Anion channel SLAH3 is a regulatory target of chitin receptor-associated kinase PBL27 in microbial stomatal closure
Figures
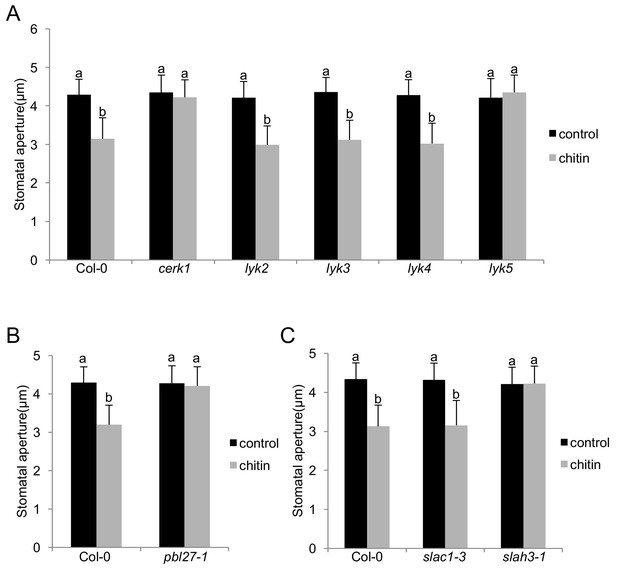
The LYK5-CERK1-PBL27 receptor complex and SLAH3 are required for chitin-induced stomatal closure.
(A–C) Stomatal aperture measurements in mutants of all members of the LysM-RLK family (A), pbl27 (B), slac1 and slah3 (C). Mature leaf discs were soaked in opening buffer (10 mM MES, 50 mM KCl, pH 6.15) and kept under light (100 μmol.m-2 s −1) for 2 hr. Stomatal apertures were measured 2 hr after treatment with 1 mg/ml chitin. Values are mean ± SD (n > 60; two-way ANOVA). Different letters indicate significantly different values at p<0.05. These experiments were repeated three times with similar results.
-
Figure 1—source data 1
Source data for stomatal measurements shown in Figure 1 and Figure 1—figure supplements 1 and 2, and for Figure 4.
- https://doi.org/10.7554/eLife.44474.005
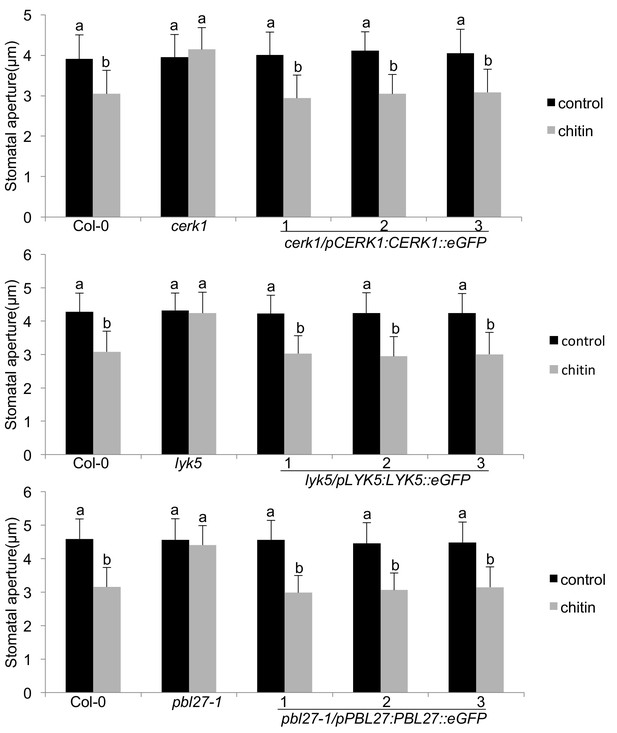
Functional complementation and guard cell expression of the LYK5-CERK1-PBL27 receptor complex.
Stomatal aperture measurements in transgenic cerk1/pCERK1::CERK1-eGFP, lyk5/pLYK5::LYK5-eGFP and pbl27-1/pPBL27::PBL27-eGFP. Mature leaf discs of six independent transgenic T1 lines were soaked in opening buffer (10 mM MES, 50 mM KCl, pH 6.15) and kept under light (100 μmol.m-2 s −1) for 2 hr. Stomatal aperture was measured 2 hr after treatment with 1 mg/ml chitin. Values are mean ± SD (n > 50; two-way ANOVA). Different letters indicate significantly different values at p<0.05. These experiments were repeated three times with similar results.
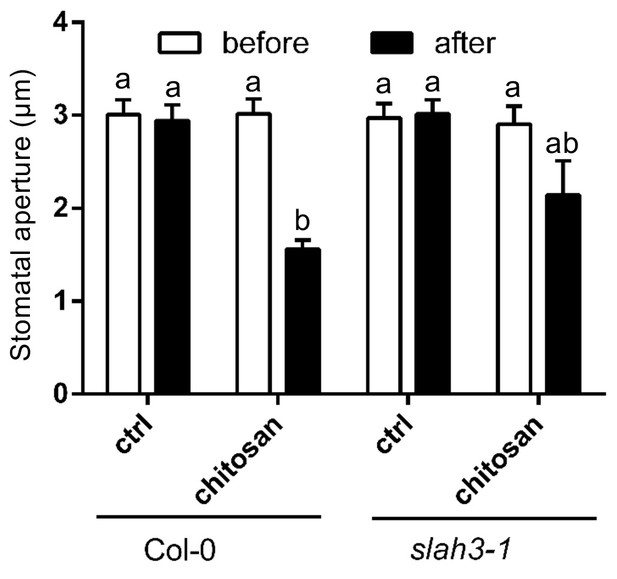
Current ejection of chitosan induces stomatal closure in Arabidopsis.
Chitosan was current ejected into the cell wall of guard cells in intact leaves via a microelectrode, by application of 1 nA positive current for 1 min. The tip of electrode was filled with 0.3 mg/ml chitosan or 10 mM MES/BTP (pH6, control), and rest with 300 mM KCl. Data are presented as average ± SE. Col-0 control, n = 8 stomata; Col-0 chitosan, n = 8 stomata; slah3-1 control, n = 3 stomata; slah3-1 chitosan, n = 8 stomata. The statistical significance indicates with different characters (one-way ANOVA, Tukey’s test, p<0.05).
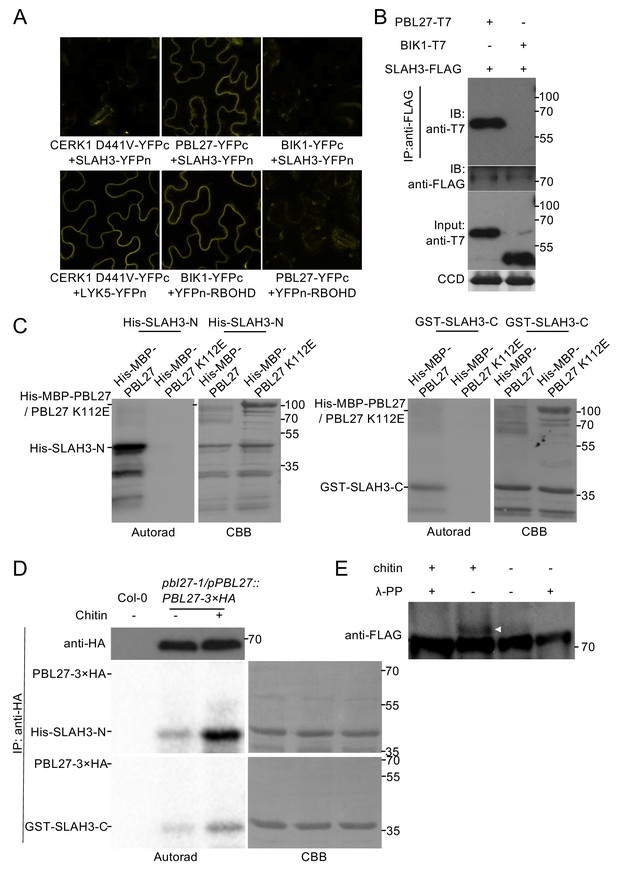
PBL27 interacts with and phosphorylates SLAH3.
(A) Confocal microscopy of N. benthamiana leaves transiently expressing the indicated split-YFP constructs. Representative images are shown. (B) Co-immunoprecipitation of PBL27 and SLAH3 transiently expressed in N. benthamiana leaves. These experiments were performed at least twice with similar results. Expected sizes of PBL27-T7 and BIK1-T7 fusion proteins correspond to 57 kDa and 46 kDa, respectively. SLAH3-FLAG used for immune-precipitation has an expected size of 73 kDa. (C) PBL27 trans-phosphorylates SLAH3-N and SLAH3-C. In vitro kinase assay incubating equal amounts of recombinant His-MBP-PBL27, His-MBP-PBL27 K112E (kinase dead) with recombinant His-SLAH3-N or GST-SLAH3-C (GST-SLAH3-C-His). Autoradiogram, left panel; Coomassie colloidal blue (CCB) stained membrane, right panel. These experiments were repeated three times with similar results. (D) Chitin-activated PBL27 trans-phosphorylates SLAH3-N. Transgenic pbl27-1/pPBL27::PBL27−3 × HA Arabidopsis seedlings were treated (+) or not (-) with 1 mg/ml chitin for 10 min. Total proteins were subjected to immunoprecipitation with anti-HA beads followed by immunoblot analysis with anti-HA to reveal PBL27−3 × HA (upper panel). Immuno-precipitated PBL27−3 × HA was incubated with recombinant His-SLAH3-N for in vitro kinase assay. Autoradiogram, left panel; Coomassie colloidal blue (CCB) stained membrane, right panel. Col-0 seedlings were used as a control. These experiments were repeated three times with similar results. (E) Chitin induces SLAH3 phosphorylation. Transgenic Arabidopsis slah3-1/35S::SLAH3−3 × FLAG transgenic were treated (+) or not (-) with 1 mg/ml chitin for 30 min. Total proteins were subjected to immunoprecipitation with anti-FLAG beads. The phosphorylated form of SLAH3−3 × FLAG was shifted upward in Phos-tag SDS-PAGE. After phosphatase treatment, the shifted band of SLAH3−3 × FLAG dispersed, indicating the SLAH3−3 × FLAG was phosphorylated after treatment with chitin. The white arrow indicates the phosphorylated form of SLAH3-FLAG. The bands were detected with an anti-FLAG antibody. This experiment was repeated three times with similar results.
-
Figure 2—source data 1
Source data for BiFC images shown in Figure 2.
- https://doi.org/10.7554/eLife.44474.008
-
Figure 2—source data 2
Source data for co-IP blots images shown in Figure 2.
- https://doi.org/10.7554/eLife.44474.009
-
Figure 2—source data 3
Source data for blots on in vitro phosphorylation shown in Figure 2.
- https://doi.org/10.7554/eLife.44474.010
-
Figure 2—source data 4
Source data for blots on in vitro-in vivo phosphorylation shown in Figure 2.
- https://doi.org/10.7554/eLife.44474.011
-
Figure 2—source data 5
Source data for blots on in vivo phosphorylation shown in Figure 2.
- https://doi.org/10.7554/eLife.44474.012
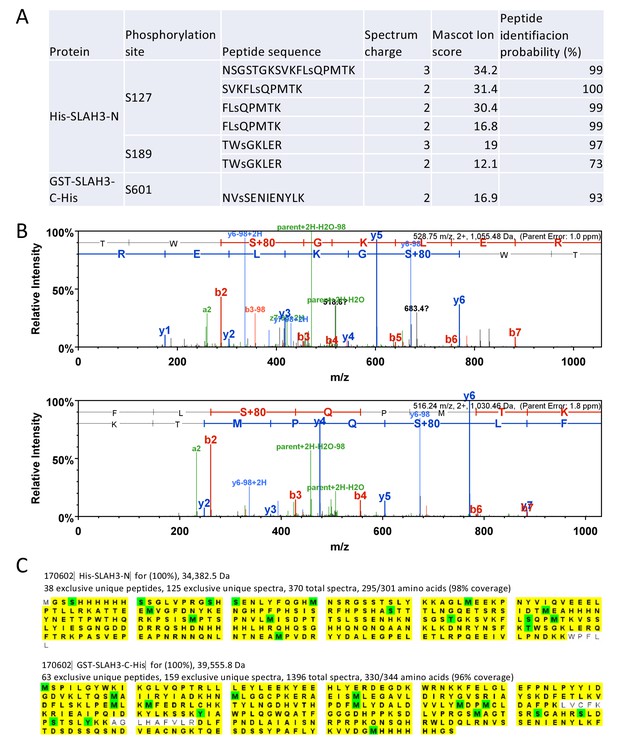
Identification of SLAH3 tryptic phospho-peptides.
(A) Summary of tryptic phospho-peptides by LC-MS/MS analysis of incubated recombinant His-MBP-PBL27 with recombinant His-SLAH3-N or GST-SLAH3-C (GST-SLAH3-C-His) in in vitro kinase assays. Phosphorylated residues are denoted as ‘‘s’’. (B) Representative spectra of phosphorylated S127 and S189 tryptic peptides. (C) Scaffold images for the samples for which the phosphopeptides were identified and at conservative settings (95% protein and peptide identity and minimum two peptides per protein). Since the samples contain recombinant protein the peptide coverage is extremely high. (96% and 98%). Peptides identified are marked in yellow and modified residues are shown in green.
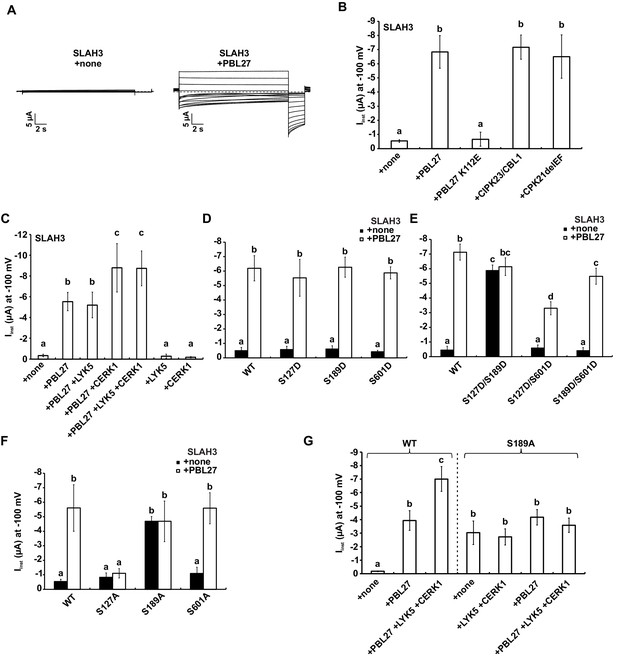
S-type anion currents are activated by co-injection of SLAH3 and PBL27 in oocytes.
(A) Macroscopic currents of Xenopus oocytes expressing SLAH3 in the presence or absence of PBL27 in response to the standard voltage protocol. Currents were recorded in 30 mM nitrate-based buffers. Representative cells are shown. (B) Instantaneous currents (Iinst) at −100 mV recorded from oocytes injected with SLAH3 alone or co-injecting SLAH3 with the indicated kinases in the presence of 100 mM nitrate (n ≥ 4; mean ± SD). PBL27 K112E represents a kinase-dead mutant. CPK21DEF represents a Ca2+-independent and thus constitutive active truncation mutant of CPK21. (C) Instantaneous currents (Iinst) at −100 mV of oocytes injected with WT SLAH3 alone or co-injected with PBL27, LYK5, CERK1 or different combinations of these components as indicated in the figure. Currents were recorded in nitrate-based buffers (100 mM) (n ≥ 4; mean ± SD). (D) and (E) Instantaneous currents (Iinst) of oocytes injected with SLAH3 WT or (D) the phospho-mimetic single mutants S127D, S189D and S601D or (E) the phospho-mimetic double mutants S127D/S189D, S127D/S601D or S189D/S601D were measured in the presence or absence of PBL27 at −100 mV. Currents were recorded in standard buffers containing 100 mM nitrate (n ≥ 4; mean ± SD). (F) Instantaneous currents (Iinst) at −100 mV of oocytes injected with SLAH3 WT or the phospho-dead mutants S127A, S189A and S601A in the presence or absence of PBL27 in nitrate-based buffers (100 mM) (n ≥ 4; mean ± SD). (G) Instantaneous currents (Iinst) at −100 mV in nitrate-based buffers of oocytes injected with WT SLAH3 or the mutant S189A alone or co-injected with PBL27, LYK5 and CERK1 as indicated in the figure (n ≥ 4; mean ± SD). (B) - (G) Significant differences (ANOVA with Tukey’s HSD test, p<0.01) between bars are denoted with different letters.
-
Figure 3—source data 1
Source data for current measurements shown in Figure 3 and Figure 3—figure supplements 1 and 2.
- https://doi.org/10.7554/eLife.44474.017
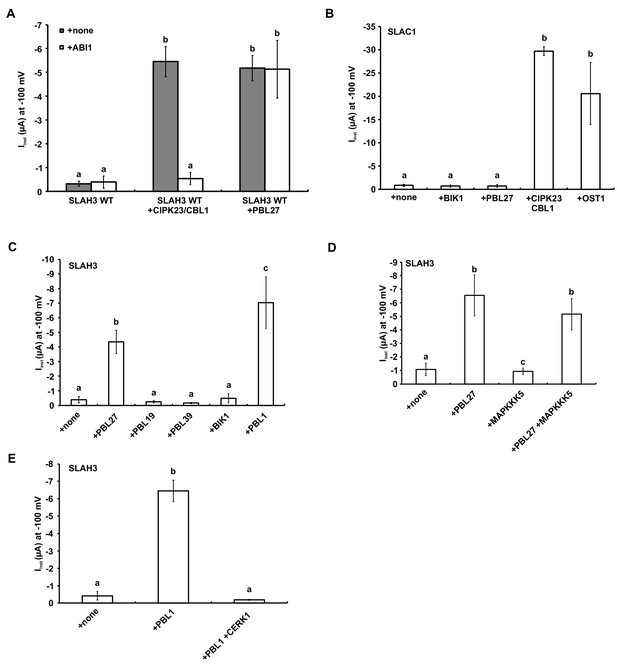
Activation of SLAH3 and its close relative SLAC1 by different kinases.
(A) Instantaneous currents of Xenopus oocytes injected with WT SLAH3 alone, with CIPK23/CBL1 or PBL27, respectively. The effect of ABI1 on the SLAH3-derived was studied by co-expression of the ABI1 phosphatase in oocytes. Currents were recorded in standard buffers containing 100 mM nitrate (n ≥ 4 experiments, mean ± SD). (B) Instantaneous currents of Xenopus oocytes injected with wild type SLAC1 alone or together with BIK1, PBL27, CIPK23/CBL1 or OST1. Currents were recorded at −100 mV in 100 mM chloride containing buffers. Note: For full activation of SLAC1 via OST1 split-YFP constructs (SLAC1-YFPc and OST1-YFPn) were used (n = 4 experiments, mean ± SD). (C) Instantaneous currents of Xenopus oocytes injected with wild type SLAH3 alone or together with PBL27, PBL19, PBL39, BIK1 or PBL1. Currents were recorded at −100 mV in 100 mM chloride containing buffers (n = 4 experiments, mean ± SD). (D) Instantaneous currents of Xenopus oocytes injected with wild type SLAH3 alone or together with PBL1, or co-injected with PBL1 and CERK1. Currents were recorded at −100 mV in 100 mM chloride containing buffers (n = 4 experiments, mean ± SD). (E) Instantaneous currents at −100 mV of Xenopus oocytes injected with WT SLAH3 alone or co-injecting PBL27, MAPKKK5 or both kinases in nitrate-based buffers (100 mM) (n ≥ 4 experiments, mean ± SD). Significant differences (ANOVA with Tukey’s HSD test, p<0.01) between bars are denoted with different letters.
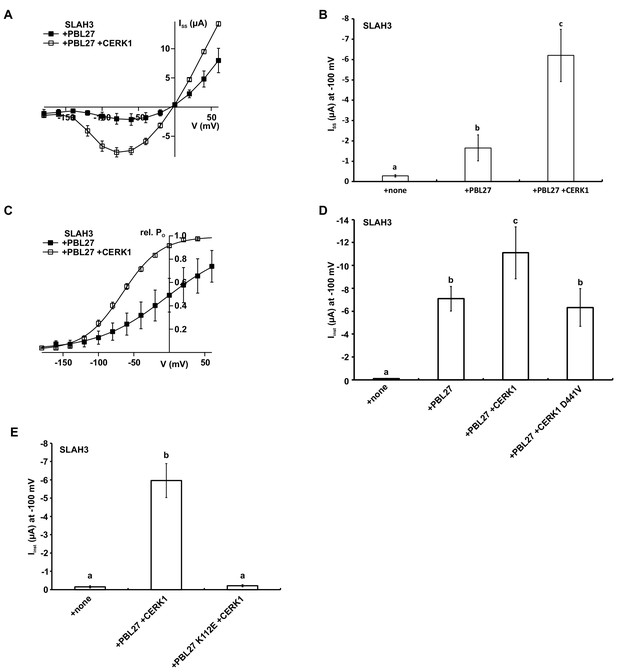
CERK1 enhances PBL27 activation of SLAH3.
(A) Steady-state currents (ISS) of oocytes co-injected with SLAH3 and PBL27 in the presence or absence of CERK1. Currents were recorded in 30 mM nitrate-based buffers (n ≥ 4 experiments, mean ± SD). (B) Steady-state currents of oocytes injected with wild type SLAH3 alone or co-injected with PBL27 in the presence or absence of wild type CERK1. Currents were recorded at −100 mV in 30 mM nitrate-based buffers (n = 4 experiments, mean ± SD) (C) Relative voltage-dependent open probabilities (rel. PO) of SLAH3 activated by either PBL27 alone or in combination with CERK1 in 30 mM nitrate. Rel. PO was calculated from a −120 mV voltage pulse following the test pulses in the voltage range of +60 to −200 mV in 20 mV decrements. Data points were fitted by a Boltzmann equation (continuous line; n ≥ 4 experiments, mean ± SD). (D) Instantaneous currents of oocytes injected with wild type SLAH3 alone or co-injected with PBL27 in the presence or absence of wild type CERK1 or the phospho-dead mutant CERK1 D441V. Currents were recorded at −100 mV in 100 mM nitrate containing buffers (n = 4 experiments, mean ± SD). (E) Instantaneous currents of oocytes injected with wild type SLAH3 alone or co-transfected with either PBL27 wild type or PBL27 K112E in the presence of CERK1. Currents were recorded at −100 mV in 100 mM nitrate containing buffers (n = 4 experiments, mean ± SD). Significant differences (ANOVA with Tukey’s HSD test, p<0.01) between bars are denoted with different letters.
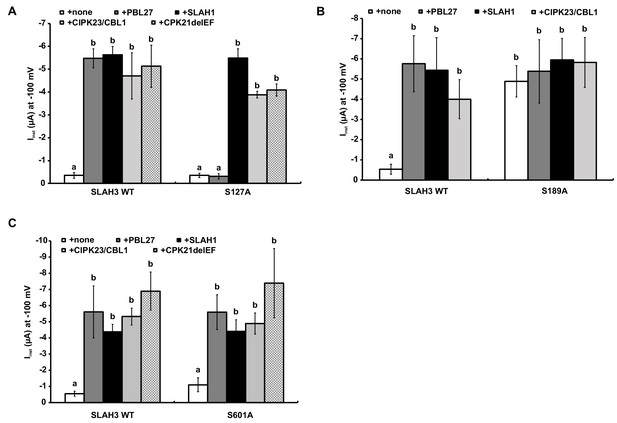
Activation of SLAH3 wild type and the SLAH3 mutants S127A, S189A and S601A.
(A), (B) and (C) Instantaneous currents of Xenopus oocytes injected with WT SLAH3 or (A) S127A, (B) S189A or (C) S601A alone or together with PBL27, CIPK23/CBL1, CPK21DEF or SLAH1. (A) As PBL27 represents the only kinase that could not activate the mutant SLAH3 S127A, this phosphorylation site seems to be specific for PBL27. Currents were recorded at −100 mV in 100 mM nitrate containing buffers (n = 4 experiments, mean ± SD). (B) None of the co-injected kinases or SLAH1 could further activate the constitutive active mutant S189A. Currents were recorded at −100 mV in 100 mM nitrate containing buffers (n = 4 experiments, mean ± SD). (C) The mutant S601A could be activated by all tested kinases as well as by SLAH1 similar to WT SLAH3. Currents were recorded at −100 mV in 100 mM nitrate containing buffers (n = 4 experiments, mean ± SD). Significant differences (ANOVA with Tukey’s HSD test, p<0.01) between bars are denoted with different letters.
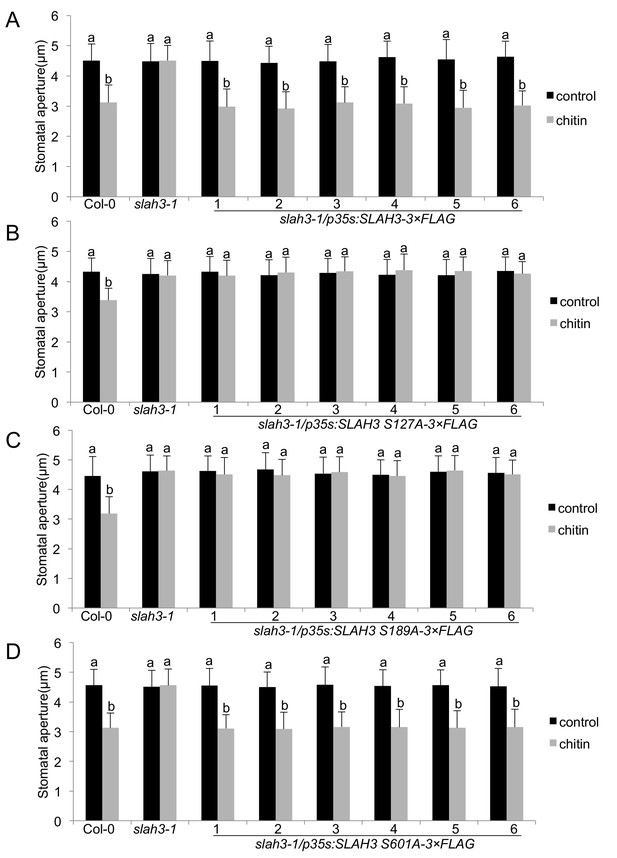
SLAH3 phospho-sites S127 and S189 are necessary for chitin-induced stomatal closure.
(A–D) Stomatal aperture measurements in transgenic slah3-1/35S::SLAH3−3 × FLAG wild type (A), S127A (B), S189A (C) and S601A (D) variants of SLAH3. Mature leaf discs of six independent transgenic T1 lines were soaked in opening buffer (10 mM MES, 50 mM KCl, pH 6.15) and kept under light (100 μmol.m-2 s −1) for 2 hr. Stomatal aperture was measured 2 hr after treatment with 1 mg/ml chitin. Values are mean ± SD (n > 50; two-way ANOVA). Different letters indicate significantly different values at p<0.05. These experiments were repeated three times with similar results.
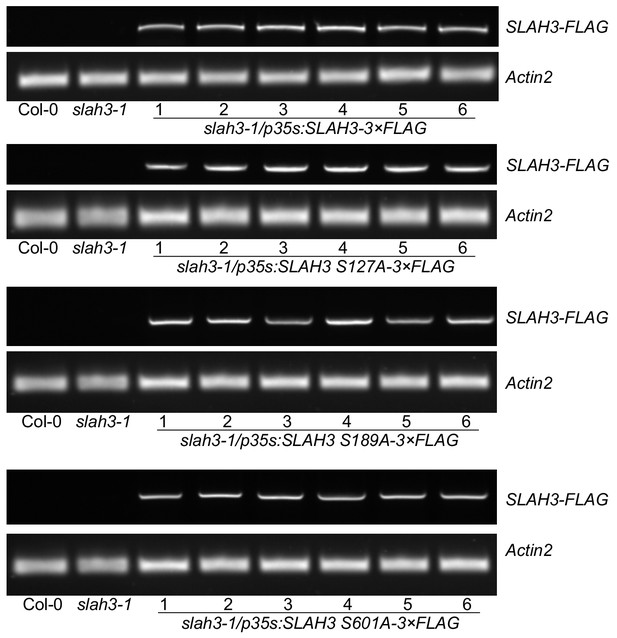
Functional complementation of SLAH3 transgenic lines.
RT-PCR analysis of T1 transgenic slah3-1/p35S::SLAH3-3xFLAG lines and respective T1 lines expressing the S127A, S189A and S601A SLAH3 variants. RNA was extracted from six independent transgenic plants used in Figure 4 for stomatal aperture measurements.
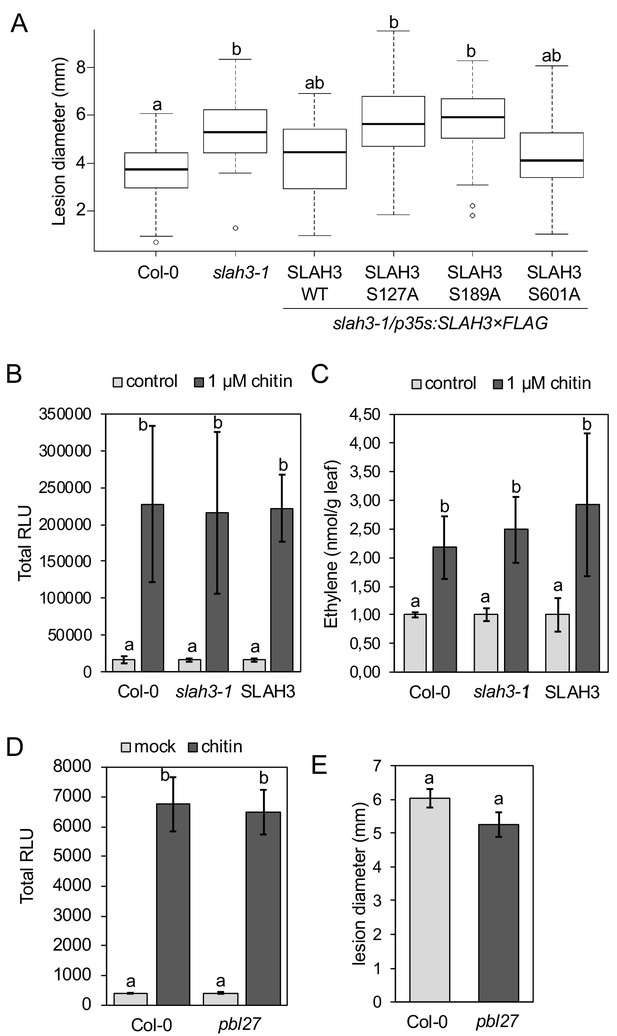
SLAH3 mediates resistance to Botrytis cinerea in leaves.
(A) Lesion diameter measurements in slah3-1 mutants and transgenic slah3-1/35S::SLAH3−3 × FLAG wild type (WT), S127A, S189A and S601A variants of SLAH3. Each four mature leaves of 8 to 10 independent transgenic T1 lines were drop-inoculated with B. cinerea. Lesion diameter was measured with callipers at 3dpi. Values are mean confidence intervals (n = 4; nested one-way ANOVA). Different letters indicate significantly different values at p<0.05. (B) ROS production measured as relative luminescence units (RLU, integral over 30 min) and (C) Ethylene accumulation in Col-0 WT, slah3-1 mutants and transgenic slah3-1/35S::SLAH3−3 × FLAG wild type (SLAH3) upon chitin treatment. Bars show mean values of 3–5 biological replicates for (B) (n = 4–8) and one for (C) (n = 3). Different letters indicate significantly different values at p<0.05. (D) ROS production measured as RLU in Col-0 and pbl27-1 mutants upon treatment with 0.1 mg/ml chitin. Bars show data from two biological replicates (n = 2; nine leaf discs per genotype) + /- SEM. (E) Lesion diameter measurements in Col-0 and pbl27-1 mutants drop-inoculated with Botrytis cinerea. Lesion diameter was measured at three dpi. Bars represent average + /- SEM (n = 5 leaves per each genotype). The experiment was repeated twice with similar results. No significant differences (t-test, p<0.05) were detected.
-
Figure 5—source data 1
Source data for Botrytis infection shown in Figure 5.
- https://doi.org/10.7554/eLife.44474.021
-
Figure 5—source data 2
Source data for ethylene measurements shown in Figure 5.
- https://doi.org/10.7554/eLife.44474.022
-
Figure 5—source data 3
Source data for ROS measurements shown in Figure 5.
- https://doi.org/10.7554/eLife.44474.023
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.44474.024





