the CRISPR/Cas9 system after clonal selection in HeLa cells. Primers used to validate gene deletion are given in Table 2. Both full length gene (FL) and the deletion (DEL) PCR products were sequenced to validate the results. (*) indicates unspecific PCR amplification containing traces of mblac1 full length gene as validated by sequencing. (B) RT-qPCR quantification (total RNA) of the relative abundance (RA) of MBLAC1 mRNA in WT and MBLAC1 CRISPR/Cas9 KD HeLa cells implies substantial reduction of MBLAC1 mRNA in the mediate KD consistent with the observed protein level reduction (Figure 3D). The MBLAC1 mRNA level was normalised to GAPDH mRNA. RT-qPCR primers used to validate the MBLAC1 stable KD are in Table 2. Sanger sequencing of RT-qPCR products confirmed MBLAC1 mRNA production in the CRISPR/Cas9-mediated KD. (C) Proliferation assay (MTS) showing the reduced growth rate of the CRISPR/Cas9-mediated KD compared to wildtype HeLa cells. Error bars represent SEM from three biological replicates. (D) Western blot analyses evaluating the effects on cyclin E and histone H3 levels in MBLAC1 CRISPR/Cas9-mediated stable knockdown after treatment with recombinant MBLAC1 in early S-phase. (E) Western blot analysis showing the cellular distribution of exogenous MBLAC1 after subcellular fractionation in early S-phase synchronised MBLAC1 CRISPR/Cas9-depleted HeLa cells compared to wild-type cells. α-Tubulin and lamin B were used as cytosolic and nuclear markers, respectively. β-Actin was used as the loading control. Arrows indicate the band corresponding to MBLAC1. Note that recombinant MBLAC1 has a higher molecular mass compared to wtMBLAC1 due to the presence of a 6xhis-tag. (F) PCR analyses (genomic DNA) evaluating the FLAG tag insertion at the 3' of the MBLAC1 gene using the CRISPR/Cas9 system after clonal selection in HeLa cells. Primers used to validate the gene insertion were the same as those used to validate the gene deletion (Table 1). PCR products were sequenced to confirm the PCR results. (G) Western blot analysis confirming the FLAG tag insertion at the MBLAC1 C-terminal at protein level using an anti-FLAG antibody. Arrows indicate bands corresponding to FLAG-positive clones.
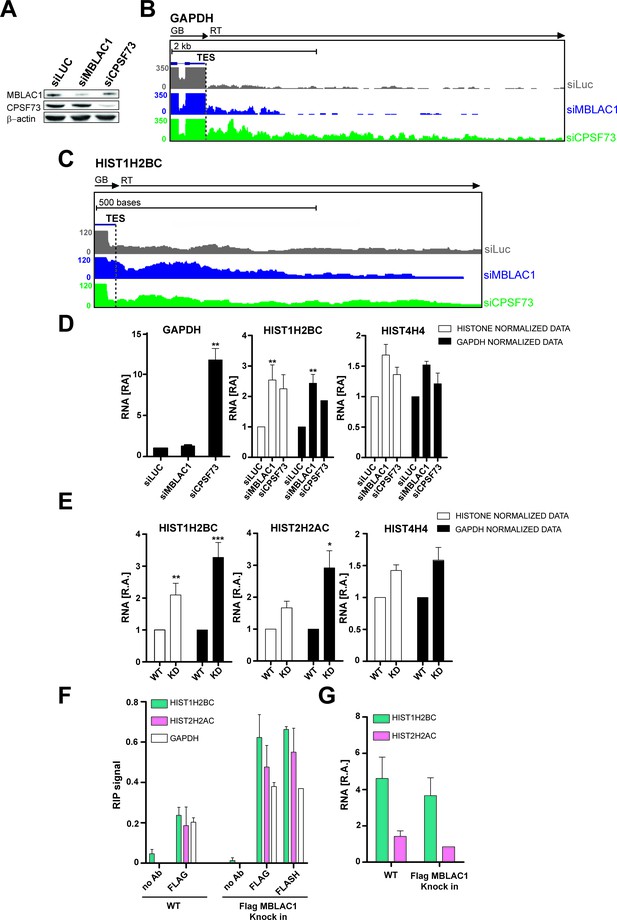
Loss of MBLAC1 leads to 3ʹ end processing defects in replication dependent histone pre-mRNA.
(A) Western blots showing the knockdown efficiency of siRNA treatments for MBLAC1 and CPSF73 in synchronised HeLa cells (using β-actin as a control). (B, C) ChrRNA-seq analyses from one of two biological replicates (Set2), on GAPDH or selected RD histone (HIST1H2BC) genes (UCSC Genome Browser) (Kent et al., 2002), following MBLAC1 or CPSF73 depletion. Arrows indicate gene bodies and transcription termination defect orientation. (D) Real-time qPCR quantification of 3ʹ end transcription termination defect following MBLAC1 and CPSF73 depletion on genes as in (b, c). Relative abundance (RA) of unprocessed RNA was normalised to the abundance of GAPDH mRNA (black), or to that of the corresponding histone gene body (white) (Table 2). Error bars represent SEM from four biological replicates. Asterisks indicate the statistical significance observed using the two-tailed t-test (**p<0.01). Abbreviations: GB, gene body; RT, read through; TES, transcription end site. (E) Real Time-qPCR quantification of 3ʹ end transcription termination defect in the MBLAC1 CRISPR/Cas9-mediated stable knockdown (KD) compared to wt HeLa cells. The relative abundance (RA) of unprocessed RNA was normalised to GAPDH mRNA (black), or to the corresponding histone gene body abundance (white) (Table 2). Error bars represent SEM from four biological replicates. Asterisks indicate the statistical significance observed using the two-tailed t-test (*,p<0.05, **, p<0.01, ***, p<0.001). Unlabeled variations were considered likely not statistically relevant. (F) RNA-Chromatin immunoprecipitation (RNA-ChIP) analyses investigating the recruitment of MBLAC1 on histone RNA. RNA-ChIP signals are shown for MBLAC1 and FLASH in the MBLAC1 C-terminal FLAG knock in HeLa cells; the FLAG background signal is shown in wild-type cells used as control for comparison. Note that both MBLAC1 and FLASH signals are comparable on normal protein coding RNA (GAPDH). Error bars represent SEM from three (MBLAC1) or two (FLASH) biological replicates. (G) RNA-ChIP input samples relative comparison. Histone mRNA relative abundance (in both wild-type and MBLAC1 C-terminal FLAG knock HeLa cells) was normalised to the abundance of GAPDH mRNA in the corresponding sample. The observed results imply that the RD histone mRNA levels are not affected (within detection limits) by the tagging of MBLAC1. Error bars represent SEM from three biological replicates.
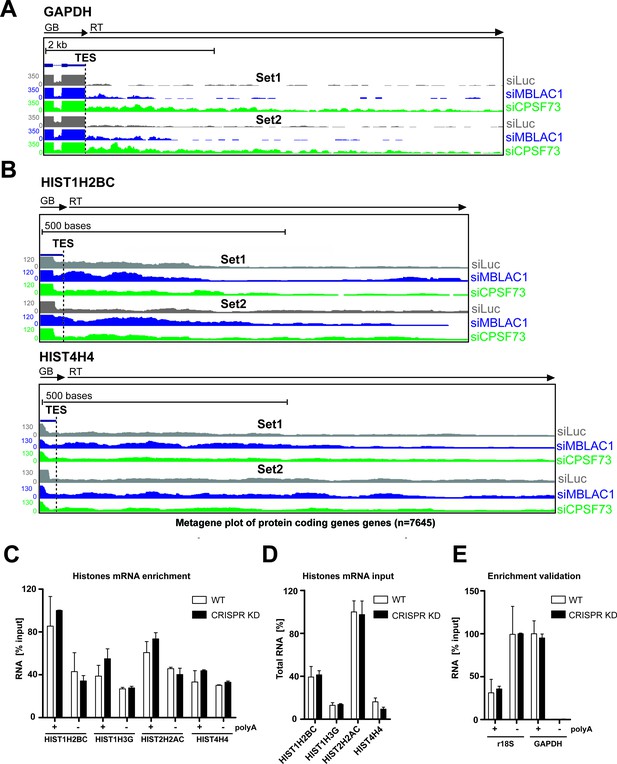
3ʹ end processing defects observed in replication-dependent histone pre-mRNA after MBLAC1 depletion and investigation of the effect of MBLAC1 depletion on the polyadenylated fraction of histone mRNA in HeLa cells.
(A, B) ChrRNA-seq analyses showing the effect of MBLAC1 or CPSF73 depletion in two biological replicates (Set1 and Set2) on GAPDH or selected RD histone (HIST1H2BC, HIST4H4) genes (UCSC Genome Browser) (Kent et al., 2002). Arrows indicate gene bodies and transcription termination defect orientation. (C) RT-qPCR quantification evaluating enrichment of polyadenylated RD histone RNA transcripts in MBLAC1 CRISPR/Cas9-mediated stable knockdown (KD) compared to wildtype (WT) cells. (D) The starting abundance of histone transcripts was quantified in total RNA by the standard curve method to ensure similar histone RNA levels before selection. (E) GAPDH and 18S rRNA were used as polya +and polyA- controls, respectively. For each primer set, amplicon abundance was calculated using the standard curve method; the relative percentage of transcripts in both, polyA ±fractions was expressed as ratios relative to the abundance of the total RNA used as the starting material for selection. Error bars represent SEM from two biological replicates. For histone genes, amplicons were selected in the gene bodies. Abbreviations: +, polyA plus; -, polyA minus.
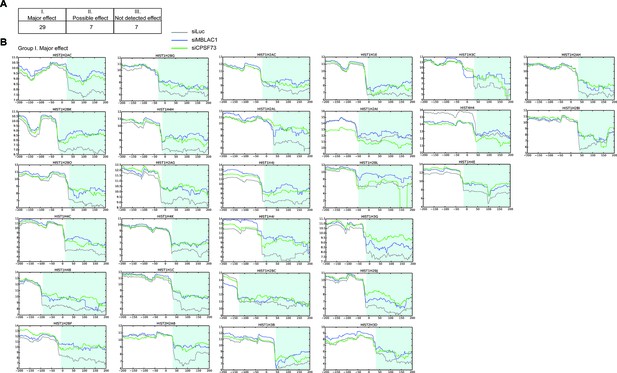
MBLAC1 and CPSF73 depletions affect RD histone pre-mRNA processing in vivo.
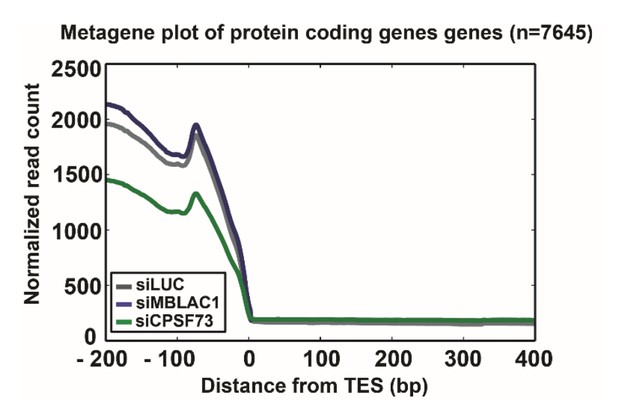
(A) Schematic grouping the apparent effects (major, possible, and no detected) of MBLAC1 depletion on processing of 43 histone genes, based on analysis of the chrRNA-seq data plots shown in B, Figure 4—figure supplements 3 and 4. The effect on the same histone genes following CPSF73 depletion is shown for comparison. (B) Log two transformed chrRNA-seq data plots illustrating extended transcripts for individual RD histone genes after normalization to the corresponding gene body following MBLAC1 or CPSF73 depletion; 43 non-overlapping RD histone genes (TES −200 bp, TES +200 bp) with an FPKM ≥1 were studied with two biological replicates (average of Set1 and Set2). Expression levels (as reads per kilobase per million mapped reads, RPKMs) were calculated for a region of 'TES −100 bp' for each gene in all the three samples, namely, LUC, MBLAC1, and CPSF73. The data are sorted into classes as described in A. (Color code: siLuc: grey, siMBLAC1: blue, siCPSF73: green. When observed, the region corresponding to the observed extended transcripts (read through) is in light blue.
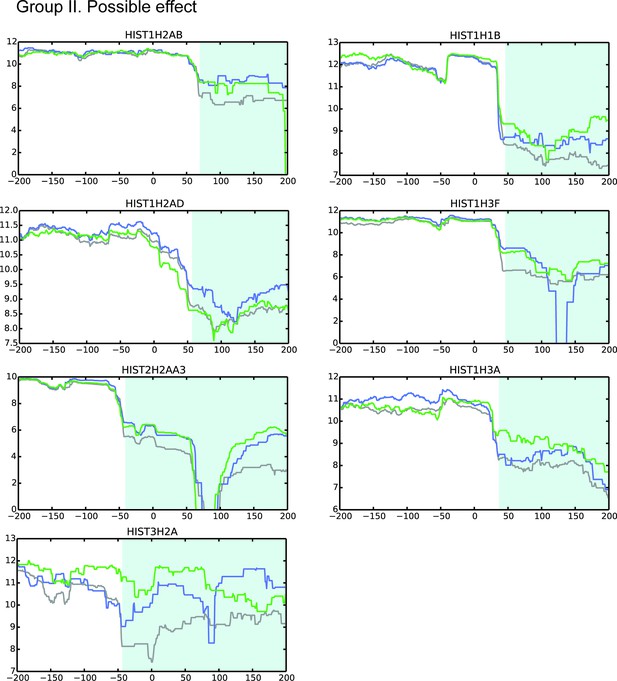
MBLAC1 and CPSF73 depletions affect RD histone pre-mRNA processing in vivo.
Refer to legend Figure 4—figure supplements 2
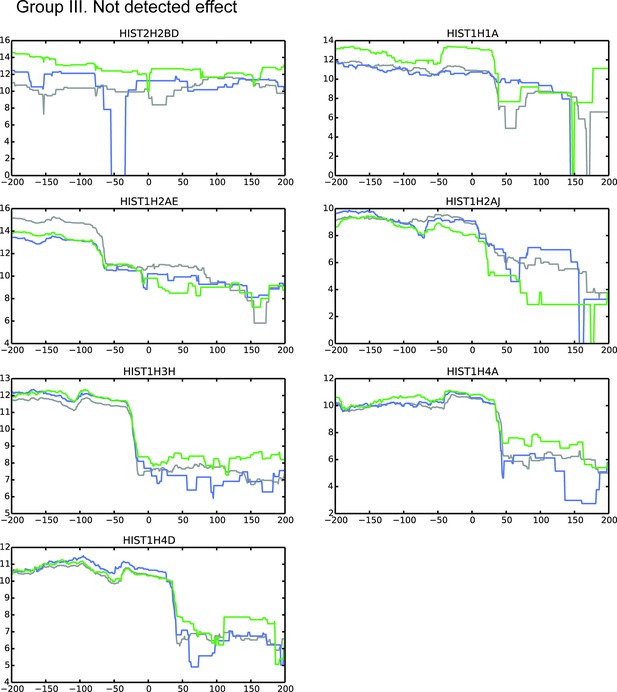
MBLAC1 and CPSF73 depletions affect RD histone pre-mRNA processing in vivo.
Refer to legend Figure 4—figure supplements 2
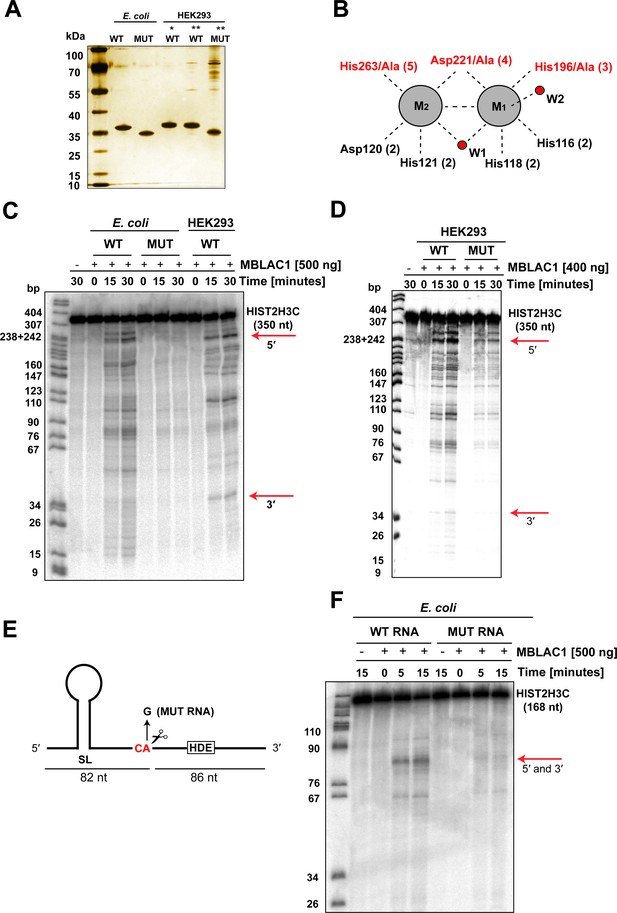
MBLAC1 has sequence-specific endoribonucleolytic activity on RD histone pre-mRNA.
(A) Silver staining analyses showing purity of MBLAC1 protein preparations produced from different sources. Note all proteins are highly purified except mutMBLAC1 produced in HEK293 which presents some contaminants. (B) Schematic view of metal binding and active site residues and the two water molecules (red spheres) observed to coordinate (black dashed lines) to the metals (grey spheres) at the MBLAC1 active site by crystallography (numbering as in Figure 1). Active site residues substituted to alanine in the MBLAC1 variant (H196A/D221A/H263A) are in red. (C) Time-dependent in vitro cleavage assay using an internally labeled [32P] histone 2H3C pre-mRNA fragment (350 nt) with E. coli produced wtMBLAC1, or an active site substituted MBLAC1 variant (H196A/D221A/H263A), or HEK293 produced wtMBLAC1 (media secreted). Note only a fraction of the total RNA is hydrolyzed. (D) Time-dependent in vitro cleavage assay using an internally labeled [32P] histone 2H3C pre-mRNA fragment (350 nt) with HEK293 produced wtMBLAC1, or an active site substituted MBLAC1 variant (H196A/D221A/H263A). The red arrows indicate the assigned position of 3' end processing in vivo. Note that both wt and mutMBLAC1 used in this experiment were purified from HEK293 cell lysates instead of from cell media due to the loss of secretion of the mutant enzyme in the cell media. (E) Schematic view of the histone 2H3C pre-mRNA fragment (168 nt) used in the cleavage assays showing preferential cleavage occurs after the CA dinucleotide located five nucleotides to the 3' side of the stem loop. The single-nucleotide substitution (A/G) at the CA cleavage site is shown. (F) Time dependence of E. coli produced wtMBLAC1 in vitro cleavage using an internally labeled [32P] unmodified (WT RNA), or with a single-nucleotide substituted (MUT RNA) histone pre-mRNA fragment (as in E). The red arrows indicate the assigned position of 3' end processing in vivo. Abbreviations: HDE, histone downstream element; SL, stem loop.
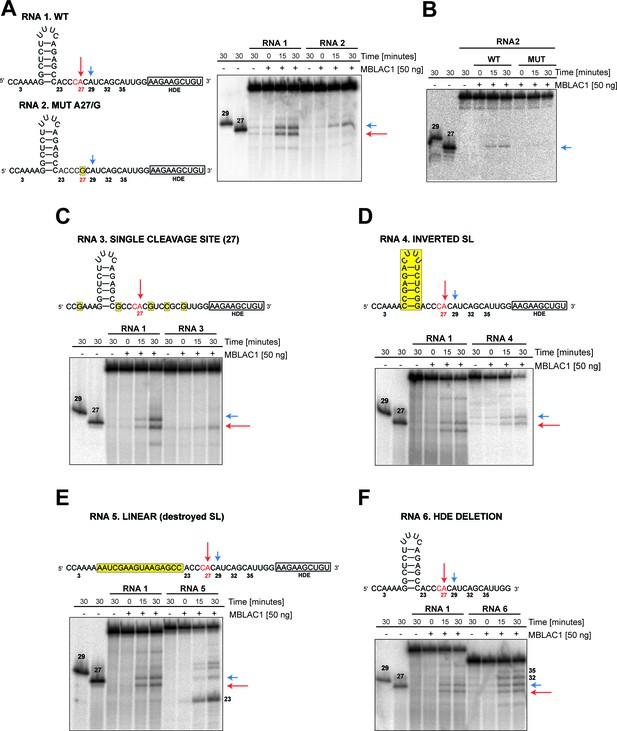
MBLAC1 shows specific endoribonucleolytic activity on histone pre-mRNA recognising specific sequence/structural RNA elements in vitro.
(A, C–F) Time-dependent in vitro cleavage assays using [γ-32P] ATP 5' labeled histone 2H3C RNA fragments (RNA 1–6) in the presence of HEK293 produced wtMBLAC1. (B) Time-dependent in vitro cleavage assays using [γ-32P] ATP 5' labeled histone 2H3C RNA 2 (50 nt) in the presence of HEK293 produced wild-type or mutMBLAC1. Cleavage products of RNA 2 after incubation with wtMBLAC1 (WT) are compared with those after incubation with mutMBLAC1 (MUT). Cleavage products of the RNA 1 are compared with those of the differently modified fragments. Schematic views of the RNA fragments are shown. The schematics show the numbering assigned to the possible cleavage sites present in each RNA; arrows indicate the physiologically major (red) and minor (blue) cleavage sites. The CA dinucleotide corresponding to the major cleavage site is in red. Sequence or structural modifications of each RNA are in yellow. The size of processed RNA products is indicated when appropriate. Unprocessed RNA fragments corresponding to nucleotides 1 to 29 and 1 to 27 of the WT RNA 1 were used as mass markers and indicated as 29 and 27, respectively. Note that the 29 and 27 markers migrate more slowly on gel electrophoresis compared to the corresponding fragments generated by MBLAC1 catalysis in positions 27 and 29. This observation is explained by the presence of an extra negatively charged phosphate group on the 5'end of the products generated by MBLAC1 catalysis, compared to the markers, as expected during maturation of histone transcripts (Furger et al., 1998).
Tables
Crystallographic data and refinement statistics PDB ID 4V0H.
https://doi.org/10.7554/eLife.39865.004| Native HSE (PDB ID: 4V0H) | |
|---|---|
| Data collection | |
| Space group | P1 |
| Cell dimensions | |
| a, b, c (Å) | 62.95, 67.13, 67.90 |
| α, β, γ (°) | 109.31, 105.40, 90.17 |
| Resolution (Å) | 45.96–1.79 (1.84–1.79 Å)* |
| Rmerge† | 0.10 (0.81) |
| I/σ(I) | 12.4 (2.6) |
| Completeness (%) | 95.5 (92.6) |
| Redundancy | 6.9 (6.7) |
| Refinement | |
| Resolution (Å) | 45.95–1.79 |
| No. reflections | 90641 |
| Rwork‡/Rfree§ | 0.182/0.211 |
| No. atoms | |
| Protein | 6235 |
| Ligand/ion | 38 |
| Water | 514 |
| B factors | |
| Protein | 27.27 |
| Ligand/ion | 44.40 |
| Water | 33.94 |
| R.m.s. deviations | |
| Bond lengths (Å) | 0.01 |
| Bond angles (°) | 1.37 |
-
* Values in parentheses are for highest-resolution shell.
†Rmerge = ∑h ∑l | Ihl - < Ih > | / ∑h ∑l < Ih > where Ihl is the lth observation of reflection h, and <Ih > is the mean intensity of that reflection.
-
‡Rwork = ∑ || Fobs |-|Fcalc||/|Fobs| x 100.
§Rfree is calculated in the same way as Rwork but using a test set containing 5.01% of the data, which were excluded from the refinement calculation.
Oligonucleotide primers and single guide RNA targeting sequences used in site directed mutagenesis, RT-qPCR experiments and CRISPR/CAS9 mediated KD studies.
Abbreviations: GB, gene body; RT, read through. Note that the standard BBL numbering system has been used (Galleni et al., 2001).
| Oligonucleotide primers used for site directed mutagenesis | |
|---|---|
| E. coli produced HSE | |
| H196A_F | GCAACACCGGGTGCTGGTGGTCAGCG |
| H196A_R | CGCTGACCACCAGCACCCGGTGTTGC |
| D221A_F | GTTGTTGCCGGTGCTGTTTTTGAACGTG |
| D221A_R | CACGTTCAAAAACAGCACCGGCAACAAC |
| H263A_F | GGTTGTTCCTGGTGCTGGTCCGCCTTTTCG |
| H263A_R | CGAAAAGGCGGACCAGCACCAGGAACAACC |
| Oligonucleotide primers used in RT-qPCR experiments | |
| GAPDH_GB_F (intron) | ACCCAGAAGACTGTGGATGG |
| GAPDH_GB _R (intron) | TTCAGCTCAGGGATGACCTT |
| GAPDH _RT_F | TCCAGCCTAGGCAACAGAGT |
| GAPDH _RT_R | TGTGCACTTTGGTGTCACTG |
| HIST1H2BC_GB_F | ACCTCCAGGGAGATCCAGAC |
| HIST1H2BC_GB_R | AGCTGGTGTACTTGGTGACG |
| HIST1H2BC_RT_F | CTCCAGGGAGATCCAGACGG |
| HIST1H2BC_RT_R | GCTCTTTTAGTGGGTATCTGGG |
| HIST4H4_GB_F | GAAGGTGCTGCGGGACAATA |
| HIST4H4_GB_R | AAGACTTTGAGGACTCCCCG |
| HIST4H4_RT_F | CGCGCAACGCAGTAGTGACC |
| HIST4H4_RT_R | CATTCAGCTTTCGGGCTTGCAGGTA |
| HIST2H2AC_GB_F | GGCTCGGGACAACAAGAAGA |
| HIST2H2AC_GB_R | AGAACGGCCTGGATGTTAGG |
| HIST2H2AC_RT_F | GAAAGCCACAAAGCCAAAAGC |
| HIST2H2AC_RT_R | GGCTTGACACCATACTCATTCACC |
| HIST1H3G_GB_F | CTGAGCTGCTGATCCGCAAG |
| HIST1H3G_GB_R | GGCGAGCGAGCTGAATGTCC |
| GAPDH_GB_F (exon-exon) | AAGGTGAAGGTCGGAGTCAA |
| GAPDH_GB _R (exon-exon) | AATGAAGGGGTCATTGATGG |
| r18S_F | GGCCCTGTAATTGGAATGAGTC |
| r18S_R | CCAAGATCCAACTACGAGCTT |
| CRISPR/Cas9 single guide RNA (sgRNA) | |
| Location | sgRNA targeting sequences |
| 5ʹ targeting | ACAGCGGCTCGGTCCGCATGAGG |
| 3ʹ targeting | TGCACTAATCAGCCTCGAGAGGG |
| CRISPR/Cas9 sequencing primers | |
| Primers sequence | |
| CRISPR/Cas9_F | CAGAGAACCGAGGCTTAGGG |
| CRISPR/Cas9_R | GAAGCTCCCACCCTTGACTG |
| CRISPR/Cas9 RT-qPCR primers | |
| Primers sequence | |
| HSE_set1_F | ATCACATCGGGAACTTGGGG |
| HSE_set1_R | CACCACGGTGCCCAGAG |
| HSE_set2_F | GCTCTGGGCACCGTGG |
| HSE_set2_R | GCGAGGCTTCCCTTAACACT |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.39865.017





