Bacterial contribution to genesis of the novel germ line determinant oskar
Figures
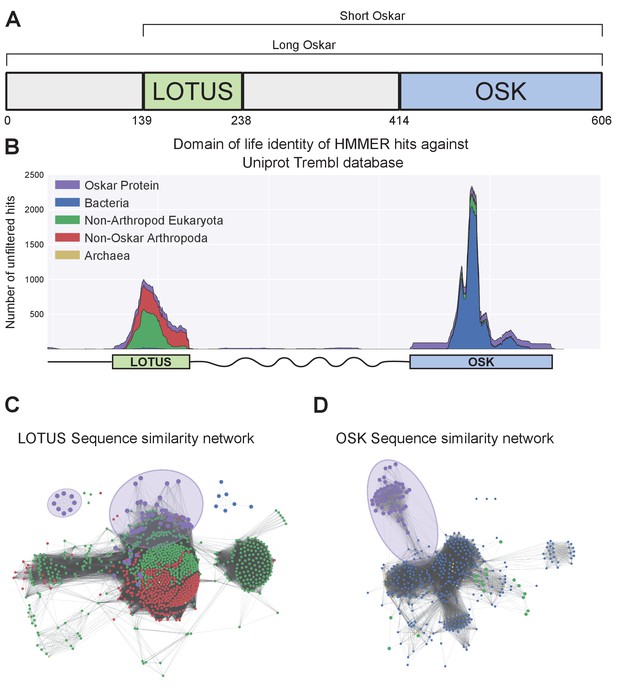
Sequence analysis of the Oskar gene.
(a) Schematic representation of the Oskar gene. The LOTUS and OSK hydrolase-like domains are separated by a poorly conserved region of predicted high disorder and variable length between species. In some dipterans, a region 5’ to the LOTUS domain is translated to yield a second isoform, called Long Oskar. Residue numbers correspond to the D. melanogaster Osk sequence. (b) Stackplot of domain of life identity of HMMER hits across the protein sequence. For a sliding window of 60 Amino Acids across the protein sequence (X axis), the number of hits in the Trembl (UniProt) database (Y axis) is represented and color coded by domain of life origin (see Materials and methods: Iterative HMMER search of OSK and LOTUS domains), stacked on top of each other. (c, d) EFI-EST-generated graphs of the sequence similarity network of the LOTUS (c) and OSK (d) domains of Oskar (Gerlt et al., 2015). Sequences were obtained using HMMER against the UniProtKB database. Most Oskar LOTUS sequences cluster within eukaryotes and arthropods. In contrast, Oskar OSK sequences cluster most strongly with a small subset of bacterial sequences.
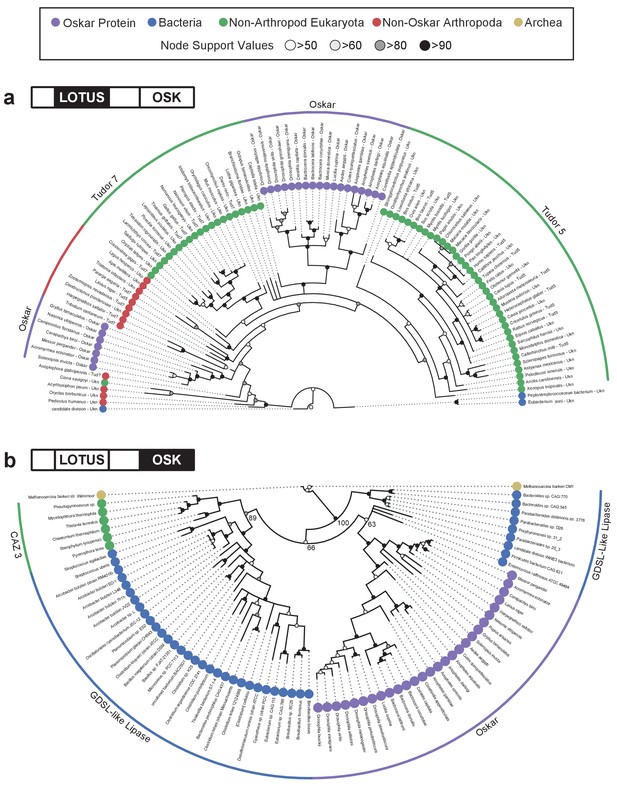
Phylogenetic analysis of the LOTUS and OSK domains.
(a) Bayesian consensus tree for the LOTUS domain. Three major LOTUS-containing protein families are represented within the tree: Tudor 5, Tudor 7, and Oskar. Oskar LOTUS domains form two clades, one containing only dipterans and one containing all other represented insects (hymenopterans and orthopterans). The tree was rooted to the three bacterial sequences added in the dataset. (b) Bayesian consensus tree for the OSK domain. The OSK domain is nested within GDSL-like domains of bacterial species from phyla known to contain germ line symbionts in insects. The ten non-Oskar eukaryotic sequences in the analysis form a single clade comprising fungal Carbohydrate Active Enzyme 3 (CAZ3) proteins. For Bayesian and RaxML trees with all accession numbers and node support values see Figure 2—figure supplements 1–4.
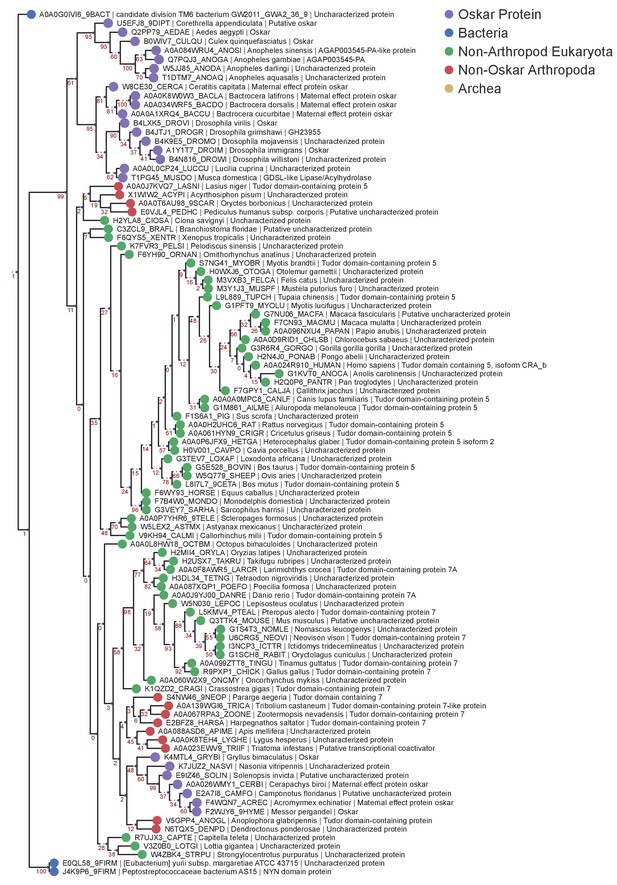
LOTUS Domain RaxML MUSCLE Tree.
Phylogenetic tree of the HMMER sequences retrieved from the UniProt database using the LOTUS alignment HMM model. The top 97 hits were selected for phylogenetic analysis, and the only three bacterial sequences found to be a match were added to the alignment manually. The resulting 100 sequences were aligned using MUSCLE with default settings. The sequences were filtered to contain only one sequence per species (best E-value kept) yielding 100 sequences for analysis. Finally, the tree was created using RaxML v8.2.4, using 1000 bootstraps and model selection performed by the RaxML automatic model selection tool. See ‘Phylogenetic Analysis’ in Materials and methods for further detail. Sequences are color-coded as follows: Purple = Oskar; Red = Non Oskar Arthropod; Green = Non Arthropod Eukaryote; Blue = Bacteria. Names following leaves display the UniProt accession number followed by the species name and the UniProt protein name.
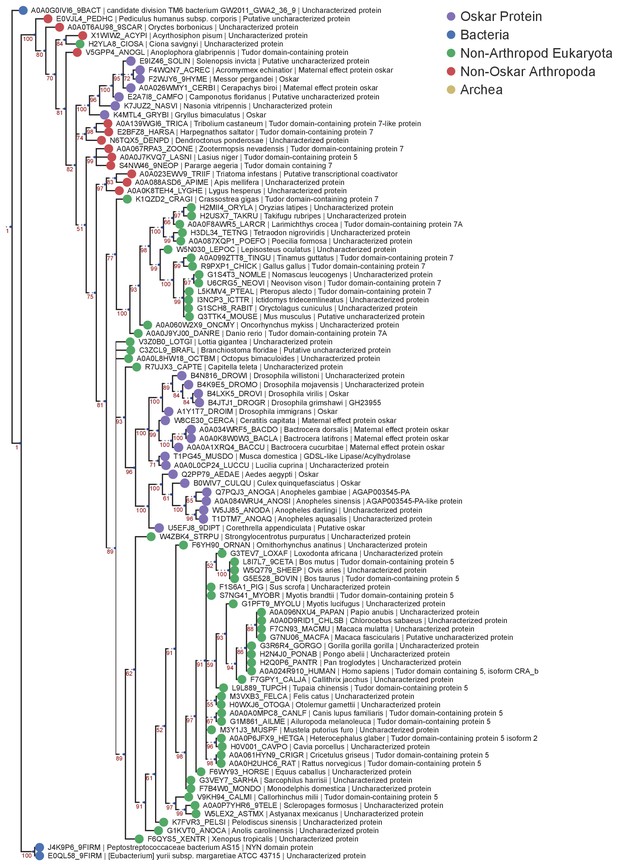
LOTUS Domain Bayesian MUSCLE Tree.
Phylogenetic tree of the HMMER sequences retrieved from the UniProt database using the LOTUS alignment HMM model. 100 sequences were chosen for analysis as described for Figure 2—figure supplement 1. The tree was created using Mr Bayes V3.2.6 using a Mixed model (prset aamodel = Mixed) and a gamma distribution (lset rates = Gamma). The algorithm was allowed to run for 3 million generations to achieve a std <0.01. See ‘Phylogenetic Analysis’ in Materials and methods for further detail. Sequences are color-coded as follows: Purple = Oskar; Red = Non Oskar Arthropod; Green = Non Arthropod Eukaryote; Blue = Bacteria. Names following leaves display the UniProt accession number followed by the species name and the UniProt protein name.
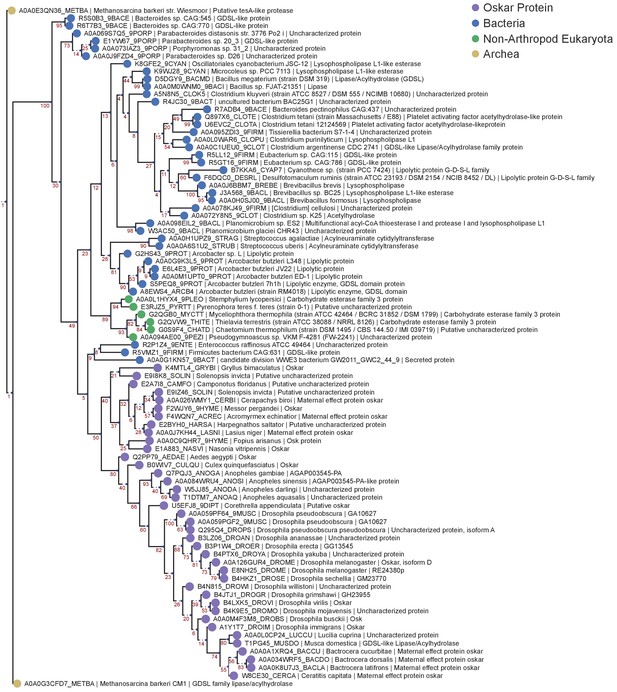
OSK Domain RaxML MUSCLE Tree.
Phylogenetic tree of the HMMER sequences retrieved from the UniProt database using the OSK alignment HMM model. The top 95 hits were selected for phylogenetic analysis, and the only five non-Oskar eukaryotic sequences found to be a match were added to the alignment manually. The resulting 100 sequences were aligned using MUSCLE with default settings. The sequences were filtered to contain only one sequence per species (best E-value kept), yielding 87 sequences for analysis. Finally, the tree was created using RaxML v8.2.4, using 1000 bootstraps and model selection performed by the RaxML automatic model selection tool. See ‘Phylogenetic Analysis’ in Materials and methods for further detail. Sequences are color-coded as follows: Purple = Oskar; Red = Non Oskar Arthropod; Green = Non Arthropod Eukaryote; Blue = Bacteria. Names following leaves display the UniProt accession number followed by the species name and the UniProt protein name.
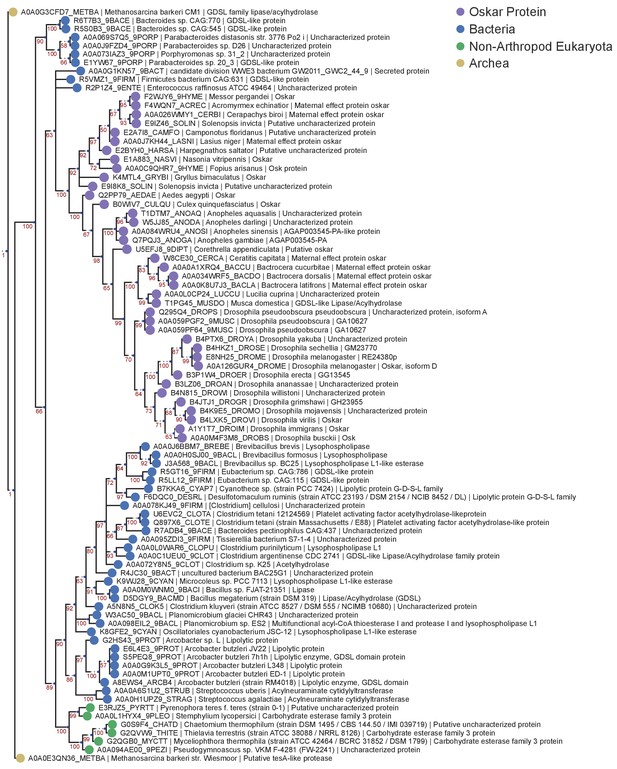
OSK Domain Bayesian MUSCLE Tree.
Phylogenetic tree of the HMMER sequences hit on the UniProt database using the OSK alignment HMM model. 87 sequences were chosen for analysis as described for Figure 2—figure supplement 3.The tree was created using Mr Bayes V3.2.6 using a Mixed model (prset aamodel = Mixed) and a gamma distribution (lset rates = Gamma). The algorithm was allowed to run for 4 million generations to achieve a std <0.01. See ‘Phylogenetic Analysis’ in Materials and methods for further detail. Sequences are color-coded as follows: Purple = Oskar; Red = Non Oskar Arthropod; Green = Non Arthropod Eukaryote; Blue = Bacteria. Names following leaves display the UniProt accession number followed by the species name and the UniProt protein name.
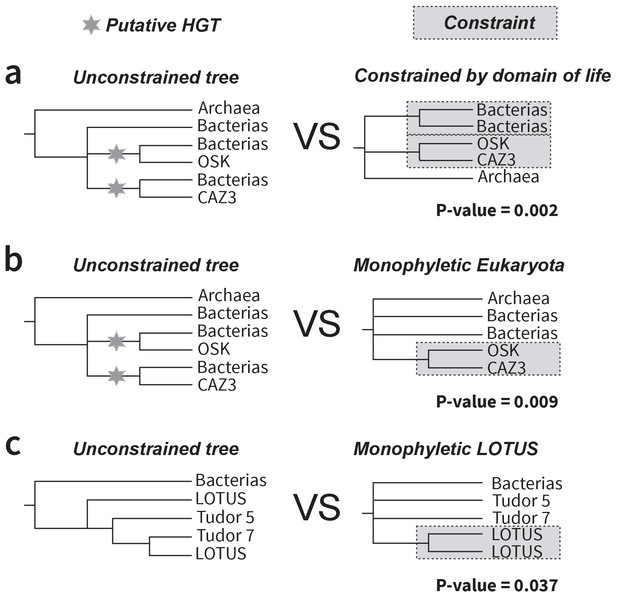
SOWHAT constrained trees and results.
Two trees constrained by alternative relationships that would be expected under vertical transmission of sequences were designed and tested against our result supporting a putative HGT event of the OSK domain. (a) The first tree (right) is constrained by domain of life, requiring bacterial and eukaryotic sequences to be monophyletic, and disallowing sister group relationships of subsets of eukaryotic sequences and bacterial sequences. Our unconstrained tree topology (left) outperformed this topology with a p-value of 0.002 (95% confidence interval upper: 0.007 lower: 0.0002). (b) The second tree requires monophyly of Eukaryota. Our unconstrained tree topology (left) outperformed this topology with a p-value of 0.009 (95% confidence interval upper: 0.017 lower: 0.004). (c) The third tree tested whether the LOTUS domain split observed in the tree generated with the MUSCLE alignment was significantly different from a tree where the LOTUS sequences formed a monophyly. The unconstrained tree (left) outperformed this topology with a p-value of 0.037 (95% confidence interval upper: 0.05 lower: 0.026).
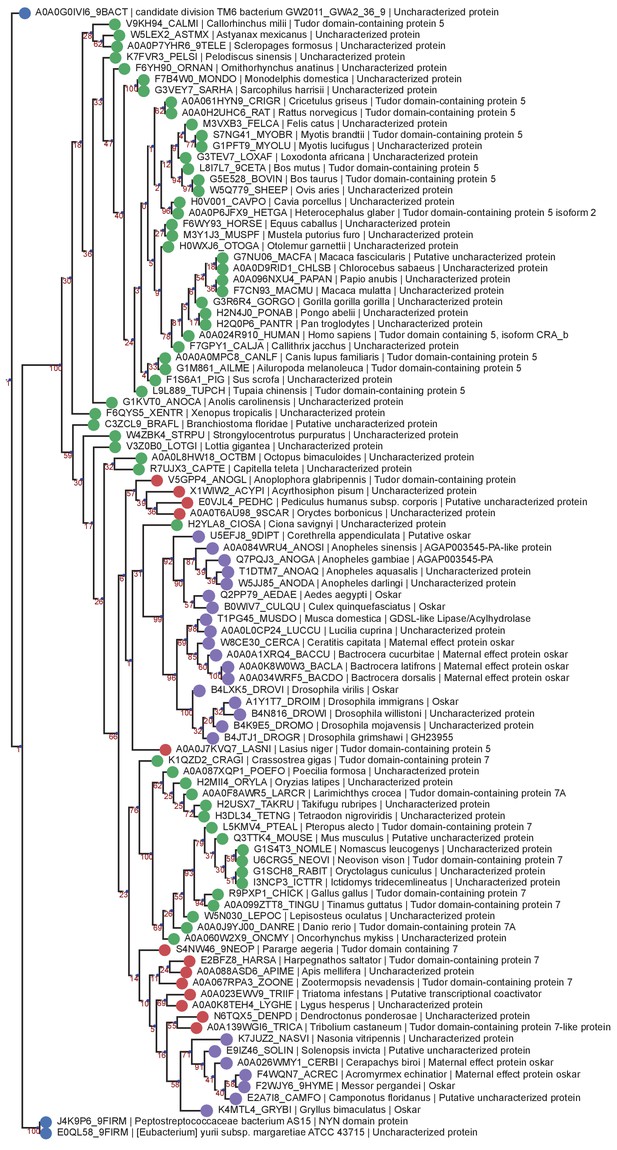
LOTUS Domain RaxML PRANK Tree.
Phylogenetic tree of the same sequences used for the previous LOTUS trees. The sequences were aligned using PRANK and the tree generated with RaxML as described in Phylogenetic Analysis Based on PRANK alignment. Sequences are color-coded as follows: Purple = Oskar; Red = Non Oskar Arthropod; Green = Non Arthropod Eukaryote; Blue = Bacteria. Names following leaves display the UniProt accession number followed by the species name and the UniProt protein name.
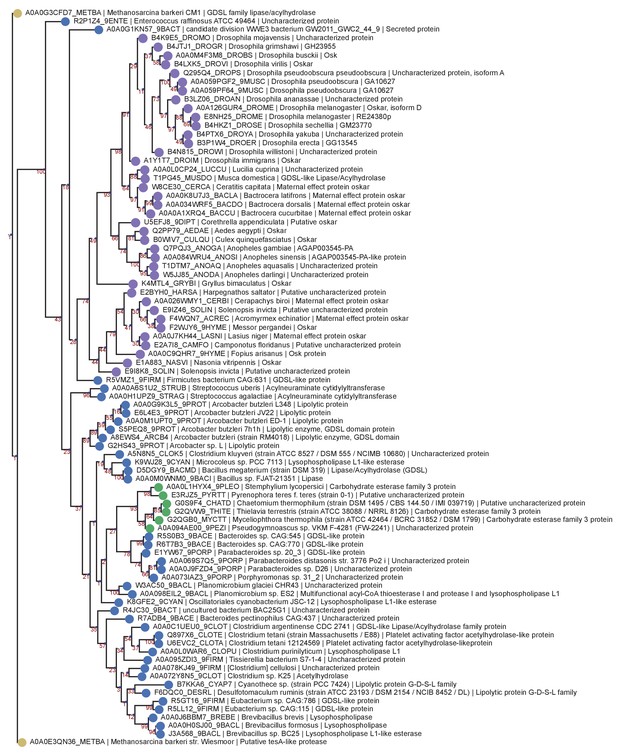
OSK Domain RaxML PRANK Tree.
Phylogenetic tree of the same sequences used for the previous OSK trees. The sequences were aligned using PRANK and the tree generated with RaxML as described in Phylogenetic Analysis Based on PRANK alignment. Sequences are color-coded as follows: Purple = Oskar; Red = Non Oskar Arthropod; Green = Non Arthropod Eukaryote; Blue = Bacteria. Names following leaves display the UniProt accession number followed by the species name and the UniProt protein name.
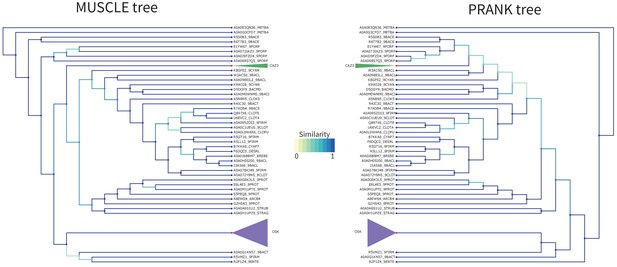
OSK Tree PRANK Comparison.
Comparison of the tree obtained with RaxML starting from the MUSCLE alignment (left) versus the PRANK alignment (right) for the OSK domain. Similarity scores for the branching events are color coded from yellow to blue (see figure color bar legend). The OSK (purple) clade and CAZ3 (green) clade have been colored and compacted for readability as they do not have any internal branching changes. Node color is blue if the leaf is a sequence, and red if this is a compacted group of sequences.
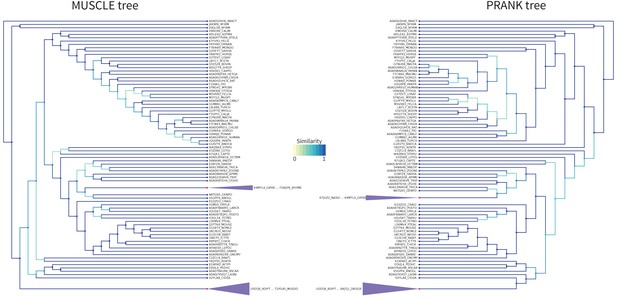
LOTUS Tree PRANK Comparison.
Comparison of the tree obtained with RaxML starting from the MUSCLE alignment (left) versus the PRANK alignment (right) for the LOTUS domain. Similarity scores for the branching events are color coded from yellow to blue (see figure color bar legend). The LOTUS (purple) clades have been colored and compacted for readability as they do not have any internal branching changes. Node color is blue if the leaf is a sequence, and red if this is a compacted group of sequences.
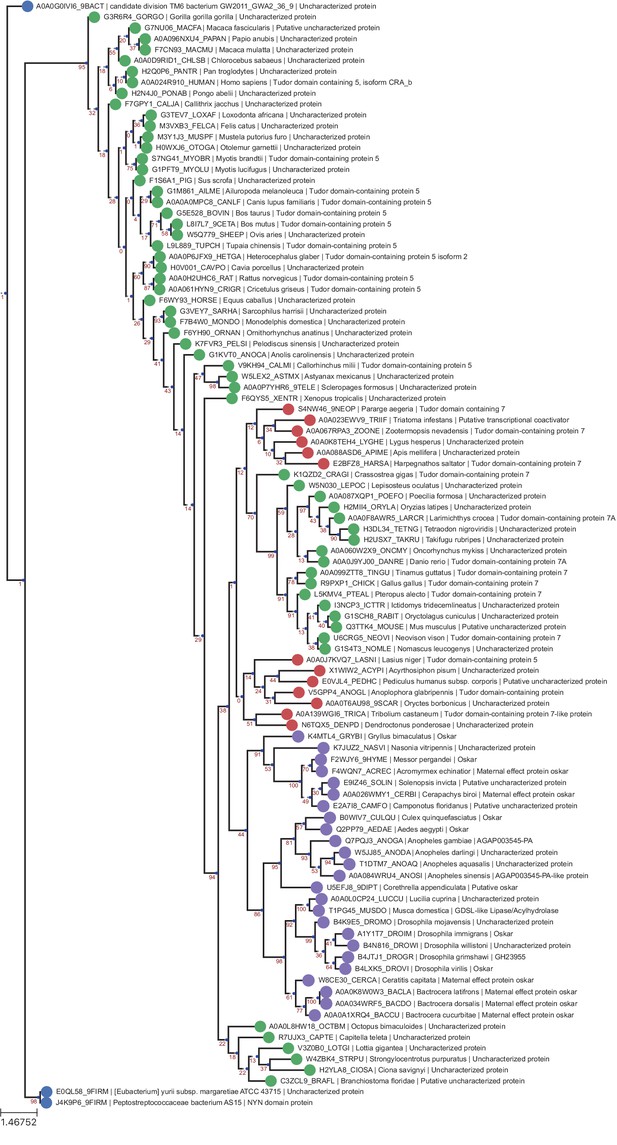
LOTUS Domain RaxML T-Coffee Tree.
Phylogenetic tree of the same sequences used for the previous LOTUS trees. The sequences were aligned using T-Coffee and the tree generated with RaxML as described in Phylogenetic Analysis Based on T-Coffee alignment. Sequences are color-coded as follows: Purple = Oskar; Red = Non Oskar Arthropod; Green = Non Arthropod Eukaryote; Blue = Bacteria. Names following leaves display the UniProt accession number followed by the species name and the UniProt protein name.
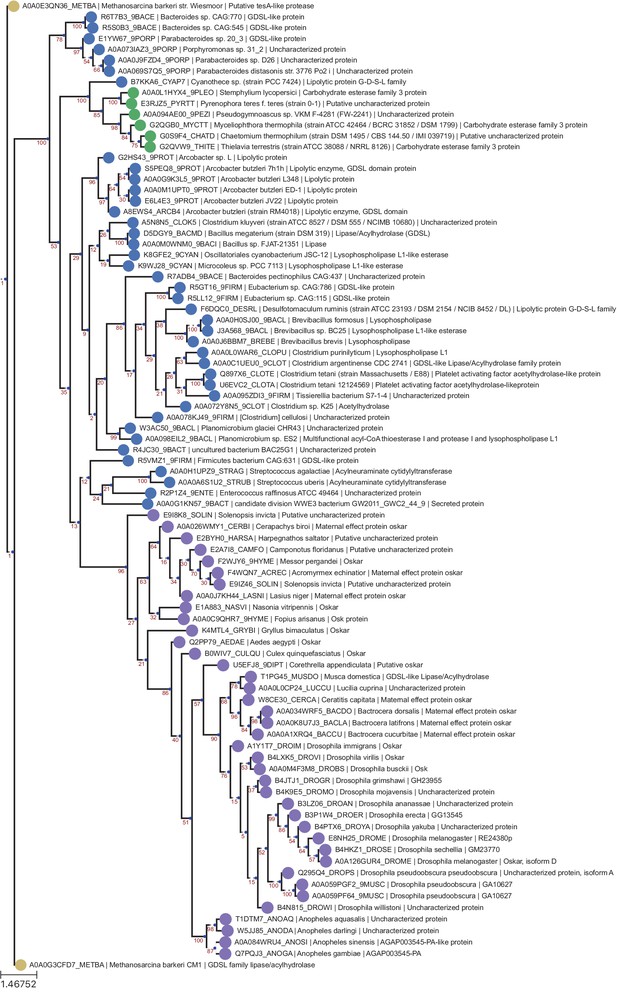
OSK Domain RaxML T-Coffee Tree.
Phylogenetic tree of the same sequences used for the previous OSK trees. The sequences were aligned using T-Coffee and the tree generated with RaxML as described in Phylogenetic Analysis Based on T-Coffee alignment. Sequences are color-coded as follows: Purple = Oskar; Red = Non Oskar Arthropod; Green = Non Arthropod Eukaryote; Blue = Bacteria. Names following leaves display the UniProt accession number followed by the species name and the UniProt protein name.
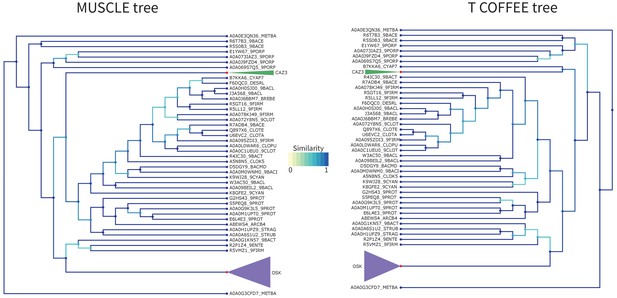
OSK Tree T-Coffee Comparison.
Comparison of the tree obtained with RaxML starting from the MUSCLE alignment (left) versus the T-Coffee alignment (right) for the OSK domain. Similarity scores for the branching events are color coded from yellow to blue (see figure color bar legend). The OSK (purple) clade and CAZ3 (green) clade have been colored and compacted for readability as they do not have any internal branching changes. Node color is blue if the leaf is a sequence, and red if this is a compacted group of sequences.
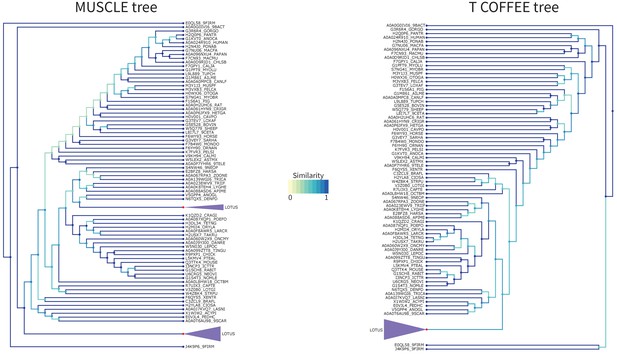
LOTUS Tree T-Coffee Comparison.
Comparison of the tree obtained with RaxML starting from the MUSCLE alignment (left) versus the T-Coffee alignment (right) for the LOTUS domain. Similarity scores for the branching events are color coded from yellow to blue (see figure color bar legend). The LOTUS (purple) clades have been colored and compacted for readability as they do not have any internal branching changes. Node color is blue if the leaf is a sequence, and red if this is a compacted group of sequences.
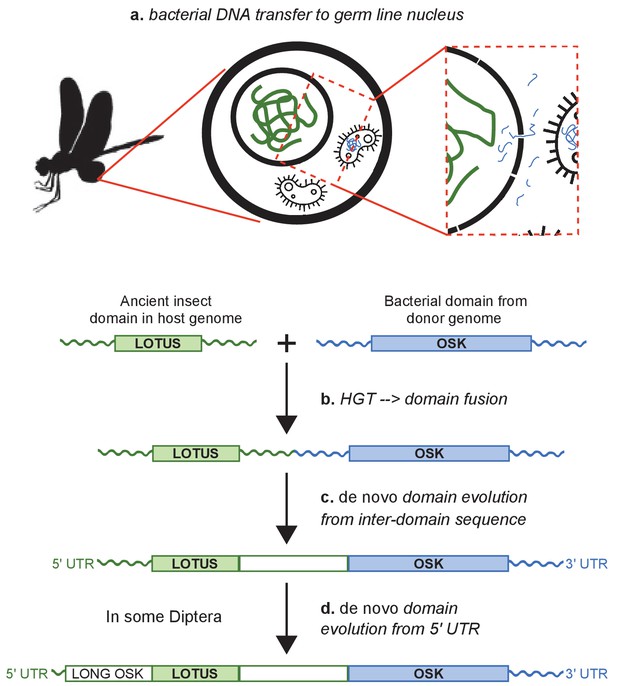
Hypothesis for the origin of oskar.
Integration of the OSK domain close to a LOTUS domain in an ancestral insect genome. (a) DNA containing a GDSL-like domain from an endosymbiotic germ line bacterium is transferred to the nucleus of a germ cell in an insect common ancestor. (b) DNA damage or transposable element activity induces an integration event in the host genome, close to a pre-existing LOTUS-like domain. (c) The region between the two domains undergoes de novo coding evolution, creating an open reading frame with a unique, chimeric domain structure. (d) In some Diptera, including D. melanogaster, part of the 5’ UTR of oskar has undergone de novo coding evolution to form the Long Oskar domain.
Additional files
-
Source data 1
Alignment and Sequence Classification Tools & Data.
Subfolder "Alignments": All sequences identified and analyzed in this study, in FASTA format and with corresponding Alignments. Subfolder BLAST search results: Results of BLASTP searches with full length Oskar, OSK or LOTUS domains as queries. Subfolder "Data": Necessary files for running the different IPython notebooks: a. Subfolder "HMM": HMM models used for iterative searching for sequences similar to full-length Oskar, LOTUS and OSK domains; b. Subfolder "Taxonomy": Conversion table for UniProt ID to taxon information (uniprot_ID_taxa.tsv); c. Subfolder "Trees": Contains the tree files obtained from i. RaxML phylogenetic analyses of the OSK and LOTUS domains aligned with MUSCLE, T-Coffee or PRANK; ii. MrBayes phylogenetic analyses of the OSK and LOTUS domains aligned with MUSCLE; iii. SOWHAT analyses.
- https://cdn.elifesciences.org/articles/45539/elife-45539-data1-v2.zip
-
Supplementary file 1
Supplementary tables.
(A) List of genomes and transcriptomes used for automated oskar search. (B) List of Oskar sequences used in the final alignment. (C) List of sequences used for phylogenetic analysis of the LOTUS domain. (D) List of sequences used for phylogenetic analysis of the OSK domain.
- https://cdn.elifesciences.org/articles/45539/elife-45539-supp1-v2.pdf
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/45539/elife-45539-transrepform-v2.docx