Muscle-specific stress fibers give rise to sarcomeres in cardiomyocytes
Figures
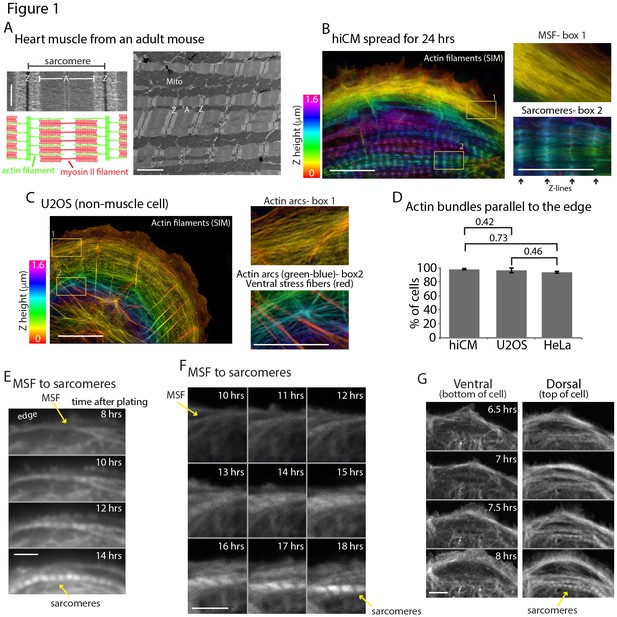
Sarcomeres arise directly from Muscle Stress Fiber (MSF) precursors.
(A) Electron microscopy (EM) schematic of a cardiac sarcomere from adult mouse. Electron dense regions on the borders of a sarcomere are Z-discs (Z), while the core of the sarcomere is composed of thin actin filaments and thick myosin II filaments (A). Multiple sarcomeres aligned adjacently form a myofibril (lower mag EM, right). (B) hiCM allowed to spread for 24 hr following plating and imaged with SIM. hiCM has been stained for actin and color coding is a representation of height (Z plane) within the cell following 3D imaging (Z-height, left). Notice the clear stress fiber and sarcomere-like actin organization at the front and rear of the cell in box 1 and 2, respectively. (C) Spread U2OS cell color coded for Z as in Figure 1B, displaying prominent actin arc stress fibers behind leading edge of cell and imaged with SIM. Box 1 shows actin arcs just behind the leading edge of cell, while box 2 shows actin arcs on dorsal surface in cell body (green and blue colored actin), while ventral stress fibers (red colored actin) are on bottom surface of cell. (D) Percentage of hiCMs, U2OS, and HeLa cells with actin arc stress fibers. hiCMs; 1372 cells over three experiments. U2OS; 37 cells over four experiments. HeLa; 186 cells over four experiments. (E) Wide-field time lapse of hiCM transfected with Lifeact-mEmerald to visualize actin. MSF at front of hiCM undergoes retrograde flow and acquires sarcomeres (yellow arrows). (F) Laser-scanning confocal microscopy of hiCM expressing Lifeact-mApple showing MSF to sarcomere transition. hiCM lacks sarcomeres at first time point, and MSF at edge of cell undergoes retrograde flow and acquires sarcomeres (yellow arrows). (G) 3D laser-scanning confocal microscopy of hiCM expressing Lifeact-mApple forming sarcomeres. Note how ventral surface (left montage) contains no sarcomere structures, while sarcomere assembly occurs on the dorsal surface of cell (right montage). Scale Bars; (A) 500 nm high mag (left), 2 µm low mag (right); (B) 10 µm low mag, 5 µm high mag insets; (C) 10 µm low mag, 5 µm high mag insets; (E), (F), (G), 10 µm. P-values denoted in graphs.
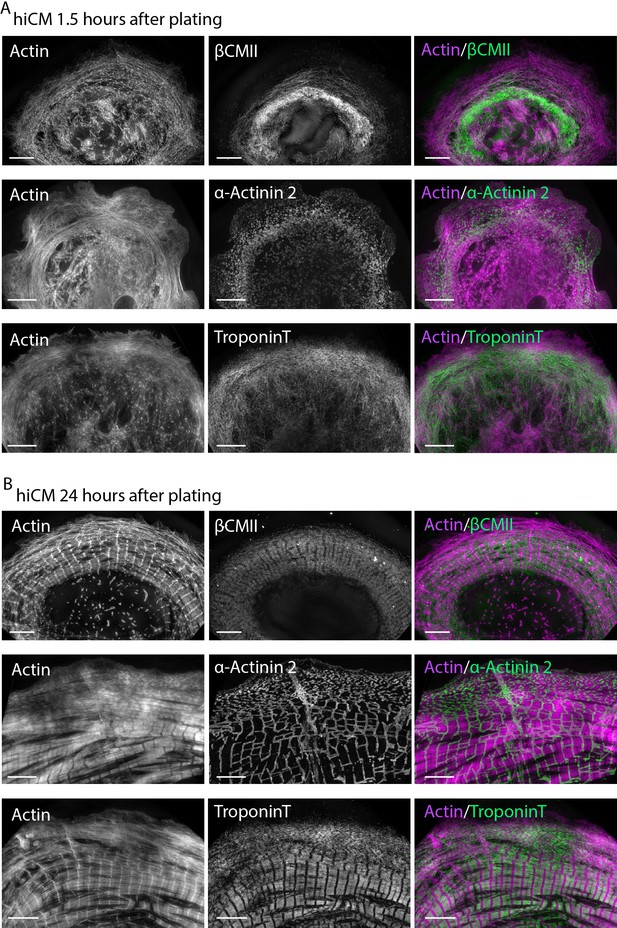
hiCMs do not contain sarcomeres at early time points post plating.
(A) hiCMs allowed to spread for 1.5 hr, fixed, and stained for actin and βCMII (top), actin and α-actinin 2 (middle), actin and TroponinT (bottom), and imaged with SIM. Note lack of sarcomere structure in all sarcomere markers shown. (B) hiCMs allowed to spread for 24 hr, fixed, and stained for actin and βCMII (top), actin and α-actinin 2 (middle), actin and TroponinT (bottom), and imaged with SIM. Note how opposed to 1.5 hr, hiCMs spread for 24 hr contain robust sarcomeres in all markers shown. Scale bars, 5 µm.
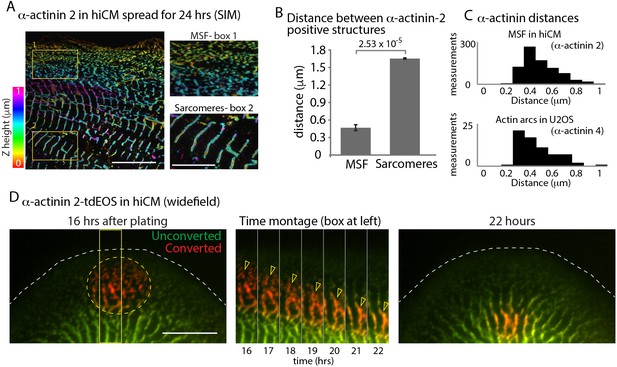
α-Actinin 2 spacing and dynamics in hiCMs.
(A) Color coded representation for Z-height of endogenous α-actinin 2 in MSFs (box 1) and sarcomeres (box 2) of hiCM imaged with SIM. Note difference in structure and spacing of α-actinin 2 in MSFs (box 1) and sarcomeres (box 2) (B) Distance between α-actinin 2 structures in MSFs and sarcomeres. MSFs; 14 cells, three experiments, 827 measurements, Sarcomeres; 15 cells, three experiments, 527 measurements. Distance between structures increases as MSFs transition to sarcomeres. (C) Histogram depicting distribution of distances between α-actinin 2 structures in MSFs in hiCMs (top) and α-actinin 4 found in actin arcs of non-muscle cells (bottom). Distribution is similar between cell types. (D) Wide-field montage of photoconversion of α-actinin 2-tdEOS in hiCM. MSFs at leading edge of the cell were photoconverted (green to red) and imaged over time. Montage (middle) depicts α-actinin 2-tdEOS puncta of MSFs (hollow yellow arrow heads) transition into sarcomere structures (middle, right). Scale Bars; (A) 10 µm low mag, 5 µm high mag insets. (B), 10 µM. P-values denoted in graphs.
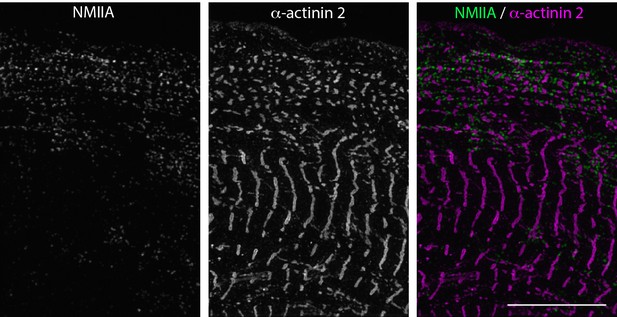
SIM of NMIIA and α-actinin 2localizations in hiCM
NMIIA localizes to MSFs but is excluded from sarcomeres (see Figure 7A and B). NMIIA is found between small α-actinin 2 puncta in MSFs at edge of hiCM (alternating green and magenta in right image), and is excluded from large α-actinin 2 Z-lines in hiCM body. Scale Bar: 10 μm.
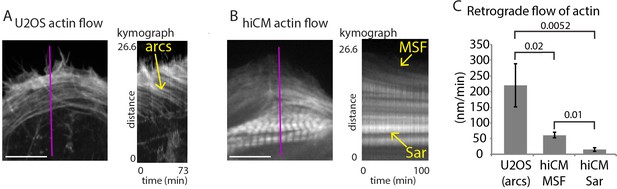
Retrograde flow of actin in non-muscle cells and hiCMs.
(A) Still of U2OS cell expressing Lifeact-mEmerald (left) imaged with spinning disk confocal. Kymograph (right) taken from purple line of left image. Note robust movement of actin arc stress fibers (yellow arrow). (B) Still of hiCM expressing Lifeact-mApple (left) imaged with spinning disk confocal. Kymograph (right) taken from purple line of left image. Note slower movement of MSF in hiCM compared to actin arcs in U2OS cell, and stationary nature of sarcomeres (Sar). Gamma image correction of 0.5 was used to display relatively bright (i.e., sarcomeres) and dim (i.e, MSF) structures. (C) Quantification of actin stress fiber translocation rates in U2OS cells and hiCMs. U2OS; 3 cells over three experiments. hiCMs; 12 cells over three experiments. Scale Bars; (A), (B), 10 µM. P-values denoted in graph.
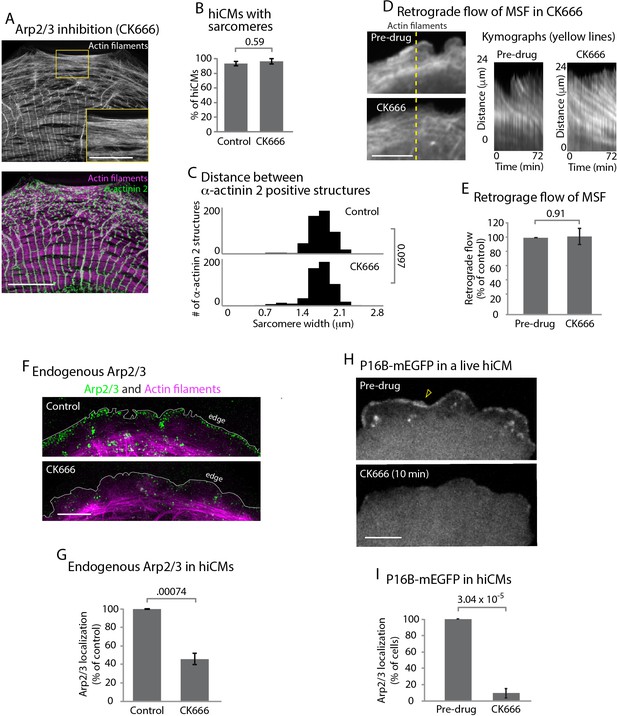
The Arp2/3 complex is not required for sarcomere assembly.
(A) hiCM allowed to spread for 24 hr in the presence of 25 µM CK666, labeled with actin and α-actinin 2 (i.e., Z-lines) and imaged with SIM. Box indicates presence of MSFs. (B) Quantification of percentage of cells with sarcomeres at 24 hr post plating in control and 25 µM CK666. Control: 76 cells, 10 experiments; 25 µM CK666: 41 cells, three experiments. (C) Histogram of distribution of distances between α-actinin 2 Z-lines. Note tight distribution of Z-lines in both conditions. Control: 14 cells, three experiments, 317 measurements. 25 µM CK666: 16 cells, three experiments, 530 measurements. (D) Stills of hiCM expressing Lifeact-mApple pre (top) and post (bottom) addition of 25 µM CK666 and imaged with spinning disk confocal. Kymographs (right) taken from dotted yellow line (left). (E) Rates of retrograde flow of hiCMs depicted as percent change in CK666 from pre-drug condition. 8 cells over three experiments. (F) Localization of the Arp2/3 complex in control (top) and 25 µM CK666 treated (bottom) hiCMs imaged with SIM. Note loss of Arp2/3 at the edge of CK666 treated hiCM. Cells spread for 24 hr in presence of 25 µM CK666 as in Figure 4A (G) Quantification of loss of the Arp2/3 complex from the leading edge of hiCMs. Control; 36 cells over three experiments. 25 uM CK666; 29 cells over three experiments. (H) Live hiCM expressing P16B-mEGFP (a component of the Arp2/3 complex) and imaged with spinning disk confocal. Localization of P16B-mEGFP at leading edge in pre-drug control (top) is acutely lost after addition of 25 µM CK666 (bottom). (I) Quantification of hiCMs displaying localization of the Arp2/3 complex (P16B-mEGFP) pre- and post-25µM CK666 in live hiCMs (as in Figure 4H). 27 cells over three experiments. Scale bars; (A) 10 µm low mag, 5 µm high mag inset. (D), (F), (H), 10 µm. P-values denoted in graphs.
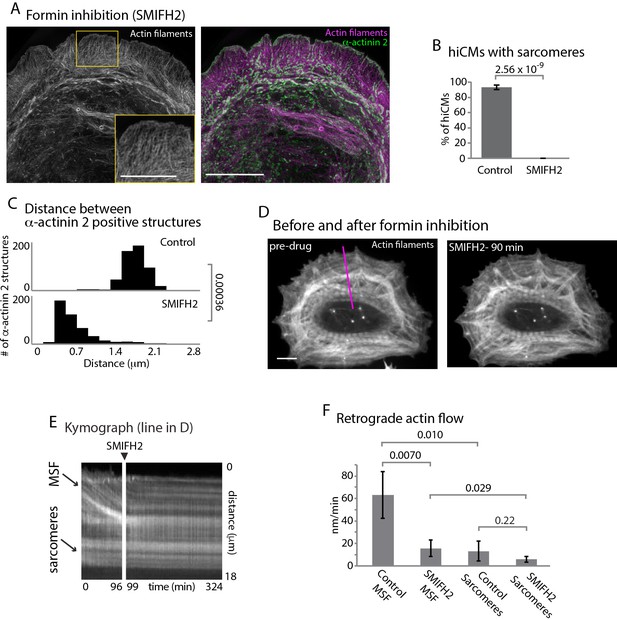
Formins are required for sarcomere assembly and MSF dynamics.
(A) hiCM allowed to spread in the presence of 25 µM SMIFH2 for 24 hr, labeled with actin and α-actinin and imaged with SIM. Box indicates loss of transverse MSFs behind leading edge of hiCM. (B) Quantification of percentage of cells with sarcomeres at 24 hr post plating. Control: 76 cells, 10 experiments; 25 µM SMIFH2, 16 cells, three experiments (C) Histogram of distribution of distances between α-actinin 2 Z-lines. Control: 14 cells, three experiments, 317 measurements, 25 µM SMIFH2: 11 cells, three experiments, 468 measurements. (D) Stills from live hiCM expressing Lifeact-mApple which were spread for 24 hr and assembled sarcomeres. hiCM before (left) and 90 min following addition of 25 µM SMIFH2 (right) (drug administered 24 hr after spreading) and imaged with spinning disk microscopy. Note how sarcomeres and overall actin architecture remains unperturbed at 90 min post 25 µM SMIFH2. (E) Kymographs of MSF and sarcomere retrograde flow taken from purple line in (D). Note immediate loss of retrograde flow following addition of 25 µM SMIFH2. (F) Quantification of actin retrograde flow in hiCMs pre and post addition of 25 µM SMIFH2. Control: 12 cells, three measurements from each cell, three experiments; 25 µM SMIFH2, 12 cells, three measurements from each cell, three experiments. Scale Bars: (A) 10 µm low mag, 5 µm high mag inset. (D), 10 µM. P-values denoted in graphs.
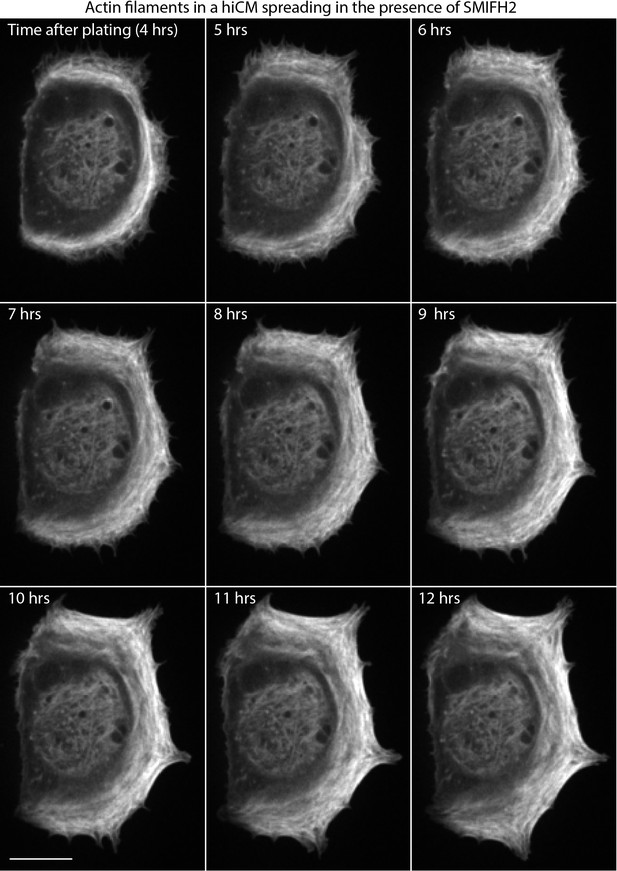
hiCM spreading in the presence of 25 µM SMIFH2 .
Spinning-disk confocal microscopy of hiCM expressing Lifeact-mApple and spreading in the presence of 25 µM SMIFH2 (added 1.5 hr post plating). Note sarcomeres do not form in the presence of 25 µM SMIFH2. Scale Bar; 20 µm.
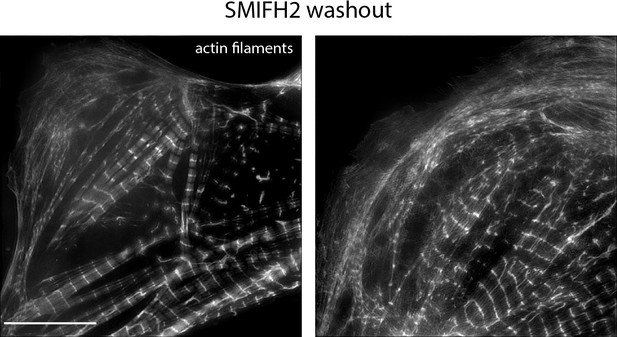
hiCMs assemble sarcomeres following washout of SMIFH2.
Examples of two hiCMs containing MSFs and sarcomeres following washout of 25 µM SMIFH2 and imaged with SIM. hiCMs were spread in 25 µM SMIFH2 (as in Figure 5A) for 24 hr, and fresh media without drug was exchanged. hiCMs were subsequently allowed to spread for an additional 24 hr before fixation. Scale Bar; 10 µm.
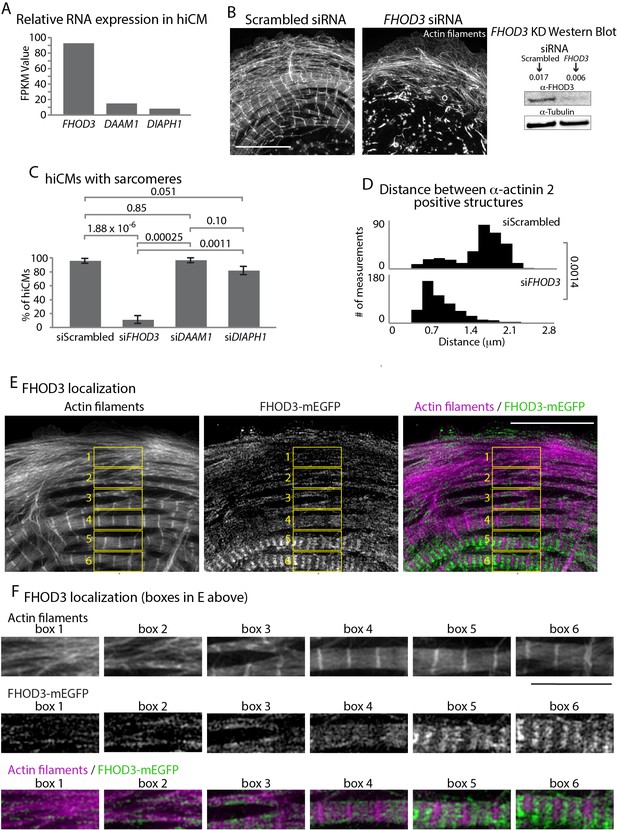
Formin FHOD3 is required for sarcomere assembly.
(A) Normalized Frames Per Kilobase Million (FPKM) of mRNA expression of the top three expressed formins in hiCMs, three experiments, three separate runs. (B) Actin of siRNA scramble control and siRNA FHOD3 hiCMs allowed to spread for 24 hr and imaged with SIM. Note loss of sarcomeres comparable to SMIFH2 treatment in FHOD3 KD hiCMs (Figure 5A). Western blot (right) denotes protein loss in FHOD3 KD. (C) Quantification of percentage of cells with sarcomeres at 24 hr post plating in scramble control, siFHOD3, siDAAM1, and siDIAHP1 hiCMs. Control: 76 cells, 10 experiments; siRNA FHOD3: 33 cells, three experiments; siRNA DAAM1: 29 cells, three experiments; siRNA Dia1: 26 cells, three experiments. (D) Histogram of distribution of distances between α-actinin 2 Z-lines. Control: 14 cells, three experiments, 317 measurements; siFHOD3: 15 cells, three experiments, 488 measurements. (E) hiCM transfected with FHOD3-mEGFP, fixed at 24 hr, stained for Actin and imaged with SIM. Boxes 1-6 depict localization of FHOD3-mEGFP along MSFs (boxes 1 and 2) and in sarcomeres (boxes 4, 5, and 6). (F) Boxes from (E) showing increased organization of FHOD3-mEGFP from MSFs to sarcomeres. Note increasingly organized structure, and localization between Z-lines of FHOD3-mEGFP. Scale bars; (B), (E), 10 µm. (F), 5 µm. P-values denoted in graphs.
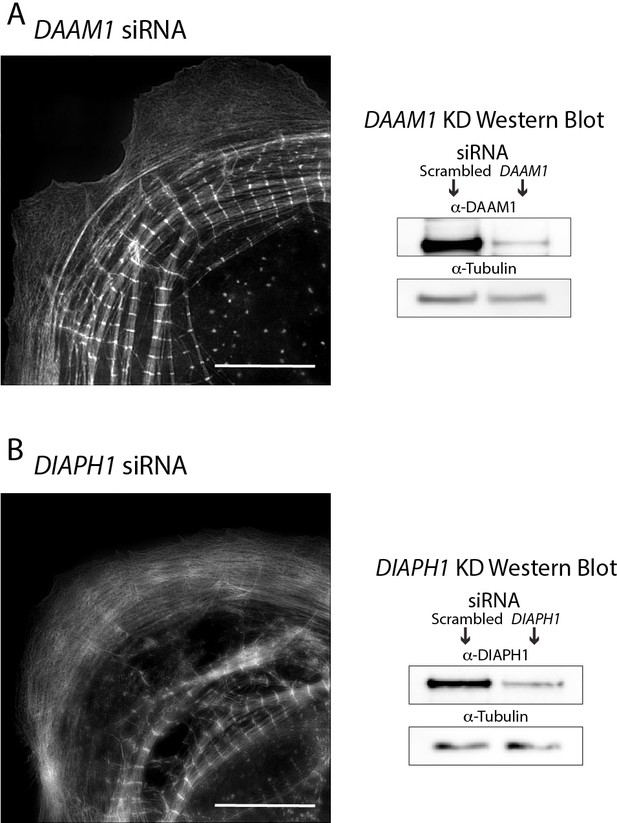
hiCMs assemble sarcomeres following knockdown (KD) of DAAM1 and DIAPH1.
(A) siRNA DAAM1 hiCM spread for 24 hr stained for actin, and imaged with SIM (left). DAAM1 KD hiCMs still assemble sarcomeres, and have extended lamellipodia. Representative western blot showing KD of DAAM1 protein (right). (B) siRNA DIAPH1 hiCM spread for 24 hr, stained for actin and imaged with SIM (left). DIAPH1 KD hiCMs appear to have fewer sarcomeres than scramble control hiCMs, but still assemble sarcomeres. Representative western blot showing KD of DIAPH1 protein (right). Scale Bars; (A), (B), 10 µm.
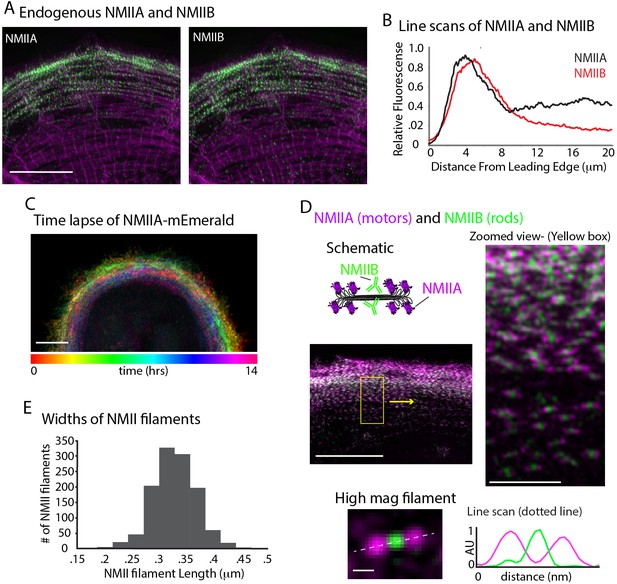
NMII Localization and Dynamics in hiCMs.
(A) Localization of endogenous NMIIA (left) and NMIIB (right) in the same hiCM and imaged with SIM. Both NMIIA and NMIIB localize to MSFs at the leading edge of hiCMs. (B) Line scans starting from edge of hiCMs showing localization of NMIIA (black) and NMIIB (red). Note NMIIA is localized slightly,~1 µm in front of NMIIB. NMIIA: 15 cells, two experiments; NMIIB: 32 cells, four experiments. (C) Color projection of time-lapse of hiCM expressing NMIIA-mEmerald and imaged with laser-scanning confocal. Note how NMIIA-mEmerald remains at the edge of hiCMs. (D) hiCM transfected with NMIIA-mEmerald (N-terminal motors), stained for endogenous NMIIB C-terminal rod domain (cartoon schematic and middle left), and imaged with SIM. High-mag views of NMIIA-NMIIB co-filaments (right) from yellow box (middle left). High mag view of single NMIIA-NMIIB co-filament (bottom) and line scan across white dotted line, from N-terminal motors (purple) and C-terminal rod domains (green). (E) Quantification of NMII co-filament length. Histogram displays the distribution of NMII co-filament lengths (motor-domain to motor-domain). Scale Bars; (A), (C), 10 µm, (D) 10 µm low mag (left), 2 µm ‘zoomed view’ inset (right), 200 nm ‘high mag filament’ inset (bottom).
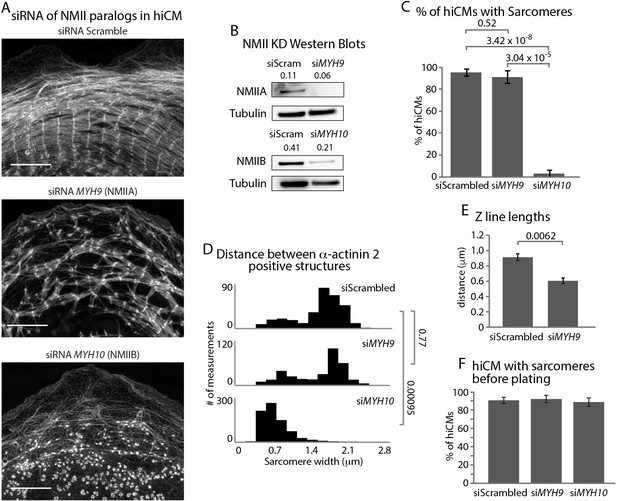
NMIIA and NMIIB are Required for Sarcomere Assembly in hiCMs.
(A) Actin of representative scramble control (top), NMIIA KD (siRNA MYH9, middle), and NMIIB KD (siRNA MYH10, bottom) hiCMs allowed to spread for 24 hr and imaged with SIM. NMIIA KD hiCMs (middle) display disorganized sarcomeres, while NMIIB KD hiCMs (bottom) display no actin-based sarcomeres. (B) Representative western blots of 2 separate experiments showing knockdown of NMIIA (siRNA MYH9, top) and NMIIB (siRNA MYH10, bottom). (C) Percentage of scramble control, NMIIA KD (siRNA MYH9), and NMIIB KD (siRNA MYH10) hiCMs with actin based sarcomeres at 24 hr spread. Control: 49 cells, six experiments; NMIIA KD: 34 cells, three experiments; NMIIB KD: 59 cells, four experiments. (D) Histogram of distribution of α-actinin 2 structures in scramble control, NMIIA KD (siMYH9) and NMIIB KD (siMYH10) hiCMs. siScrambled: 554 measurements, 14 cells, three experiments; siMYH9: 332 measurements, 15 cells, three experiments; siMYH10: 772 measurements, 15 cells, three experiments. (E) Quantification of Z-line lengths in scramble control and NMIIA KD (siMYH9) hiCMs. Control: 22 cells, four experiments; NMIIA: 14 cells, three experiments. (F) Quantification of hiCMs with sarcomeres in scramble control, NMIIA KD (siMYH9), and NMIIB KD (siMYH10) hiCMs before re-plating. siScrambled: 772 cells, four experiments; siMYH9: 642 cells, two experiments; siMYH10: 385 cells, two experiments. Scale bar; (A) 10 µm. P-values denoted in graphs.
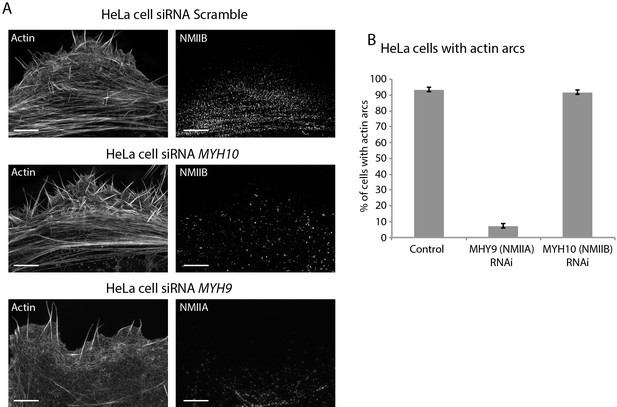
NMIIA but not NMIIB is required for actin arc formation in HeLa cells.
(A) HeLa cells treated with control, anti myh10, or anti myh9 (top, middle, bottom, respectively) siRNA, stained for actin, and imaged with SIM. Note how control and NMIIB KD cells display prominent actin arc stress fibers, but NMIIA KD cells contain no actin arcs. (B) Quantification of HeLa cells with actin arc stress fibers in indicated conditions. Scale Bars: 5 µm.
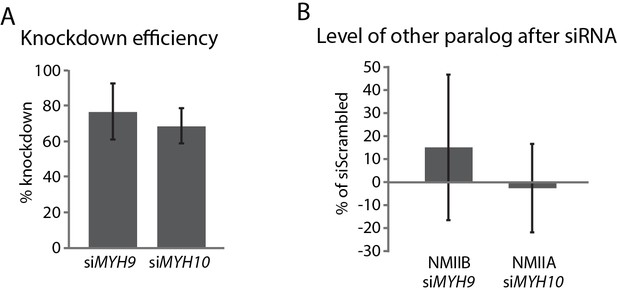
Knockdown of NMIIA and NMIIB in hiCMs.
(A) Quantification from western blots showing KD of NMIIA (siMYH9) and NMIIB (siMYH10) in hiCMs compared to scramble control. N = 3 for each condition. (B) Quantification of NMIIB levels in siMYH9 hiCMs (left) and NMIIA levels in siMYH10 hiCMs (right) compared to scramble control. N = 3 for each condition.
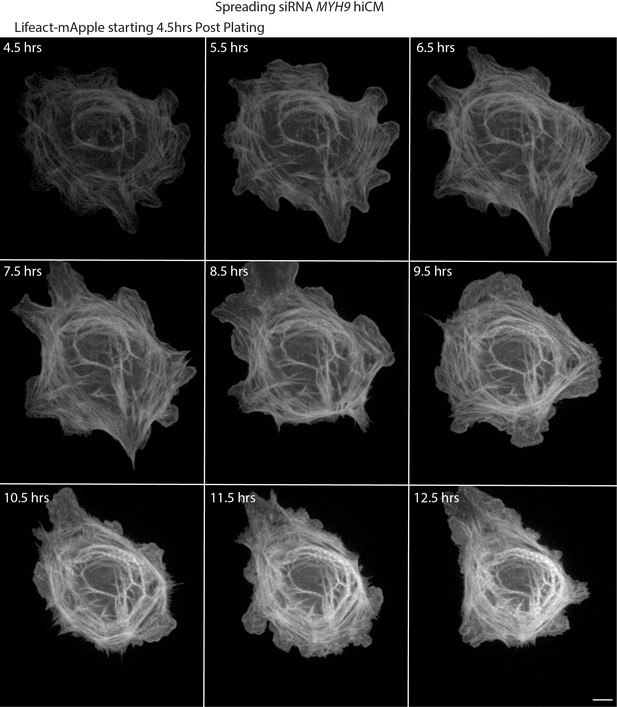
Live montage of spreading NMIIA KD hiCM.
Laser-scanning confocal of spreading NMIIA KD hiCM expressing Lifeact-mApple. NMIIA KD hiCM is able to assemble sarcomere structures. Scale Bar; 10 µm.
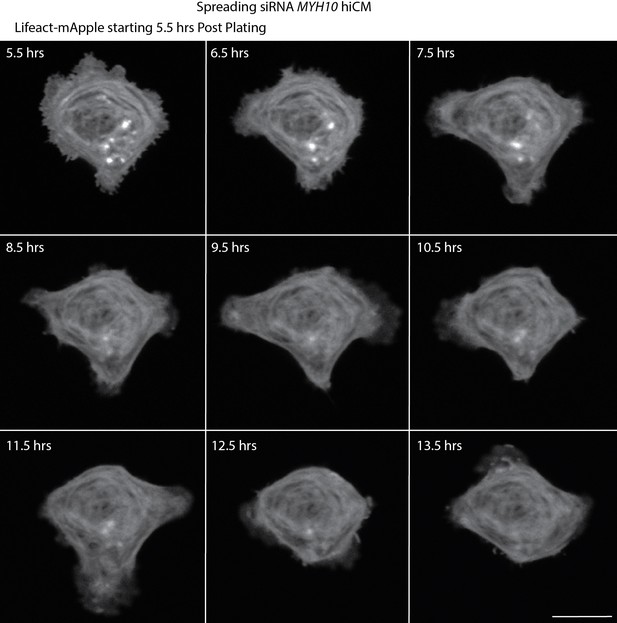
Live montage of spreading NMIIB KD hiCM.
Laser-scanning confocal of spreading NMIIB KD hiCM expressing Lifeact-mApple.
NMIIB KD hiCM does not form sarcomeres. Scale Bar; 20 µm.
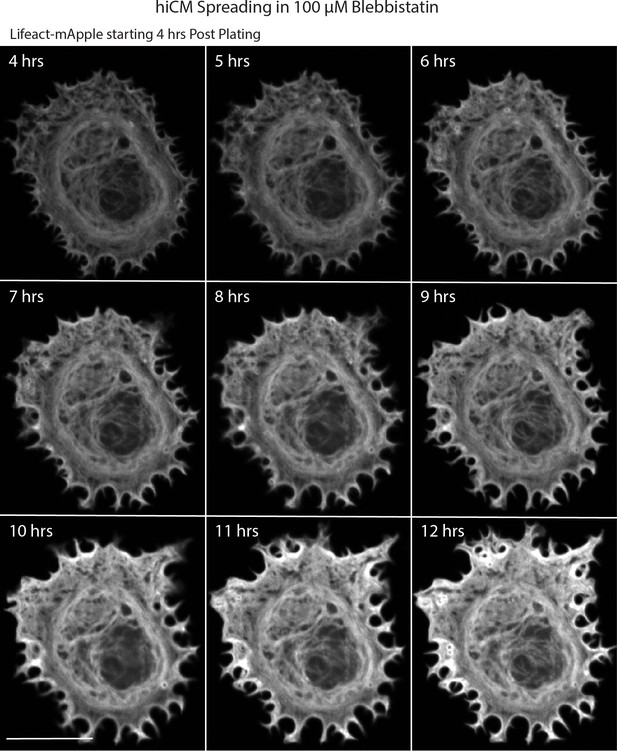
Live montage of hiCM spreading in 100 µM blebbistatin.
Laser-scanning confocal of hiCM spreading in the presence of 100 µM blebbistatin added 1.5 hr post plating. hiCM does not assemble sarcomeres in the presence of 100 µM blebbistatin. Scale Bar; 20 µm.
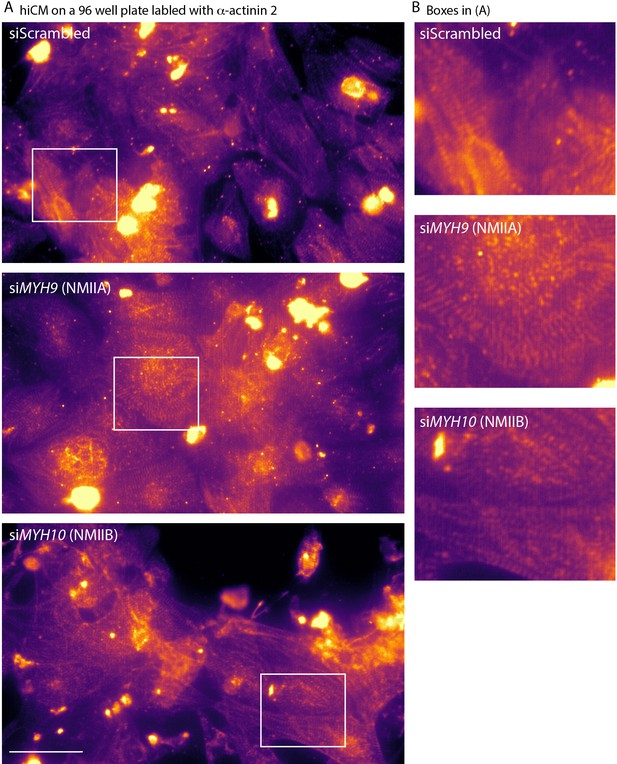
NMIIA KD and NMIIB KD hiCMs contain sarcomeres before plating.
(A) siScrambled (top) siMYH9 (NMIIA KD, middle), and siMYH10 (NMIIB KD, bottom) hiCMs treated in 96 well plates, fixed, and localized for α-actinin 2 imaged with wide-field microscopy. hiCMs were allowed to attach for 4 days (as per manufacturer’s recommendation, see Materials and methods), ensuring robust sarcomere assembly before siRNA treatment began. Note all conditions contain sarcomeric staining of α-actinin 2. Look up table is mpl-magma in Fiji.
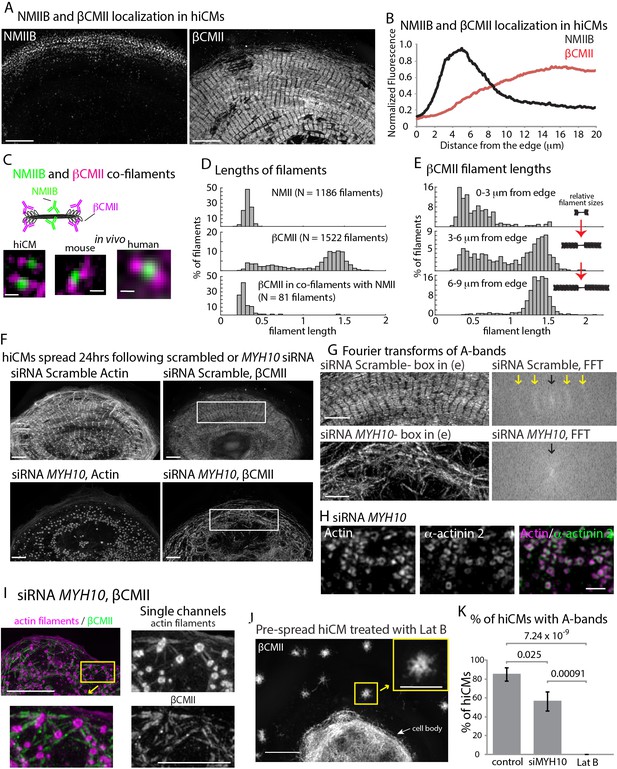
β Cardiac Myosin II (βCMII) Filament Assembly in hiCMs.
(A) Endogenous localization of NMIIB (left) and βCMII in the same hiCM and imaged with SIM. (B) Averaged line-scans of NMIIB (Figure 7B) and βCMII localization in hiCMs spread for 24 hr. Note the peak fluorescence of βCMII is more towards the cell body than peak fluorescence of NMIIB. 23 cells from four experiments were used for βCMII localization. (C) Schematic (top) of NMIIB-βCMII co-filaments. High-mag views of βCMII-NMIIB co-filaments (bottom). Endogenous staining of hiCM (bottom, left) for βCMII (N-terminal motors) and NMIIB (rod domain) and imaged with SIM. Mouse and human tissue (bottom middle, and bottom right, respectively) stained for βCMII (motors) and NMIIB (rod domain) and imaged with SIM and Zeiss 880 with Airyscan, respectively. (D) Histograms displaying width of NMII filaments (top), βCMII filaments (middle), and NMIIB-βCMII co-filaments in hiCMs. Measurements made from motor-domain to motor domain as in Figure 7D and E. (E) Histograms displaying distribution of βCMII filaments widths with respect to their location in hiCMs. Note βCMII filaments tend to grow larger as they move towards the center of the cell. Measurements were not taken from ‘mature’ sarcomere structures in highly organized A-bands. (F) Actin and βCMII of scramble control hiCM (top) and NMIIB KD (siMYH10) hiCM (bottom) spread for 24 hr. Note loss of organized A-bands but presence of βCMII filaments in NMIIB KD hiCM. (G) Fourier transforms of βCMII signal from white boxes in Figure 11F from scramble control and NMIIB KD hiCMs (above and below respectively). Yellow arrows indicate sarcomeric periodicity in scramble control hiCMs, which is lacking in NMIIB KD cells. (H) Actin and α-actinin 2 localized in a hiCM after NMIIB KD. (I) High mag views of actin and βCMII in NMIIB KD (siRNA MYH10) hiCM imaged with SIM. βCMII filaments localize to residual actin filaments. (J) βCMII in hiCM spread for total of 24 hr, with the final 6 hr in 5 µM Latrunculin B and imaged with SIM. Notice lack of βCMII A-bands in the periphery of hiCM, and large βCMII filament aggregates (yellow box). (K) Percentage of scramble control, NMIIB KD (siRNA MYH10), and 5 µM Latrunculin B hiCMs with βCMII A-bands. Control: 26 cells, three experiments; NMIIB KD: 26 cells, two experiments; Latrunculin B: 11 cells, three experiments. Cell bodies were not analyzed in the latrunculin experiment due to the density of βCMII localization. Scale Bars; (A) 10 µm, (C) 200 nm, (F), (G) 5 µm, (H) 1 µm. (I) 10 µm low mag, 5 µm high mag insets. (J) 10 µm low mag, 5 µm high mag inset. P-values denoted in graphs.
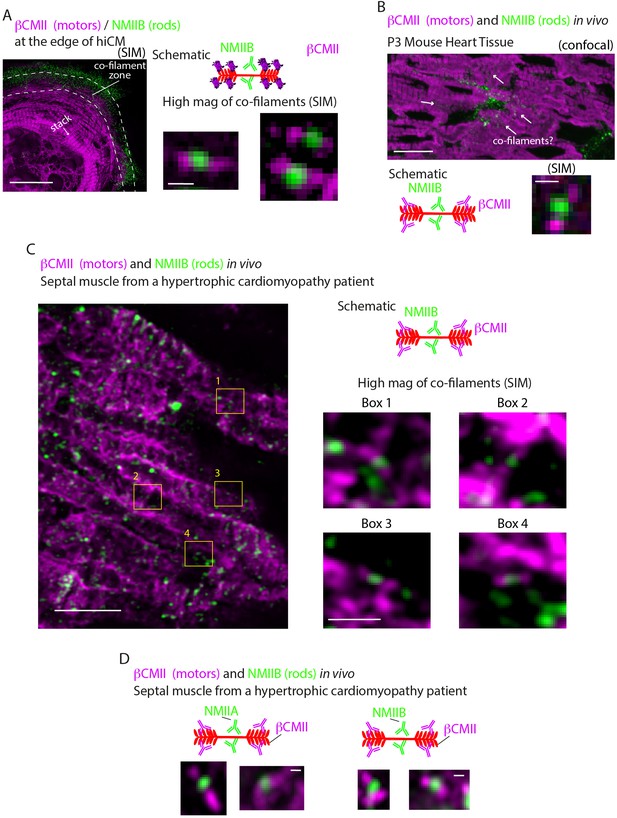

Myosin II co-filaments in hiCMs and in vivo.
(A) hiCM transfected with βCMII-mEGFP (N-terminal motors), stained for endogenous NMIIB rods and imaged with SIM. Schematic depicts visualization strategy. Low mag of hiCM left, and high mag examples bottom right. (B) Low mag view of P3 mouse heart tissue stained for βCMII (motors) and NMIIB (rods), and imaged with laser-scanning confocal (top). As in hiCMs, NMIIB is restricted from sarcomere structures but is localized adjacently to sarcomeres. Arrows indicates areas of possible co-filaments. High mag example (bottom) taken from P3 mouse tissue imaged with SIM from similar area indicated by arrow on low mag image. Schematic indicates visualization strategy. (C) Localization of βCMII and NMIIB in human septal muscle from a hypertrophic cardiomyopathy patient. Low-mag image (left) of βCMII (magenta) and NMIIB (green) imaged on Zeiss 880 with AiryScan. High-mag examples (right) from Boxes in low-mag image (left) indicate βCMII-NMIIB co-filaments. Schematic indicates visualization strategy. (D) High-mag examples of NMIIA (left) and NMIIB-βCMII (right) co-filaments from a human hypertrophic cardiomyopathy patient imaged on Zeiss 880 with AiryScan. Cartoon schematic indicates imaging strategy. Scale bars: (A, left), 10 μm; (A, right), 200 nm, (B, top), 20 µm: (B, bottom), 200 nm: (C, left), 5 µm: (C, right), 1 µm: (D), 200 nm.
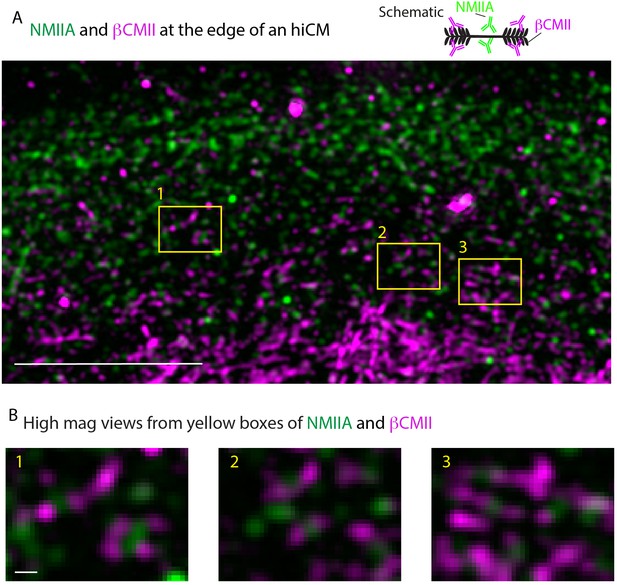
Endogenous localization of NMIIA and βCMII in hiCMs.
(A) SIM of endogenous localization of NMIIA (green) and βCMII (magenta) in hiCM. Yellow boxes denote areas of overlap between NMIIA and βCMII. (B) High-mag views of NMIIA-βCMII co-filaments taken from yellow boxes in (A).
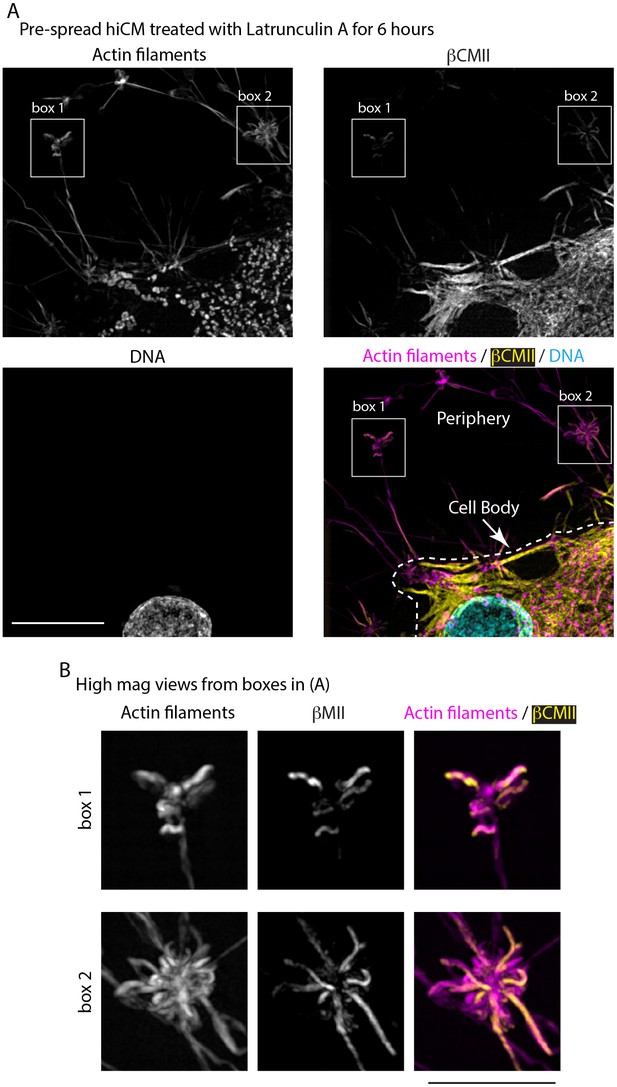
Actin, βCMII, and DNA localization in hiCMs treated with Latrunculin A.
(A) hiCM spread for a total 24 hr and treated with 5 μM Latrunculin A in the final 6 hr of spreading before fixation and imaged with SIM. Note loss of sarcomeric actin and βCMII. Following treatment with Latrunculin A, βCMII is found adjacent to the nucleus (DNA) and in large aggregates in cell periphery (box 1 and 2). (B) High-mag views from (A), showing βCMII filament aggregates localized to actin aggregates in cell periphery following treatment with 5 μM Latrunculin A. Scale bars: 10 μm in (A), 5 μm in (B).
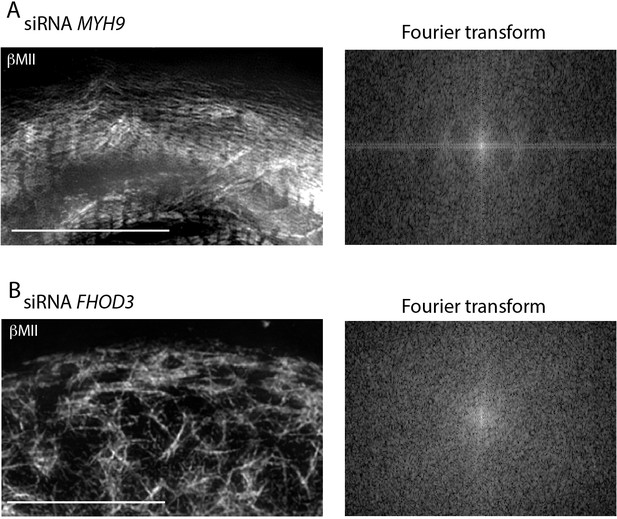
βCMII filament assembly is perturbed in NMIIA and FHOD3 KD hiCMs.
(A) Endogenous βCMII localized in NMIIA KD (siRNA MYH9) hiCMs (left) imaged with SIM. Fourier transform (right) of hiCM shows some periodicity of βCMII which is not as readily apparent compared to scramble control hiCMs (Figure 9G). (B) Endogenous βCMII localized in FHOD3 KD (siRNA FHOD3) hiCM (left) imaged with SIM. Note loss of βCMII A-bands. Fourier transforms (right) show loss of sarcomeric periodicity. Scale Bars: 10 μm.
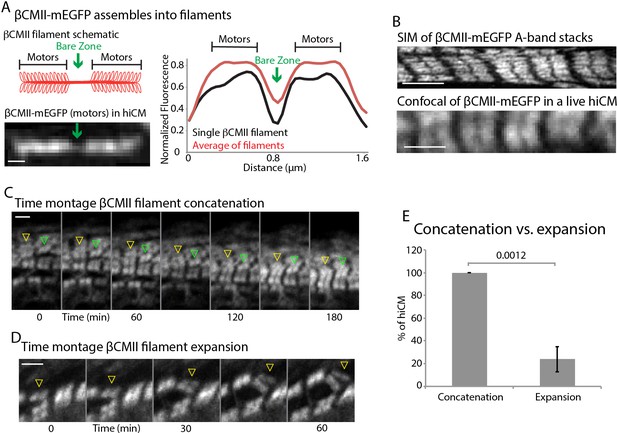
βCMII filaments concatenate to form larger A-band structures.
(A) Cartoon of βCMII filament (left, above). N-terminal tagged human βCMII-mEGFP filament expressed in hiCM (left, below). Gap in signal represents bare-zone lacking motors (green arrows). βCMII single filament and A-band filament (βCMII filaments found within organized A-bands) widths measured by line scans (right). βCMII Filaments: 16 filaments, three experiments. βCMII myofibrils: 28 myofibrils, three experiments. Note more level ‘plateau’ of signal from motors in A-band βCMII filaments. (B) SIM of representative βCMII-mEGFP myofibril in hiCM (top) and laser scanning confocal (bottom). (C) Representative montage showing two separate concatenation events. Yellow arrowhead denotes a large stack of βCMII-mEGFP filaments concatenating with a smaller stack of βCMII-mEGFP filaments as they undergo retrograde flow. Green arrowhead denotes smaller βCMII-mEGFP filament concatenating with larger βCMII-mEGFP stack as they undergo retrograde flow. Both events result in larger and more organized βCMII-mEGFP filament stack (i.e. the A-band). (D) Example of βCMII-mEGFP filament splitting event. Note how small βCMII-mEGFP stack splits to create two smaller βCMII-mEGFP filaments and does not result in larger or more organized βCMII-mEGFP filament stacks. (E) Quantification of % of hiCMs which display concatenation or expansion events of βCMII filaments. Scale Bars; (A) 200 nm, (B), (C) and (D) 2 µm. P-values denoted in graph.
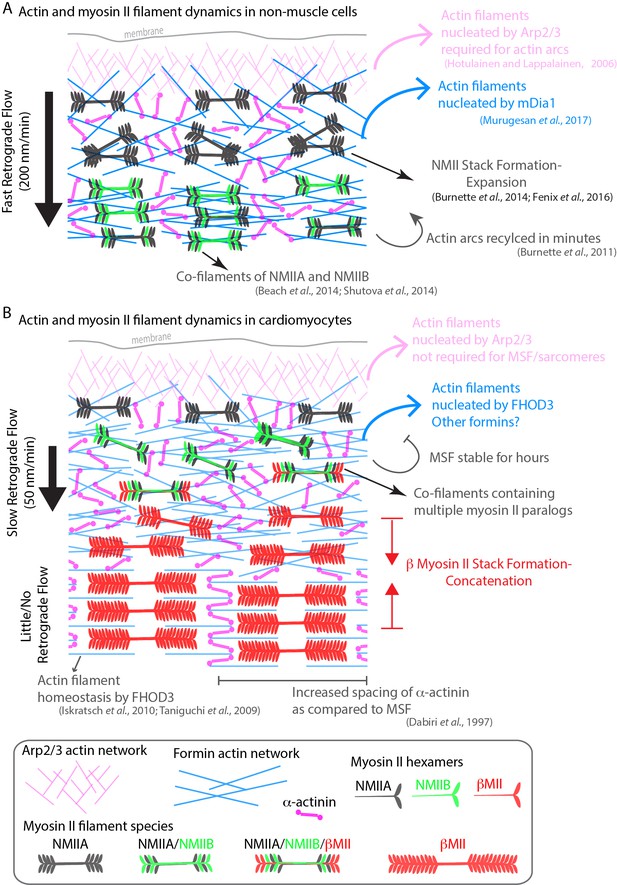
Model of actomyosin stress fiber formation in non-muscle cells and human cardiomyocytes.
(A) Actin and myosin II stress fiber formation in non-muscle cells. Actin stress fibers are formed via the Arp2/3 complex and the formin mDia1. NMIIA is the predominant isoform at the leading edge of non-muscle cells, and stress fiber formation is NMIIA dependent. Non-muscle cells display robust retrograde flow of actin stress fibers and display rapid turnover. Large NMIIA stacks are formed via growth and expansion of smaller NMIIA filaments. Citations leading to this model are presented in the cartoon. (B) Model of actin and myosin II stress fiber formation in human cardiomyocytes. Sarcomeres are templated by Muscle Stress Fibers (MSFs). MSFs do not require the Arp2/3 complex, and require the formin FHOD3. MSFs display slow retrograde flow compared with non-muscle stress fibers. Both NMIIA and NMIIB are localized to the edge of hiCMs, and display prominent NMII co-filaments. NMIIB-βCMII co-filaments are also present with MSFs. Large βCMII filament stacks form via concatenation and stitching of individual βCMII filaments.
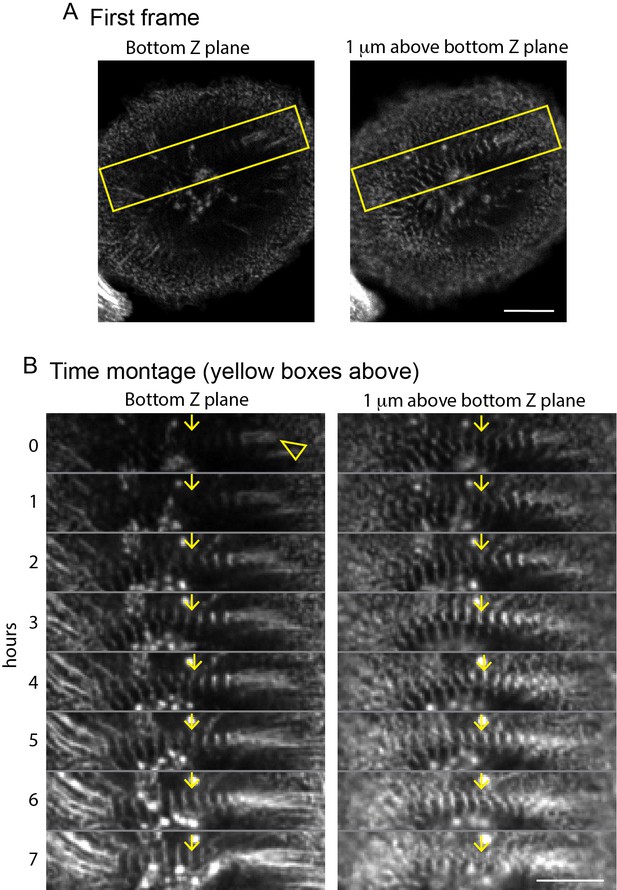
Sarcomeres form on dorsal surface of hiCMs and subsequently move towards ventral surface.
(A) hiCM expressing α-actinin 2-mCherry plated for 24 hr and imaged with laser-scanning confocal. Note that when viewing the ventral (bottom) surface (left image), no sarcomeric pattern of α-actinin 2 can be seen. However, when viewing the dorsal (top) surface of the cell (right image), sarcomeres are clearly present. (B) Time montage from yellow box in (A). Myofibril from top of the cell (right montage, yellow arrow) moves towards the bottom of the cell (left image, yellow arrow). Arrow head (left montage) denotes α-actinin 2 containing adhesion.
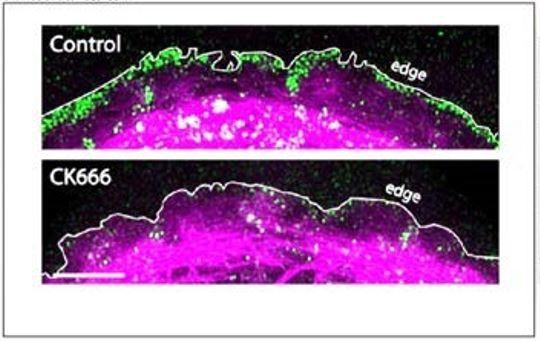
Figure 4F stretched.
https://doi.org/10.7554/eLife.42144.036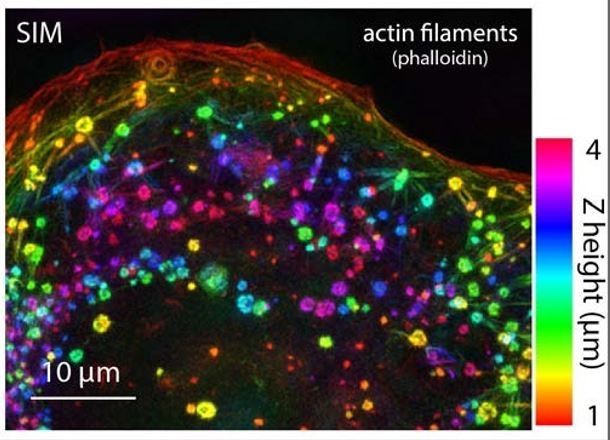
3D projection of actin- NMIIB KD hiCM.
https://doi.org/10.7554/eLife.42144.037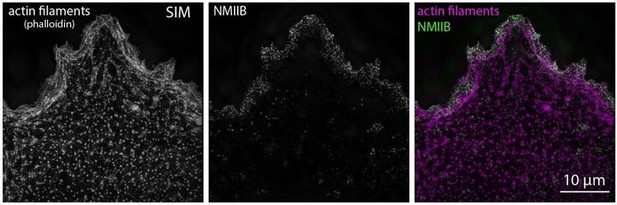
hiCM spread in presence of 100µM Blebbistatin; 24 hours after plating.
https://doi.org/10.7554/eLife.42144.038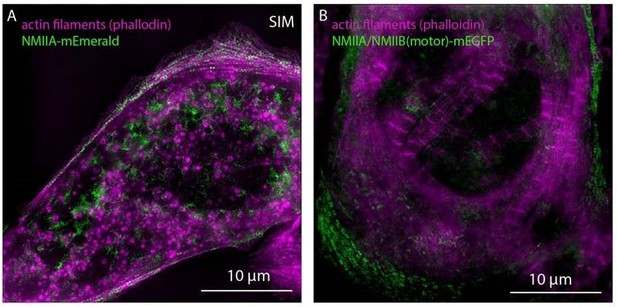
Expression of NMIIA and NMIIA/NMIIB(motor) chimera in NMIIB KD hiCM.
https://doi.org/10.7554/eLife.42144.039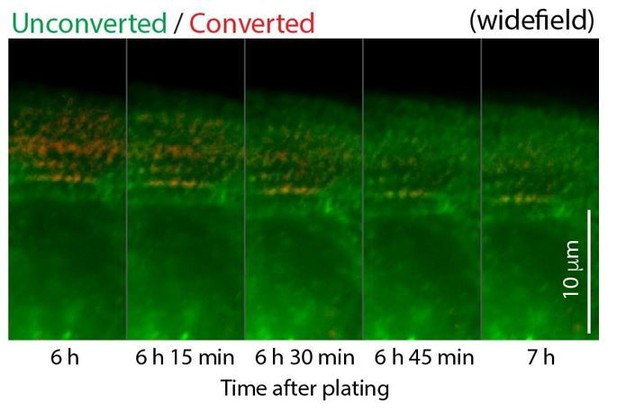
Time montage of Myosin IIB-/mEOS2 in a hiCM.
https://doi.org/10.7554/eLife.42144.040Videos
Actin filaments in a hiCM assembling sarcomeres. hiCM transfected with Lifeact-mApple and imaged with SIM.
MSFs undergo retrograde flow and transition to sarcomere containing myofibrils towards cell body. Lookup table: orange hot. 30.5 by 20.9 µm. Video length: 9.5 hr.
NMIIA filament dynamics during sarcomere assembly. hiCM transfected with NMIIA-mEmerald and Lifeact-mApple and imaged with 3D laser-scanning confocal.
Note how NMIIA-mEmerald filaments form at the edge of hiCMs, are localized to MSFs, but are restricted from sarcomeres during sarcomere assembly. 28 by 35 μm. Video length: 14 hr.
NMIIB filament dynamics during sarcomere assembly. hiCM transfected with pHalo-NMIIB and α-actinin 2-mEmerald and imaged with wide-field microscopy.
Note how NMIIB-Halo filaments form at the edge of hiCMs, are localized to MSFs, but are restricted from sarcomeres during sarcomere assembly. 47 by 27 μm. Video Length: 7 hr.
βCMII filaments in a hiCM assembling sarcomeres. hiCM transfected with βCMII-mEGFP and imaged with SIM.
Note filaments concatenating. Lookup table: orange hot. 30.7 by 29.7 μm. Video length: 7.5 hr.
Sarcomeres traveling from dorsal to ventral surface of hiCM hiCM plated for 24 hr and expressing α-actinin 2-mCherry.
Four views from four Z-planes imaged using laser-scanning confocal microscopy. In the first frame, no sarcomeres can be visualized on the bottom of the cell (left most frame), but can clearly be seen in the 2 μm Z plane (middle right frame). As the video continues, the sarcomeres in the 2 μm Z plane travel towards the ventral (bottom Z plane) surface of the hiCM, which can be readily seen as the myofibril comes into focus first in the 1 μm Z plane (middle left), and then in the Bottom Z plane (left). Height: 38.2 μm. Video length: 17.5 hr.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (human) | iCell Cardiomyocytes^2 | Cellular Dynamics International | ipsc derived cardiac myocytes | |
| Cell line (human) | Cor.4U Cardiomyocytes | Ncardia (formerly Axiogenesis) | ipsc derived cardiac myocytes | |
| Cell line (human) | HeLa | American Type Culture Collection (ATCC) | CCL-2 | |
| Cell line (human) | U2-OS | American Type Culture Collection (ATCC) | HTB-96 | |
| Recombinant DNA reagent | pHalo-NMHC IIA-C18 | this paper | see Materials and methods for details on creation | |
| Recombinant DNA reagent | pHalo-NMHC IIB-C18 | this paper | see Materias and methods for details on creation | |
| Recombinant DNA reagent | α-actinin-2-mEmerald | Addgene | 53988 | |
| Recombinant DNA reagent | α-actinin-2-tdEOS | Addgene | 57577 | |
| Recombinant DNA reagent | NMIIA-(N-terminal)-mEGFP | Addgene | 11347 | |
| Recombinant DNA reagent | Lifeact-mEmerald | Gift from Michael Davidson | ||
| Recombinant DNA reagent | Lifeact-mApple | Gift from Michael Davidson | ||
| Recombinant DNA reagent | NMIIB-(N-terminal)-mEmerald | Addgene | 54192 | |
| Recombinant DNA reagent | FHOD-mEGFP (lacking T(D/E)5XE exon) | Gift from Dr. Elizabeth Ehler | Iskratsch et al. (2010) | |
| Recombinant DNA reagent | βCMII-mEGFP | Genescript (Piscataway, NJ, USA) this paper | see Materials and methods for details on creation | |
| Recombinant DNA reagent | p16b-eGFP | Gift from Dr. Jenniffer Lippincott-Schwartz and Dr. Pekka Lappalainen | Lai, F. P., et al 2008.Koestler et al., 2013. | |
| Antibody | rabbit polyclonal IgG anti-NMIIA | Biolegend | P909801 | Used at 1:1000 |
| Antibody | rabbit polyclonal IgG anti-NMIIB | Cell Signaling | P3404S | Used at 1:200 |
| Antibody | rabbit polyclonal IgG anti-NMIIB | Biolegend | 909901 | Used at 1:200 |
| Antibody | mouse monoclonal IgG anti-βCMII | Iowa Hybridoma Bank | A4.1025 | Used at 1:2 |
| Antibody | rabbit polyclonal IgG anti-α-actinin-2 | Sigma-Aldrich | clone EA-53 | Used at 1:200 |
| Antibody | rabbit polyclonal IgG anti-p34-Arc/ArpC2 (Arp2/3 complex) | Millipore Sigma | 07–227 | Used at 1:100 |
| Antibody | rabbit polyclonal IgG anti-DAAM1 | Bethyl Laboratories | HPA026605 | Used at 1:100 |
| Antibody | rabbit polyclonal IgG anti-DIAPH1 | Sigma-Aldrich | A300-078A | Used at 1:100 |
| Antibody | mouse monoclonal IgGanti-cardiac troponin T | Santa Cruz Biotechnology | CT3, sc-20025 | Used at 1:50 |
| Antibody | goat anti mouse IgG Alexa Fluor 568 | ThermoFisher Scientific | A-11004 | Used at 1:100 |
| Antibody | goat anti rabbit IgG Alexa Fluor 568 | ThermoFisher Scientific | A-11011 | Used at 1:100 |
| Antibody | goat anti mouse IgG Alexa Fluor 488 | ThermoFisher Scientific | AA28175 | Used at 1:100 |
| Antibody | goat anti rabbit IgG Alexa Fluor 647 | ThermoFisher Scientific | A-21244 | Used at 1:100 |
| Antibody | goat anti mouse IgM Alexa Fluor 488 | ThermoFisher Scientific | A-21042 | Used at 1:100 |
| Antibody | mouse monoclonal IgG anti-FHOD3 | Santa Cruz Biotechnology | G-5, sc-374601 | Used at 1:100 |
| Sequence-based reagent | SmartPool siRNA DIAPH1 | GE Dharmacon | E-010347-00-0005 | |
| Sequence-based reagent | SmartPool siRNA DAAM1 | GE Dharmacon | E-012925-00-0005 | |
| Sequence-based reagent | SmartPool siRNA FHOD3 | GE Dharmacon | E-023411-00-0005 | |
| Sequence-based reagent | SmartPool siRNA MYH10 (NMIIB) | GE Dharmacon | E-023017-00-0010 | |
| Sequence-based reagent | SmartPool siRNA MYH9 (NMIIA) | GE Dharmacon | E-007668-00-0005 | |
| Commercial assay or kit | Rneasy Mini Kit | Qiagen | 74104 | |
| Commercial assay or kit | QIAprep spin Miniprep Kit (250) | Qiagen | 27106 | |
| Chemical compound, drug | Blebbistatin | Sigma-Aldrich | B0560 | |
| Chemical compound, drug | LatrunculinB | Sigma-Aldrich | L5288 | |
| Chemical compound, drug | LatrucluinA | Sigma-Aldrich | L5163 | |
| Chemical compound, drug | CK666 | Sigma-Aldrich | SML0006 | |
| Chemical compound, drug | SMIFH2 | Sigma-Aldrich | S4826 | |
| Chemical compound, drug | TransIT-TKO | Mirus Bio | MIR 2150 | |
| Chemical compound, drug | FuGENE HD | Promega | E2311 | |
| Chemical compound, drug | Alexa Fluor 488 Phalloidin | ThermoFisher Scientific | A12379 | |
| Chemical compound, drug | Viafect | promega | E4981 | |
| Software, algorithm | (Fiji Is Just) ImageJ | NIH (open source) |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.42144.034





