Binding and transport of D-aspartate by the glutamate transporter homolog GltTk
Abstract
Mammalian glutamate transporters are crucial players in neuronal communication as they perform neurotransmitter reuptake from the synaptic cleft. Besides L-glutamate and L-aspartate, they also recognize D-aspartate, which might participate in mammalian neurotransmission and/or neuromodulation. Much of the mechanistic insight in glutamate transport comes from studies of the archeal homologs GltPh from Pyrococcus horikoshii and GltTk from Thermococcus kodakarensis. Here, we show that GltTk transports D-aspartate with identical Na+: substrate coupling stoichiometry as L-aspartate, and that the affinities (Kd and Km) for the two substrates are similar. We determined a crystal structure of GltTk with bound D-aspartate at 2.8 Å resolution. Comparison of the L- and D-aspartate bound GltTk structures revealed that D-aspartate is accommodated with only minor rearrangements in the structure of the binding site. The structure explains how the geometrically different molecules L- and D-aspartate are recognized and transported by the protein in the same way.
https://doi.org/10.7554/eLife.45286.001Introduction
Mammalian excitatory amino acid transporters (EAATs) are responsible for clearing the neurotransmitter glutamate from the synaptic cleft (for review see Grewer et al., 2014; Takahashi et al., 2015; Vandenberg and Ryan, 2013). EAATs are secondary transporters that couple glutamate uptake to co-transport of three sodium ions and one proton and counter-transport of one potassium ion (Levy et al., 1998; Owe et al., 2006; Zerangue and Kavanaugh, 1996). EAATs transport L-glutamate, L- and D-aspartate with similar affinity (Arriza et al., 1994).
D-aspartate is considered as a putative mammalian neurotransmitter and/or neuromodulator (Brown et al., 2007; D'Aniello et al., 2011; Spinelli et al., 2006) (reviewed in D'Aniello, 2007; Genchi, 2017; Ota et al., 2012). Such a role is also proposed for L-aspartate (Cavallero et al., 2009), however this is still a matter of debate (Herring et al., 2015). Both stereoisomers bind to and activate N-methyl-D-aspartate receptors (NMDARs) (Patneau and Mayer, 1990) and might be involved in learning and memory processes (reviewed in Errico et al., 2018; Errico and Usiello, 2017; Katane and Homma, 2011; Ota et al., 2012).
Although it is well established that EAATs take up D-aspartate (Arriza et al., 1994; Gundersen et al., 1993), structural insight in the binding mode of the enantiomer is lacking. The best structurally characterized members of the glutamate transporter family are the archeal homologs GltPh and GltTk (Akyuz et al., 2015; Boudker et al., 2007; Guskov et al., 2016; Jensen et al., 2013; Reyes et al., 2013; Reyes et al., 2009; Scopelliti et al., 2018; Verdon et al., 2014; Verdon and Boudker, 2012; Yernool et al., 2004), which share 32–36% sequence identity with eukaryotic EAATs (Jensen et al., 2013; Slotboom et al., 1999; Yernool et al., 2004). In contrast to EAATs, GltPh and GltTk are highly selective for aspartate over glutamate, and couple uptake only to co-transport of three sodium ions (Boudker et al., 2007; Groeneveld and Slotboom, 2010; Guskov et al., 2016). Despite these differences, the amino acid residues in the substrate-binding sites of mammalian and prokaryotic glutamate transporters are highly conserved (Boudker et al., 2007; Jensen et al., 2013). The first structures of human members of the glutamate transporter family (Canul-Tec et al., 2017; Garaeva et al., 2018), showed that the substrate-binding sites are indeed highly similar among homologs (Figure 2—figure supplement 1).
Here, we present the structure of GltTk with the enantiomeric substrate D-aspartate. The crystal structure was obtained in the outward-facing state with the substrate oriented in a very similar mode as L-aspartate, showing that the two enantiomers bind almost identically regardless of the mirrored spatial arrangement of functional groups around the chiral Cα atom.
Results
Affinity of D-aspartate and stoichiometry of sodium binding to GltTk
Using Isothermal Titration Calorimetry (ITC), we determined the binding affinities of D-aspartate to GltTk in the presence of varying concentrations of sodium ions (Figure 1A, Table 1). The affinity of the transporter for D-aspartate was strongly dependent on the concentration of sodium, similar to what has been reported for L-aspartate binding to GltPh and GltTk (Boudker et al., 2007; Hänelt et al., 2015; Jensen et al., 2013; Reyes et al., 2013). At high sodium concentration (500 mM), the Kd values of GltTk for D- and L-aspartate binding level off to 374 ± 30 nM and 62 ± 3 nM, respectively. The ΔH values for binding of both substrates were favorable, with a more negative value of ~1 kcal mol−1 for L-aspartate, indicating a better binding geometry for L- than for D-aspartate. For both substrates, the ΔS contribution was unfavorable (Table 1). When plotting the observed Kd values for L- and D-aspartate against the sodium concentration (on logarithmic scales), the slopes of both curves in the lower limit of the sodium concentration are close to −3, indicating that binding of both compounds is coupled to the binding of three sodium ions (Boudker et al., 2007; Lolkema and Slotboom, 2015) (Figure 1B).

Binding and transport of D-aspartate by GltTk.
(A) ITC analysis of D-aspartate binding to GltTk in presence of 300 mM NaCl (Kd of 0.47 ± 0.17 µM). Insets show no D-aspartate binding in absence of NaCl. (B) Sodium and aspartate binding stoichiometry. Logarithmic plot of Kd values (nM) for L-aspartate (black squares; slope is −2.8 ± 0.4; taken for reference from Guskov et al., 2016) and D-aspartate (gray circles; slope is −2.9 ± 0.2) against logarithm of NaCl concentration (mM). The negative slope of the double logarithmic plot (red line) in the limit of low sodium concentrations indicates the number of sodium ions that bind together with aspartate. Error bars represent the ±SD from at least three independent measurements. (C) GltTk transport rate of D-aspartate in presence of 100 mM NaCl. The solid line reports the fit of the Michaelis-Menten model to the data revealing a Km value of 1.1 ± 0.11 µM. Error bars represent the ±SD from duplicate experiments. (D) Determination of Na+ : aspartate coupling stoichiometry in GltTk using equilibrium potential measurement. The uptake or efflux of radiolabeled aspartate was determined by comparing the lumenal radioactivity associated with the liposomes after 2 min of incubation with the radioactivity initially present (Δcpm). Gray circles and black squares show the measurements for D- and L-aspartate, respectively. The solid and dashed lines are the best linear regression for the D- and L-aspartate data, respectively. The 95% confidence interval for D-aspartate is displayed by gray curves. Numbers in parentheses are the coupling stoichiometries expected to give zero flux conditions for each membrane voltage. Error bars represent the ± SD obtained in five replicates.
-
Figure 1—source data 1
Final concentrations of internal and external buffer used in each reversal potential experiment after diluting the proteoliposomes.
Proteoliposomes were loaded with 20 mM HEPES/Tris, pH 7.5, 200 mM NaCl, 10 μM L- or D-aspartate, 50 mM KCl and diluted 20 fold in buffer containing 20 mM HEPES/Tris, pH 7.5, 200 mM NaCl, 35.0/26.4/19.2 mM CholineCl, 0/11.1/18.4 mM KCl in the presence of 3 µM valinomycin.
- https://doi.org/10.7554/eLife.45286.003
Thermodynamic parameters of D- and L-aspartate binding at high (300 mM) and low (75 mM) Na+ concentration.
https://doi.org/10.7554/eLife.45286.004| Substrate/ Na+ | Kd (µM) | ΔH (cal mol−1) | ΔS (cal mol−1 K−1) |
|---|---|---|---|
| L-aspartate/300 mM NaCl | 0.12 ± 0.04 | −1.61 (±0.08) x 104 | −22.1 ± 2.2 |
| D-aspartate/300 mM NaCl | 0.47 ± 0.17 | −1.48 (±0.11) x 104 | −20.6 ± 3.6 |
| L-aspartate/75 mM NaCl | 1.04 ± 0.39 | −1.22 (±0.13) x 104 | −13.2 ± 5.2 |
| D-aspartate/75 mM NaCl | 5.66 ± 1.59 | −1.14 (±0.41) x 104 | −14.3 ± 14.3* |
-
*At low Na+ concentrations high errors prevented accurate measuring of ΔS values.
To test whether D-aspartate is a transported substrate, purified GltTk was reconstituted into proteoliposomes and uptake of [3H]-D-aspartate was assayed. GltTk catalyzed transport of the radiolabeled substrate into the proteoliposomes. The Km for transport was 1.1 ± 0.11 μM at a sodium concentration of 100 mM (Figure 1C). This value is comparable to the Km for L-aspartate uptake under the same conditions (0.75 ± 0.17 μM). The stoichiometry Na+: D-aspartate was determined by flux measurements of radiolabeled D-aspartate at different membrane voltages (Fitzgerald et al., 2017). Depending on the concentrations of Na+ and D-aspartate on either side of the membrane, the imposed voltages either lead to flux of radiolabeled D-aspartate across the membrane (accumulation into or depletion from the lumen), or does not cause net flux (when the voltage equals the equilibrium potential) (Fitzgerald et al., 2017). The equilibrium potentials for different possible stoichiometries are calculated by:
where n and m are the stoichiometric coefficients for Na+ and substrate S, respectively. Membrane voltages were chosen that would match the equilibrium potential for stoichiometries of 2:1 (−78 mV), 3:1 (−39 mV) or 4:1 (−26 mV), and flux of radiolabeled D-aspartate was measured (Figure 1D). At −78 mV D-aspartate was taken up into the lumen; at −26 mV it was released from the liposomes; and at −39 mV there was little flux. From these data, we conclude that D-aspartate is most likely symported with three sodium ions. However, the flux was not exactly zero at the calculated equilibrium potential of −39 mV for 3:1 stoichiometry. This small deviation could be caused by systematic experimental errors, or by leakage or slippage (Parker et al., 2014; Shlosman et al., 2018). To exclude that it was caused specifically by D-aspartate, we repeated the experiment using radiolabeled L-aspartate. The equilibrium potentials for the experiments using D- and L-aspartate were identical, showing that the two stereoisomers use the same coupling stoichiometry.
Similar mode of enantiomers binding
We determined a crystal structure of GltTk in complex with D-aspartate at 2.8 Å resolution (Figure 2A,B). The obtained structure is highly similar to the previously described GltTk and GltPh structures with the transport domains in the outward-oriented occluded state. Comparison of the GltTk structures in complex with L- and D-aspartate revealed a highly similar binding mode of the substrates with analogous orientation of amino and carboxyl groups. Despite the impossibility to superimpose two enantiomers, D- and L-aspartate are capable of forming almost identical hydrogen bonding networks with conserved amino acid residues of the substrate-binding site (Figure 2C). There are only small changes in the positions of the Cα atoms and Cβ carboxyl groups due to the constitutional differences. However, this divergence leads to only minor changes in the interaction network, consistent with the comparable Kd and ΔH values determined by ITC (Table 1).
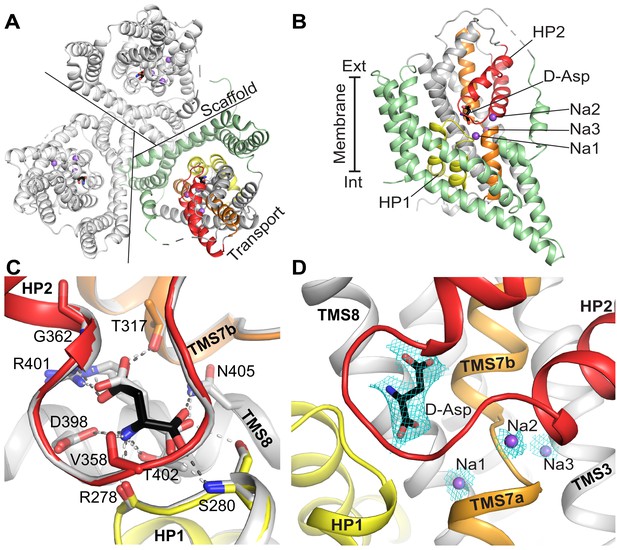
The crystal structure of GltTk with D-aspartate.
The model contains one protein molecule in the asymmetric unit with the substrate present in each protomer of the homotrimer. (A) Cartoon representation of the homotrimer viewed from the extracellular side of the membrane. Lines separate protomers. Each protomer consists of the scaffold domain (pale green) and the transport domain. In the transport domain HP1 (yellow), HP2 (red), TMS7 (orange) are shown. D-aspartate is shown as black sticks and Na+ ions as purple spheres. Like in most GltPh structures a part of the long flexible loop 3–4 between the transport and scaffold domain is not visible. It is indicated by a dashed connection. (B) A single protomer is shown in the membrane plane. (C) Comparison of the substrate-binding site of GltTk in complex with L-aspartate (gray; PDB code 5E9S) and D-aspartate (black). Cartoon representation; substrates and contacting amino acid residues are shown as sticks; hydrogen bonds are shown as dashed lines. The GltTk structures with D- and L-aspartate can be aligned with Cα-RMSD = 0.38 Å for the three transport domains. (D) Composite omit map (cyan mesh) for D-aspartate (contoured at 1σ) and sodium ions (2σ) calculated using simulated annealing protocol in Phenix (Terwilliger et al., 2008). Color coding in all panels is the same.
Three peaks of electron density (Figure 2D; Figure 2—figure supplement 2) located at the same positions as three sodium ions in the GltTk complex with L-aspartate (Guskov et al., 2016) most probably correspond to sodium ions, consistent with a 3:1 Na+: D-aspartate coupling stoichiometry (Figure 1B,D).
Discussion
Most proteins selectively bind a single stereoisomer of their substrates (for a review see Nguyen et al., 2006). On the other hand, some proteins are able to bind different stereoisomers of a ligand, which is believed to be possible due to different binding modes, because enantiomers cannot be superimposed in the three-dimensional space and thus cannot interact with the binding site identically.
Based on three- and four-point attachment models (Easson and Stedman, 1933; Mesecar and Koshland, 2000; Ogston, 1948) it has been suggested that stereoisomers can bind in the same site but with significant differences. This hypothesis was supported by crystal structures of enzymes with different enantiomeric substrates (Brem et al., 2016; Sabini et al., 2008), including enantiomeric amino acids (Aghaiypour et al., 2001; Bharath et al., 2012; Driggers et al., 2016; Temperini et al., 2006). In contrast, the binding poses of enantiomers in some other enzymes are remarkably similar, for instance in aspartate/glutamate racemase EcL-DER, where active site forms pseudo-mirror symmetry (Liu et al., 2016).
To our knowledge GltTk is the first amino acid transporter for which the binding of enantiomeric substrates has been characterized. The only other transporter for which structures have been determined in the presence of D- and L-substrates is the sodium-alanine symporter AgcS. However, in that case, limited resolution prevented determination of the absolute orientation of bound enantiomers (Ma et al., 2019). In the substrate-binding site of GltTk, L- and D-aspartate take similar poses leading to almost identical networks of contacts. Since mirror imaged substrates inevitably have differences in angles between donors and acceptors of hydrogen bonds, the binding affinities are not identical, with 4–6 times higher Kd of the GltTk-D-aspartate complex in comparison with L-aspartate (Table 1). Similar differences in binding affinities between these enantiomers were also found for the GltPh homologue (Boudker et al., 2007). The higher Kd values for the D-aspartate enantiomer might be explained by a higher dissociation rate (koff) in comparison with L-aspartate, that was shown in kinetic studies of sodium and aspartate binding on GltPh (Ewers et al., 2013; Hänelt et al., 2015). GltTk couples binding and transport of three sodium ions to one D-aspartate molecule (Figure 1B,D), the same number as for L-aspartate. Although the affinity for D-aspartate is lower than for L-aspartate, the binding of D-aspartate is not accompanied by a loss of sodium binding sites, which is in line with the observation that none of the sodium binding sites are directly coordinated by the substrate L-aspartate. In the crystal structure of GltTk with D-aspartate peaks of density were resolved at positions corresponding to the three sodium ions in the L-aspartate bound GltTk structure (Figure 2D) (Guskov et al., 2016). Altogether our data suggest that the mechanism of D- and L-aspartate transport in GltTk is most probably identical.
Mammalian glutamate transporters take up D-aspartate, L-glutamate and L-aspartate with similar micromolar affinity, but have significantly lower affinity (millimolar) for D-glutamate (Arriza et al., 1997; Arriza et al., 1994). In the absence of the structures of human SLC1A transporters with different stereoisomeric substrates, one can only speculate why EAATs can readily bind and transport both L- and D-aspartate, but only L-glutamate. It seems that the extra methylene group in D-glutamate compared to D-aspartate could cause sterical clashes within the binding site (Figure 2— Figure 2—figure supplement 3), which might affect affinity of binding.
Materials and methods
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene | TK0986 | UniProt database | Q5JID0 | |
| Strain, strain background (E. coli) | MC1061 | Casadaban and Cohen, 1980 | ||
| Biological sample (Thermococcus kodakarensis KOD1) | ATCC BAA-918/JCM 12380/KOD1 | |||
| Recombinant DNA reagent | pBAD24-GltTk-His8 | Jensen et al., 2013 | Expression plasmid for C-terminally His8-tagged GltTk. | |
| Chemical compound | D-Asp | Sigma-Aldrich | 219096–25G | ReagentPlus99% |
| Software | Origin 8 | OriginLab | ||
| Other | GltTk-D-aspartate coordinate file and structural factors | This paper | accession number PDB ID code 6R7R | Crystal structure of the glutamate transporter homologue GltTk in complex with D-aspartate |
Protein purification and crystallization
Request a detailed protocolGltTk was expressed and purified as described previously (Guskov et al., 2016). It was shown that L-aspartate binds to GltTk only if sodium ions are present, and the protein purified in absence of sodium ions is in the apo state (Jensen et al., 2013). For crystallization with D-aspartate the apo protein was purified by size exclusion chromatography (SEC) on a Superdex 200 10/300 GL (GE Healthcare) column equilibrated with buffer containing 10 mM Hepes KOH, pH 8.0, 100 mM KCl, 0.15% DM. Crystals of GltTk with D-aspartate were obtained in presence of 300 mM NaCl, 300 µM D-aspartate (Sigma-Aldrich, 99%) by the vapour diffusion technique (hanging drop) at 5°C by mixing equal volumes of protein (7 mg ml−1) and reservoir solution (20% glycerol, 10% PEG 4000, 100 mM Tris/bicine, pH 8.0, 60 mM CaCl2, 60 mM MgCl2, 0.75% n-octyl-b-D-glucopyranoside (OG)).
Data collection and structure determination
Request a detailed protocolCrystals were flash-frozen without any additional cryo protection and data sets were collected at 100K at the beamline ID23-1 (ESRF, Grenoble). The data were indexed, integrated and scaled in XDS (Kabsch, 2010) and the structure was solved by Molecular Replacement with Phaser (McCoy et al., 2007) using structure of GltTk (PDB ID 5E9S) as a search model. Manual model rebuilding and refinement were carried out in COOT (Emsley et al., 2010) and Phenix refine (Afonine et al., 2012). Data collection and refinement statistics are summarized in Table 2. Coordinates and structure factors for GltTk have been deposited in the Protein Data Bank under accession codes PDB 6R7R. All structural figures were produced with an open-source version of PyMol.
Data collection and refinement statistics.
https://doi.org/10.7554/eLife.45286.009| GltTk D-Asp | |
|---|---|
| Data collection | |
| Space group | P3221 |
| Cell dimensions | |
| a, b, c (Å) | 116.55, 116.55, 314.77 |
| α, β, γ (°) | 90.00, 90.00 120.00 |
| Resolution (Å) | 48.06-2.80 (2.87-2.80)* |
| Rmeas | 0.11 (>1) |
| CC1/2 | 99.9 (11.7) |
| I / σI | 8.40 (0.98) |
| Completeness (%) | 99.3 (98.9) |
| Redundancy | 5 (4) |
| Refinement | |
| Resolution (Å) | 2.80 |
| No. reflections | 301,077 |
| Rwork/Rfree (%)s | 23.4/27.2 |
| No. of atom | |
| Protein | 9262 |
| PEG/detergent | 181/33 |
| Ligand/ion | 27/9 |
| Water | - |
| B-factors | |
| Protein | 127 |
| PEG/detergent | 147/174 |
| Ligand/ion | 114/117 |
| Water | - |
| R.m.s. deviations | |
| Bond lengths (Å) | 0.008 |
| Bond angles (°) | 1.162 |
-
*Values in parentheses are for the highest-resolution shell.
Isothermal titration calorimetry
Request a detailed protocolITC experiments were performed at a constant temperature of 25°C using an ITC200 calorimeter (MicroCal). Varying concentrations of the indicated substrates (in 10 mM Hepes KOH, pH 8.0, 100 mM KCl, 0.15% DM and indicated sodium concentrations) were titrated into a thermally equilibrated ITC cell filled with 250 μl of 3–20 μM GltTk supplemented with 0 to 1000 mM NaCl. Data were analyzed using the ORIGIN-based software provided by MicroCal.
Reconstitution into proteoliposomes
Request a detailed protocolA solution of E. coli total lipid extract (20 mg ml−1 in 50 mM KPi, pH 7.0) was extruded with a 400-nm-diameter polycarbonate filter (Avestin, 11 passages) and diluted with the same buffer to a final concentration of 4 mg ml−1. The lipid mixture was destabilized with 10% Triton-X100. Purified GltTk and the destabilized lipids were mixed in a ratio of 1:1600 or 1:250 (protein: lipid) and incubated at room temperature for 30 min. Bio-beads were added four times (25 mg ml−1, 15 mg ml−1, 19 mg ml−1, 4 mg ml−1 lipid solution) after 0.5 hr, 1 hr, overnight and 2 hr incubation, respectively, on a rocking platform at 4°C. The Bio-beads were removed by passage over an empty Poly-Prep column (Bio-Rad). The proteoliposomes were collected by centrifugation (20 min, 298,906 g, 4°C), subsequently resuspended in 50 mM KPi, pH 7.0 to the concentration of the protein 33.4 µg ml−1 and freeze-thawed for four cycles. The proteoliposomes were stored in liquid nitrogen until subsequent experiments.
Uptake assay
Request a detailed protocolStored proteoliposomes with reconstitution ratio of 1:1600 were thawed and collected by centrifugation (20 min, 298,906 g, 4°C), the supernatant was discarded and the proteoliposomes were resuspended in buffer containing 10 mM KPi, pH 7.5, 300 mM KCl. The internal buffer was exchanged by three cycles of freezing in liquid nitrogen and thawing, and finally extruded through a polycarbonate filter with 400 nm pore size (Avestin, 11 passages). The proteoliposomes were finally pelleted by centrifugation (20 min, 298,906 g, 4°C) and resuspended to the concentration of the protein 625 ng µl−1. 2 µl of proteoliposomes were diluted 100 times in reaction buffer containing 10 mM KPi, pH 7.5, 100 mM NaCl, 200 mM Choline-Cl, 3 µM valinomycin and 0.2–15 µM D-aspartate (each concentration point contained 0.2 µM [3H]-D-aspartate). After 15 s the reaction was quenched by adding 2 ml of ice-cold buffer (10 mM KPi, pH 7.5, 300 mM KCl) and immediately filtered on nitrocellulose filter (Protran BA 85-Whatman filter), finally the filter was washed with 2 ml of quenching buffer. The filters were dissolved in scintillation cocktail and the radioactivity was measured with a PerkinElmer Tri-Carb 2800RT liquid scintillation counter.
Measuring transporter equilibrium potentials
Request a detailed protocolStored proteoliposomes with reconstitution ratio of 1:250 were thawed and collected by centrifugation (20 min, 298,906 g, 4°C), the supernatant was discarded and the proteoliposomes were resuspended to a concentration of 10 mg ml−1 of lipids in buffer containing 20 mM Hepes/Tris, pH 7.5, 200 mM NaCl, 50 mM KCl, 10 μM D-aspartate (containing 1 μM [3H]-D-aspartate). The internal buffer was exchanged by freeze-thawing and extrusion as described above. The experiment was started by diluting the proteoliposomes 20 times into a buffer containing 20 mM Hepes/Tris, pH 7.5, 200 mM NaCl, 3 μM valinomycin, varying concentrations of KCl and Choline Cl were added in order to obtain the desired membrane potential as shown in (Figure 1—source data 1).
After 1, 2 and 3 min the reaction was quenched with ice-cold quenching buffer containing 20 mM Hepes/Tris, pH 7.5, 250 mM Choline Cl and immediately filtered on nitrocellulose filter (Protran BA 85-Whatman filter), finally the filter was washed with 2 ml of quenching buffer. The initial amount of radiolabeled aspartate was measured by filtering the proteoliposomes immediately after diluting them in quenching buffer. The filters were dissolved in scintillation cocktail and the radioactivity was measured with a PerkinElmer Tri-Carb 2800RT liquid scintillation counter. The equilibrium, or reversal, potential, Erev, for each condition was calculated as described in Fitzgerald et al. (2017).
Data availability
Diffraction data and the derived model have been deposited in PDB under accession number 6R7R.
-
Protein Data BankID 6R7R. Diffraction data and the derived model.
References
-
Towards automated crystallographic structure refinement with phenix.refineActa Crystallographica Section D Biological Crystallography 68:352–367.https://doi.org/10.1107/S0907444912001308
-
Functional comparisons of three glutamate transporter subtypes cloned from human motor cortexThe Journal of Neuroscience 14:5559–5569.https://doi.org/10.1523/JNEUROSCI.14-09-05559.1994
-
Structural basis of Metallo-β-Lactamase inhibition by captopril stereoisomersAntimicrobial Agents and Chemotherapy 60:142–150.https://doi.org/10.1128/AAC.01335-15
-
Analysis of gene control signals by DNA fusion and cloning in Escherichia coliJournal of Molecular Biology 138:179–207.https://doi.org/10.1016/0022-2836(80)90283-1
-
D-Aspartic acid: an endogenous amino acid with an important neuroendocrine roleBrain Research Reviews 53:215–234.https://doi.org/10.1016/j.brainresrev.2006.08.005
-
D-Aspartic acid is a novel endogenous neurotransmitterThe FASEB Journal 25:1014–1027.https://doi.org/10.1096/fj.10-168492
-
Structure-Based insights into the role of the Cys-Tyr crosslink and inhibitor recognition by mammalian cysteine dioxygenaseJournal of Molecular Biology 428:3999–4012.https://doi.org/10.1016/j.jmb.2016.07.012
-
Studies on the relationship between chemical constitution and physiological actionBiochemical Journal 27:1257–1266.https://doi.org/10.1042/bj0271257
-
Features and development of cootActa Crystallographica. Section D, Biological Crystallography 66:486–501.https://doi.org/10.1107/S0907444910007493
-
BookD-Aspartate, an Atypical Amino Acid with NMDA Receptor Agonist Features: Involvement in SchizophreniaIn: Hashimoto K, editors. The NMDA Receptors. Humana Press, Cham. pp. 83–101.https://doi.org/10.1007/978-3-319-49795-2_5
-
Cryo-EM structure of the human neutral amino acid transporter ASCT2Nature Structural & Molecular Biology 25:515–521.https://doi.org/10.1038/s41594-018-0076-y
-
SLC1 glutamate transportersPflügers Archiv - European Journal of Physiology 466:3–24.https://doi.org/10.1007/s00424-013-1397-7
-
Low affinity and slow na+ binding precedes high affinity aspartate binding in the Secondary-active transporter GltPhThe Journal of Biological Chemistry 290:15962–15972.https://doi.org/10.1074/jbc.M115.656876
-
Is aspartate an excitatory neurotransmitter?Journal of Neuroscience 35:10168–10171.https://doi.org/10.1523/JNEUROSCI.0524-15.2015
-
Crystal structure of a substrate-free aspartate transporterNature Structural & Molecular Biology 20:1224–1226.https://doi.org/10.1038/nsmb.2663
-
Integration, scaling, space-group assignment and post-refinementActa Crystallographica Section D Biological Crystallography 66:133–144.https://doi.org/10.1107/S0907444909047374
-
d-Aspartate—An important bioactive substance in mammals: A review from an analytical and biological point of viewJournal of Chromatography B 879:3108–3121.https://doi.org/10.1016/j.jchromb.2011.03.062
-
The hill analysis and co-ion-driven transporter kineticsThe Journal of General Physiology 145:565–574.https://doi.org/10.1085/jgp.201411332
-
Phaser crystallographic softwareJournal of Applied Crystallography 40:658–674.https://doi.org/10.1107/S0021889807021206
-
Chiral drugs: an overviewInternational Journal of Biomedical Science : IJBS 2:85–100.
-
The ionic stoichiometry of the GLAST glutamate transporter in salamander retinal gliaThe Journal of Physiology 577:591–599.https://doi.org/10.1113/jphysiol.2006.116830
-
Binding thermodynamics of a glutamate transporter homologNature Structural & Molecular Biology 20:634–640.https://doi.org/10.1038/nsmb.2548
-
Structural basis for substrate promiscuity of dCKJournal of Molecular Biology 378:607–621.https://doi.org/10.1016/j.jmb.2008.02.061
-
The prokaryotic na+/Ca2+ exchanger NCX_Mj transports Na+ and Ca2+ in a 3:1 stoichiometryThe Journal of General Physiology 150:51–65.https://doi.org/10.1085/jgp.201711897
-
Structural features of the glutamate transporter familyMicrobiology and Molecular Biology Reviews : MMBR 63:293–307.
-
D-aspartic acid in the nervous system of aplysia limacina: possible role in neurotransmissionJournal of Cellular Physiology 206:672–681.https://doi.org/10.1002/jcp.20513
-
Glutamate transporter EAAT2: regulation, function, and potential as a therapeutic target for neurological and psychiatric diseaseCellular and Molecular Life Sciences 72:3489–3506.https://doi.org/10.1007/s00018-015-1937-8
-
Carbonic anhydrase activators. activation of isoforms I, II, IV, VA, VII, and XIV with L- and D-phenylalanine and crystallographic analysis of their adducts with isozyme II: stereospecific recognition within the active site of an enzyme and its consequences for the drug designJournal of Medicinal Chemistry 49:3019–3027.https://doi.org/10.1021/jm0603320
-
Iterative-build OMIT maps: map improvement by iterative model building and refinement without model biasActa Crystallographica Section D Biological Crystallography 64:515–524.https://doi.org/10.1107/S0907444908004319
-
Mechanisms of glutamate transportPhysiological Reviews 93:1621–1657.https://doi.org/10.1152/physrev.00007.2013
-
Crystal structure of an asymmetric trimer of a bacterial glutamate transporter homologNature Structural & Molecular Biology 19:355–357.https://doi.org/10.1038/nsmb.2233
Article and author information
Author details
Funding
Nederlandse Organisatie voor Wetenschappelijk Onderzoek (865.11.001)
- Dirk J Slotboom
European Research Council (282083)
- Dirk J Slotboom
Nederlandse Organisatie voor Wetenschappelijk Onderzoek (723.014.002)
- Albert Guskov
The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Acknowledgements
This work is funded by the Netherlands Organisation for Scientific Research (Vici grant 865.11.001 to DJS and Vidi grant 723.014.002 to AG) and European Research Council starting grant 282083 to DJS. We thank A Garaeva and M Ejby for synchrotron data collection. The European Synchrotron Radiation Facility beamlines ID23-1 and ID29 (Grenoble, France) and EMBL beamlines P13 and P14 (Hamburg, Germany) are acknowledged for beamline facilities. This work has been supported by iNEXT, grant number 653706, funded by the Horizon 2020 programme of the European Commission.
Copyright
© 2019, Arkhipova et al.
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 2,642
- views
-
- 297
- downloads
-
- 40
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Citations by DOI
-
- 40
- citations for umbrella DOI https://doi.org/10.7554/eLife.45286






