Synapse-specific opioid modulation of thalamo-cortico-striatal circuits
Figures
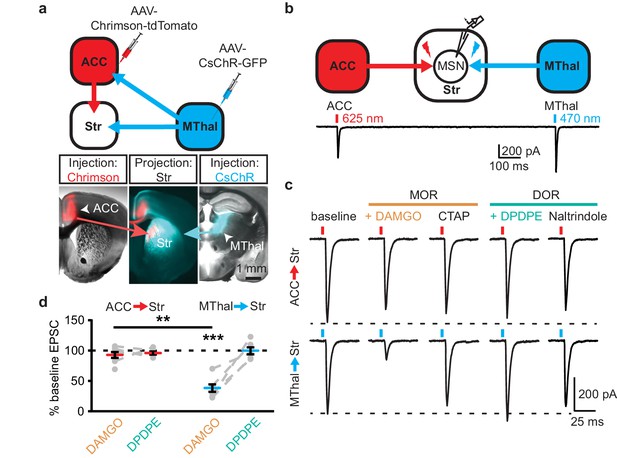
Mu-opioid agonists suppress thalamic but not cortical inputs to single MSNs in the striatum.
(a) Schematic (upper panel) and an acute mouse brain slice example (lower panel) of viral injection design and the axonal projections to the striatum, respectively. Overlaid brightfield and epifluorescent images showing the injection site of Chrimson (left image, red) in the ACC, and CsChR (right image, cyan) in the MThal, and convergent axonal projections from both injections to the DMS (center image). (b) Schematic (upper panel) and representative recordings (lower panel) for optical excitation. (c) Example oEPSCs of individual MSNs evoked by 625 nm (from the ACC, upper traces), and by 470 nm (from the MThal, lower traces) light pulses. The MOR (orange label) agonist DAMGO (1 µM) was perfused followed by the MOR antagonist CTAP (1 µM). Following CTAP, the DOR (teal label) agonist DPDPE (1 µM) was perfused followed by the moderately-selective DOR antagonist naltrindole (0.3 µM). Red bars: 3 ms of 625 nm light stimulation; blue bars: 1 ms of 470 nm light stimulation. (d) Summary data of dual wavelength excitation of the ACC and MThal input oEPSCs recorded from single MSNs. Data are plotted as the percentage of baseline current following exposure to DAMGO or DPDPE for inputs from the ACC and MThal (N = 5, n = 8, Linear mixed model: 3-way interaction, opioid type (mu vs. delta opioid) x input source (ACC vs. MThal) x drug condition (baseline vs. agonist vs. antagonist), F (4,8)=2.938, p=0.091; MThalbaseline x DAMGO: z = 4.738, p<0.001; MThalbaseline x DAMGO vs. ACCbaseline x DAMGO; z = −3.026, p<0.01). Mean ± standard error of the mean. Str: striatum.
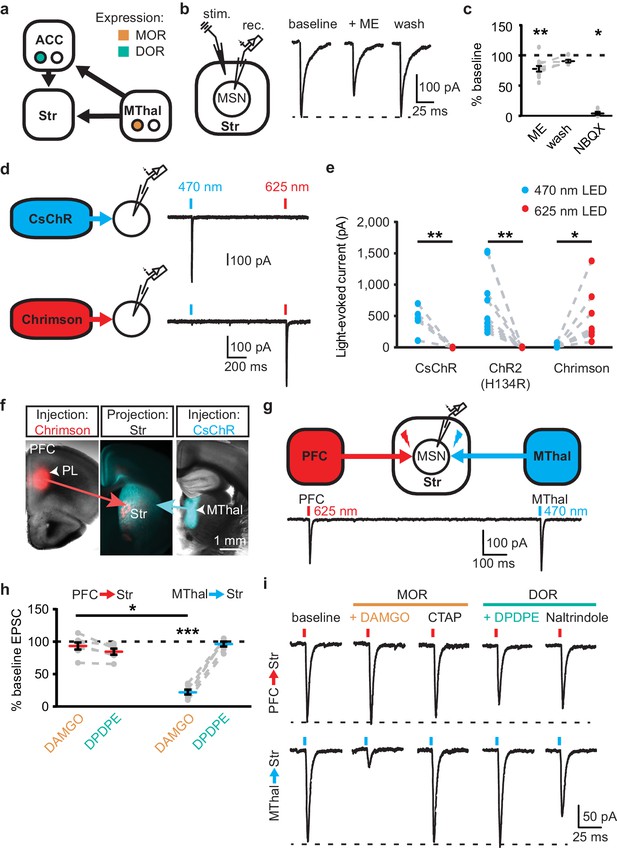
Mu-opioid agonists suppress thalamic, but not cortical inputs.
(a) Schematic of thalamo-cortico-striatal circuits and putative expression pattern of MORs (orange) and DORs (teal) (Erbs et al., 2015; Hunnicutt et al., 2016). (b) Schematic and example traces of electrically-evoked EPSCs from an MSN in the DMS. ME: non-selective opioid agonist [Met5]-enkephalin (3 µM). (c) Summary data of opioid inhibition of EPSCs, as shown in (b), plotted as a percent of peak current in the drug condition relative to the baseline. ME: N = 6, n = 11; wash: N = 3, n = 6, SM = 9.712, p<0.01; baseline vs. ME: W(11) = 1, p<0.01; baseline vs. wash: W(6) = 1, p=0.063; NBQX: N = 4, n = 5; baseline vs. NBQX: W(5) = 0, p<0.05. (d) Independent activation of CsChR and Chrimson using two-wavelength optical excitation. CsChR (blue): virus expressing CsChR injected into the MThal; Chrimson (red): virus expressing Chrimson injected into the ACC. Blue bars: optical excitation with 470 nm light (1 ms, 0.5 mW); red bars: 625 nm light (3 ms, 1.4 mW). (e) Quantification of spectral selectivity of CsChR, ChR2(H134R), and Chrimson plotted as peak optically-evoked EPSC (oEPSC) amplitude following 470 nm (blue) or 625 nm (red) LED optical excitation (CsChR: N = 2, n = 5, I625nm:I470nm = -0.0002 ±. 0081, paired t-test p<0.01; ChR2 (H134R): N = 4, n = 10, I625nm:I470nm = 0.0009 ±. 002, paired t-test p<0.01: Chrimson: N = 5, n = 8, I470nm:I625nm = 0.0423 ± 0.014, paired t-test p<0.05). (f) Overlaid brightfield and epifluorescent images of coronal brain slices showing expression of CsChR in the MThal and axonal projections to the DMS (blue), and expression of Chrimson (red) in the PFC and axonal projections to the DMS (red). (g) Two-wavelength optical excitation elicited EPSCs in response to 470 and 625 nm light in single MSNs when CsChR was injected into the MThal and Chrimson was injected into the PFC. (h) Summary data of DAMGO (1 µM, orange) and DPDPE (1 µM, teal) effects on oEPSCs elicited from the MThal (blue) and PFC (red) using two-wavelength optical excitation protocol. Linear mixed model: 3-way interaction; opioid type (mu vs. delta opioid) x source (PFC vs. MThal) x condition (baseline vs. agonist vs. antagonist): F(4,7) = 2.171, p=0.174, N = 5, n = 7; DAMGO: IMThal; 22.2 ± - 3.6% of baseline, z = 3.497, p<0.001, IPFC; 93.9 ± 5.6%, z = 0.052, p=0.958; DPDPE: IMThal; 96.4 ± 3.9% of baseline, z = 0.087, p=0.931; IPFC: 81.6 ± 3.9%, z = 0.672, p=0.502. MThalbaseline x DAMGO vs. ACCbaseline x DAMGO; z = −2.436, p<0.05. (i) Example traces for summary data shown in (h). *p≤0.05; **p≤0.01; ***p≤0.001. Mean ± standard error of the mean. N: number of animals; n: number of recorded cells; Str: striatum; PL: prelimbic cortex.
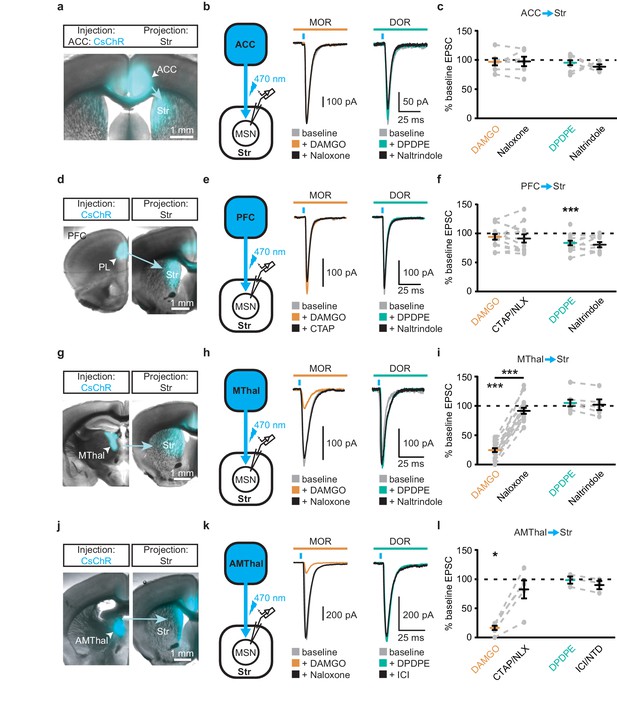
Single channelrhodopsin injections reproduced specific effect of mu-opioid-mediated inhibition of thalamic but not cortical inputs.
Virus expressing CsChR was injected into the ACC (a–c), the PFC (d–f), the MThal (g–i), or the anteromedial thalamus (AMThal) (j–l) as four sets of the experiments. (a, d, g, j) Representative images of the injection and projection for the above four sets of experiments, respectively. (b, e, h, k) Viral infection and optical stimulation schematic and example recordings of four sets of experiments, respectively. oEPSCs were evoked under baseline conditions (gray) and in the presence of DAMGO (1 µM, orange) followed by CTAP or naloxone (NLX) (1 µM, black), or DPDPE (1 µM, teal) followed by naltrindole (NTD) or ICI 174,864 (ICI) (0.3 µM, black). (c, f, i, l) Summary data of effects of DAMGO, CTAP/NLX, DPDPE, and ICI/NTD on oEPSC amplitude for the above four sets of experiments, respectively. (c) DAMGO/naloxone: N = 4/4, n = 7/6, SM = 0.054, p=0.974; DPDPE/naltrindole: N = 5/4, n = 9/5, SM = 4.539, p=0.103. (f) DAMGO/naloxone: N = 9/8, n = 14/12, SM = 2.286, p=0.319. DPDPE/naltrindole: N = 9/6, n = 14/10, SM = 8.227, p<0.05, IDPDPE: 83.4 ± 3.9% of baseline, W(13) = 4, p<0.01. (i) DAMGO/naloxone: N = 15/13, n = 19/16, SM = 25.660, p<0.001, IDAMGO: 24.5 ± 3.3% of baseline, W(19) = 0, p<0.001, DAMGO vs. naloxone, W(12) = 0, p<0.001; DPDPE/naltrindole: N = 7/5, n = 8/5, SM = 2.522, p=0.283. (l) DAMGO/naloxone: N = 5/4, n = 6/5, SM = 9.399, p<0.01, IDAMGO: 16.5 ± 3.7% of baseline, W(6) = 0, p<0.05; DPDPE/naltrindole: N = 2/2, n = 3/3, SM = 4.667, p=0.097. Blue bars: 1 ms of 470 nm light stimulation. Skillings-Mack test followed by paired Wilcoxon signed-ranks test post-hoc analysis. *p≤0.05; **p≤0.01; ***p≤0.001. Mean ± standard error of the mean. SM: Skillings-Mack statistic.
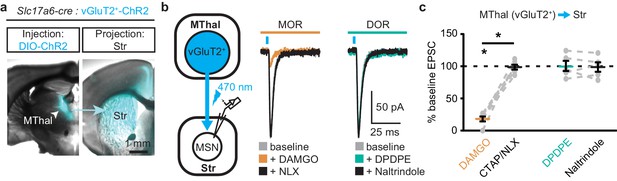
Mu-opioid agonists selectively suppress thalamic inputs from vGluT2-positive thalamic neurons.
(a) Example overlaid brightfield and epifluorescent images showing Cre-dependent expression of ChR2(H134R)-EYFP (cyan) in the MThal (left panel) and axonal projections into the DMS (right panel) following injection of AAV-DIO-ChR2(H134R)-EYFP in the MThal of Slc17a6-Cre mice, which express Cre-recombinase in vGluT2-positive cells. (b) Experimental schematic showing optical stimulation of glutamate inputs in the DMS (left panel). Representative traces of oEPSCs showing effects of MOR agonist DAMGO (1 µM, middle panel, orange) and antagonist naloxone (NLX, 1 µM, middle panel, black), and the DOR agonist DPDPE (1 µM, right panel, teal) and DOR antagonist naltrindole (0.3 µM, right panel, black). Blue bars: 1 ms of 470 nm light stimulation. (c) Summary data of oEPSCs showing effects of MOR agonist DAMGO and antagonist CTAP or NLX (1 µM), and the DOR agonist DPDPE and DOR antagonist naltrindole. DAMGO/(CTAP/NLX): N = 3, n = 6, SM = 9.33, p<0.01; DPDPE/naltrindole, N = 3, n = 5, SM = 0, p=1.0. Skillings-Mack test followed by paired Wilcoxon signed-ranks test post-hoc analysis. Mean ± standard error of the mean. *p≤0.05; SM: Skillings-Mack statistic; Str: striatum.
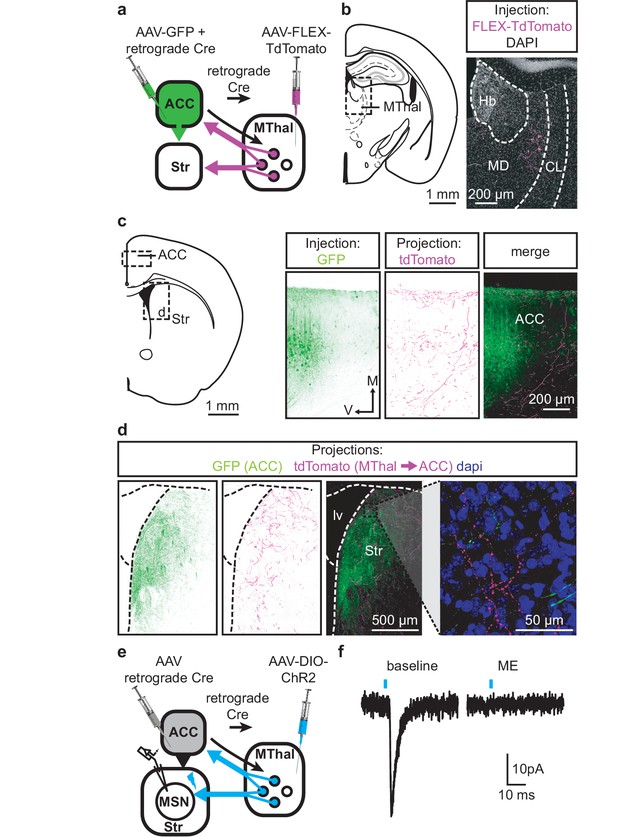
Individual mediodorsal thalamic neurons project to both the DMS and ACC.
(a) Schematic of the rAAV-retro-Cre and Cre-dependent AAV-DIO-TdTomato injections. (b) Fluorescent image of cell bodies expressing TdTomato following injections shown in (a) in the MThal (magenta, right panel), with corresponding mouse brain atlas section (left panel). (c) Representative mouse brain atlas section showing approximate origin of images taken from ACC (left panels) and DMS as shown in (d). Images of the ACC showing AAV-GFP injection site and axons from the MThal (magenta). (d) Images of the DMS showing overlapping axons from both the ACC (green) and MThal (magenta). Rightmost panel shows high magnification image taken at the black box demarcation in the left panel. Cell nuclei are stained with DAPI (blue). (e) Schematic of retrograde rAAV-retro-Cre and Cre-dependent AAV-DIO-ChR2(H134R)-EYFP injections, and recordings of MSNs in the DMS. (f) An example trace of oEPSCs of a MSN in the DMS from a mouse injected as shown in (e). ME: opioid agonist [Met5]-enkephalin. Blue bars: 1 ms of 470 nm light stimulation. V: ventral; M: midline; Hb: habenula; CL: centrolateral thalamus; MD: mediodorsal thalamus; Str: striatum; lv: lateral ventricle. Mouse brain atlas sections from Franklin and Paxinos (2001).
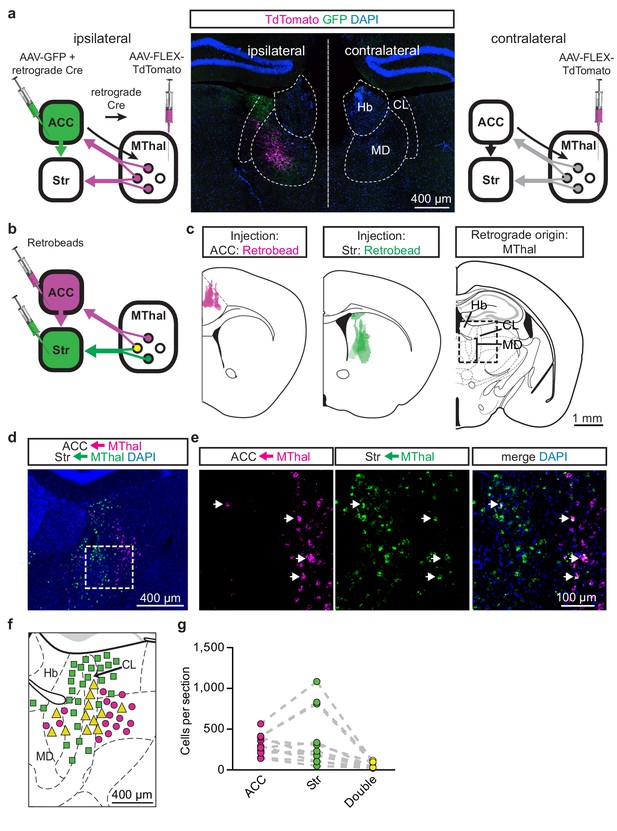
A subset of mediodorsal thalamic neurons send collaterals to both the ACC and DMS.
(a) Experimental diagram in which AAV-FLEX-TdTomato was injected bilaterally into the MThal (left and right panels) and an example image of the labeling (middle panel). A combination of AAV-GFP and rAAV-retro-Cre (retrograde Cre) was injected into the ACC (ipsilateral, left panel). DAPI-stained (blue) coronal brain slice with retrograde labeled thalamic neurons expressing TdTomato (red, amplified with anti-DsRed primary and Alexa 594 secondary antibodies) and GFP-positive axons (green, amplified with anti-GFP primary and Alexa 488 secondary antibodies) originating from the ACC (middle panel). (b) Experimental diagram of fluorescent retrograde bead (retrobead) injections into the ACC and the striatum. (c) Averaged retrobead injection sites are shown in relationship to the mouse brain atlas (left panel, magenta, and center panel, green) as described in (b). The region of interest in which fluorescent retrobeads were imaged and counted in the MThal is shown as the squared box on an mouse brain atlas section (right panel). (d) Example image of retrobeads in the Mthal in ACC- (magenta) and Str-projecting (green) neurons from the area outlined in (c). (e) High magnification images of retrobeads in the MThal from the box outlined in (d). Arrowheads: neurons doubled labeled with both retrobeads (green and magenta). (f) Locations of retrograde labeled neurons surrounding the MThal were mapped onto the mouse brain atlas with estimation. Magenta circles: striatum-projecting neurons; green squares: ACC-projecting neurons; yellow triangles: striatum-projecting and ACC-projecting neurons. Note that the size of the markers is not to scale. (g) Summary graph showing the number of ACC (magenta), Str (green), or double labeled (yellow) retrobead clusters quantified per slice. Hb: habenula; CL: central lateral thalamus; MD: mediodorsal thalamus.
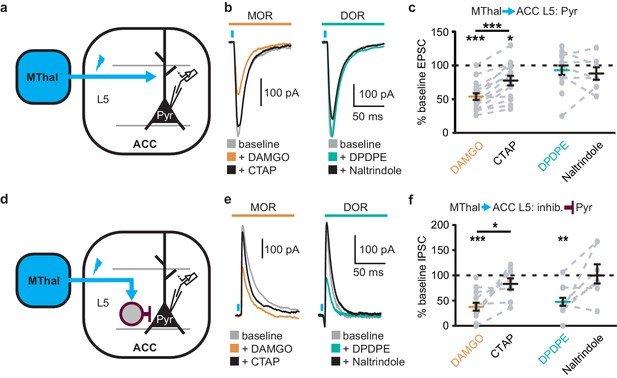
Mu-opioid agonists suppress thalamic inputs to pyramidal neurons in the ACC while delta-opioid agonists suppress cortical feed-forward inhibition in the ACC.
(a–c) oEPSCs of the pyramidal neurons in the ACC elicited by optical stimulation of MThal input. (a) Schematic of ChR2 injection, MThal optical stimulation, and recording of oEPSCs of the layer 5 (L5) pyramidal neurons (Pyr) in the ACC. Blue: ChR2 expression and optical stimulation. (b) Example traces of oEPSCs elicited from optical stimulation of the MThal terminals in the ACC during baseline (gray), application of DAMGO (1 µM, left panel, orange), followed by CTAP (1 µM, left panel, black), or application of DPDPE (1 µM, right panel, teal,) followed by naltrindole (0.3 µM, right panel, black). Blue bars: 1 ms of 470 nm light stimulation. (c) Summary data of oEPSCs of all recording as shown in (b) with responses plotted as a percent of the baseline. DAMGO: N = 13, n = 17; CTAP: N = 12, n = 14; SM = 22.85, p<0.001; DPDPE: N = 11, n = 15; naltrindole: N = 5, n = 7, SM = 0.989, p=0.610. (d–f) oIPSCs of the pyramidal neurons from the MThal optical stimulation, via inhibition through interneurons in the ACC. (d) Schematic of ChR2 injection, MThal optical stimulation, and recording of the pyramidal neurons in the ACC via feed-forward inhibition. Blue: ChR2 expression and optical stimulation; magenta: outline of an interneuron. (e) Example traces of oIPSCs elicited from optical stimulation of the MThal terminals in the ACC during baseline (gray), application of DAMGO (1 µM, left panel, orange) followed by CTAP (1 µM, left panel, black), or application of DPDPE (1 µM, right panel, teal) followed by naltrindole (0.3 µM, right panel, black). (f) Summary data of oIPSCs for all recordings as in (b) with responses plotted as a percent of the baseline. DAMGO: N = 9, n = 14; CTAP, N = 7, n = 8, SM = 15.68, p<0.001; DPDPE: N = 6, n = 12; naltrindole: N = 5, n = 7, SM = 7.426, p<0.05. Skillings-Mack test followed by paired Wilcoxon signed-ranks test post-hoc analysis. *p≤0.05; **p≤0.01; ***p≤0.001. Mean ± standard error of the mean. SM: Skillings-Mack statistic; Blue bars: 1 ms of 470 nm light stimulation.
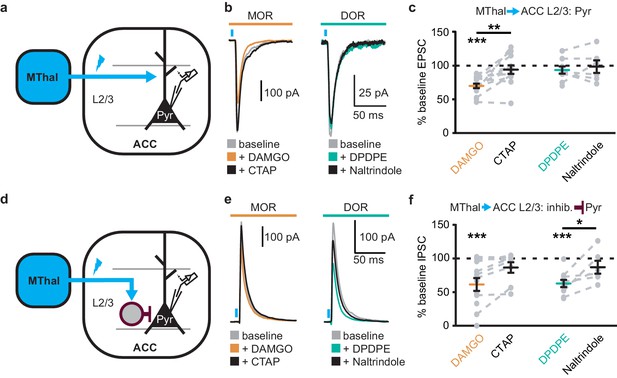
Opioid inhibition of synaptic currents onto layer 2/3 pyramidal neurons in the ACC.
(a) Experimental diagram in which ChR2(H134R) (blue) was injected in the MThal, and oEPSCs were recorded from layer 2/3 (L2/3) pyramidal neurons (Pyr) in the ACC. (b) Example recordings of oEPSCs elicited from MThal terminals in the ACC. oEPSC amplitudes in response to DAMGO (1 µM, orange) followed by CTAP (1 µM, black) (left panel), and oEPSC amplitudes in response to DPDPE (1 µM, teal) followed by naltrindole (0.3 µM, black) (right panel). (c) Summary data of oEPSC amplitudes as described in (b), plotted as a percent of baseline. DAMGO: N = 8, n = 15; CTAP: N = 8, n = 12, SM = 18.148, p<0.001, IDAMGO: 69.3 ± 3.3% of baseline, W(15) = 0, p<0.001; DAMGO vs. CTAP, W(12) = 5, p<0.01; DPDPE: N = 7, n = 9; naltrindole: N = 6, n = 6, SM = 1.866, p=0.393. (d) Experimental diagram in which ChR2(H134R) (blue) was injected in the MThal, and oIPSCs were recorded from L2/3 pyramidal neurons in the ACC. (e) Example recordings of oIPSCs elicited from MThal terminals in the ACC under baseline conditions, in response to DAMGO and CTAP, or DPDPE and naltrindole as in (b). (f) Summary data of oIPSC amplitudes as described in (c, e), plotted as a percent of baseline. DAMGO: N = 7, n = 12; CTAP: N = 6, n = 8; SM = 8.882, p<0.05, IDAMGO: 61.2 ± 9.5% of baseline, W(12) = 1, p<0.001; DPDPE: N = 7, n = 10; naltrindole: N = 5, n = 6, SM = 13.329, p<0.01, IDPDPE: 62.7 ± 5.4% of baseline, W(10) = 0, p<0.01, DAMGO vs. CTAP, W(6) = 0, p<0.05. Skillings-Mack test followed by paired Wilcoxon signed-rank test post-hoc analysis. *p≤0.05; **p≤0.01; ***p≤0.001. Mean ± standard error of the mean. inhib.: inhibitory neuron; SM: Skillings-Mack statistic.
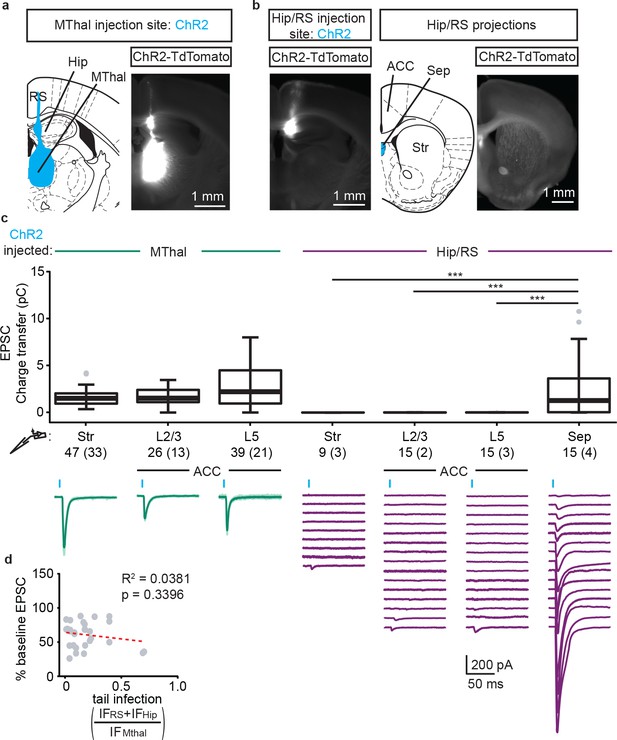
Verification of origins of optically-evoked response.
(a) Representative fluorescent image (right panel) of a typical MThal ChR2 injection in which virus was injected in the MThal while also infecting a ‘tail’ region including hippocampus (Hip) and retrosplenial cortex (RS). The fluorescence at the injection site is saturated to reveal the ‘tail’ region. Schematic of ChR2(H134R)-TdTomato expression shown in blue on a section of the mouse brain atlas (left). (b) Example fluorescent image of ChR2(H134R)-TdTomato expressed only in the ‘tail’ region and corresponding image of projection areas (right) that highlight projections in the septum (Sep) but lack of visible projections in the ACC and striatum following Hip/RS ‘tail’ injection. (c) Charge transfer of EPSCs (upper panel) recorded in striatal MSNs (Str), layer 2 and 3 (L2/3) or layer 5 (L5) pyramidal neurons following optical stimulation in mice injected with ChR2 in either MThal or Hip/RS, as well as Sep as a positive control for the Hip/RS injections. Hip/RS ‘tail’ injections resulted in only occasional small amplitude oEPSCs in the Str and ACC. Kruskal-Wallis test, H = 69.654, df = 6, p<0.001; post-hoc Dunn’s test, Bonferonni-corrected. ***p≤0.001. Box plot with median, 1st and 3rd quartile, and whiskers represent 1.5 times interquartile range. Representative average recordings (lower panel, dark green, average of 10 single trials) from the MSNs, L2/3 and L5 neurons upon optical stimulation of ChR2(H134R) injected in the MThal as shown in (a). Averaged oEPSCs (lower panel, purple, average of 10 single trials) of all individual neurons recorded in Hip/RS injected mice, as shown in (b). Blue bars: 1 ms of 470 nm light stimulations. (d) Plot of the DAMGO effect on baseline oEPSC amplitudes recorded in all L2/3 and L5 pyramidal neurons, as a function of the ratio of integrated fluorescence intensity (IF) of ChR2(H134R)-TdTomato fluorescence in RS +Hip (IFRS +IFHip) relative to MThal (IFMThal) in all the Mthal injected groups. There was no correlation between the amount of ‘tail’ infection and the degree of oEPSC inhibition by DAMGO. Linear regression model, R2 = 0.034, F (1, 27)=0.945, p=0.340.
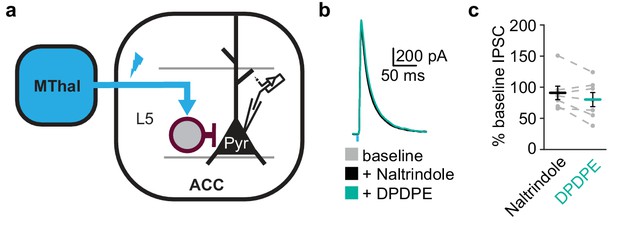
Opioid agonists induced suppression of inhibitory synaptic currents onto layer 5 pyramidal neurons in the ACC is mediated via delta-opioid receptors.
(a) Experimental diagram in which ChR2(H134R) (blue) was injected in the MThal, and feed-forward GABAergic transmission (oIPSCs) was recorded from layer 5 (L5) pyramidal neurons (Pyr) in the ACC. (b) Example recordings of oIPSCs elicited from MThal terminals in the ACC during baseline (gray), in response to naltrindole (1 µM, black) and followed by DPDPE (1 µM, teal). Blue bar: 1 ms of 470 nm light stimulation. (c) Summary data of oIPSCs as described in a, plotted as percent of baseline (N = 3, n = 7), SM = 3.714, p = 0.156; Inaltrindole: 90.9± 10.8 % of baseline, IDPDPE: 80.1 ± 11.4 % of baseline; baseline vs. naltrindole, W(7) = 6, p = 0.219, naltrindole vs. DPDPE, W(7) = 7, p = 0.297. Skillings-Mack test followed by paired Wilcoxon signed-rank test post-hoc analysis. Mean ± standard error of the mean. SM: Skillings-Mack statistic.
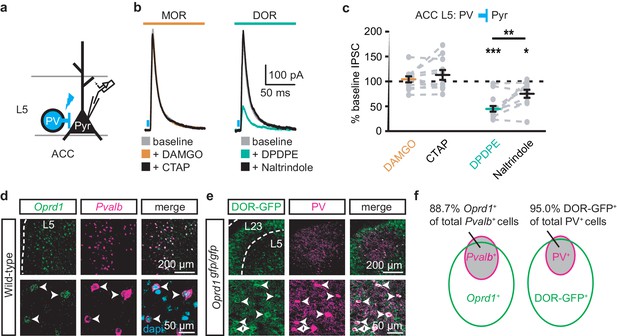
DORs expressed on PV-positive interneurons suppress oIPSCs onto layer five pyramidal neurons.
(a) Schematic of ChR2 expression and recording of oIPSCs of parvalbumin-positive interneurons (PV) to layer 5 (L5) pyramidal neurons (Pyr) in the ACC of Pvalb-cre+/-;Ai32+/- mice. Blue: ChR2 expression and optical stimulation. (b) Example traces of oIPSCs during baseline (gray), application of DAMGO (1 µM, left panel, orange) and followed by CTAP (1 µM, left panel, black), or application of DPDPE (1 µM, right panel, teal) followed by naltrindole (0.3 µM, right panel, black). Blue bars: 1 ms of 470 nm light stimulation. (c) Summary data of PV interneurons to pyramidal neuron oIPSCs for all recording as in (b). Responses plotted as a percent of the baseline. DAMGO/CTAP: N = 5, n = 11, SM = 0.005, p=0.998; DPDPE: N = 8, n = 15; naltrindole: N = 5, n = 10, SM = 19.60, p<0.001. (d) In-situ hybridization in wild-type mouse brain sections containing the ACC stained with probes against mRNA coding for Oprd1 (Oprd1, left panel, green) and parvalbumin (Pvalb, middle panel, magenta) and overlaid with DAPI (right panel, cyan). (e) Immunohistochemistry of brain sections of the ACC from a DOR-GFP knockin mouse and stained with anti-GFP antibodies (left panel, green), and anti-parvalbumin antibodies (middle panel, magenta), and overlaid (right panel). (f) Venn diagram quantifying overlap of Oprd1-positive and Pvalb-positive (left panel, N = 2, n = 8), and DOR-GFP-positive and PV-positive cells in the mouse ACC (right panel, N = 2, n = 15). Skillings-Mack test followed by paired Wilcoxon signed-ranks test post-hoc analysis. *, p<0.05; **, p<0.01; ***p<0.001. Mean ± standard error of the mean. SM: Skillings-Mack statistic.
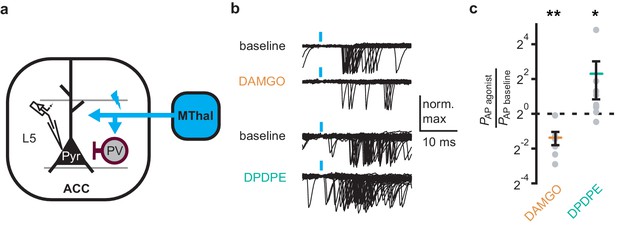
DOR activation results in increased cortical excitability.
(a) Schematic of ChR2 expression and recording of optically-evoked action potentials (APs, recorded as action currents using voltage-clamp mode) using a loose cell-attached recording configuration. Blue: ChR2 expression and optical stimulation; magenta: outline of a parvalbumin-positive interneuron (PV). (b) Example traces of 50 trails from a single layer 5 (L5) pyramidal neuron (Pyr) in which APs were evoked by optical stimulation (blue bars) under baseline conditions, or in the presence of DAMGO (1 µM, orange), or DPDPE (1 µM, teal). (c) Summary data plotted on a log2 scale for action potential firing probability (PAP) in the presence of drugs (PAP agonist) relative to baseline (PAP baseline). Paired t-test; *p≤0.05; **p≤0.01. DAMGO: N = 3, n = 7; DPDPE: N = 5, n = 8.
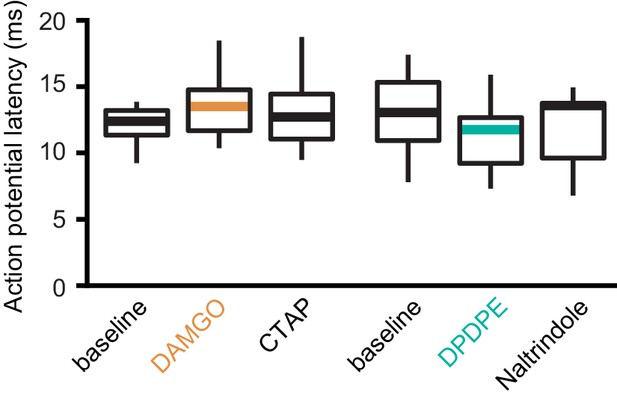
Latencies of thalamocortical-evoked action potentials in ACC pyramidal neurons.
The average latency to the first action potential following optical stimulation of the MThal inputs recorded in layer 5 (L5) pyramidal neurons (Pyr), using loose cell-attached configuration. There were no significant differences in the latency to the first spike. DAMGO: N = 3, n = 7; CTAP: N = 3, n = 7, SM = 2, p = 0.368; DPDPE: N = 4, n = 8; naltrindole: N = 4, n = 8, SM = 1.75, p = 0.417. Box plot with median, 1st and 3rd quartile, and whiskers represent 1.5 times interquartile range. SM: Skillings-Mack statistic.
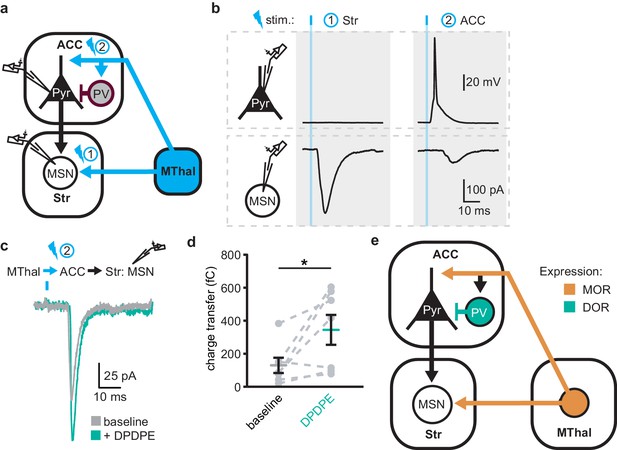
DOR activation disinhibits thalamo-cortico-striatal circuits.
(a) Schematic of ChR2 expression and dual recordings in layer 5 (L5) pyramidal neurons (Pyr) in the ACC and MSNs in the DMS. Blue: ChR2 expression and optical stimulation; magenta: outline of a parvalbumin-positive interneuron (PV). 470 nm light stimulation locations are shown as 1 and 2. (b) Example traces of current-clamp recording from a L5 pyramidal neuron in the ACC (upper panels), and voltage-clamp recording from a MSN in the DMS (lower panels) in response to light stimulations at location 1 (Str) and location 2 (ACC). (c) Example traces of poly-synaptic current evoked by optical stimulation of MThal terminals in the ACC (location 2) while recording from an MSN in the DMS during baseline (gray), and in presence of DPDPE (1 µm, teal). (d) Summary data of the charge transfer of poly-synaptic oEPSCs in the MSNs evoked by optical stimulation of location 2 in the ACC. Paired Wilcoxon signed-ranks test. *p≤0.05; Gray: baseline; teal: DPDPE. (e) Summary schematic depicting a part of the affective and motivational pain circuit consisting of the MThal, ACC and DMS. Str: striatum.
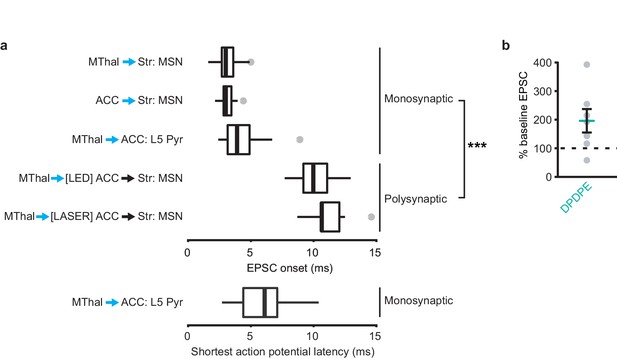
EPSCs and action potential latencies within the thalamo-cortico-striatal circuits.
(a) Average onset time for monosynaptic and polysynaptic EPSCs using either full field (LED stimulation: all monosynaptic data, and [LED] ACC polysynaptic data), or focal (laser stimulation: [LASER] ACC polysynaptic data) optical stimulation. Mann-Whitney U test, U = 15, p<0.001. ***p≤0.001. Box plot with median, 1st and 3rd quartile and whiskers represent 1.5 times interquartile range. Lower panel: average of shortest action potential onset latency in L5 pyramidal neurons following stimulation of MThal terminals. Stimulation intensity was set to elicit ~50% spike probability out of 50 trials. (b) EPSC amplitudes measured in MSNs in the striatum (Str) evoked by optical stimulation of MThal terminals in the ACC in the presence of DPDPE, plotted as a percentage of baseline EPSC. N = 6, n = 7, paired Wilcoxon signed-rank test, W(7) = 3, p=0.078. Mean ± standard error of the mean.
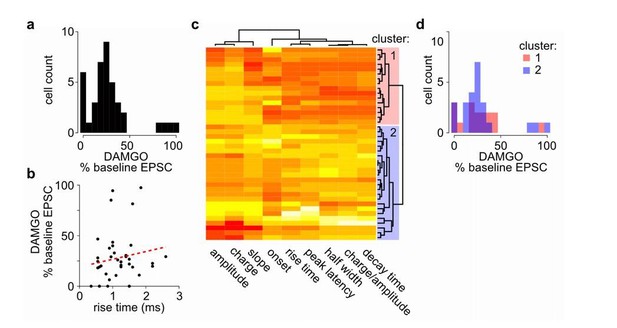
No clear bi-modal or two population effects detected in MOR-mediated suppression of thalamostriatal inputs to the MSNs in the DMS.
(a) Histogram of cell distribution based on effects of mu-opioid agonist DAMGO on optically-evoked excitatory postsynaptic current (EPSC) amplitude recorded in the MSNs; n = 40 neurons from 25 mice. The observed DAMGO effects had a non-normal distribution (Shapiro-Wilk normality test, W = 0.8219, p < 0.001), but showed no clear bi-modal distribution of two roughly equal populations. (b) Plot of the DAMGO effects on baseline oEPSC amplitude recorded from MSNs as a function of risetime of the baseline EPSC. There was no correlation between EPSC risetime and degree of EPSC inhibition by DAMGO. Linear regression model (red dash line), R2 = 0.018, F (1, 38) = 0.677, p = 0.416. (c-d) In case two populations exist within the recorded MSNs, we wanted to test whether the effect of DAMGO on baseline EPSC amplitude was significantly different in the two populations defined based on baseline EPSC parameters. (c) Agglomerative hierarchical cluster analysis based on Pearson correlation between baseline EPSC parameters (columns), and Euclidean distance between recorded MSNs (rows), with heatmap representing centered/standardized values (x-x-σ). Two main clusters were defined (cluster 1: red n = 16 neurons/11 mice, cluster 2: blue n = 23 neurons/17 mice) based on baseline EPSC parameters, representing putative subpopulations among the recorded MSNs. (d) Histograms of cell distributions based on DAMGO effects on baseline recorded from the MSNs defined in cluster 1 or 2, red or blue, respectively. Mann-Whitney U test, U = 204, p = 0.751.
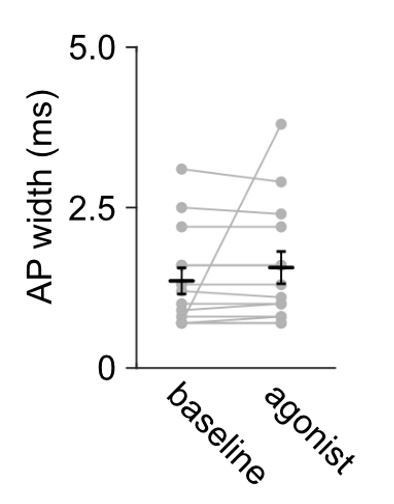
Action potential widths remain stable during cell-attached recording and they are comparable to other studies in the L5 pyramidal neurons on the cortex.
Summary data of median full width at half-maximum of action potential currents recorded in L5 pyramidal neurons of the ACC; n = 14 neurons from 5 mice. The agonists (DAMGO or DPDPE) did not change the recorded action potential current width; Mann-Whitney U test U = 189, p = 0.5320.
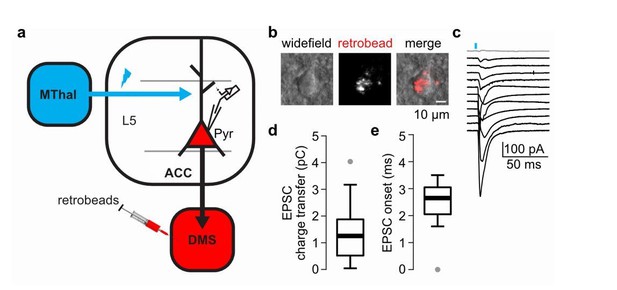
The DMS-projecting L5 pyramidal neurons in the ACC receive direct inputs from the MThal.
(a) Experimental schematic showing retrogradely-transported beads (retrobeads) injection in the DMS to retrogradely-label DMS-projecting pyramidal neurons (Pyr) in the ACC, and AAV2/1-CAG-ChR2(H134R)-tdTomato injection in the MThal to allow for optical stimulation of specific thalamic inputs to the ACC. (b) Representative brightfield (left panel) and fluorescent (center panel) images of an ACC layer 5 (L5) with retrobeads present in the soma (merge, right panel). (c) Averaged optically-evoked excitatory postsynaptic currents (EPSCs, average of 12 single trials) of all individual bead-positive L5 pyramidal neurons recorded in the DMS/MThal injected mice (n = 12 neurons from 4 mice). 11 out of 12 recorded cells (black traces) received inputs from the MThal, defined as peak amplitude > (mean baseline + 5 x standard deviation). Grey trace: non-responding L5 pyramidal neuron. Blue bar: 1 ms of 470 nm light stimulation. (d) Quantification of the EPSC charge transfer of all recorded EPSCs as shown in B. (e) Quantification of EPSC onset time of all recorded EPSCs as shown in B. Box plot with median, first and third quartile and whiskers represent 1.5 times interquartile range.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (M. musculus, C57BL/6J) | wildtype | Jackson Laboratories | Stock # 000664 RRID: IMSR_JAX:000664 | |
| Genetic reagent (M. musculus) | Ai32 (B6;129S-Gt(ROSA)26Sortm32(CAG-COP4*H134R/EYFP)Hze/J) | Jackson Laboratories | Stock # 012569 RRID: IMSR_JAX:012569 | PMID: 22446880 |
| Genetic reagent (M. musculus) | Ai9 (B6.Cg-Gt(ROSA)26Sor tm9(CAG-TdTomato)Hze/J) | Jackson Laboratories | Stock # 007909 RRID: IMSR_JAX:007909 | PMID: 20023653 |
| Genetic reagent (M. musculus) | Pvalb-IRES-Cre (B6.129P2-Pvalbtm1(cre)Arbr/J) | Jackson Laboratories | Stock # 008069 RRID: IMSR_JAX:007909 | PMID: 15836427 |
| Genetic reagent (M. musculus) | Slc17a6-IRES-Cre (B6.Slc17a6tm2(cre)Lowl/J) | Jackson Laboratories | Stock # 016963 RRID: IMSR_JAX:016963 | PMID: 21745644 |
| Genetic reagent (Dependoparvovirus) | AAV2-syn-hChR2 (H134R)-EYFP | UNC virus vector core | NA | |
| Genetic reagent (Dependoparvovirus) | AAV2/1-CAG-hChR2 (H134R)-TdTomato | UPenn Vector Core | Addgene plasmid # 28017 | PMID: 21982373 |
| Genetic reagent (Dependoparvovirus) | AAV2-syn-ChrimsonR-TdTomato (Klapoetke et al., 2014) | UNC virus vector core | NA | PMID: 24509633 |
| Genetic reagent (Dependoparvovirus) | AAV2-syn-CsChR-GFP (Klapoetke et al., 2014) | UNC virus vector core | NA | PMID: 24509633 |
| Genetic reagent (Dependoparvovirus) | AAVrg-pmSyn1-EBFP-cre (Madisen et al., 2015; Tervo et al., 2016) | Addgene | Cat# 51507 | PMID: 25741722, 27720486 |
| Genetic reagent (Dependoparvovirus) | AAV2-FLEX-CAG-TdTomato | UNC virus vector core | NA | |
| Genetic reagent (Dependoparvovirus) | AAV2-SSpEMBOL-Chicken-beta-actin (CBA)-GFP | UNC virus vector core | NA | |
| Genetic reagent (Dependoparvovirus) | AAV1-CAG-hChR2 (H134R)-TdTomato | UPenn Vector Core | Addgene plasmid # 28017 | PMID: 21982373 |
| Genetic reagent (Dependoparvovirus) | AAV2-DIO-EF1alpha-ChR2 (H134R)-EYFP | UNC virus vector core | NA | |
| Antibody | Living Colors DsRed Polyclonal Anibody | Takara Bio/Clontech | Cat# 632496 RRID: AB_10013483 | dilution: 1:500 |
| Antibody | Anti-GFP chicken Polyclonal IgY fraction | Thermo Fisher | Cat# A10262 RRID: AB_2534023 | dilution: 1:500 |
| Antibody | Alexa Fluor 488 goat anti-chicken Polyclonal IgG | Thermo Fisher | Cat# A11039 RRID: AB_2534096 | dilution: 1:750 |
| Antibody | Alexa Fluor 594 goat anti-rabbit Polyclonal IgG (H + L) | Thermo Fisher | Cat# A11012 RRID: AB_2534079 | dilution: 1:750 |
| Antibody | Anti-parvalbumin goat Polyclonal | Swant | Cat# PVG 214 RRID: AB_10000345 | dilution: 1:1000 |
| Antibody | Anit-GFP chicken Polyclonal | Abcam | Cat# ab13970 RRID: AB_300798 | dilution: 1:1000 |
| Chemical compound, drug | Mecamylamine | R and D Systems/Tocris | Cat# 2843 | |
| Chemical compound, drug | Scopolamine | Sigma Aldrich | Cat# S1013 | |
| Chemical compound, drug | SR95531 | Hello Bio | Cat# HB0901 | |
| Chemical compound, drug | Picrotoxin | Hello Bio | Cat# HB0506 | |
| Chemical compound, drug | [Met5]-enkephalin | Sigma Aldrich | Cat# M6638 | |
| Chemical compound, drug | DAMGO | Sigma Aldrich | Cat# E7384 | |
| Chemical compound, drug | CTAP | R and D Systems/Tocris | Cat# 1560 | |
| Chemical compound, drug | Naloxone | Abcam | Cat# ab120074 | |
| Chemical compound, drug | DPDPE | Sigma Aldrich | Cat# E-3888 | |
| Chemical compound, drug | Naltrindole | Sigma Aldrich | Cat# N-115 | |
| Chemical compound, drug | ICI 174,864 | R and D Systems/Tocris | Cat# 0820 | |
| Chemical compound, drug | MK801 | Hello Bio | Cat# HB0004 | |
| Chemical compound, drug | CPP | R and D Systems/Tocris | Cat# 0173 | |
| Chemical compound, drug | CGP 55845 | R and D Systems/Tocris | Cat# 1248 | |
| Chemical compound, drug | Bestatin | Sigma Aldrich | Cat# B8385 | |
| Chemical compound, drug | Thiorphan | Sigma Aldrich | Cat# T6031 | |
| Chemical compound, drug | MPEP | R and D Systems/Tocris | Cat# 1212 | |
| Chemical compound, drug | Red Retrobeads IX | Lumafluor Inc | Cat# R180 | |
| Chemical compound, drug | Green Retrobeads IX | Lumafluor Inc | Cat# R180 | |
| Chemical compound, drug | DNQX | Sigma Aldrich | Cat# D0540 | |
| Chemical compound, drug | QX314 | R and D Systems/Tocris | Cat# 2313 | |
| Commercial assay, kit | RNAscope Multiplex Fluorescent Assay | Advanced Cell Diagnostics | Cat# 320850 | |
| Commercial assay, kit | RNAscope Probe- Mm-Oprd1 | Advanced Cell Diagnostics | Cat# 427371 | |
| Commercial assay, kit | RNAscope Probe- Mm-Pvalb-C2 | Advanced Cell Diagnostics | Cat# 421931-C2 | |
| Software, algorithm | FIJI 1.49b | Wayne Rasband, NIH | PMID: 22743772 | |
| Software, algorithm | AxoGraph 1.4.4 | Axograph | ||
| Software, algorithm | Microsoft Excel 2011 | Microsoft Corp. | ||
| Software, algorithm | Illustrator CS5 | Adobe Systems | ||
| Software, algorithm | Photoshop CS5 | Adobe Systems | ||
| Software, algorithm | Prism 6 | GraphPad | ||
| Software, algorithm | Imaris | Bitplane | ||
| Software, algorithm | Zen | Zeiss | ||
| Software, algorithm | Chart 5 | AD Instruments | ||
| Software, algorithm | Matlab r2007b and r2018a | Mathworks | ||
| Software, algorithm | R (3.5.0) | R Development Core Team | ||
| Software, algorithm | Ephus | Vidrio Technologies, LLC | ||
| Software, algorithm | Custom data analysis software | https://gitlab.com/maolab/opi_syn_circuit.git | Data and analysis scripts related to quantification in manuscript and rebuttal |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.45146.016





