Global DNA methylation remodeling during direct reprogramming of fibroblasts to neurons
Figures
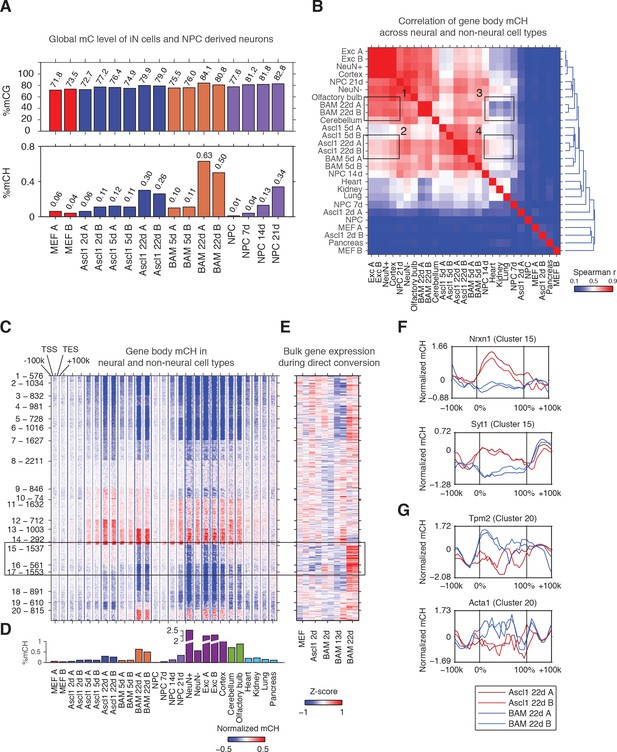
The Ascl1, Brn2 and Myt1l reprogramming factors induce an authentic, neuron-specific CH methylation pattern in fibroblasts.
(A) Global mCG (top panel) and mCH (bottom panel) levels of iN cells and NPC-derived cells. mCG and mCH accumulates in both maturing iN cells and differentiating NPCs. (B) Correlation of gene body mCH levels of iN cell reprogramming samples, NPC derived cells, neural and non-neural tissues. Mature cortical neurons clustered more closely to BAM 22d than Ascl1 22d iN cells. NeuN +and NeuN- indicate neuronal and glial nuclei separated using anti-NeuN antibody, respectively (Lister et al., 2013). Exc indicates purified nuclei from excitatory neuron expressing Camk2a+ (Mo et al., 2015). (C) Left panel - K-means clustering of gene body mCH normalized to flanking regions (100 kb surrounding the gene). Reprogramming with BAM produces a global mCH profile more similar to cortical neurons compared to Ascl1 alone. TSS - Transcription Start Site. TES - Transcription End Site. (D) Global mCH level of samples compared in (C). (E) Relative expression (Z-score) of bulk RNA-seq analysis of iN cell reprogramming. Gene expression is inversely correlated to gene body mCH. (F–G) mCH pattern at genes with neuronal (F) and myogenic functions (G) in Ascl1 22d and BAM 22d iN cells. BAM 22d cells show greater depletion of mCH at synaptic genes and accumulation of mCH at myogenic genes, suggesting more efficient activation of neuronal genes and suppression of the alternate myogenic program compare to Ascl1 22d.
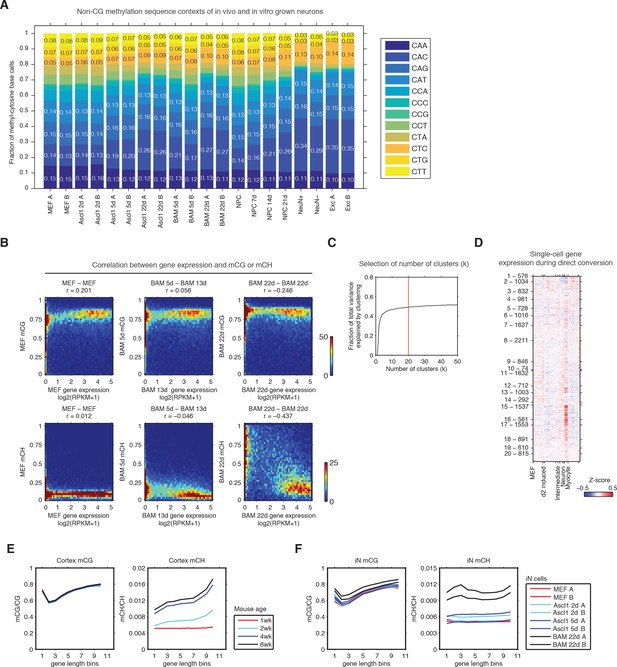
Fully reprogrammed iN cells accumulate CH methylation.
(A) The frequency of tri-nucleotide contexts in mCH base calls for iN cells and cortical neuron samples. (B) Correlations between iN cell gene body mCG or mCH levels and gene expressions. (C) The fraction of total variance explained plotted as a function of the number of clusters for the kmeans clustering shown in Figure 1C. (D) Single-cell gene expressions plotted for gene clusters shown in Figure 1C. (E) Genes with greater length are associated with high levels of mCH in mouse brain cortex. (F) iN cells show a less pronounced gene length- associated elevation of mCH level than observed in mouse cortex.
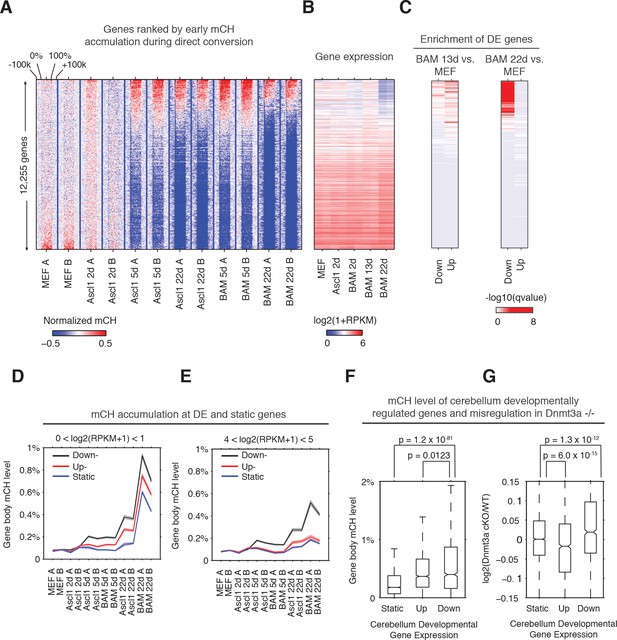
Early gene body mCH accumulation predicts later transcriptional downregulation.
(A and B) Normalized gene body mCH (A) and transcript abundance (B) for genes ranked by early mCH accumulation at BAM 5d. Early mCH accumulation is strongly correlated to gene repression in BAM 22d iN cells, and both upregulated and downregulated genes in BAM 13d iN cells. (C) Significance (hypergeometric test) of the enrichment in down- and up- regulated genes for BAM 13d and BAM 22d iN cells. (D and E) Gene body mCH dynamics of static, down- and up- regulated genes with different transcripts abundances - log2(RPKM +1) between 0 and 1 (D), between 4 and 5 (E) during iN cell reprogramming. (F) Gene body mCH level of cerebellum developmentally (P60 vs. P7) regulated genes that were actively expressed (RPKM >3). (G) Alteration of cerebellum gene expression by Nes-Cre;Dnmt3a-/- for developmentally regulated genes.
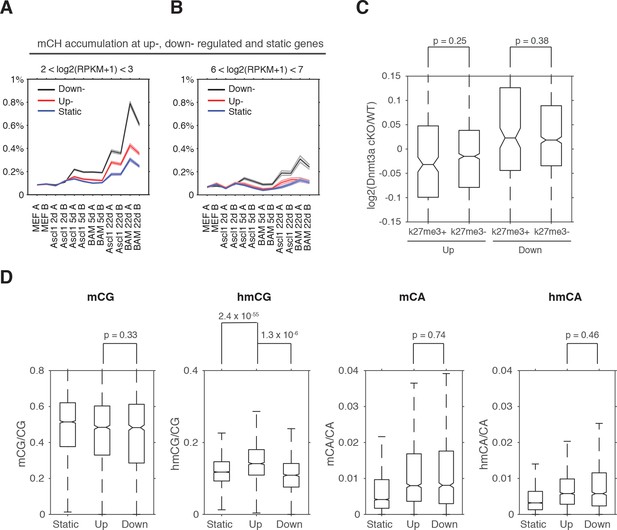
Dynamically regulated genes are enriched in CH methylation during direct reprogramming and cerebellum development.
(A and B) Gene body mCH dynamics of developmentally static, down- and up- regulated genes with different transcripts abundances - log2(RPKM +1) between 2 and 3. (A), between 6 and 7 (B). (C) Genes under the regulation of polycomb repressive complex (H3K27me3+) show similar misregulation in Dnmt3a cKO as genes not regulated by polycomb complex (H3K27me3-). (D) Developmentally upregulated genes in cerebellum accumulate more hydroxymethylcytosine in the CG dinucleotide context than developmentally downregulated genes and static genes.
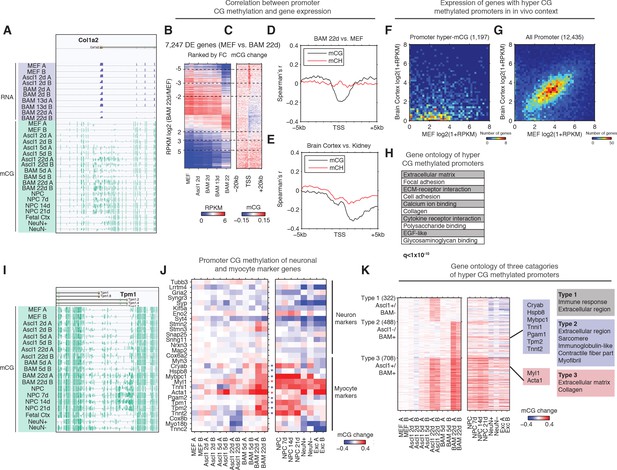
Silencing of key non-neuronal programs is associated with promoter hyper CG methylation.
(A) Browser view of fibroblast marker gene Col1a2 promoter methylation dynamics and corresponding gene expression during reprogramming. The promoter is strongly methylated throughout NPC differentiation, but is unmethylated in fetal cortex, cortical neurons and glia. However, it gradually accumulates mCG through the course of reprogramming using either Ascl1 alone or BAM factors. (B) Differentially expressed genes between BAM 22d iN and MEF were ranked by their relative fold-changes. (C) mCG change between MEF and BAM 22d in ±20 kb regions surrounding TSS of differentially expressed genes. Hyper CG methylation was found at TSS for downregulated genes. (D) Correlation between mCG change and differential gene expression between BAM 22d iN and MEF in ±5 kb region surrounding TSS. (E) Correlation between mCG change and differential gene expression between brain cortex and kidney in ±5 kb regions surrounding TSS. (F–G) Scatter plots comparing the expression of genes with hyper CG methylation promoter in BAM 22d iN cells (F) or all genes (G) between MEF and brain cortex. Hyper CG methylation genes in BAM 22d cells are strongly repressed compared to all genes in the in vivo context. (H) Gene ontology term enrichment of genes with hyper CG methylation promoters in BAM 22d iN cells. (I) Browser view of promoter mCG dynamics of myocyte marker gene Tpm1. The promoter is methylated through NPC differentiation, but is unmethylated in fetal cortex and cortical neurons and glia. However, it remains unmethylated when reprogramming with Ascl1 alone, and only accumulates mCG in the presence of Brn2 and Myt1l in BAM. (J) Promoter mCG dynamics of neuron and myocyte marker genes during reprogramming. Blue asterisks indicate promoters hyper-methylated in BAM 22d iN, but not in Ascl1 22d iN. Red asterisks indicate promoters hyper-methylated in both BAM 22d iN and Ascl1 22d iN. A majority of myocyte genes are only hyper methylated by BAM and not Ascl1 alone. (K) Left: Heatmaps showing promoter mCG during neuronal reprogramming, NPC differentiation and in primary neurons. Promoters were categorized into three groups showing hyper-mCG (mCG increase >0.1) in Ascl1 22d only, BAM 22d only or both type of iN cells. Right: Gene ontology enrichment of promoters for the three groups. Promoters of fibroblast genes were hyper CG methylated in both Ascl1 alone and BAM iN cells, but only BAM 22d cells exhibit strong hyper CG methylation at myogenic genes.
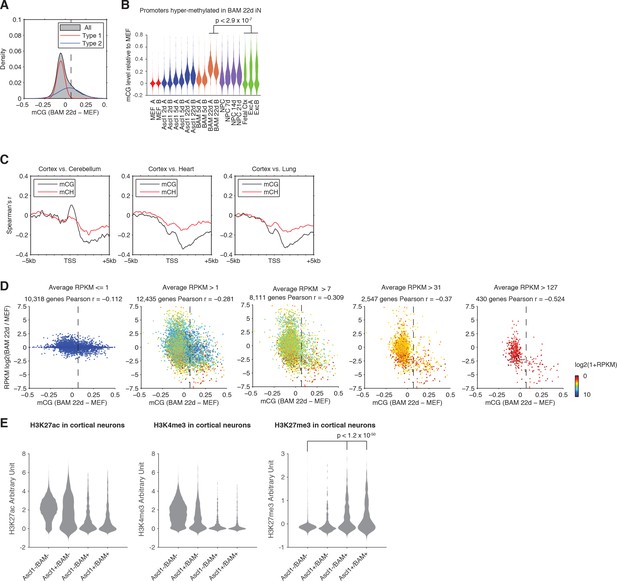
Direct reprogramming induces distinct promoter CG methylation accumulation.
(A) Probability density of normalized mCG changes (BAM 22d – MEF) for 1 kb regions surrounding TSS. A Gaussian mixed model with two components was fitted to the data and resulted in two distributions (type A and B) with μ values of −0.0551 and 0.0485, σ values of 0.0021 and 0.0169 and weights of 0.5559 and 0.4441, respectively. The threshold of hyper-mCG TSS was determined to be 0.0691. (B) mCG level of promoters showing hyper-methylation during direct reprogramming induced by BAM compared to in vivo grown cell. mCG levels were normalized to that of MEF. (C) Correlation between mCG change and differential gene expression between cortex, cerebellum, heart, and lung in ±5 kb regions surrounding TSS. (D) Scatter plots of promoter mCG change against differential gene expression between BAM 22d iN cells and MEF for genes with average RPKM <= 1,>1,>7,>31 and>127. The vertical dashed line indicates the threshold of promoters showing significant hyper CG methylation. There is a strong inverse correlation between promoter hyper CG methylation and gene expression change. (E) All promoters were separated into four categories depending on their hyper-mCG states in Ascl1 22d iN or BAM 22d iN cells. Violin plots show H3K27ac, H3K4me3 and H3K27me3 levels of the four types of promoters in cortical neurons to explore their chromatin states in in vivo grown cells.
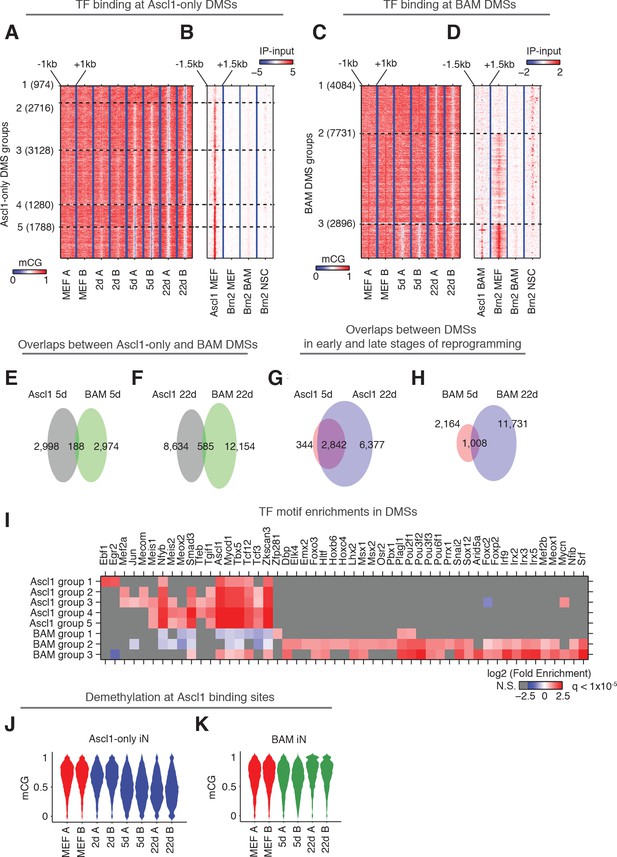
Ascl1 and BAM factors induce distinct CG methylation reconfigurations.
(A and C), DMSs identified and combined from all pairwise comparisons during reprogramming driven by Ascl1 alone (A) or BAM (C) were ordered by the kinetics of mCG remodeling (see Materials and methods). The plots show clusters containing more than 100 DMSs, which are statistically more robust. (B and D) Signal intensity of Ascl1 and Brn2 ChIP-seq were plotted for 3 kb regions surrounding the DMSs. (E–H) The overlaps of DMSs identified at different stages of reprogramming driven by Ascl1 or BAM expression. (I) TF binding motif enrichment in DMS groups shown in (A) and (C). Insignificant enrichments (q > 1×10−5) were shown as gray. (J) The distribution of mCG level at Ascl1 peaks during Ascl1-only induced cells during reprogramming. (K) The distribution of mCG level at Ascl1 peaks in BAM induced cells during reprogramming.
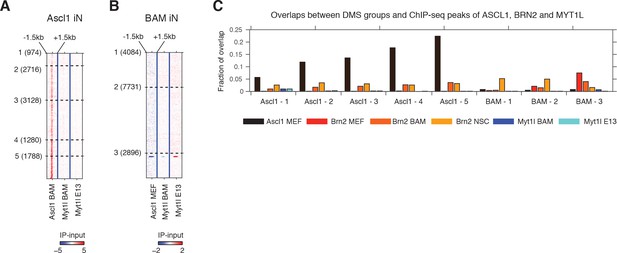
Differentially methylated sites (DMSa) induced by direct reprogramming overlap with ASCL1 and BRN2, but not MYT1L binding sites.
(A) Signal intensity of Ascl1 and Myt1l ChIP-seq were plotted for 3 kb regions surrounding DMSs induced by Ascl1-only reprogramming. (B) Signal intensity of Ascl1 and Myt1l ChIP-seq were plotted for 3 kb regions surrounding DMSs induced by BAM reprogramming. (C) The overlap between DMS groups and Ascl1, Brn2 and Myt1l binding sites.
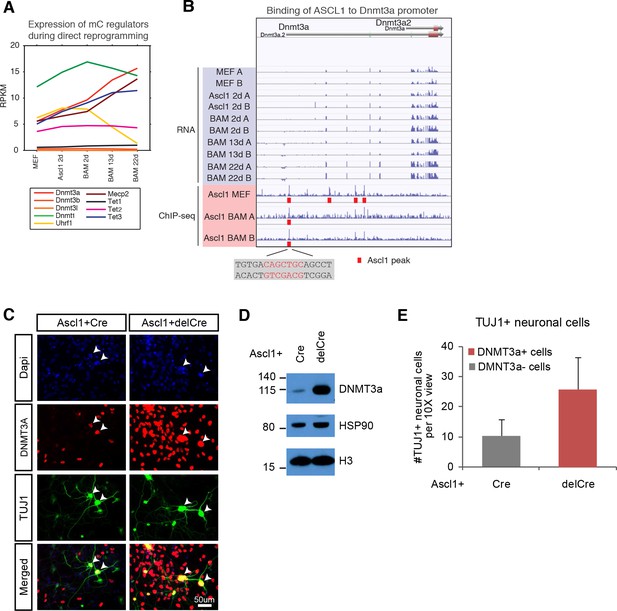
de novo DNA methylation is required for efficient reprogramming from fibroblasts to neurons.
(A) Expression of DNA methylation regulators and readers during neuronal reprogramming. (B) Ascl1 ChIP-seq peak is located proximal to Dnmt3a promoter. (C) Immunostaining of Dnmt3a/3bfl/fl cells 14d post-Ascl1 induction with dox. There appears to be a similar reprogramming efficiency when comparing the CreGfp (left) and delCreGfp (right) conditions. However, most TUJ1 +iN cells in the CreGfp condition were still expressing low levels of DNMT3A (white arrows) despite co-infection with CreGfp. (D) Western blot showing efficient knock-out of DNMT3A in Ascl1 +Cre expressing cells compared to Ascl1 +delCre control 13 days post induction of Ascl1. (E) Average counts of TUJ1 +iN cells per 10X field of view 14d post-Ascl1 induction that are co-infected with 1) CreGfp and Dnmt3a- (gray) or 2) delCreGfp and Dnmt3a+ (red) (error bars are stderr, n = 3).
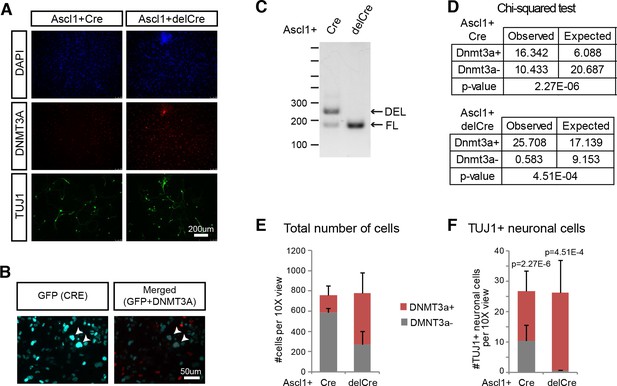
Deletion of Dnmt3a/3b in fibroblasts leads to reduced reprogramming efficiency.
(A) 10X view of immunostaining in Ascl1 +CreGfp/delCreGfp samples. (B) Corresponding GFP staining (for CREGFP fusion protein) for Figure 5C. Even though iN cells are expressing high levels of CRE, which should loop out the Dnmt3a locus, cells are still expressing DNMT3A protein. (C) Genotyping of Dnmt3a/b fl/fl cells. Floxed mutant bands (FL) appear at ~200 bp while deleted bands (DEL) appear at ~250 bp. (D) Chi-squared test for significance for Figure 5—figure supplement 1E–F. The null hypothesis is that if Dnmt3a- cells reprogram as well as the Dnmt3a + cells, there should be no significant difference in the frequency of Dnmt3a- cells and the frequency of cells that actually reprogram into neurons. There is a significant difference for both Cre (p=2.27E-6) and delCre (p=4.51E-4) conditions, indicating that Dnmt3a- cells are less efficient in reprogramming into iN cells. (E and F) Average counts of DAPI +cells (E) or TUJ1 +iN cells (F) 14d post-Ascl1 induction that are either DNMT3A positive (red) or negative (gray) per 10X field of view. Using a Chi-squared test (Figure 5D; Figure 5—figure supplement 1D), there is a significant difference between the frequencies of total number DNMT3A- cells compared to the number of DNMT3A- iN cells in both Cre and delCre conditions.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Antibody | Mouse monoclonal anti-TUBB3 (Tuj1) | Covance | MMS-435P; RRID:AB_2313773 | (1:1000) IF |
| Antibody | Rabbit polyclonal anti-DNMT3A (H-295) | Santa Cruz Biotech | sc-20703; RRID:AB_2093990 | (1:1000) WB, (1:200) IF |
| Antibody | Chicken polyclonal anti-GFP | Abcam | ab13970; RRID:AB_300798 | (1:1000) IF |
| Antibody | rabbit polyclonal anti-HSP90 | Cell signaling | 4874; RRID:AB_2121214 | (1:1000) WB |
| Antibody | rabbit polyclonal anti-H3 | Cell signaling | 9712 | (1:10000) WB |
| Other | Dapi | ThermoFisher Scientific | D1306 | 1 ug/ml |
| Genetic reagent (M. musculus, female) | C57BL/6J | The Jackson Laboratory | 000664; RRID:IMSR_JAX:000664 | |
| Genetic reagent (M. musculus, male) | Mapt-tm1(EGFP)Klt | The Jackson Laboratory | 004779 | Homozygous TauEGFP |
| Strain, strain background (M. musculus) | Dnmt3a fl/fl | PMID: 15215868,25130491,15757890 | ||
| Strain, strain background (M. musculus) | Dnmt3b fl/fl | PMID: 15215868,25130491,15757890 | ||
| Commercial assay or kit | DNeasy Blood and Tissue Kit | Qiagen | 69506 | |
| Commercial assay or kit | MethylCode Bisulfite Conversion Kit | Invitrogen | MECOV50 | |
| Software, algorithm | Methylpy | PMID: 26030523 | ||
| Software, algorithm | Bowtie2 | PMID: 22388286 | RRID:SCR_016368 | |
| Software, algorithm | STAR | PMID: 23104886 | RRID:SCR_015899 | |
| Software, algorithm | edgeR | PMID: 19910308 | RRID:SCR_012802 | |
| Software, algorithm | DSS | PMID: 24561809 | RRID:SCR_002754 |
Additional files
-
Supplementary file 1
Summary of MethylC-seq experiments.
- https://doi.org/10.7554/eLife.40197.013
-
Supplementary file 2
Enriched gene ontology terms in gene clusters defined by gene body mCH.
- https://doi.org/10.7554/eLife.40197.014
-
Supplementary file 3
Differentially methylated sites (DMSs) identified during direct reprogramming driven by Ascl1-alone or BAM.
- https://doi.org/10.7554/eLife.40197.015
-
Transparent reporting form
- https://doi.org/10.7554/eLife.40197.016





