Delayed inhibition mechanism for secondary channel factor regulation of ribosomal RNA transcription
Figures
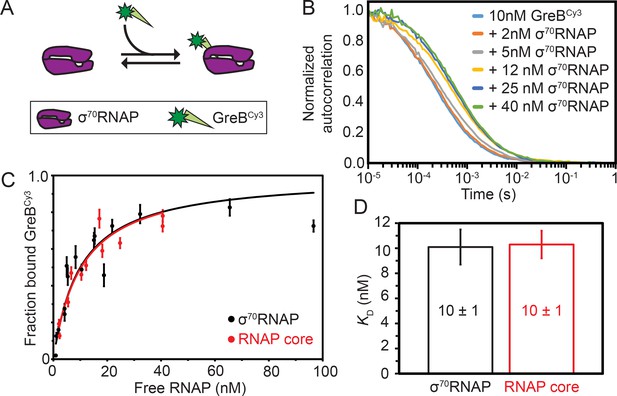
Thermodynamic and kinetic stability of GreB•RNAP complexes.
(A) Experimental scheme for measuring by FCS the affinity with which Cy3- (green star) labeled GreB binds σ70RNAP. (B) Examples of normalized FCS autocorrelation curves obtained after adding the indicated amount of σ70RNAP to GreBCy3. (C) Fraction of GreBCy3 bound to RNAP determined at different σ70RNAP (black) and RNAP core (red) concentrations in the presence of 10 nM GreBCy3, determined by fitting FCS autocorrelation curves including those shown in (B). Fraction bound measurements (points; ±SE) were fit to a Langmuir binding equation (lines). (D) Dissociation equilibrium constants KD (±SE) determined from the fits in (C).
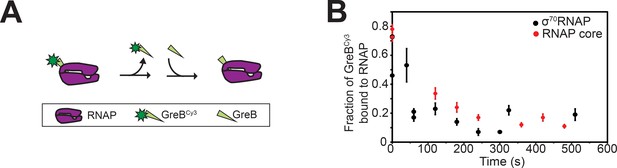
Kinetics of GreB dissociation from RNAP and σ70RNAP.
(A) Experimental scheme for observing dissociation of GreBCy3-RNAP complexes by FCS. (B) GreBCy3 remaining bound (±SE) to σ70RNAP (black) or RNAP (red) at different times after the addition of 100-fold excess unlabeled GreB, calculated from fits to FCS autocorrelation curves.
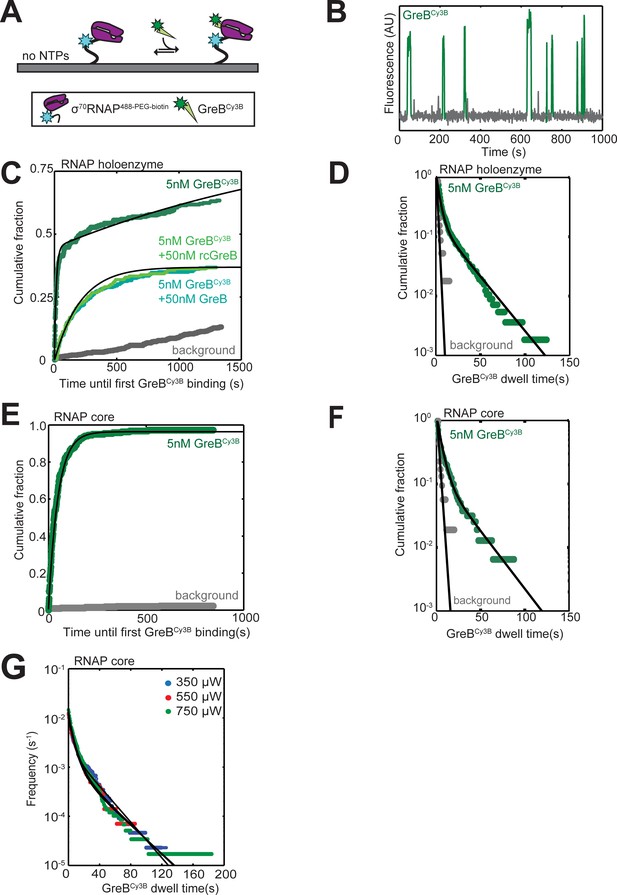
Kinetics of GreB binding to and dissociation from RNAP and σ70RNAP.
(A) Experimental scheme for single-molecule measurement of the kinetics of GreB binding to and dissociation from surface-tethered σ70RNAP. σ70RNAP488-PEG-biotin was tethered to the slide surface via a neutravidin linkage and the co-localization of GreBCy3B (5 nM, 93% labeled) was monitored. (B) Example GreBCy3B intensity record from the location of a single σ70RNAP488-PEG-biotin molecule. An objective image analysis algorithm (see Materials and methods) detected a spot of GreBCy3B fluorescence at the times shown in green. (C) Cumulative distribution of the time until the first GreBCy3B appearance at the location of each σ70RNAP488-PEG-biotin molecule (dark green). Exponential fit (black) is corrected for non-specific binding of GreBCy3B to the chamber surface measured at control locations where no σ70RNAP488-PEG-biotin was detected (gray), yielding kon = 1.5 ± 0.9×107 M−1 s−1. Also shown are data from experiments with added 50 nM GreB or rcGreB (light green curves) demonstrating that either species effectively competed with 5 nM GreBCy3B, indicating that GreBCy3B binding is specific. The curve shown (black) is the predicted cumulative distribution of times until the first GreBCy3B binding event at 50 nM GreB or rcGreB based on a competition model which assumes that GreB and rcGreB exhibit the same kinetics of binding to and release from σ70RNAP488-PEG-biotin as GreBCy3B (i.e., the same kon, Af, and τ1 reported in Table 1). (D) Cumulative distribution of GreBCy3B dwell times on surface-tethered σ70RNAP488-PEG-biotin molecules. Data (green) were fit to a biexponential model (black) with the parameters listed in Table 1. (E and F) Same as (C) and (D), but with core RNAP488-PEG-biotin instead of holoenzyme. Fit parameters and numbers of observations distributions in C-F are given in Table 1. (G) GreBCy3B photobleaching control. Cumulative frequency distribution (Friedman and Gelles, 2015) of GreBCy3Bdwell times on surface-tethered RNAP core measured as in (F) but at different 532 nm laser excitation powers. Data show no evidence for shorter lifetimes at higher power, suggesting that there is no appreciable photobleaching under the conditions of the experiments.
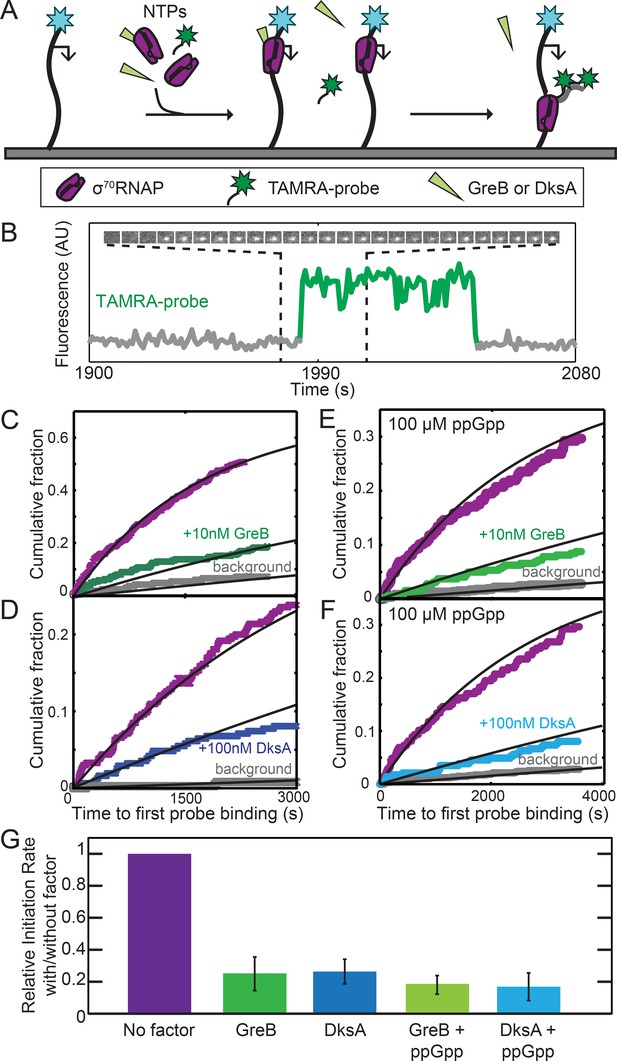
GreB and DksA equally inhibit transcription initiation from rrnB P1 at equal occupancies of σ70RNAP.
(A) Schematic of CoSMoS transcription initiation rate measurement in the presence of GreB or DksA. DNA molecules (314 bp) containing the rrnB P1 promoter (bent arrow) and labeled with AF488 (blue star) were tethered to the chamber surface via a biotin-neutravidin linkage. Initiation in the presence or absence of DksA, GreB, and/or ppGpp is detected as co-localization of two oligonucleotide probes labeled with TAMRA (green star) that hybridize near the 5′ end of the nascent transcript. (B) Example transcript probe fluorescence images (1.1 × 1.1 µm) taken at 5 s intervals (top) and corresponding intensity record (bottom) from the location of a single template DNA molecule. An objective image analysis algorithm (see Materials and methods) detected a spot of probe fluorescence at the times shown in green. (C–F) Cumulative distributions of the time until first probe detection on each DNA molecule. Only probe colocalization events 5 s or longer in duration were scored. Measurements were made in the absence (purple) or presence (green and blue) of secondary channel factors GreB (C, E) or DksA (D, F), without (C, D) or with (E, F) 100 μM ppGpp. Exponential fits (lines) are corrected (see Materials and methods) for non-specific binding of probe to the chamber surface measured at control locations that lack DNA molecules (gray). (G) Inhibition of initiation by secondary channel factors with and without 10 nM GreB, 100 nM DksA, and/or 100 μM ppGpp derived from experiments in (C–F), taken from the fit parameters (Table 2). Graph indicates the rate of initiation in the presence of the indicated factors (blue and green) relative to that measured in the same experiment without the factors (purple). Error bars: S.E.
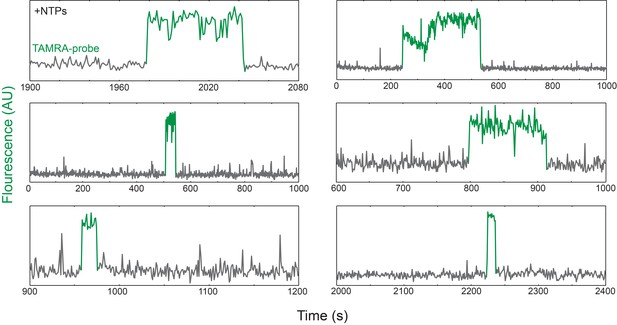
Additional examples of time records of transcript probe (green-excited) fluorescence co-localized at single rrnB P1 DNA locations.
Data are plotted as in Figure 2B.
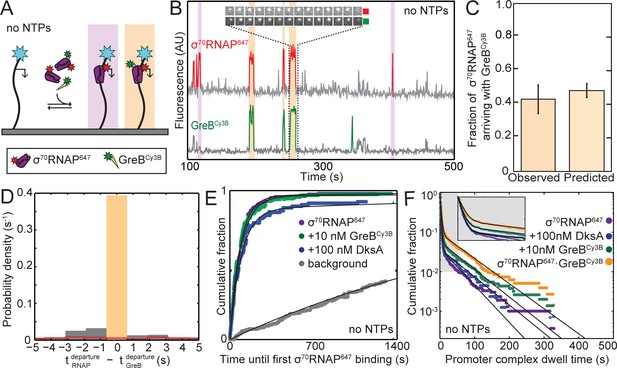
GreB and RNAP arrive together at rrnB P1 and depart together.
(A) Experiment design. Experiments were conducted in the absence of NTPs to monitor colocalization of σ70RNAP647 (1 nM) and GreBCy3B (10 nM) with the same surface-tethered rrnB P1 DNA used in Figure 2. (B) Recording excerpt of fluorescence co-localized with a single DNA molecule. Colored portions of the traces indicate detection of a DNA-colocalized spot of σ70RNAP647 (red) or GreBCy3B (green) fluorescence. Events in which molecules of σ70RNAP647 and GreBCy3B simultaneously arrive at and later simultaneously depart from the DNA molecule (orange) and events in which σ70RNAP647 arrives alone (purple) are highlighted. Inset shows time series images (0.5 s per frame; each 1.1 × 1.1 µm) of one of these events. (C) Fraction (±SE) of σ70RNAP647 DNA binding events (N = 2,408) in which a GreBCy3B arrived simultaneously (±1 frame) with σ70RNAP647 (left) and the fraction predicted by an equilibrium model (see Materials and methods) in which σ70RNAP and GreB·σ70RNAP bind to the promoter with equal association rate constants (right). (D) Histogram of time differences between σ70RNAP647 departure and GreBCy3B departure from σ70RNAP647•GreBCy3B•DNA complexes (solid bars). Only complexes that formed by simultaneous arrival (C; N = 1,053) were analyzed. Departure was simultaneous (±1 frame; orange highlight) in 91% of complexes. In a control analysis of the same data (red, open bars) in which departure events from different molecules were randomly paired (see Materials and methods), less than 3% were simultaneous. Plot shows the center of the distributions; 6% of experimental data and 61% of control fall outside the plotted range. In (C) and (D), data are aggregated from three replicate experiments. (E) Cumulative fraction of DNA molecules to which the first binding of σ70RNAP647 occurred at or before the indicated time. Experiments were conducted with or without SCFs as indicated. Background curve (gray) is collected from randomly selected locations without detected DNA molecules. (For clarity, only the background curve from the experiment without SCFs is shown). Exponential fits (lines), each corrected using background data taken from the same recording yielded RNAP-DNA association rate constants (Table 3). (F) Cumulative distributions of σ70RNAP647 dwell times on rrnB P1 DNA in the absence or presence of added SCFs (points). Also shown is the distribution of the subset of σ70RNAP647 dwell times observed in the presence of 10 nM GreBCy3B for which GreBCy3B arrived and departed simultaneously with σ70RNAP647 (orange). Inset: magnified view. Fits to a biexponential dwell time distribution (black) yielded parameters reported in Table 3.
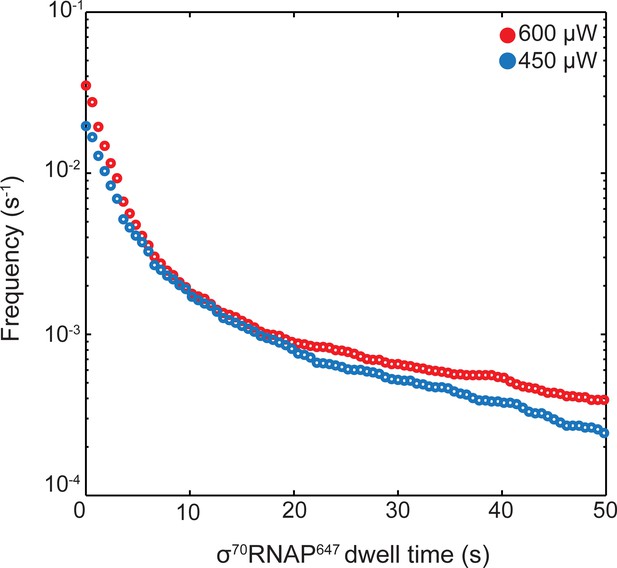
Photobleaching control for σ70RNAP647.
Cumulative frequency distributions (Friedman and Gelles, 2015) of σ70RNAP647dwell times recorded at 450 μW (blue; N = 2,143) and 600 μW (red; N = 2,576) 633 nm laser excitation powers using the experimental design shown in Figure 3A. Data show no evidence for systematically shorter lifetimes at higher power, suggesting that there is not significant photobleaching under the conditions of the experiments.
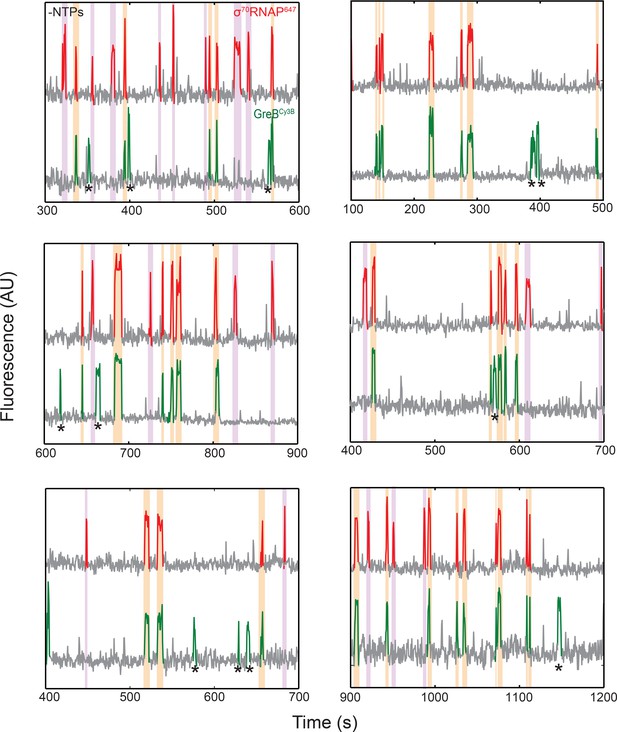
Fluorescence intensity records from six additional DNA locations demonstrating σ70RNAP647 and GreBCy3B co-localization in the absence of NTPs.
Data are plotted as in Figure 3B. Events in which molecules of σ70RNAP647 and GreBCy3B simultaneously arrive at and later simultaneously depart from the DNA molecule (orange) and events in which σ70RNAP647 arrives alone (purple) are highlighted. Occasional events in which GreBCy3B is present without σ70RNAP647 (examples marked by asterisks) are presumed to be due to the presence of a fraction of unlabeled σ70RNAP molecules, since little GreB binding is observed in the absence of RNAP (see text).
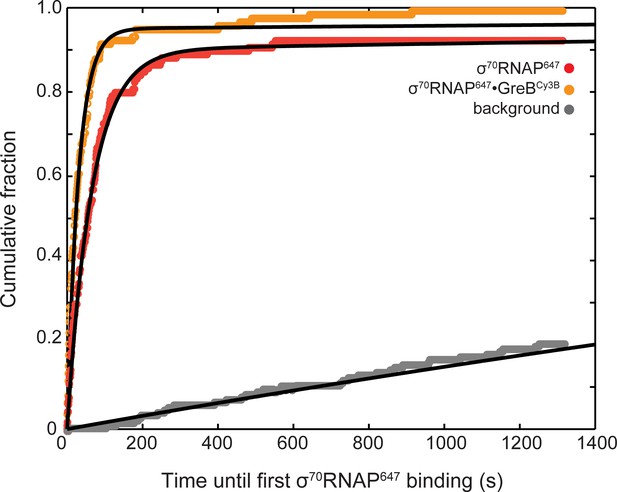
Effect of bound GreBCy3B on the kinetics of σ70RNAP647 association with DNA.
Data are taken from the same experiment and analysis as Figure 3E, green curve, except that the set of σ70RNAP647 binding events were separated into two disjoint subsets: σ70RNAP647 that arrived at DNA without (red) and with (orange) GreBCy3B. Background curve (gray) was collected from randomly selected locations without detected DNA molecules in the same experiment. Data were fit to background-corrected exponential models as in Figure 3E; fit parameters and number of observations are reported in Table 3.
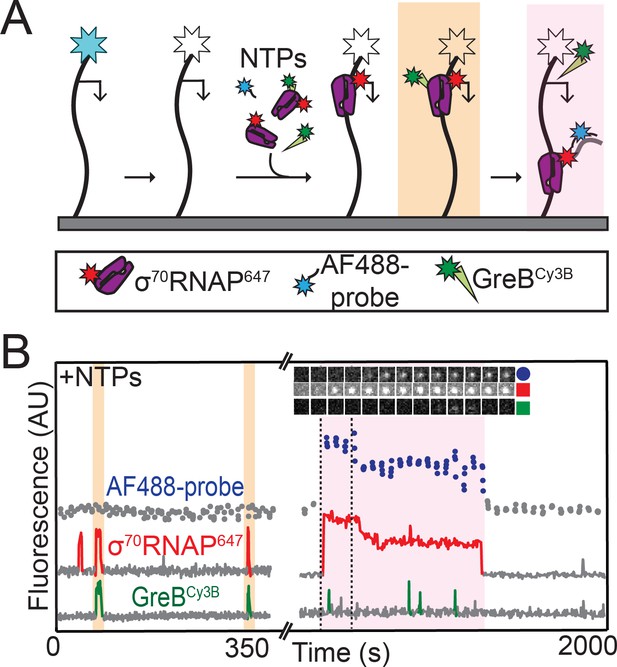
RNAP does not initiate transcription when GreB is bound.
(A) Experiment design. Template and labeled proteins were the same as in Figure 3A. DNA was first imaged, then the AF488 dye (blue star) was bleached (white star). Then AF488 oligonucleotide probes and 1 mM NTPs were introduced as in Figure 2, along with 1 nM σ70RNAP647 and 10 nM GreBCy3B. (B) Two excerpts from the record of fluorescence intensities colocalized with one template DNA molecule. Two events in which σ70RNAP647 bound to DNA accompanied by GreBCy3B (orange) and one in which σ70RNAP647 arrived alone (purple) are highlighted. Only the last resulted in transcription initiation as judged by AF488-probe colocalization. Time series images (1.1 × 1.1 µm) from the time interval delimited by dashed lines confirm appearance of co-localized AF488-probe and σ70RNAP647.
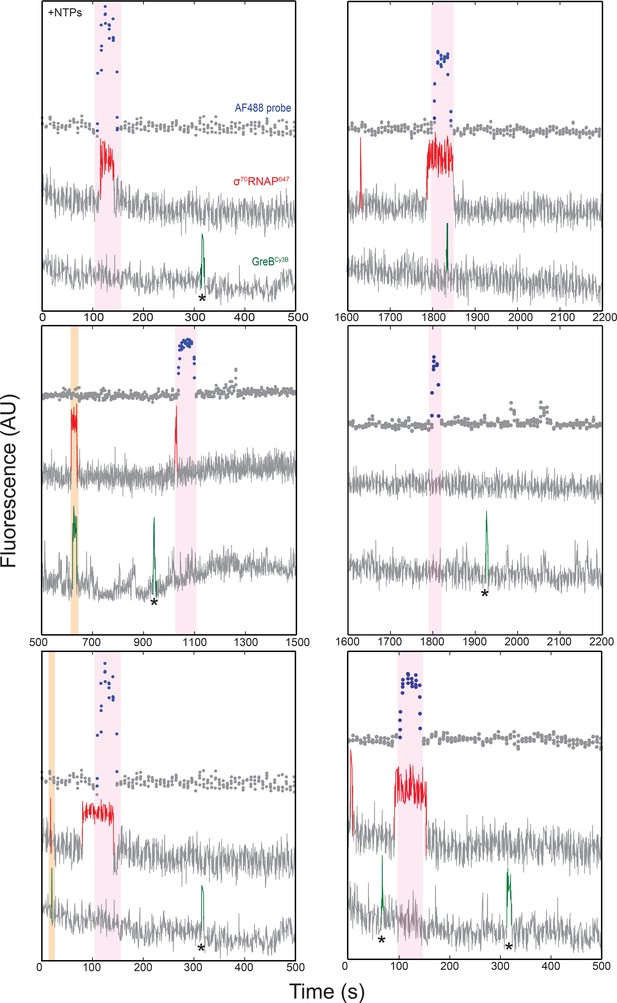
Additional fluorescence intensity records demonstrating σ70RNAP647, GreBCy3B, and AF488-oligo probe co-localization at single DNA locations in the presence of NTPs, as in Figure 4B.
Examples of simultaneous σ70RNAP647•GreBCy3B co-localization events (orange) and initiation events (pink) are highlighted.
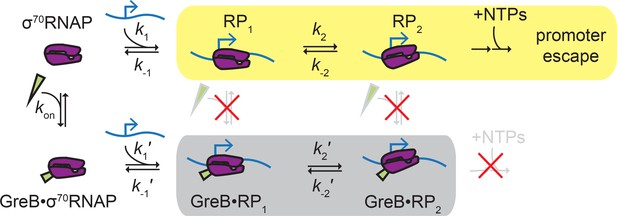
Proposed ‘delayed inhibition’ mechanism for growth-dependent regulation of rRNA initiation.
σ70RNAP (purple) binds promoter DNA (blue) and forms closed complexes RP1 and RP2 before isomerization to open complex, initial transcript synthesis, and promoter escape in the productive initiation pathway (yellow). GreB (green) binds rapidly and reversibly to free σ70RNAP (Table 1), and GreB does not alter the kinetics of closed complex formation or decay (Table 4). However, GreB blocks a later step in initiation and GreB dissociation from RNAP cannot compete with RNAP dissociation from the promoter; thus, promoter binding by GreB•σ70RNAP is a non-productive pathway (gray) that does not initiate RNA synthesis. DksA likely regulates rRNA initiation by a similar mechanism.
Tables
Kinetics of GreBCy3B interactions with RNAPs.
| Experiment | Species | Time to first binding measurement | Dwell time distribution fit parameters† | KD (nM) | ||
|---|---|---|---|---|---|---|
| kbkgnd (107 M−1 s−1) | kon (107 M−1 s−1) | Af‡ | ||||
| FCS (Figure 1) | RNAP core | N.D. | N.D. | N.D. | N.D. | 10 ± 1 |
| FCS (Figure 1) | σ70RNAP | N.D. | N.D. | N.D. | N.D. | 10 ± 1 |
| CoSMoS (Figure 1—figure supplement 2) | RNAP core | 0.00023 ± 0.00009 (N = 12) | 0.94 ± 0.06 (N = 163) | 0.87 ± 0.13 | a = 91 ± 4% τ1 = 5.3 ± 0.5 τ2 = 27 ± 3 s (N = 872) | 18 ± 2* |
| CoSMoS (Figure 1—figure supplement 2) | σ70RNAP | 0.0017 ± 0.0008 (N = 29) | 1.5 ± 0.9 (N = 152) | 0.43 ± 0.11 | a = 89 ± 7% τ1 = 5.6 ± 0.6 τ2 = 25 ± 7 (N = 943) | 11 ± 1* |
| CoSMoS§ | EC | N.R. | 2.7 ± 1.0 | 0.80 ± 0.06 | τ = 0.4 ± 0.1 s | 93 ± 36 |
| CoSMoS§ | Static EC−6 | N.R. | 1.9 ± 0.1 | 1 | τ = 0.29 ± 0.02 s | 184 ± 8 |
-
*Calculated as KD = (kon τ1)−1.
†From single- or bi-exponential fit.
-
‡Active fraction of surface-tethered molecules (Friedman and Gelles, 2015).
§Data from Tetone et al. (2017).
-
N.D., not determined; N.R., not reported.
Fit parameters for single molecule initiation experiments*.
| Conditions | Active fraction | N† | kbkgnd (10−5 s−1) | kinit (10−4 s−1) | Avg kinit (10−4 s−1) (Fold inhibition)‡ |
|---|---|---|---|---|---|
| σ70RNAP | 0.63 ± 0.12 | 42/225 | 1.1 ± 0.4 | 6.1 ± 0.6 | 5.8 ± 0.6 (1) |
| 0.37 ± 0.08 | 55/253 | 1.3 ± 0.2 | 6.4 ± 0.3 | ||
| 0.29 ± 0.05 | 90/373 | 0.9 ± 0.4 | 5.6 ± 0.5 | ||
| 0.47 ± 0.09 | 143/307 | 0.9 ± 0.6 | 4.9 ± 0.8 | ||
| 0.68 ± 0.10 | 185/298 | 0.6 ± 0.3 | 3.1 ± 0.5 | ||
| 0.62 ± 0.14 | 124/201 | 1.1 ± 0.9 | 7.0 ± 0.9 | ||
| 0.87 ± 0.13 | 279/322 | 7.7 ± 0.2 | 8.5 ± 0.3 | ||
| σ70RNAP +10 nM rcGreB | 0.63 ± 0.12 | 19/278 | 1.1 ± 0.4 | 1.5 ± 0.3 | 1.3 ± 0.4 (4.4 ± 0.7) |
| 0.47 ± 0.09 | 24/252 | 0.9 ± 0.6 | 1.2 ± 0.6 | ||
| σ70RNAP +10 nM GreB (Figure 2C) | 0.62 ± 0.14 | 21/246 | 2.6 ± 0.6 | 1.2 ± 0.2 | 1.9 ± 0.5 (3.1 ± 0.9) |
| 0.87 ± 0.13 | 16/198 | 7.7 ± 0.2 | 2.3 ± 0.4 | ||
| σ70RNAP +10 nM GreB +100 μM ppGpp (Figure 2E) | 0.37 ± 0.08 | 18/276 | 1.3 ± 0.2 | 1.1 ± 0.4 | 1.0 ± 0.7 (5.8 ± 1.1) |
| 0.47 ± 0.09 | 14/231 | 0.9 ± 0.6 | 0.9 ± 0.3 | ||
| σ70RNAP +100 nM DksA (Figure 2D) | 0.29 ± 0.05 | 25/302 | 0.9 ± 0.4 | 0.6 ± 0.3 | 1.7 ± 0.6 (3.4 ± 1.0) |
| 0.68 ± 0.10 | 36/305 | 0.6 ± 0.3 | 1.7 ± 0.8 | ||
| σ70RNAP +100 nM DksA + 100 μM ppGpp (Figure 2F) | 0.37 ± 0.08 | 14/202 | 1.3 ± 0.2 | 1.3 ± 0.5 | 1.1 ± 0.6 (5.3 ± 1.0) |
| 0.68 ± 0.10 | 24/299 | 0.6 ± 0.3 | 1.0 ± 0.6 | ||
| σ70RNAP647 | 0.71 ± 0.19 | 265/377 | 0.7 ± 0.4 | 5.3 ± 0.8 | 5.5 ± 0.7 (0.95 ± 0.6) |
| 0.49 ± 0.13 | 104/215 | 0.8 ± 0.3 | 5.8 ± 0.5 | ||
| σ70RNAP647 +10 nM GreBCy3B | 0.71 ± 0.19 | 29/306 | 0.7 ± 0.4 | 1.2 ± 0.9 | 1.4 ± 0.7 (3.9 ± 1.0) |
| 0.49 ± 0.13 | 14/147 | 0.8 ± 0.3 | 1.6 ± 0.6 | ||
| σ70RNAP647 +10 nM GreBCy3B +100 μM ppGpp | 0.71 ± 0.19 | 12/228 | 0.7 ± 0.4 | 0.9 ± 0.5 | 0.9 ± 0.5 (6.1 ± 0.7) |
| 0.49 ± 0.13 | 17/311 | 0.8 ± 0.3 | 0.9 ± 0.4 |
-
*At least two replicates per experimental condition were performed due to variability in active fraction between trials. The rate of TAMRA-probe binding to AF488-DNA locations (kinit) and rate of TAMRA-probe binding to locations without AF488-DNA molecules (kbkgnd) were obtained for each individual replicate. The mean initiation rate (Avg kinit) was then calculated for each set of experimental conditions.
†The two numbers given for each replicate are the number of rrnB P1 DNA molecules on which a TAMRA-probe binding lasting >5 s was observed and the total number of DNA molecules in that replicate.
-
‡ Fold-inhibition was determined by dividing the Avg kinit for the relevant RNAP (σ 70RNAP or σ70RNAP647) by the calculated Avg kinit in the presence of the specified SCF/ppGpp. For the σ70 RNAP647 alone condition, fold inhibition is calculated relative to unlabeled σ70RNAP.
Fit parameters for time to first RNAP binding (Figure 3E and Figure 3—figure supplement 3) and RNAP dwell time (Figure 3F) distributions on rrnB P1 DNA*.
| Experimental conditions | Time to first binding measurement | Dwell time measurement | ||
|---|---|---|---|---|
| kbkgnd (106 M−1s−1) | ka (106 M−1s−1) | Af | ||
| σ70RNAP647 (Figure 3E) | 0.037 ± 0.006 (N = 34) | 2.2 ± 0.6 (N = 227) | 0.95 ± 0.04 | a = 91 ± 4% τ1 = 2.6 ± 0.4 s τ2 = 29.7 ± 0.2 s (N = 2,576) |
| +10 nM GreBCy3B (Figure 3E) | 0.016 ± 0.002 (N = 19) | 2.0 ± 0.4 (N = 269) | 0.93 ± 0.06 | a = 91 ± 6% τ1 = 3.1 ± 0.7 s τ2 = 42.2 ± 0.4 s (N = 2,408) |
| +100 nM DksA (Figure 3E) | 0.0093 ± 0.0005 (N = 8) | 1.9 ± 0.5 (N = 187) | 0.89 ± 0.05 | a = 92 ± 7% τ1 = 2.5 ± 0.4 s τ2 = 29.9 ± 0.8 s (N = 1,381) |
| +10 nM GreBCy3B; σ70RNAP647· GreBCy3B subset† (Figure 3—figure supplement 3) | 0.016 ± 0.002 (N = 19) | 2.4 ± 0.7 (N = 124) | 0.94 ± 0.05 | a = 90 ± 5% τ1 = 3.4 ± 0.8 s τ2 = 44.6 ± 0.7 s (N = 1,053) |
| +10 nM GreBCy3B; σ70RNAP647 subset† (Figure 3—figure supplement 3) | 0.016 ± 0.002 (N = 19) | 1.8 ± 0.6 (N = 145) | 0.91 ± 0.08 | N.D. |
-
*Figure 3E and F each show curves from an individual experimental replicate; fit parameters here represent pooled data from all three replicates.
†Data in 10 nM GreBCy3B experiment were divided into two disjoint subsets of RNAP binding events σ70RNAP647 that arrived at DNA 1) without GreBCy3B and 2) as a σ70RNAP647•GreBCy3B complex (Figure 3—figure supplement 3).
Rate constants (±SE*) for kinetic scheme depicted in Figure 5.
| k1 or k1′ (M−1s−1)† | k-1 or k-1′ (s−1)‡ | k2 or k2′ (s−1)‡ | k-2 or k-2′ (s−1) ‡ | |
|---|---|---|---|---|
| σ70RNAP | 2.2 ± 0.6 × 106 | 0.48 ± 0.12 | 1.4 ± 0.6 × 10−2 | 2.4 ± 0.8 × 10−2 |
| σ70RNAP·GreB | 2.4 ± 0.7 × 106 | 0.30 ± 0.16 | 2.3 ± 0.7 × 10−2 | 1.7 ± 0.4 × 10−2 |
-
*Determined by bootstrapping (Friedman and Gelles, 2012).
†Identical to the corresponding ka values in Table 3.
-
‡Calculated as described (Friedman and Gelles, 2012) from the dwell time fit parameters reported in Table 3.
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Software | CoSMoS analysis software | https://github.com/gelles-brandeis/CoSMoS_Analysis | RRID:SCR_01689 |