Length regulation of multiple flagella that self-assemble from a shared pool of components
Figures
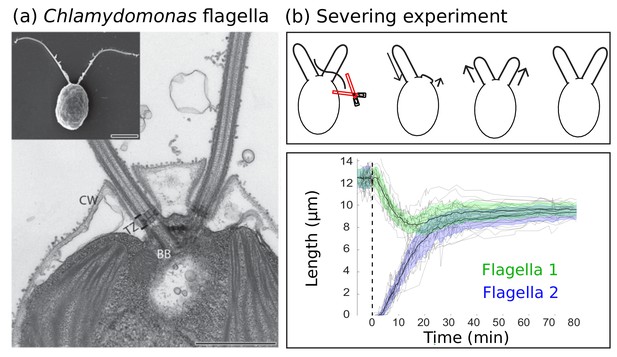
Experimental background.
(a) Electron microscopy images of the biflagellate green algae Chlamydomonas and its flagella captured by Elisa Vannuccini and Pietro Lupetti (University of Siena, Italy) and reproduced from Morga and Bastin (2013) under the Creative Commons Attribution License CC BY 2.0 [http://creativecommons.org/licenses/by/2.0]). The inset shows the whole organism (scale bar 5 µm) and the close-up shows the flagellar basal body (BB), transition zone (TZ), and cell wall (CW) (scale bar 1 µm). (b) Severing experiments: after one flagellum is severed, the two flagella equalize at a shorter length and then grow together to the original steady-state length. This is demonstrated in experimental data of 20 severing experiments from Ludington et al. (2012) provided by the authors. The green and blue shaded regions show the mean plus or minus one standard deviation.
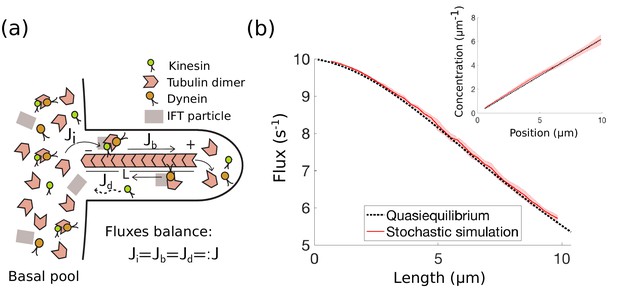
IFT in Chlamydomonas with diffusive return of kinesin-2 to the base.
(a) Kinesin-2, dynein, and tubulin combine to form a complex with other IFT components in the basal pool and are injected into the flagellum. The kinesin-2 motors move toward the tip of the flagellum, where the complex eventually breaks down into its constituent parts. Dynein motors move in the retrograde direction carrying back some, but not all IFT components. Specifically, kinesin motors are not part of the retrograde IFT complex and they diffuse back to the base. In steady-state, the injection flux , ballistic flux , and diffusive flux are all equal. (b) IFT flux vs. length, comparison between the quasi-steady state approximation (Equation 4) and stochastic simulations (Appendix 1). The shaded red area represents the mean and standard deviation computed from 10 realizations of the stochastic simulations. (inset) Kinesin-2 concentration vs. position along the flagellum (steady-state approximation (Equation 31)).
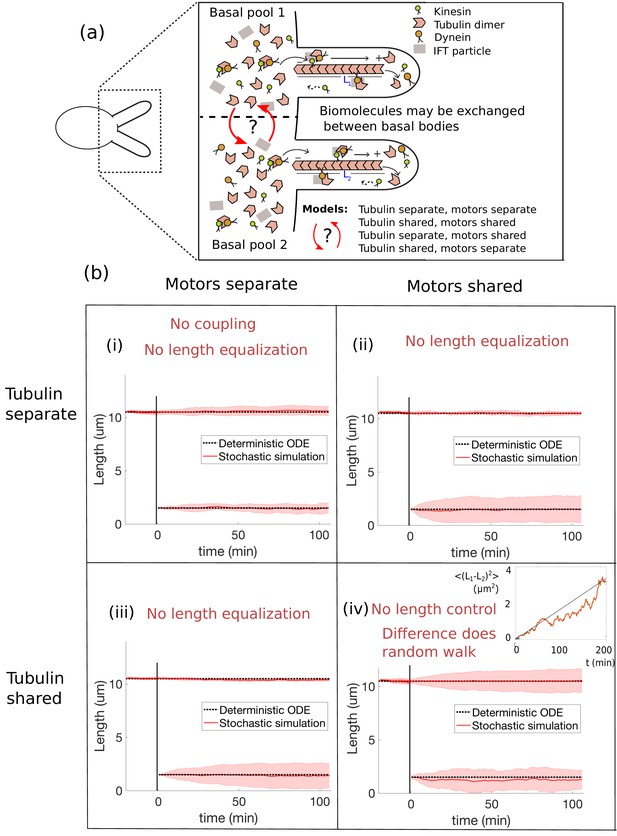
Length dynamics of two flagella assembling from shared pools of building blocks.
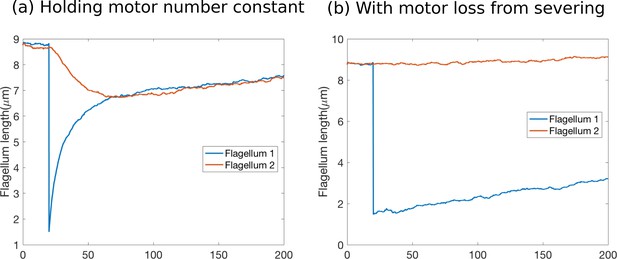
(a) Flagellar assembly in Chlamydomonas reinhardtii and modes of coupling between basal proteins pools. (b) Simulations of the severing experiment using different modes of coupling between basal pools of proteins. In all cases severing occurs at time zero. (i) In the case of separate pools of both motors and tubulin (Equations 44 and 45), the unsevered flagellum does not decrease in length. (ii) When only motors are shared (Equations 49 and 50), the flagellar lengths do not equalize after severing. (iii) When only tubulin is shared (Equations 56 and 57), the flagellar lengths do not equalize after severing. (iv) When both tubulin and motors are shared (Equations 58 and 59), there is no unique steady-state. The difference between the two flagellar lengths undergoes a random walk, as shown by the mean square displacement (inset, average over 10 runs).
The constant disassembly model with tubulin separate and motors shared does not yield length equalization.
The constant disassembly model with tubulin shared and motors separate does not yield length equalization.
The constant disassembly model with both tubulin and motors shared does not yield length control.
Replenishing protein pools in the constant disassembly model with tubulin separate and motors shared does not yield length control.
Replenishing protein pools in the constant disassembly model with tubulin shared and motors separate does not yield length control.
Replenishing protein pools in the constant disassembly model with both tubulin and motors shared does not yield length control.
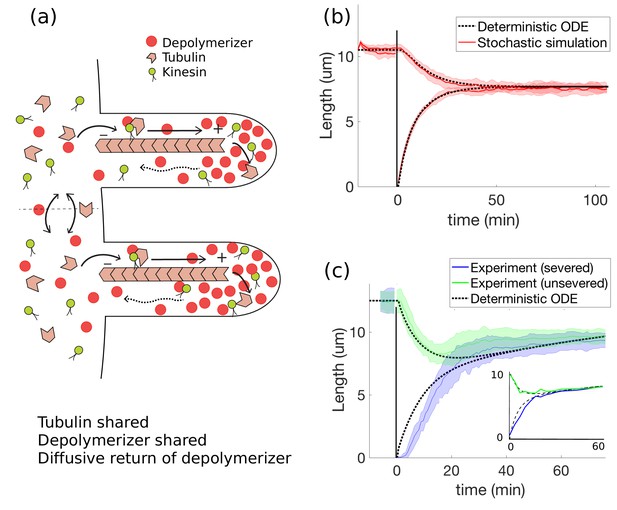
Concentration-dependent disassembly model: simultaneous length control is achieved using shared tubulin and shared depolymerizers.
(a) The depolymerizer moves ballistically to the flagellar tip and diffuses back, (b) The model Equations 20 and 21 captures rapid length equalization, (c) Protein replenishment with timescale min is included through Equations 18 and 19 and the model is fit to severing data from Ludington et al. (2012) (and to Rosenbaum et al., 1969, see inset). For the fit to Ludington et al. (2012), we use fitting parameters of , , , , and ; all other parameters are as in Table 1. For the fit to Rosenbaum et al. (1969), we use the same fitting parameters except for , , and .
The active disassembly model with both tubulin and motors shared but without protein replenishment exhibits rapid length equalization.
Replenishing protein pools in the active disassembly model with both tubulin and motors shared exhibits rapid length equalization and slow recovery.
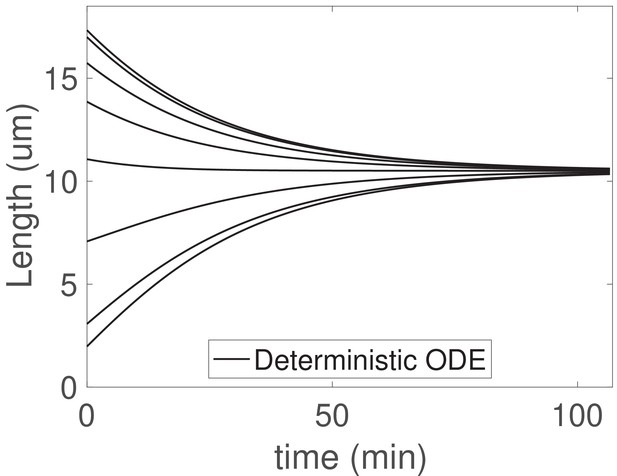
The concentration-dependent disassembly model generalizes to arbitrary flagellar number .
We solve (Equations 25) with flagella and the larger shared pool , , and ; otherwise all parameters are as in Table 1.
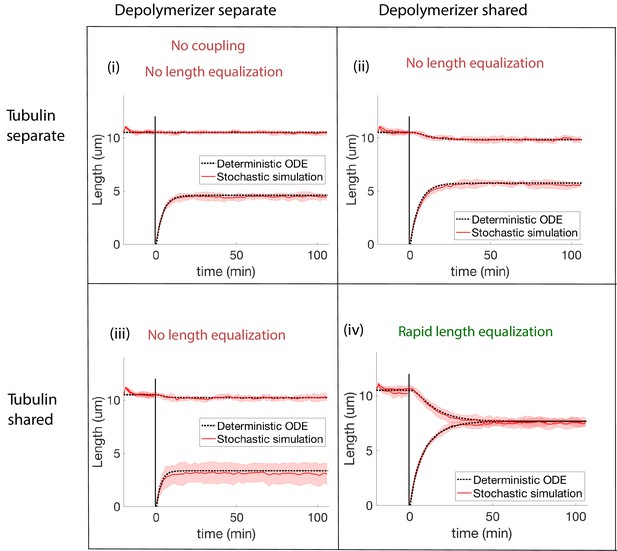
Concentration-dependent disassembly model: simulations of severing experiment with different modes of coupling between basal pools (and no replenishment of protein levels).
We conclude that biomolecule pools are fully shared after ruling out all models that disagree with the rapid length equalization that occurs in severing experiments. (i) In the case of separate pools of both tubulin and depolymerizers, the unsevered flagellum does not decrease in length. (ii) When only depolymerizers are shared, the flagellar lengths do not equalize after severing. (iii) When only tubulin is shared, the flagellar lengths do not equalize after severing. (iv) When both tubulin and depolymerizers are shared, the flagella lengths rapidly equalize after severing, as observed in experiments.
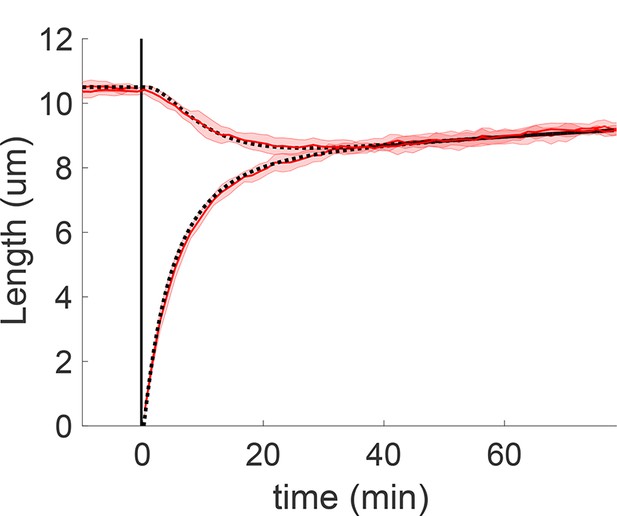
Concentration-dependent disassembly model Equation 68 and 69 with tubulin in excess yields rapid length equalization consistent with severing experiments, here with replenishment timescale .
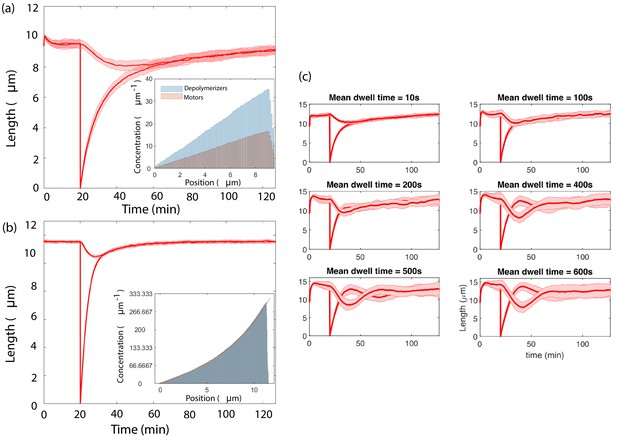
Simultaneous length control is achieved by different versions of the concentration-dependent disassembly model.
(a) Results from agent-based simulations with different populations of depolymerizer and rate-limiting IFT protein, using capacity , (inset) Concentrations along the flagellum of diffusing IFT proteins and depolymerizers, (b) An exponential profile is generated by motile proteins that bind and unbind from the flagellum, move ballistically in the anterograde direction when bound, and undergo diffusion when unbound, (inset, fit to Equation 89), (c) Agent-based simulations of processive depolymerization, in which depolymerizers diffusing near the flagellar tip have some probability of binding and removing tubulin at a fixed rate. We perform simulations of severing over a range of mean depolymerization times from 10 s to 600 s.
Tables
Parameter values and definitions.
| Symbol | Definition | Value | Units | References |
|---|---|---|---|---|
| Parameters | ||||
| Steady-state length | 10–12 | µm | Marshall and Rosenbaum, 2001; Rosenbaum et al., 1969 | |
| Tubulin pool per flagellum | 38–47 | µm | Marshall et al., 2005 | |
| Disassembly speed | 0.5 | µm/min | Marshall and Rosenbaum, 2001; Ludington et al., 2012 | |
| IFT speed | 2.5–3 | µm/s | Kozminski et al., 1993; Buisson et al., 2013 | |
| Diffusion coefficient | 1.7 | µm2/s | Chien et al., 2017 | |
| Assembly rate per tubulin | 2.3 × 10-2 –3.6 × 10-2 | min−1 | Fit | |
| Injection rate constant | 0.8–4 | min−1 | Fit | |
| Prefactor in Equation 5 | 2.5 × 10-4 | Estimate (Appendix 2) | ||
| Variables | ||||
| Number of flagella | ||||
| Free tubulin | µm | |||
| Total motors | ||||
| Free motors | ||||
| Ballistic motors | ||||
| Diffusing motors | ||||
| Flux | min−1 | |||
| Motor concentration | µm−1 | |||
| Average concentration | µm−1 |