CtIP forms a tetrameric dumbbell-shaped particle which bridges complex DNA end structures for double-strand break repair
Figures
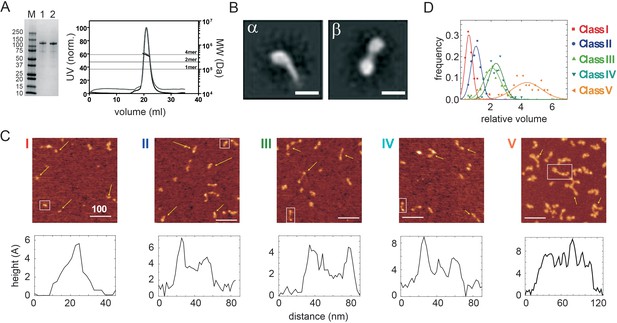
Wild type CtIP is a tetrameric protein that forms a dumbbell-shaped particle.
(A) SDS-PAGE and SEC-MALS analysis of wild type CtIP as prepared (lane 1, black trace) and following dephosphorylation post-purification (lane 2, grey trace). Horizontal lines on the SEC-MALS graph show the expected molecular weights for monomeric, dimeric and tetrameric CtIP species. (B) 2D class averages for wild type CtIP observed by negative stain EM. Scale bar = 20 nm. Further examples are shown in the Figure Supplements. (C) AFM imaging of purified wild type CtIP protein. Five classes of particles (I–V) were detected based on morphology and volume analysis (see text for details). Scale bar = 100 nm. Examples are marked with yellow arrows and further images at higher magnification are shown in the Figure Supplements. Representative height profiles are shown which are derived from the boxed particle. (D) Relative volume distributions for the five AFM particle classes shown in C. The volume is determined using a technique in which a fiducial DNA marker is used as a reference (Fuentes-Perez et al., 2012). There exists a linear relationship between the relative volume and absolute molecular weight for a wide range of proteins. The molecular weight of the large class V particle imaged here is 380 kDa (i.e. close to the value expected for a tetramer) based on a mean relative volume of 4.4x the marker.
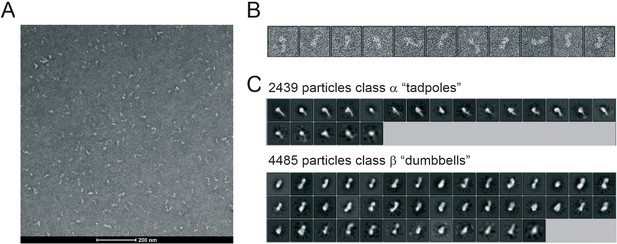
Negative stain electron microscopy.
(A) An example of a micrograph obtained with negative staining of CtIP at 42,000 X magnification. (B) Examples of individual CtIP particles. (C) Further examples of 2D classes for CtIP particles classified as class α or β as indicated.
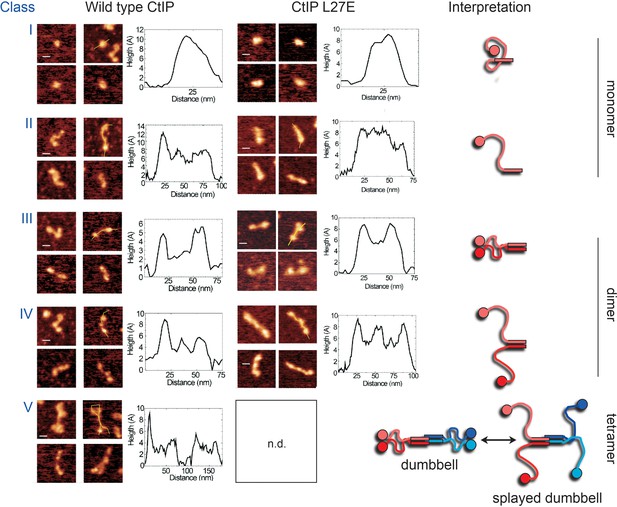
Further examples and structural interpretation of AFM particle classes.
Class V particles are thought to represent tetrameric CtIP (see main text and Figure 1D for justification), consisting of a dimer-of-dimers arrangement (blue and red) of parallel coiled coils (zig-zags) meeting at the centre with polar globular domains (circles) that either appear together or in a splayed arrangement. Based on their morphology and relative volume, the cartoons below show speculative interpretations of the other particle classes observed in AFM experiments.
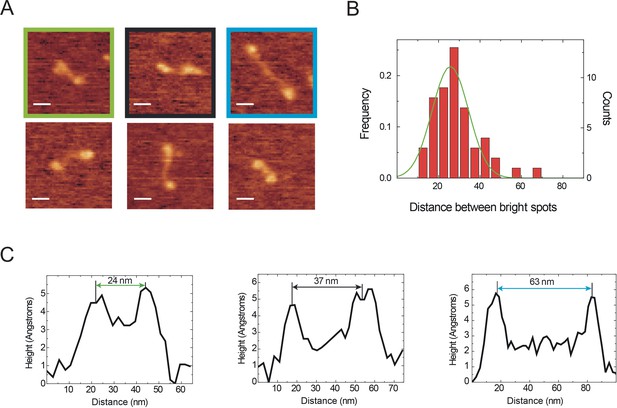
The rod between the globular domains is flexible and variable in length.
(A) Examples of CtIP particles displaying significantly different intervening lengths between spots. (B) The distribution of the distances between bright spots in CtIP particles is centred at ~ 30 nM but is very broad, ranging from < 20 to>60 nm. (C) Examples of distance measurements extracted from height profiles for three of the particles shown in A, as indicated by the colouring scheme.

CtIP purified from insect cells is hyper-phosphorylated.
Human wild type CtIP was expressed in insect cells, purified in the presence of dephosphorylase inhibitors, and analysed for post-translational modifications by Orbitrap LC-MS/MS mass spectrometry. (A) Coverage of the CtIP polypeptide (91% total) is indicated at different confidence thresholds; less than 1% false discovery rate (green) or less than 5% false discovery rate (yellow). (B) Predicted phosphorylation sites are indicated by the letter P above the CtIP polypeptide at the confidence levels indicated.
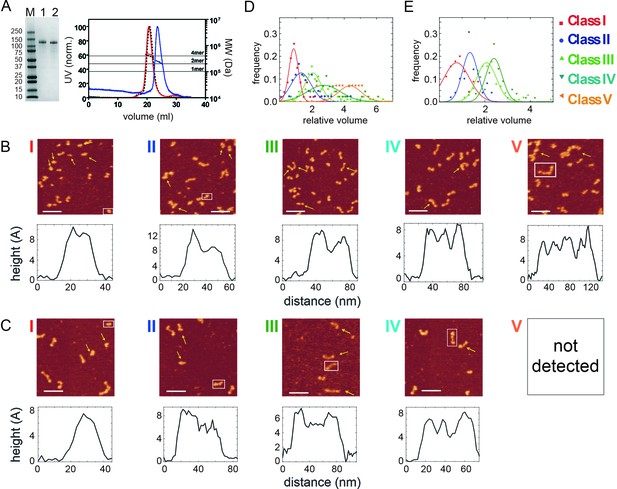
Mutation in the N-terminal coiled-coil domain prevents CtIP tetramerisation.
(A) SDS-PAGE and SEC-MALS analysis of CtIP L27E (lane 1, blue trace) and CtIP R839A (lane 2, red trace). Data for wild type CtIP is also shown for comparison (dotted black lines). (B) AFM imaging of purified CtIP R839A protein. Five classes of particles (I–V) were detected based on morphology and volume analysis. Representative height profiles are shown below the images which are derived from the boxed particle. (C) AFM imaging of purified CtIP L27E protein. Four classes of particles (I–IV) were detected based on morphology and volume analysis. Representative height profiles are shown which are derived from the boxed particle. (D) Relative volume distributions for the AFM particle classes detected in the CtIP R839A and (E) CtIP L27E preparations. Note that the class V particle is completely absent from the CtIP L27E preparation.
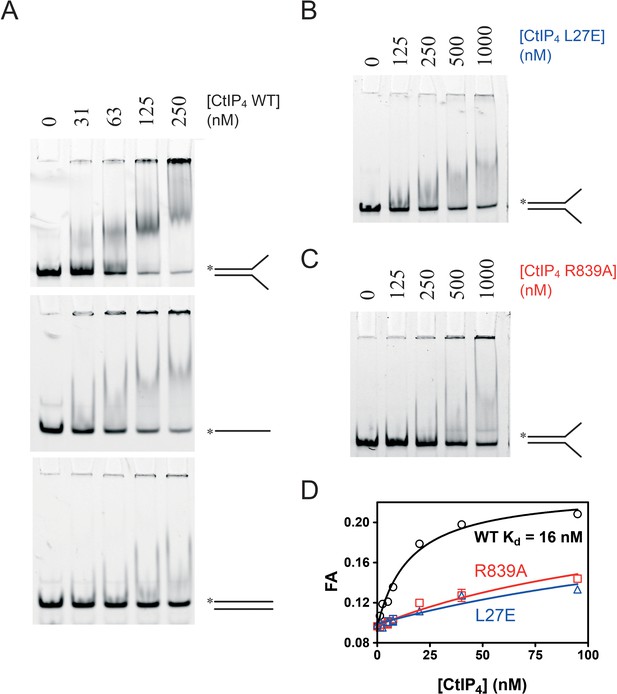
CtIP binds preferentially to ss-dsDNA Y-junctions in a manner dependent on both the N-terminal tetramerisation and C-terminal DNA binding motifs.
(A) Electrophoretic mobility shift assay. Radiolabelled DNA molecules with the different structures (indicated) were incubated with increasing concentrations of CtIP tetramer and run on non-denaturing PAGE gels as described in the Materials and methods section. (B) Binding of the fork DNA substrate by the L27E mutant protein. (C) Binding of the fork DNA substrate by the R839A mutant protein. (D) Quantitative comparison of wild type and mutant CtIP proteins binding to a HEX-labelled DNA fork structure using fluorescence anisotropy.
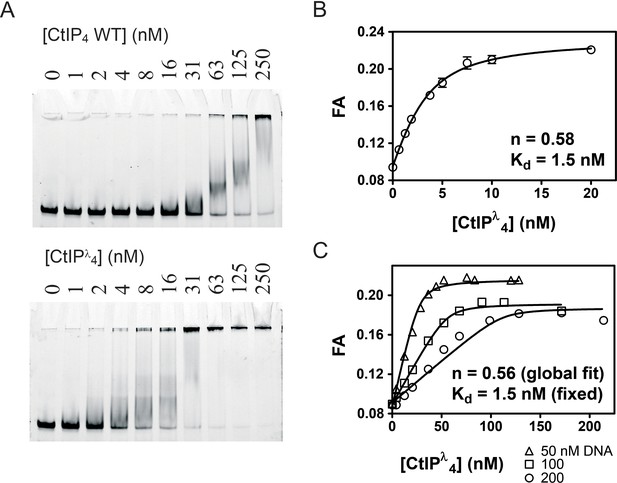
Dephosphorylation of CtIP potentiates DNA binding and facilitates determination of the DNA binding stoichiometry.
(A) EMSA assays comparing the binding of wild type CtIP as prepared and following treatment with λ phosphatase (denoted CtIPλ) as described in the Materials and methods. (B) Fluorescence anisotropy assay monitoring the binding of CtIP to 5 nM HEX-labelled fork DNA. The data were fit to the tight binding equation. (C) Assay as in B, but using 50, 100 and 200 nM HEX-labelled fork DNA as indicated. The data were fit globally to the Equation 5 to determine the binding stoichiometry (n) which measures the number of CtIP tetramers bound to each DNA fork.
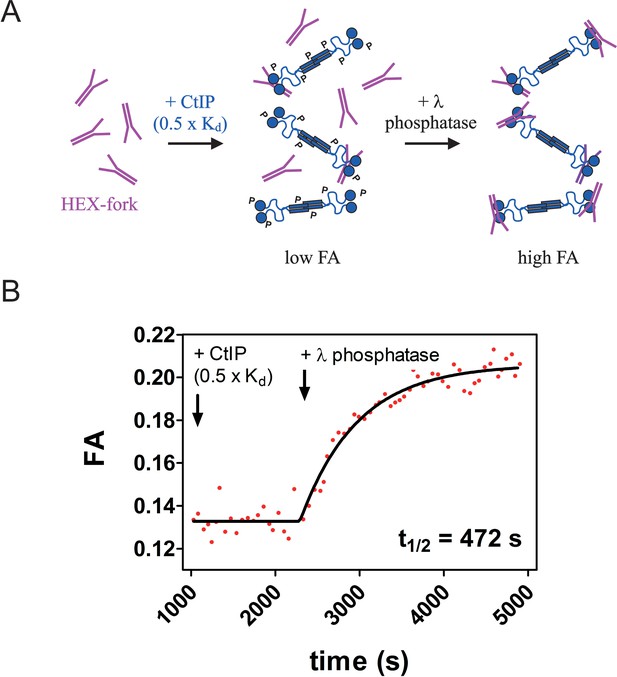
Real time measurement of DNA binding potentiation upon treatment of CtIP with λ phosphatase.
(A) Principle of the assay. CtIP (blue) was added to 5 nM HEX-labelled fork DNA (purple) at a concentration equal to ~ 0.5 x Kd resulting in a low fluorescence anisotropy reading (~0.13). A small volume of λ phosphatase was added to the mixture, resulting in increased anisotropy due to an improved binding affinity. Note that CtIP as prepared in insect cells is hyperphosphorylated and the letter P is intended to represent a phosphorylated amino acid. (B) Representative trace showing potentiation of DNA binding by dephosphorylation of CtIP. The data were well-fitted to a single exponential to yield the half-life shown.
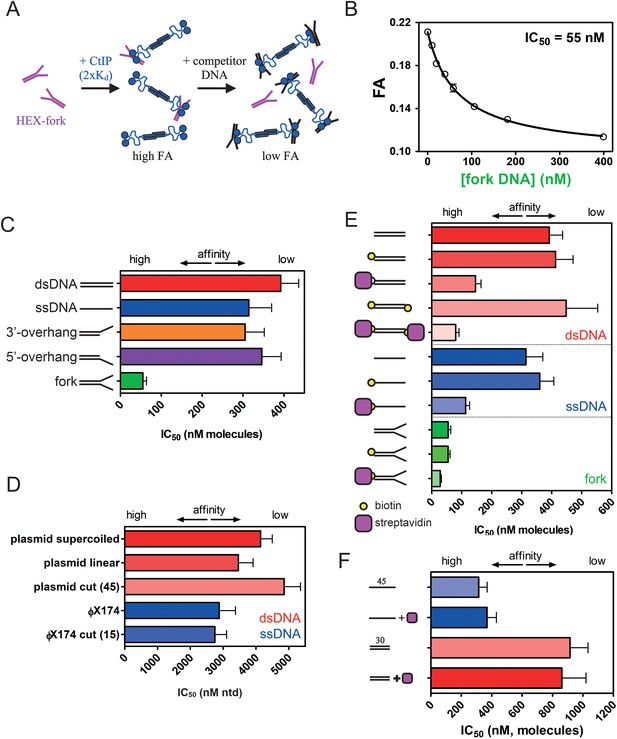
CtIP binding to DNA is stabilised by DNA Y-junctions and DNA end blocks.
(A) Principle of the competition DNA unbinding assay monitored by fluorescence anisotropy. HEX-labelled and unlabelled DNA fork molecules are shown in purple and black respectively. CtIP is in blue. (B) Increasing concentrations of unlabelled fork DNA were titrated into a pre-formed complex between CtIP and a HEX-labelled DNA. The data were fit to a hyperbolic decay curve to yield an IC50 value for the inhibition of binding. (C) Comparison of the IC50 values for different DNA structures. Full details of the DNA sequences and structures used in these experiments can be found in the Supplementary Information. (D) Comparison of IC50 values for single- and double-stranded DNA competitors with different topologies and containing different densities of DNA breaks (bracketed number). Note that the IC50 value is quoted in terms of the total nucleotide concentration. (E) Comparison of the IC50 values for duplex (red), single-stranded (blue) and forked (green) DNA substrates with and without the ends blocked by biotin:streptavidin. (F) Comparison of IC50 values for single- or double-stranded DNA substrates which lack biotin, determined both in the presence and absence of streptavidin. Details for the substrate construction and data for a wider range of substrates are available in the Supplementary Information.
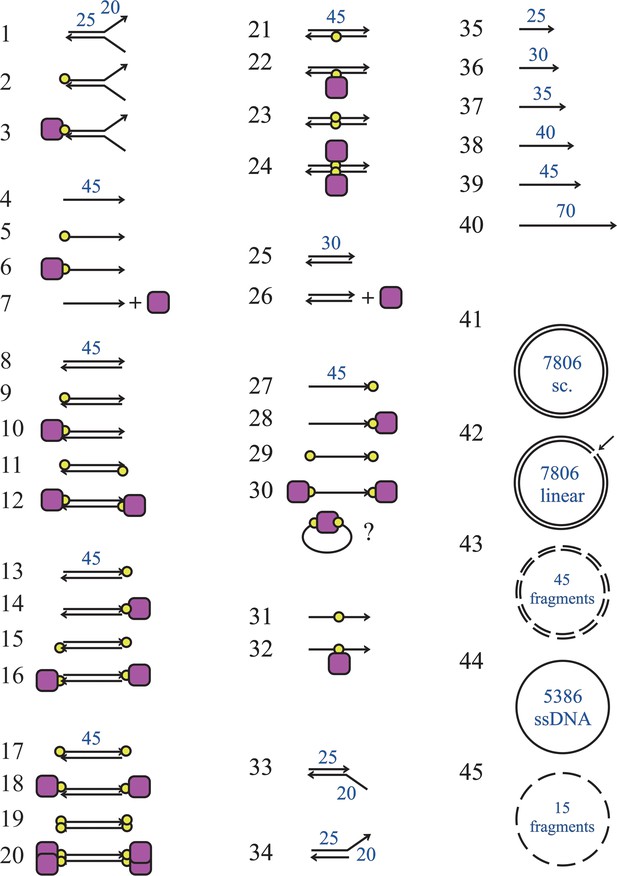
Structures of competitor DNA molecules used in this study.
The substrates are numbered as in Supplementary file 1. The arrows represent the 3′-ends of DNA, a yellow circle indicates biotin and the purple square is streptavidin. The blue numbers represent the substrate lengths (in bp or bases) with the exception of substrates 43 and 45, where they represent the number of fragments of the circular DNA they originated from. Substrate 30 may form adopt a circularised structure due to the low persistence length of single-stranded DNA as indicated. For substrate 41, sc indicates that the substrate is supercoiled.
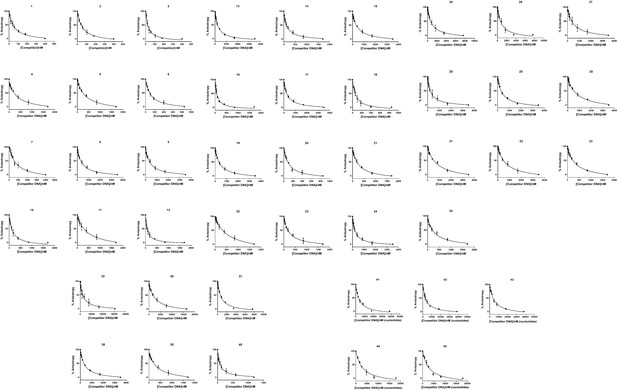
Raw data for all IC50 measurements.
Numbers above each graph indicate the competitor DNA used as per Supplementary file 1.
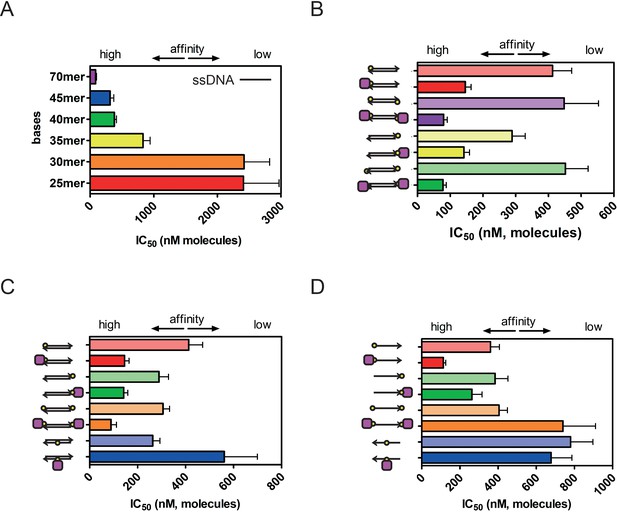
IC50 values for CtIP binding measured for a wider range of competitor DNA molecules.
(A) Comparison of IC50 values for ssDNA molecules of different length as indicated (B) Comparison of how streptavidin binding affects IC50 values for duplex DNA molecules containing different numbers and attachment polarities of the biotin moieties (C) Comparison of how streptavidin binding affects IC50 values for duplex DNA molecules with biotin in different positions along the duplex (D) Comparison of how streptavidin binding affects IC50 values for single-stranded DNA molecules with biotin in different positions. In all figures the arrow represents the 3’-end of the DNA molecule, the yellow circle is biotin, and the purple square is streptavidin.
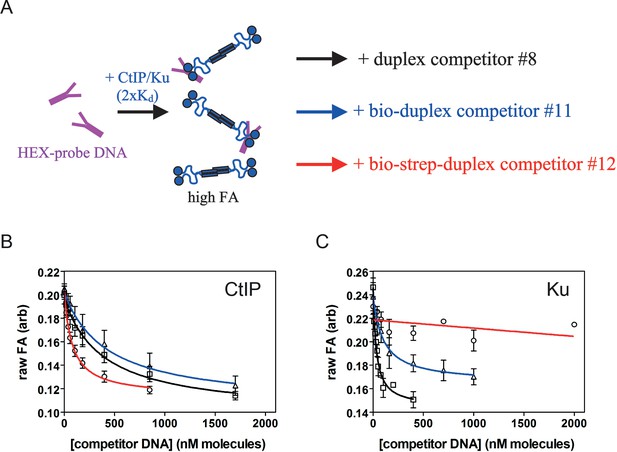
CtIP and Ku display distinctive DNA binding modes.
(A) Principle of experiment to compare the effects of DNA end blocks on DNA binding by CtIP and Ku. CtIP and Ku are pre-bound at 2xKd to HEX-labelled probe DNA (a fork for CtIP and an oligoduplex for Ku). Different competitor DNAs are then added to test their efficacy as competitors including an oligoduplex, a biotinylated oligoduplex, or a biotinylated oligoduplex bound to streptavidin. The numbering is as in the cartoons shown in Figure 5—figure supplement 1. (B) Competition unbinding curves for CtIP using the three competitor DNA molecules. Note that the presence of streptavidin improves binding to the oligoduplex competitor. (C) Equivalent data for Ku. Note that, in direct contrast to the behaviour for CtIP, biotin moieties reduce the efficacy of DNA binding and the additional presence of streptavidin almost completely eliminates the ability of the duplex DNA to compete.
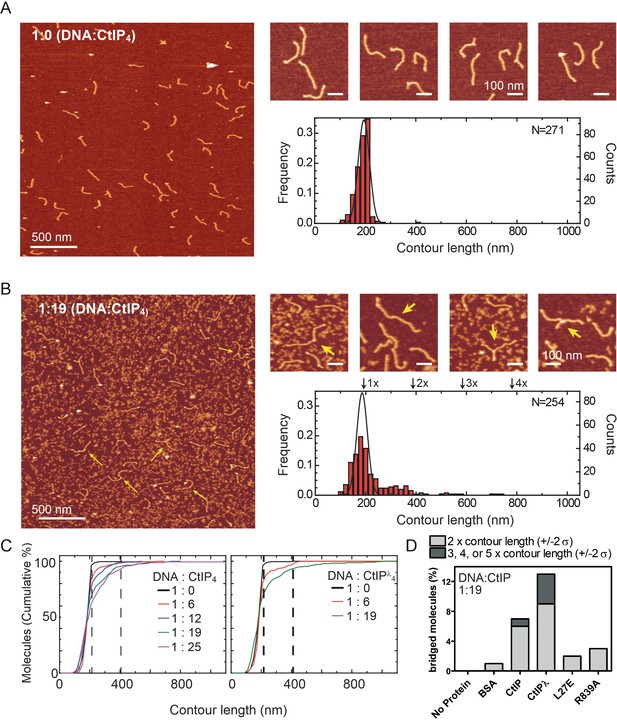
CtIP promotes intermolecular DNA bridging.
(A) Representative AFM images of forked DNA substrates in the absence of CtIP. The contour length histogram shows a single gaussian peak centred on a value equivalent to a single contour length. (B) Representative AFM images of forked DNA substrates in the presence of CtIP. The contour length histogram shows multiples peaks for single-, double-, triple and even quadruple contour length peaks (indicated). The solid line shows the gaussian fit for data collected in the absence of CtIP for comparison. Note that the size of the single-contour peak is substantially reduced. The zoomed-in panels show examples of bridged DNA molecules (yellow arrows). (C) Cumulative frequency of DNA molecules longer than a given contour length, highlighting the CtIP-dependent increase in 2mer and 3mer DNA substrates. The vertical dashed lines are equivalent to theoretical single- and double- contour lengths. (D) Histogram showing percentage of bridged molecules for wild type, dephosphorylated and mutant CtIP proteins at a single fixed CtIP concentration. The light grey data shows the percentage of molecules (within error) at 2x contour length, whereas the dark grey data is the sum of the percentage of molecules at 3x, 4x and 5x contour lengths.
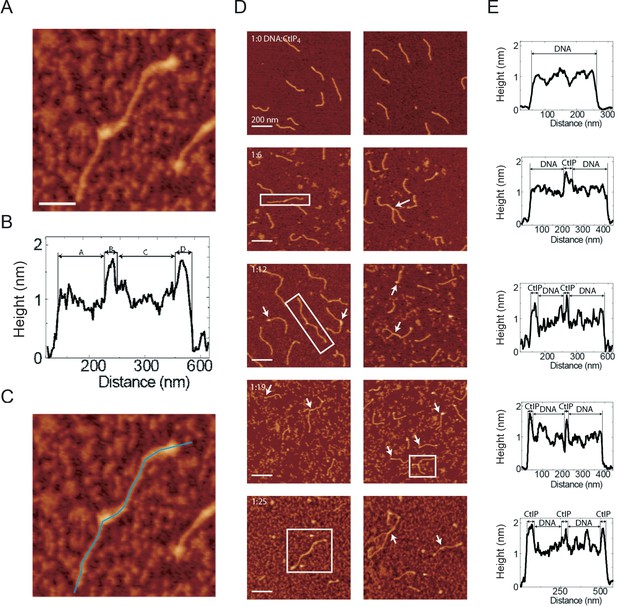
Evidence for CtIP bound to DNA at bridging interfaces.
(A) An example of a bridged DNA molecule bound to CtIP obtained at a 1:19 DNA:CtIP4 ratio (B) Height profile along the molecule of interest. Section A is interpreted as DNA (188 nm contour length), section B as DNA bound CtIP (32 nm), Section C as a second DNA molecule (222 nm), and section D as CtIP (44 nm). (C) The same image as above showing the tracing that was used to create the height profile. (D) Several further examples of bridged DNA molecules with evidence for CtIP binding at their interface, obtained at a variety of different DNA:CtIP4 ratios as indicated. (E) Height profile trace for the boxed molecule shown in D.
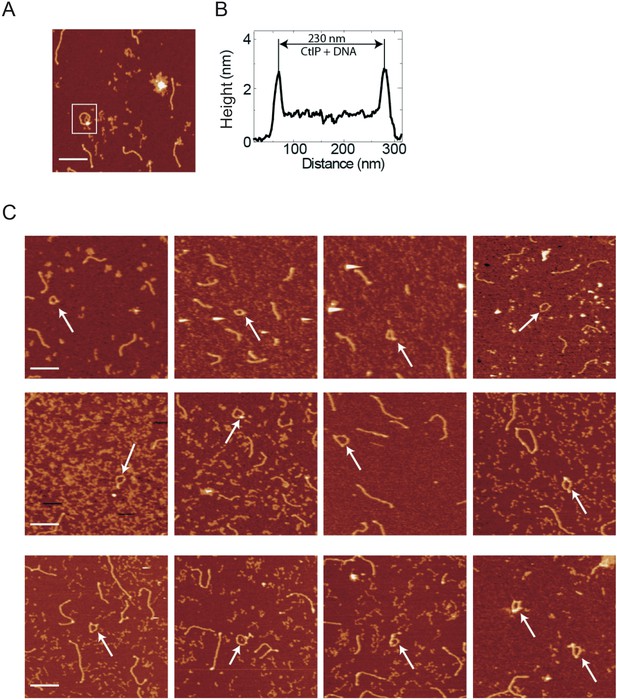
CtIP promotes intramolecular DNA bridging.
(A) Example of an AFM image showing an internally-bridged (circularised) DNA molecule. (B) Height profile for the boxed molecule shown in A reveals additional height associated with the circularised, single contour length, DNA molecule which we interpret as CtIP. (C) Several further examples of DNA molecules with evidence for internal bridging, highlighted with white arrows.
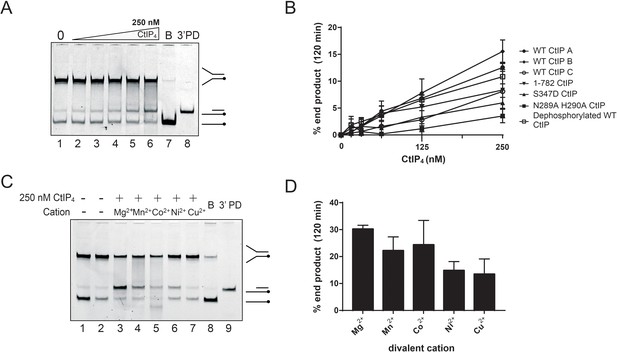
Assessment of nuclease activity in CtIP preparations.
It is currently controversial as to whether purified CtIP and its orthologues possess intrinsic nuclease activity (Andres and Williams, 2017). Therefore, to test our CtIP preparations for nuclease activity we incubated CtIP at high concentrations with a forked DNA substrate in the presence of various divalent cations and for extended times. (A) Representative nuclease assay for a wild type CtIP preparation (concentrations as indicated in (B)) incubated with 5 nM 5′-Cy5-labelled forked DNA for two hours. (B) Quantified nuclease assays using several different wild type CtIP preparations (labelled A-C) and different CtIP mutants (as indicated) titrated against 5 nM forked DNA. (C) Representative nuclease assay using 250 nM wild type CtIP against 5 nM forked DNA in the presence of different divalent cations. (D) Quantified nuclease assay using 250 nM wild type CtIP against 5 nM forked DNA in the presence of different divalent cations. Nuclease activity was observed which was consistent with cleavage of the 5′-terminated ssDNA flap as reported previously. This nuclease activity may explain the small proportion of less than single contour length substrates observed in our AFM bridging experiments. However, at the highest concentrations tested (which are >> Kd for the binding interaction with the DNA) the half-life of the DNA fork substrate was on the order of several hours. The specific nuclease activity varied between wild type preparations and did not correlate with the ability of CtIP to bind DNA. For example, the C-terminal deletion mutant CtIP1-782 does not bind DNA (data not shown) while dephosphorylated CtIP has ~ 10 fold greater affinity for DNA than wild type (Figure 4, main text), but both preparations retain apparent nuclease activity comparable to wild type. Mutations designed to inactivate a putative nuclease active site (N289A and H290A) did reduce the observed nuclease activity in the preparation as reported previously, but a mutation thought to result in hyperactivated nuclease activity (S347D) was less active than any of our wild type preparations (Makharashvili et al., 2014). The dependence of the observed activity on divalent cation identity was qualitatively different for our preparation compared to that reported previously, with Mg2+ being the marginally favoured co-factor as opposed to a substantial preference for Mn2+ (Makharashvili et al., 2014). Taken together, these observations suggest to us that the nuclease activity in the preparation most likely belongs to a contaminant or co-purified protein rather than to the CtIP polypeptide. However, we are unable to exclude the possibility that CtIP (either as prepared or dephosphorylated) possesses a very weak and/or tightly-regulated nuclease activity.
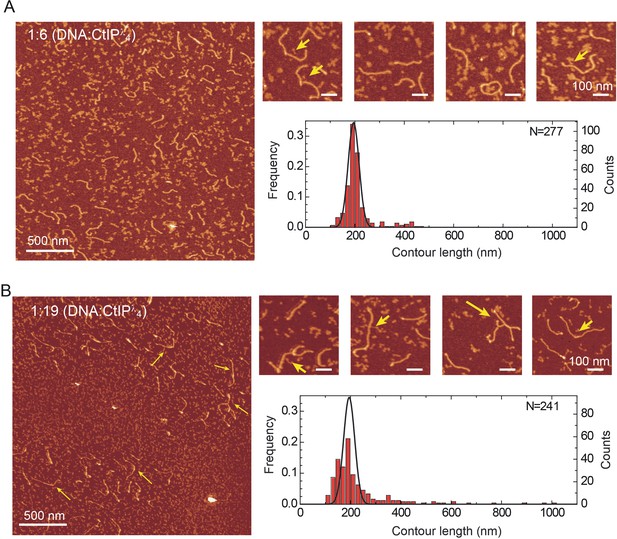
Dephosphorylated CtIP promotes intermolecular DNA bridging.
(A) Representative AFM images of forked DNA substrates in the absence of dephosphorylated CtIP (CtIPλ). The contour length histogram shows a single gaussian peak centred on a value equivalent to a single contour length. (B) Representative AFM images of forked DNA substrates in the presence of CtIPλ. The contour length histogram shows multiples peaks for single-, double-, triple and even quadruple contour length peaks. The solid line shows the gaussian fit for data collected in the absence of CtIPλ for comparison. Note that the size of the single-contour peak is substantially reduced.
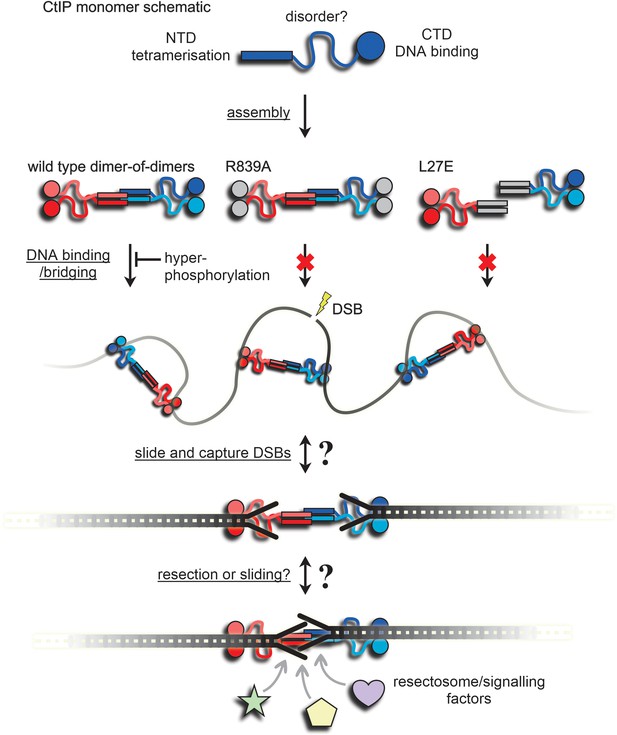
A speculative model for binding and bridging of broken DNA ends by CtIP.
The CtIP monomer comprises at least three functional regions; an N-terminal tetramerization domain, a central region of predicted disorder, and a C-terminal DNA binding domain. This assembles to form a dumbbell-shaped tetramer (evidence in Figure 1). Wild type CtIP binds tightly to two DNA molecules via C-terminal RHR motifs, but this is disfavoured by hyperphosphorylation (Figures 3 and 4). Mutation within this motif (R839A) prevents DNA binding and bridging (Figures 3 and 6). Mutation within the N-terminal coiled-coil domains prevents assembly into a tetramer, DNA binding and DNA bridging (Figures 2, 3 and 6). The grey coloured domains are intended to indicate their inactivation by mutation. The inability of L27E to bind to DNA implies that mutations in the coiled coils have effects on the distal DNA binding domains (see main text for discussion). Wild type CtIP binds DNA without preference for ends per se (i.e. internally) and bridges DNA segments either intra- and inter-molecularly (Figures 5 and 6, and the Figure Supplements). Based on the preferential binding to DNA with forked or blocked ends (Figure 5), we speculate that CtIP may then slide and capture DNA ends possessing forked structures or nucleoprotein conjugates, because they prevent dissociation from the free ends (see main text for discussion). This arrangement might then allow the dimeric sub-assemblies within CtIP to slide away from each DNA end to which they are bound, facilitating resection and/or access to the DSBs for resection and signalling factors while maintaining their pairing. This proposal will be the subject of future work.
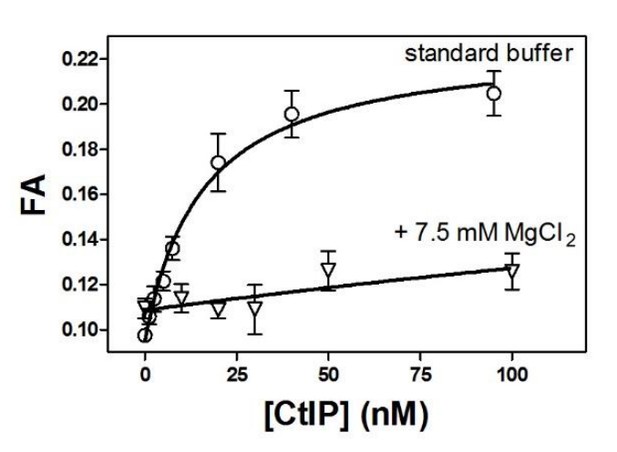
Moderately elevated MgCl2 concentrations dramatically reduce the affinity of CtIP for DNA.
https://doi.org/10.7554/eLife.42129.025Additional files
-
Supplementary file 1
Comprehensive data for DNA competition assay and details of substrate construction.
Supplementary Table 1: IC50 values for competitor DNA molecules used in DNA unbinding assays. The reported error is the error associated with the fit to a hyperbolic unbinding curve as shown in Figure 5—figure supplement 2. Supplementary Table 2: Assembly/Source of DNA substrates. Small DNA substrates were prepared by annealing different combinations of short oligonucleotides (A-T). The sequences for the oligonucleotides are presented in Supplementary Tables 3. Supplementary Table 3: Sequences of oligonucleotides used to assemble competitor DNA molecules.
- https://doi.org/10.7554/eLife.42129.022
-
Transparent reporting form
- https://doi.org/10.7554/eLife.42129.023





