A Y-chromosome shredding gene drive for controlling pest vertebrate populations
Figures
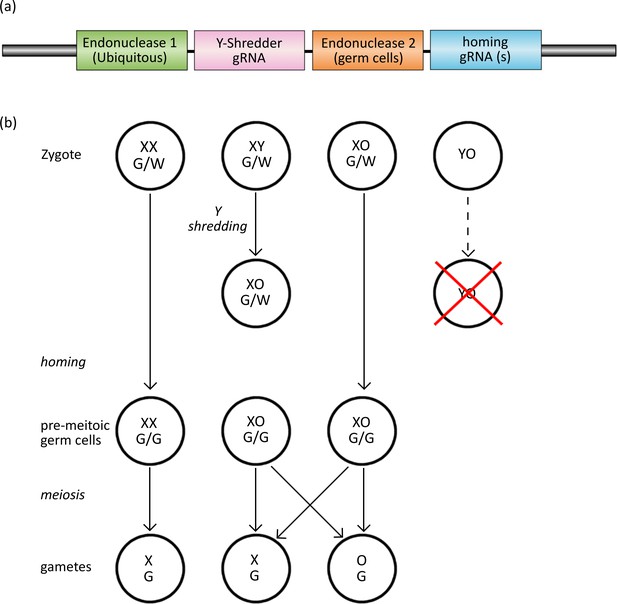
The Y-CHOPE drive.
(a) The proposed Y-shredder gene drive construct. Y-shredder activity is mediated by a ubiquitously expressed programmable endonuclease (Endonuclease 1; e.g. Cas12a) and a gRNA targeting a Y chromosome repeat sequence. Homing activity is controlled by an orthogonal endonuclease (Endonuclease 2; e.g. Cas9) with restricted expression in the premeiotic germ cells, in combination with one or more gRNAs targeting the integration site of the transgene. (b) Chart of zygotic development depicting timing of Y-shredding, homing and meiosis. YO embryos fail to develop as indicated by the red cross.
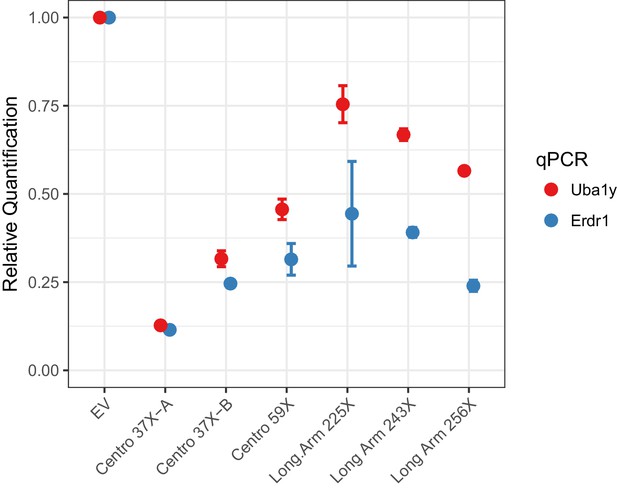
Empirical performance of the Y-shredding technology, using qPCR of genomic DNA to quantify Y-chromosome dosage.
Controls were transfected with empty vector (EV). Sox1 qPCR was used as the internal reference control. Data are presented as mean ±95% confidence intervals from n ≥ 3 biological replicates.
-
Figure 2—source data 1
Empirical performance of the Y-shredding technology.
- https://doi.org/10.7554/eLife.41873.003
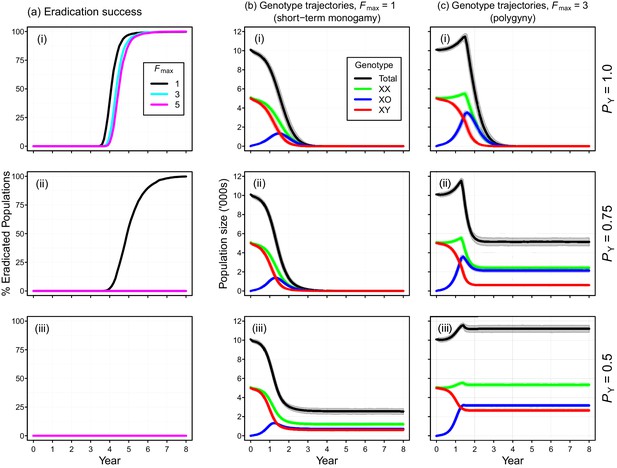
Performance of the Y-chromosome shredding gene drive for different Y-shredding efficiencies (PY), assuming resistant alleles cannot evolve.
(a) The percentage of eradicated mouse populations over time, and different levels of polygyny (Fmax). (b) Total population and allosome genotype trajectories (mean ±95% confidence intervals), assuming a short-term monogamous mating system for mice (Fmax = 1). (c) As for (b), but assuming a polygynous mating system (Fmax = 3). All results were based upon 1000 iterations per scenario, and assumed an initial population size of 10,000 mice that was inoculated with 100 gene-drive carrying XX females.
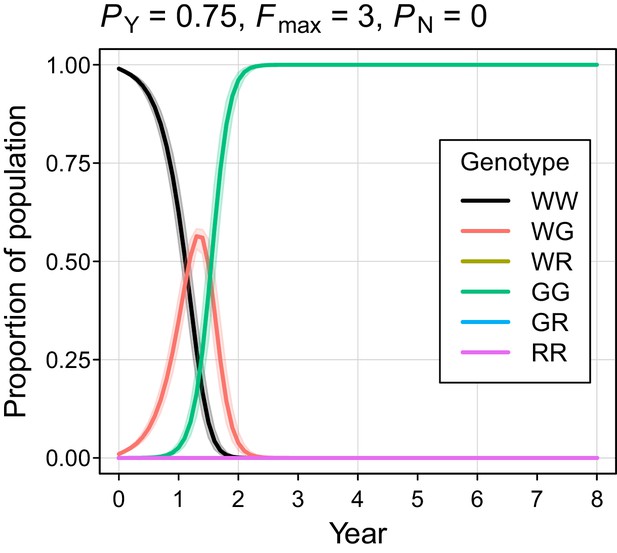
Autosome genotype trajectories (mean ±95% confidence intervals) for an example parameterization that resulted in permanent suppression (but not eradication) of the mouse population.
Abbreviations denoting different alleles are: W, wildtype; G, gene drive, R, resistant. (Note that resistant alleles cannot be created when PN = 0.).
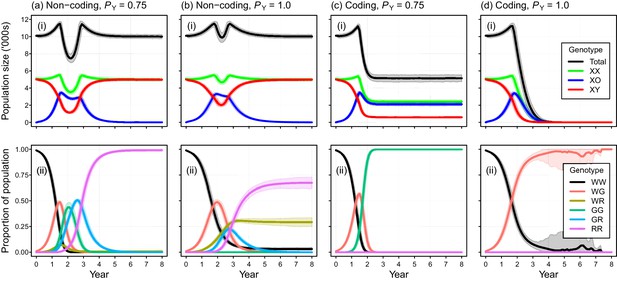
Simulated efficacy of Y-chromosome shredding gene drive when resistant alleles are created by non-homologous end-joining during homing.
Simulated trajectories (mean ±95% confidence intervals) are shown for two different Y-shredding efficiencies (PY), assuming the drive was positioned within a non-coding (a, b) or coding region (c, d). Total population and allosome genotype trajectories (i), and the proportion of different autosome genotypes (ii), are shown for each scenario. In the bottom right panel, all simulated populations were eradicated within 8 years. All simulations assumed an initial population size of 10,000 mice, inoculation with 100 gene-drive carrying XX females, polygynous breeding (Fmax = 3), and substantial opportunity for the creation of resistant alleles through non-homologous end-joining (PN = 0.1). In (c,d), the gene drive is assumed to be positioned within a haploinsufficient critical gene, such that resistance alleles cause early embryonic lethality (Pnf = 1). Abbreviations denoting different autosome alleles are: W, wildtype; G, gene drive, R, resistant.
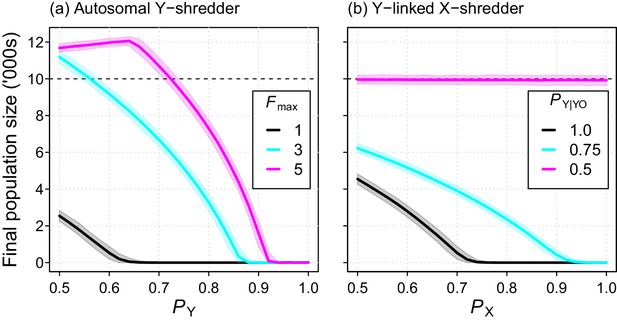
Final population size (mean ±95% confidence intervals) after 10 year simulations for two different chromosome-shredding gene-drive strategies.
(a) An autosomally integrated Y-CHOPE drive. Final population size is shown as a function of the Y-shredding efficiency (PY) and the level of polygyny assumed for mice (Fmax), assuming placement of the gene drive within the coding region of a haploinsufficient developmental gene, and the baseline parameterisation (see Table 1). (b) An X-shredding Y-drive. Results are shown as a function of the X-shredding efficiency (PX), and the probability of offspring inheriting a Y-chromosome from male gene-drive carriers that is germline YO (PY|YO). The initial population size of 10,000 mice is indicated (horizontal dashed line).
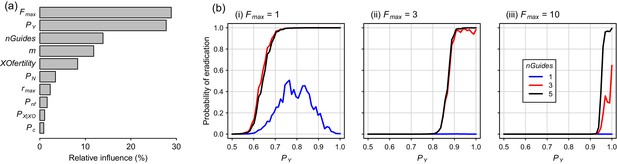
Boosted regression tree (BRT) summary of the sensitivity analysis for the Y-CHOPE gene drive.
(a) Relative influence of the ten simulation parameters included in the sensitivity analysis (parameter abbreviations are provided in Table 1). (b) Predicted probabilities of eradication derived from the BRT model for three different levels of polygyny ranging from a maximum of one to 10 females mates per male (i-iii). These partial dependency plots assume the following seven less influential parameters were set as follows: m = 6, XOfertility = 0.6, Pn = 0.1, rmax = 7.76, Pnf = 0.66, nGuides = 3, PX|XO = 0.66, and PC = 0.95.
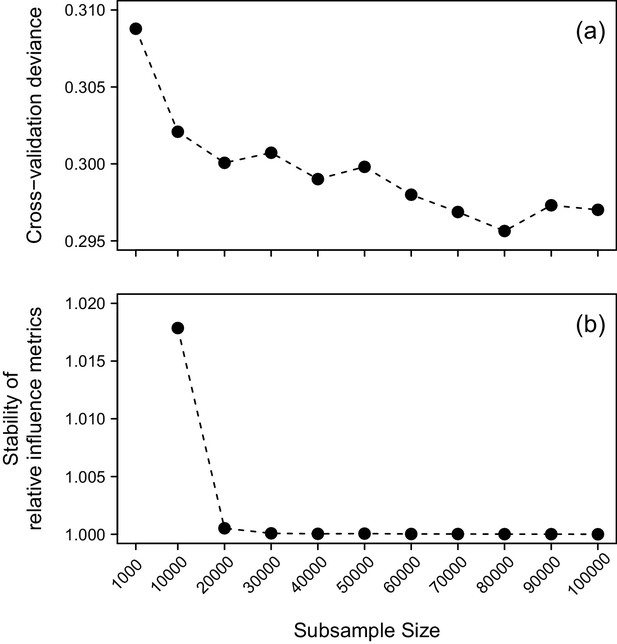
The cross-validation deviance (a) and stability of sensitivity measures (b) derived from BRT emulators as the number of parameter samples used for sensitivity analysis is increased.
Results are shown for emulation of the probability of simulated population eradication within 10 years. Note that in panel (b), a stability of 1 indicates perfect agreement between the relative influence metrics obtained from a given sample size and the next smallest sample size tested.
Tables
Details of the baseline parameterization of the individual-based model, and the parameter ranges tested through sensitivity analysis using Latin hypercube sampling.
https://doi.org/10.7554/eLife.41873.011| Parameter | Baseline | Sensitivity Analysis |
|---|---|---|
| Probability of Cas9 cutting (PC) | 0.95 | U(0.7, 1) |
| Probability of NHEJ (PN) | 0, 0.1 | U(0, 0.5) |
| Number of guide RNAs (nGuides) | 3 | U(1, 5)* |
| Probability of mutation causing non-functionality (Pnf) | 1 | U(0.66, 1) |
| Y-shredding efficiency (PY) | 0.5–1 | U(0.5, 1) |
| Fertility multiplier for XO females (XOfertility) | 0.6 | U(0, 1) |
| X:O bias in inheritance (PX|XO) | 0.66 | U(0.5, 1) |
| Maximum number of female mates per male (Fmax) | 1, 3 | U(1, 10)* |
| Mean litter size (m) | 6 | U(2, 10) |
| Maximum annual population growth rate (rmax) | 7.97 | U(6, 9) |
-
*For the sensitivity analysis, these parameters were sampled from discrete uniform distributions.
Additional files
-
Source code 1
R code to simulate Y- and X-shredding gene drives in mice.
- https://doi.org/10.7554/eLife.41873.012
-
Supplementary file 1
Details of the gRNA sequences (a) and qPCR primers (b) used for the empirical study of Y-shredding efficiency.
- https://doi.org/10.7554/eLife.41873.013
-
Transparent reporting form
- https://doi.org/10.7554/eLife.41873.014





