Large protein organelles form a new iron sequestration system with high storage capacity
Figures
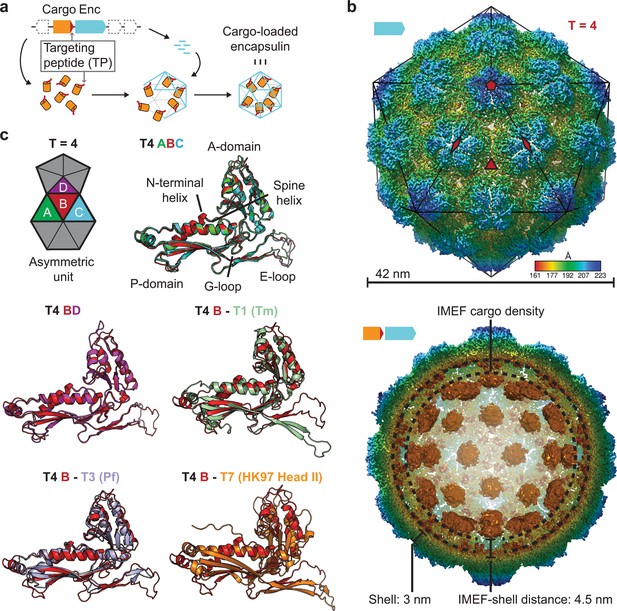
Overall architecture of the cargo-loaded T = 4 encapsulin.
(a) Schematic diagram of a core encapsulin operon and targeting peptide (TP)-dependent cargo encapsulation. (b) Surface view of the cryo-EM map of the Qs T = 4 encapsulin shell (top) and inside view of cargo-loaded encapsulin (bottom). 5-, 3- and 2-fold symmetry axes are indicated by red symbols. The overall icosahedral symmetry is highlighted by black lines representing icosahedral facets. Cargo-densities are shown in orange while the shell is radially colored. To depict the complete cargo-loaded compartment, a 10 Å filtered map highlighting the cargo was combined with the 3.85 Å map of the shell. (c) Asymmetric unit of the T4 encapsulin shell and structural alignment of the four unique T4 shell monomers with one another and with the T. maritima (Tm) T = 1 monomer (3DKT), the P. furiosus (Pf) T = 3 monomer (2E0Z) and the HK97 bacteriophage Head II T = 7 monomer (2FT1).
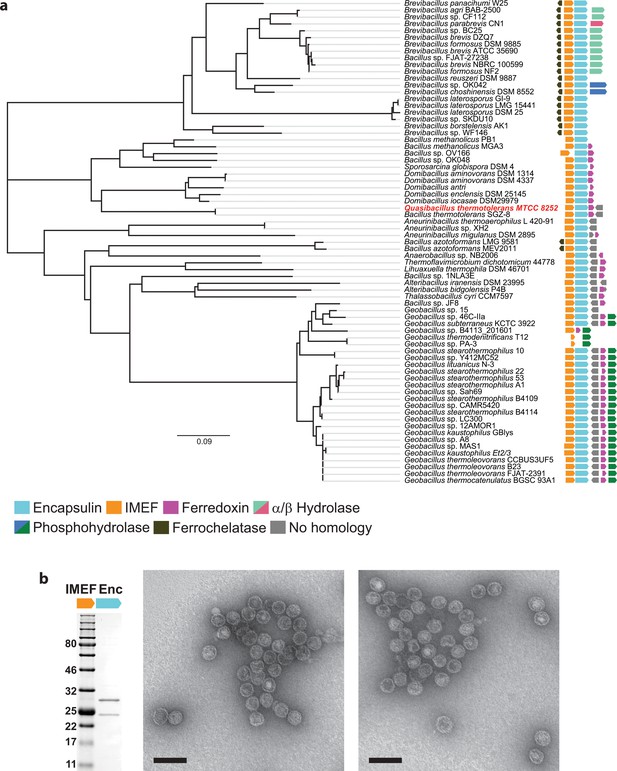
Identification of IMEF operons and confirmation of protein compartment formation.
(a) The Qs operon is highlighted in red. The nearest-neighbor phylogenetic tree is based on a ClustalO alignment of IMEF cargo proteins. IMEF cargo accessions are shown in Supplementary Table 1. Evolutionary distances were estimated as the number of amino acid substitutions per site. The scale bar represents 0.09 expected amino acid residue substitutions per site. b, Shown are negative stain (uranyl formate) TEM micrographs of purified compartments (right) and a representative SDS-PAGE gel (left). The SDS-PAGE gel shows that the IMEF cargo protein (22.6 kDa) co-purifies with the encapsulin capsid protein (32.2 kDa). The SDS-PAGE gel is the same as shown for comparison in Fig. 3e of the main text. Scale bars in micrographs correspond to 100 nm.
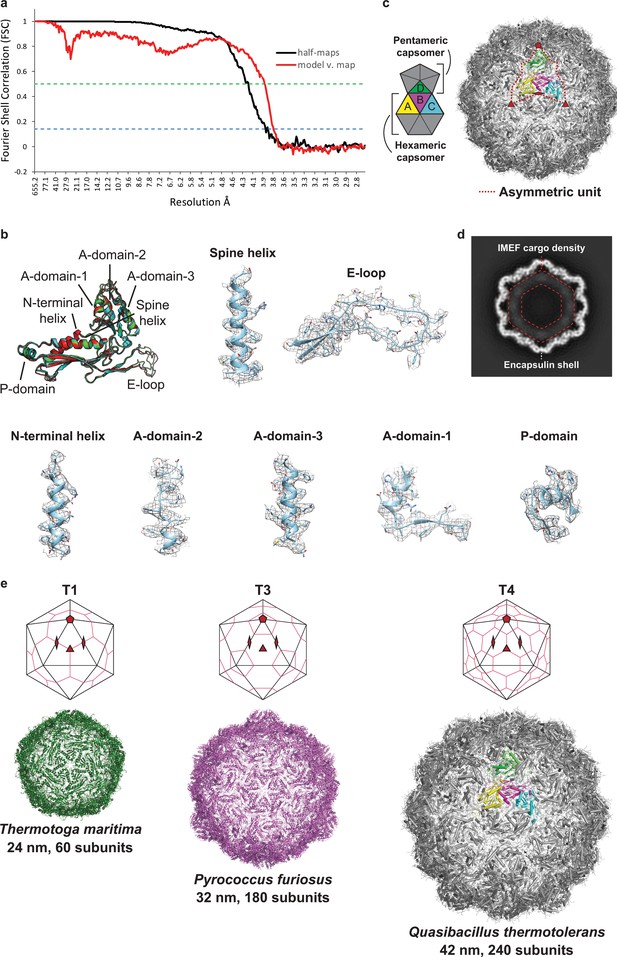
Supplementary cryo-EM data.
(a) FSC curves calculated from independent half-maps (black line) and a comparison of the atomic model of the IMEF encapsulin versus the experimental map (red line). The green dashed line represents an FSC cutoff of 0.5 and the blue dashed line represents an FSC cutoff of 0.143. (b) Annotated monomer alignment of the IMEF capsid protein monomers with local cryo-EM maps. The different structural elements of the T = 4 encapsulin monomer are shown. Representative density maps (mesh) and atomic models (model B of the asymmetric unit) illustrating side chain features and overall fit of the atomic model with the cryo-EM map are shown. (c) T = 4 IMEF encapsulin capsid with highlighted asymmetric unit (right) and annotation (left) defining the 4 capsid monomers of the asymmetric unit and the hexameric and pentameric capsomers. (d) Central slice of cargo-loaded IMEF encapsulin capsid. Less defined lower resolution density can clearly be seen in the interior (red dotted lines). (e) Top: Drawings illustrating the icosahedral symmetry and tiling of capsids with different triangulation numbers. Bottom: Capsid models of T = 1, T = 3 and the newly discovered T = 4 encapsulin shells including maximum external diameter and subunit number.
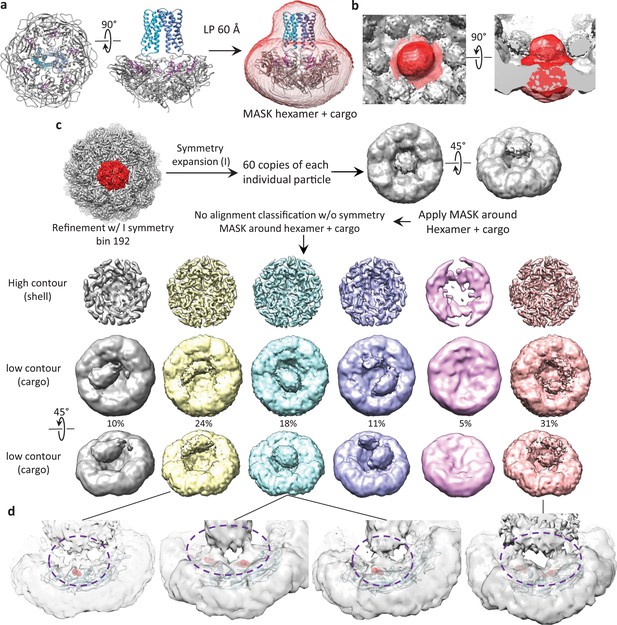
Symmetry expansion classification of cargo density.
(a) Model of a hexameric unit with cargo colored blue, encapsulin shell colored gray and targeting peptide colored magenta. A simulated map of the model low-pass filtered to 60 Å was used for signal subtraction and classification. (b) View of the low-pass filtered mask encompassing a hexameric unit of the encapsulin shell and associated symmetry averaged cargo density. (c) Data processing scheme to expand icosahedral symmetry and perform focused classification on a hexameric nit with associated cargo proteins. (d) Examples of low-resolution features showing a connection between cargo density and general areas of the targeting peptide in the encapsulin shell. Targeting peptides are shown as red spheres, encapsulin shell is shown as blue ribbon. Areas circled in purple dashed lines indicate densities for a flexible tether between the targeting peptide and cargo.
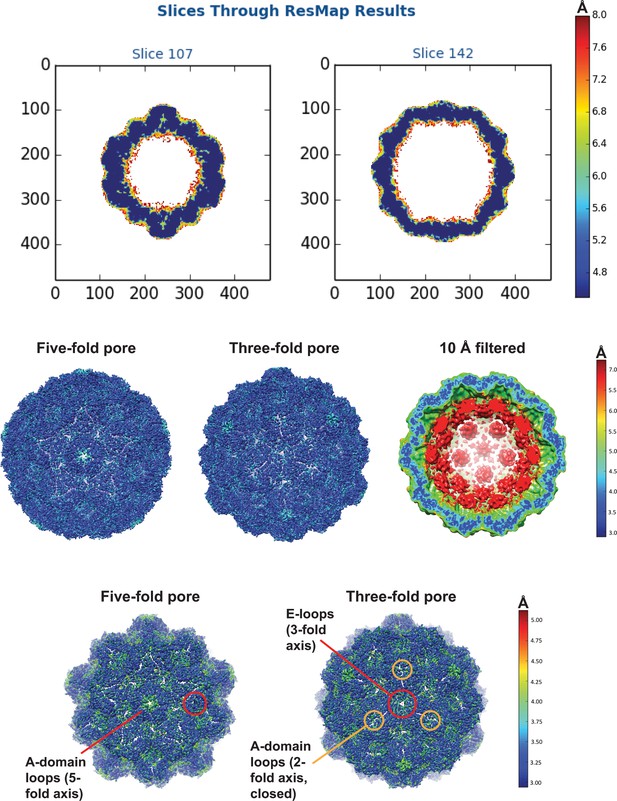
Local resolution maps of the cargo-loaded T = 4 IMEF encapsulin.
Top: central slices through the T = 4 shell. Middle: Comparison of shell and internal cargo resolution. Bottom: Focus on shell resolution. The lowest resolution parts of the T = 4 shell are highlighted and include 5- and 3-fold pore residues corresponding to A-domain loops and E-loops, respectively, and A-domain loops forming the closed ‘pore’ at the 2-fold symmetryaxis.
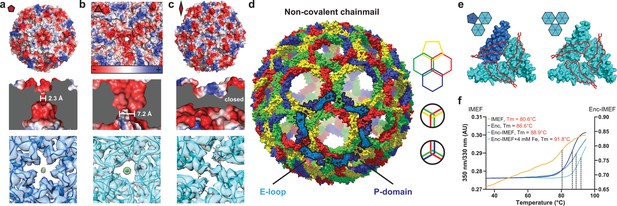
Non-covalent chainmail topology, thermal stability and pores of the T = 4 encapsulin shell.
(a, b and c) Electrostatic surface representation of the 5-fold (d) and 3-fold (e) T = 4 shell pores and the 2-fold symmetry axis (f). Outside views showing negatively charged pores (top) with no pore opening observed at the two-fold symmetry axis, cutaway side view highlighting the narrowest point of the pores (middle) and cryo-EM maps with fitted monomer models in ribbon representation (bottom). Additional cryo-EM density is observed at the center of both pores in interaction distance with the side chains of pore residues (5-fold: Asn200, 3-fold: Asp9, Asp71, Glu251 and Glu252, shown in stick representation). (d) Chainmail network mediated by E-loop and P-domain interactions. Only E-loops and P-domains are shown. E-loops and P-domains of the outlined ring belonging to the same monomer are located next to one another and are shown in light and dark blue, respectively. (e) Extended E-loop interactions interlock neighboring capsid monomers at the two unique three-fold interfaces. Each E-loop interacts with two P-domains. (f) Representative thermal unfolding curves for Qs T = 4 encapsulin components determined via differential scanning fluorimetry. Tm: midpoint of the thermal unfolding curve.
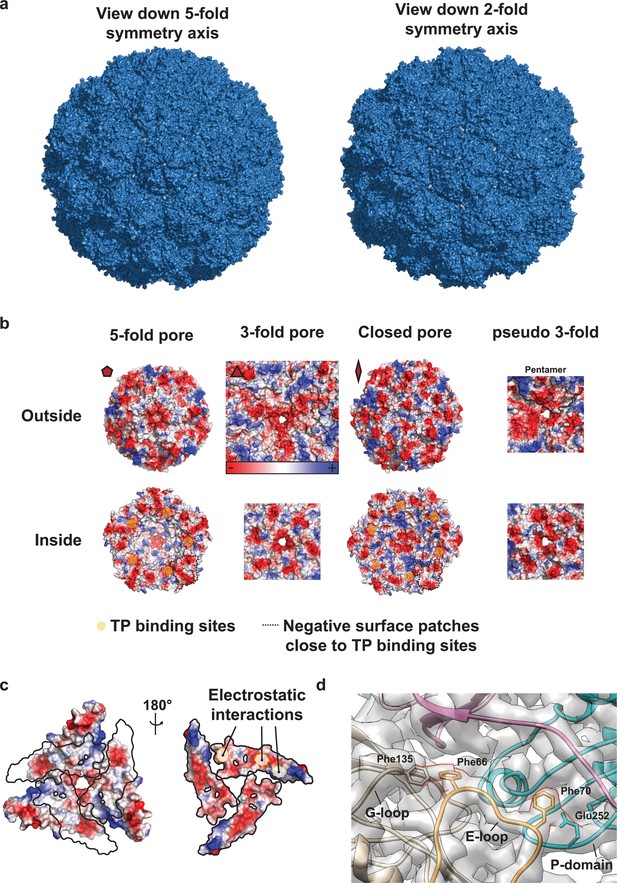
Structural details of the T4 encapsulin shell.
(a) Left: View down the 5-fold symmetry axis highlighting the 5-fold and pseudo 3-fold pores. Right: View down the 2-fold symmetry axis highlighting the 3-fold pores and the closed 2-fold ‘pore’. This surface representation indicates tight packing of monomers resulting in the 5- and (pseudo) 3-fold pores being the only conduits to the internal space created by the encapsulin shell. Pseudo 3-fold pores are defined as pores at the interface of 2 hexameric and one pentameric capsomer that in contrast to actual 3-fold pores (the interface of 3 hexameric capsomers) do not coincide with an icosahedral 3-fold symmetry axis. (b) Electrostatic surface representations of both the inside and outside views are shown. 5- and (pseudo) 3-fold pores are clearly negatively charged on both sides and all the way through the pore itself. The potential pore at the 2-fold symmetry axis is positively charged (outside) right at the pore entrance due to the presence of two asparagine residues closing off the pore while the inside of the 2-fold ‘pore’ is strongly negatively charged similar to the other pores. TP binding sites around the 5- and 2-fold symmetry axes are indicated with yellow circles and large negatively charged surface patches close to TP binding sites are outlined by black dotted lines. These patches might be involved in increasing cargo affinity for the interior encapsulin shell close to the binding site due to ionic interactions with positively charged residues of the long IMEF cargo linker. (c) Electrostatic surface models of P-domains and E-loops (outlines on the left and surface on the right) highlighting complementary electrostatic interactions around 3-fold pores. d, Specific interactions observed at the interface of 3 capsid monomers. Strong cryo-EM density observed connecting subunits are outlined with red dotted lines. This density suggests aromatic interactions for Phe135 and Phe66 and potential anion-π interactions for Phe70 and Glu252. The interactions between the G-loop, E-loop and P-domain of 3 different subunits are likely one of the factors responsible for the observed thermal stability of this system. It is of note that the proposed Phe70-Glu252 interaction in the T = 4 IMEF encapsulin is located at the same location as the isopeptide bond observed in the HK97 bacteriophage capsid.
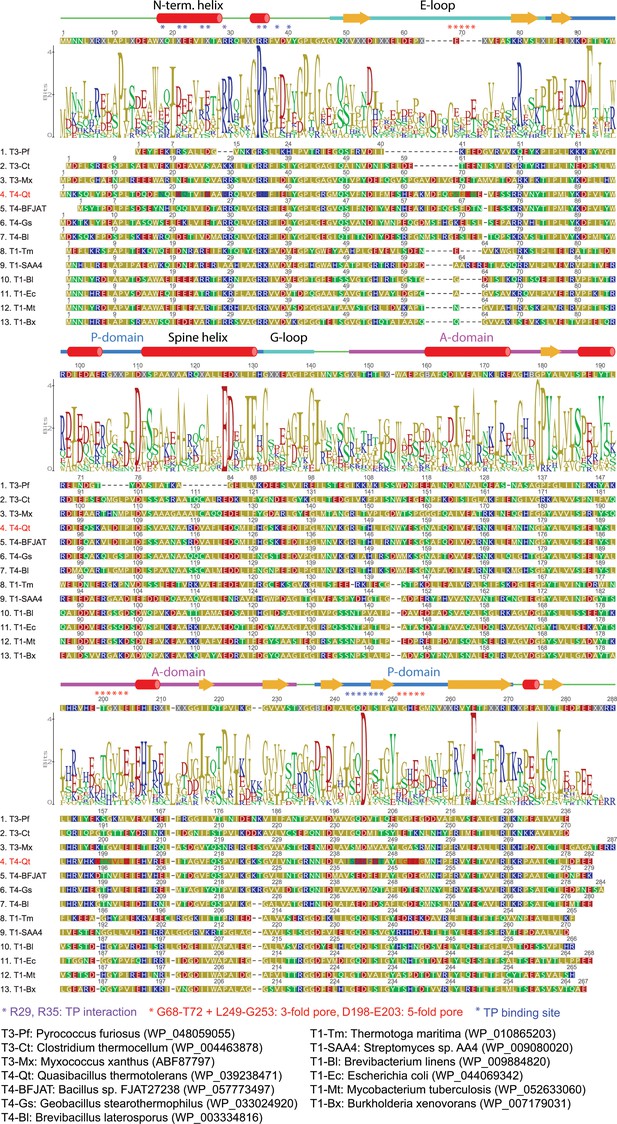
Sequence alignment of representative T = 1, T = 3 and T = 4 encapsulin capsid proteins.
Annotated secondary structural elements, the consensus sequence and a sequence logo are shown above the sequences. Organisms and protein sequences used are shown below the alignment. Residues important for TP interaction (purple), formation of the 3- and 5-fold pores (red) and in forming the overall TP binding site (blue) are indicated with asterisks.
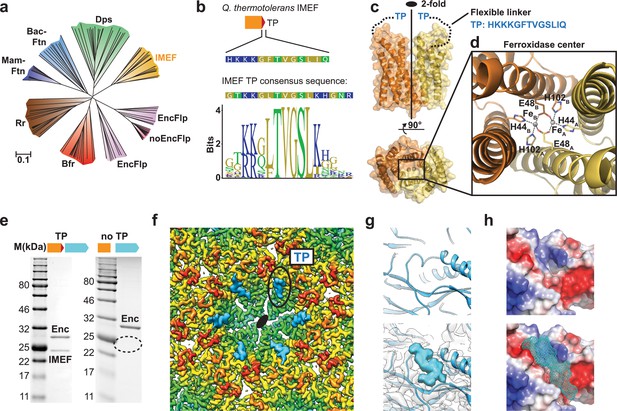
Structure and analysis of the IMEF cargo protein and TP-mediated cargo-shell co-assembly.
(a) Neighbor-joining phylogeny (cladogram) of protein classes involved in iron metabolism that are part of the Flp superfamily. Scale bar: amino acid substitutions per site. EncFlp: Flps found within encapsulin operons containing TPs, noEncFlp: Flps found outside encapsulin operons not containing TPs, Rr: rubrerythrins, Mam-Ftn: mammalian ferritins, Bac-Ftn: bacterial ferritins. (b) TP sequence of the Qs IMEF cargo protein and TP sequence logo highlighting strong sequence conservation. (c) X-ray crystal structure of the Qs IMEF cargo. (d) Di-iron ferroxidase active site of the IMEF cargo. The iron-coordinating residues are shown in stick representation. (e) SDS-PAGE gels of purified encapsulins showing that co-purification is dependent on the presence of the TP. (f) Cryo-EM map interior view of the 2-fold symmetry axis with TP density shown in cyan. (g) Close-up of additional cryo-EM density observed around the 2-fold symmetry axis. (h) Electrostatic surface representation of the TP binding site without (top) and with (bottom) TP. The 7 C-terminal IMEF residues are shown as a surface mesh.
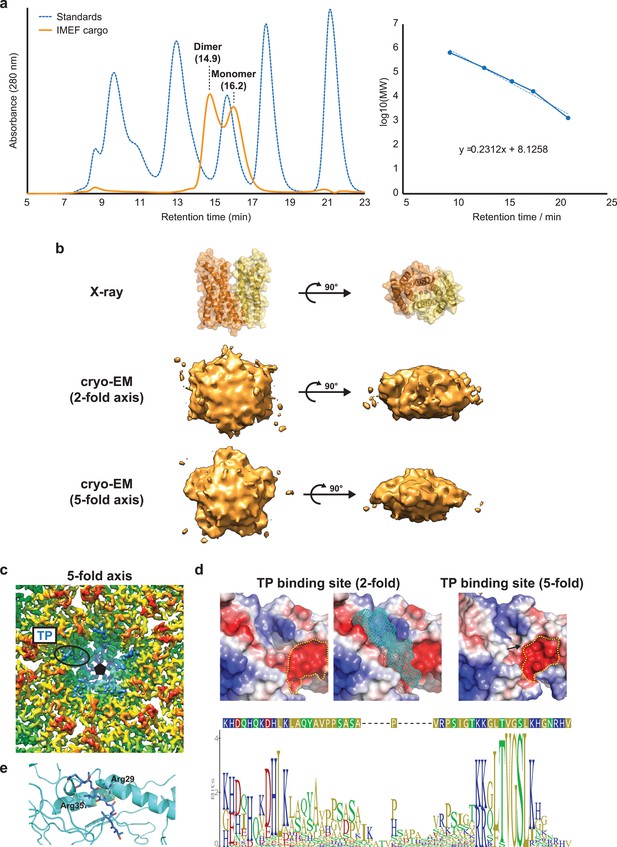
Biochemical and structural analysis of IMEF cargo loading.
(a) Left: Chromatogram indicating that His-tagged IMEF exists as a mixture of dimer and monomer in solution (in the absence of encapsulin). Right: Calibration curve used to estimate molecular weights and oligomerization state. (b) Top: X-ray structure of the dimeric IMEF cargo. Middle: Cargo density observed above hexameric capsomers (2-fold symmetry axis). Bottom: Cargo density observed above pentameric capsomers (5-fold symmetry axis). The symmetry of cryo-EM cargo densities indicates averaging during cryo-EM reconstruction. The sizes of the densities observed in combination with x-ray and gel filtration experiments suggests that the IMEF cargo is present in a dimeric form when encapsulated. Due to steric hinderance it is very unlikely that on average more than one dimer per hexameric or pentameric capsomer is present in a fully loaded IMEF encapsulin. This means that when fully loaded 42 IMEF dimers are present per T = 4 IMEF encapsulin. (c) TP densities observed around the 5-fold symmetry axis. The densities are weaker than densities observed around 2-fold symmetry axes (Figure 3f). (d) Top: 2- and 5-fold TP binding sites. A mesh model of the modeled TP is shown in cyan. Yellow dotted lines highlight large negative surface patches. Conformational changes lead to a less pronounced surface groove for the 5-fold binding site (black arrow). Overall the 2- and 5-fold binding sites are different due to the different conformations of capsid monomers when present in hexameric vs. pentameric capsomers, explaining the different binding affinities and observed differing TP density strengths. Bottom: Sequence logo and consensus sequence of the flexible linker and TP of all identified IMEF cargo proteins indicating the presence of many positively charged residues at both ends of the linker connecting IMEF and TP. e, TP binding site (2-fold) highlighting key ionic interactions. The shell monomer is shown as ribbons, the TP is shown in stick representation.
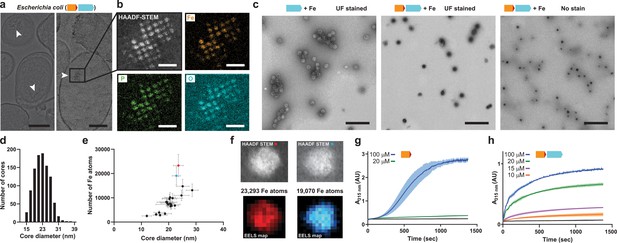
Mineralization of large iron-rich particles by the T = 4 encapsulin.
(a) Thin section micrographs of E. coli heterologously expressing the Qs IMEF core operon. Electron-dense particles often cluster together in regular arrays. Scale bars: 500 nm (left), 400 nm (right). (b) Close-up high angle angular dark field (HAADF) scanning TEM and EDS maps of a cluster of particles showing Fe, P and O as the main particle constituents. Scale bars: 100 nm. (c) Micrographs of uranyl formate (UF)-stained encapsulins produced in and isolated from E. coli grown in high iron media expressing the capsid protein alone (left) or the core operon (middle and right). Without UF stain, electron-dense particles are clearly visible (right). Scale bars: 250 nm. (d) Size distribution of electron-dense particles in unstained micrographs. (e) Electron energy loss spectroscopy (EELS) of 22 select cores carried out on isolated encapsulin particles. (f) HAADF-STEM micrographs and EELS maps of the two highlighted cores from (E). (g) In vitro ferroxidase assay of purified IMEF cargo at different Fe2+ concentrations. Mean values resulting from technical triplicates and error bands using standard deviation are shown. (h) Ferroxidase assay of cargo-loaded T = 4 encapsulin at different Fe2+ concentrations.
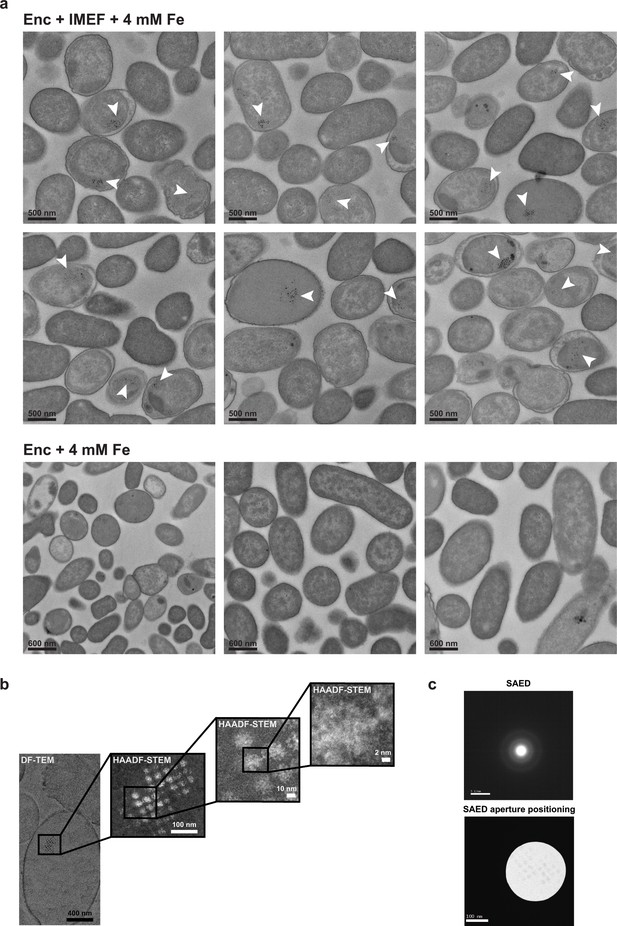
Heterologus IMEF cargo-dependent in vivo minearlization and characterization of iron-rich particles.
(a) Top: Representative fields of view for E. coli expressing the core IMEF operon (IMEF cargo + capsid protein) under high iron conditions (4 mM Fe(NH4)2(SO4)2).Clusters of electron-dense particles are highlighted with white arrows. Bottom: E. coli cells expressing only the IMEF encapsulin capsid protein without the IMEF cargo. No clusters of electron-dense particles were observed. (b) HAADF-STEM images at high resolution show irregular non-crystalline material. (c) Selected area electron diffraction (SAED) of electron-dense particles was carried out on thin sections of E. coli expressing the core IMEF operon targeting clusters of electron-dense particles. No diffraction spots and thus no crystallinity could be observed meaning that the electron-dense material deposited inside IMEF encapsulin shells is amorphous (supported by the presence of faint concentric rings indicative of an amorphous phase). HAADF-STEM: high angle angular dark field scanning TEM. DF-TEM: dark field TEM.
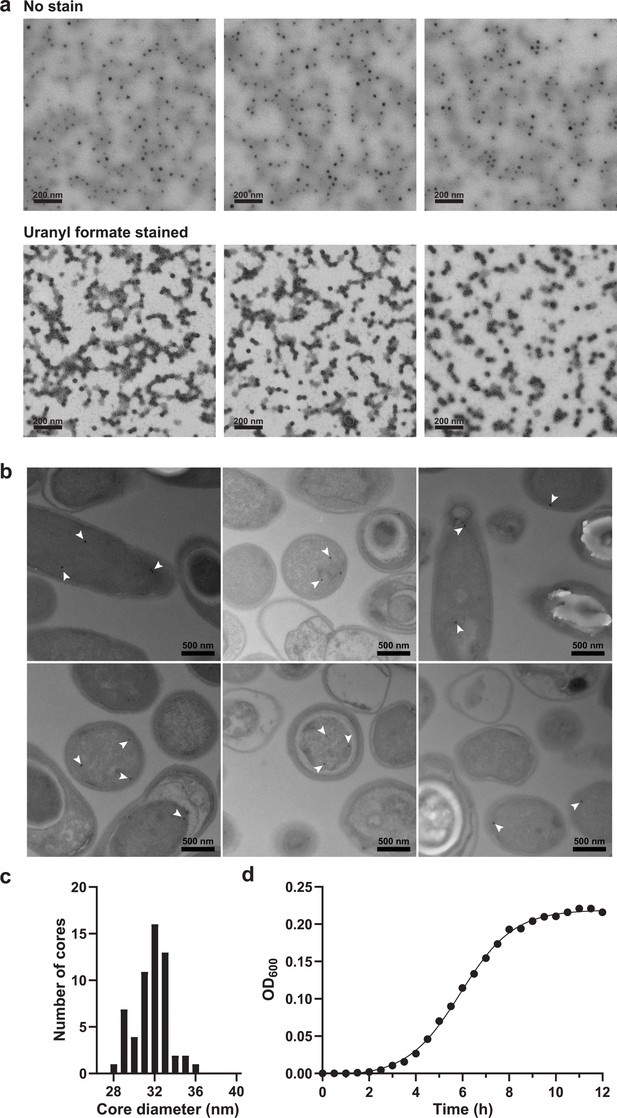

Purification of iron-loaded T4 encapsulins from E. coli and identification of electron-dense particles in Geobacillus.
(a) Top: Representative fields of unstained particles. Electron-dense cores are clearly visible even without stain. Bottom: Uranyl formate stained representative fields of the same particles. Images as shown on top were used to determine the core size distribution using ImageJ. (b) Geobacillus stearothermophilus ATCC 7953 was grown in high iron medium (4 mM) and samples prepared in early stationary phase (after 9 h). Representative fields of cells are shown with electron-dense cores highlighted by white arrows. Substantially fewer cores were observed in this strain that natively encodes the IMEF operon compared with recombinant E. coli heterologously expressing the IMEF operon. Consequently, no clustering of cores was observed. These thin section micrographs were not suitable for more detailed core analyses like EDS and EELS due to rapid carbon build-up and very high background. Thus, materials characterization of cores was done on purified iron-loaded particles that could be isolated in high quantity from recombinant E. coli resulting in less carbon build-up and better signal-to-noise. (c) Size distribution of native electron-dense cores formed in G. stearothermophilus. (d) Representative growth curve of G. stearothermophilus.
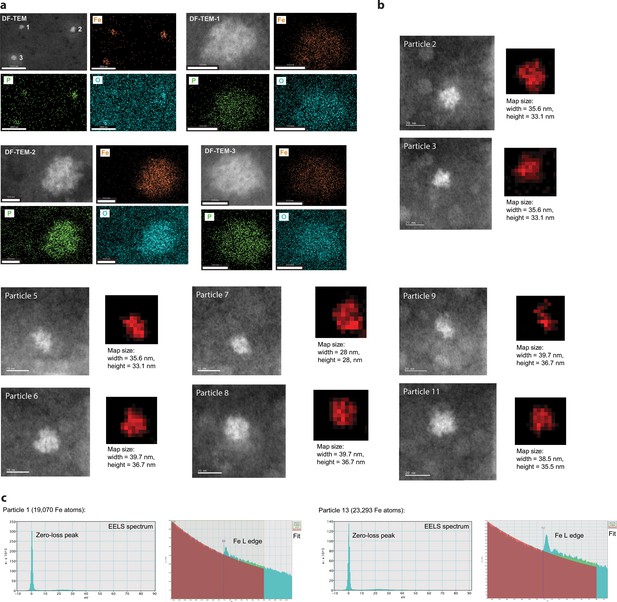
EDS and EELS analysis of purified iron-loaded T4 encapsulins.
(a) Three representative particles are shown with individual elemental EDS maps for Fe, P and O shown in orange, green and cyan, respectively. (b) Representative particles (HAADF-STEM images) and corresponding EELS maps are shown. (c) EELS spectra and fit for particle 1 and particle 13 (shown in Figure 4e and f in the main text, see also: Supplementary Table 5).
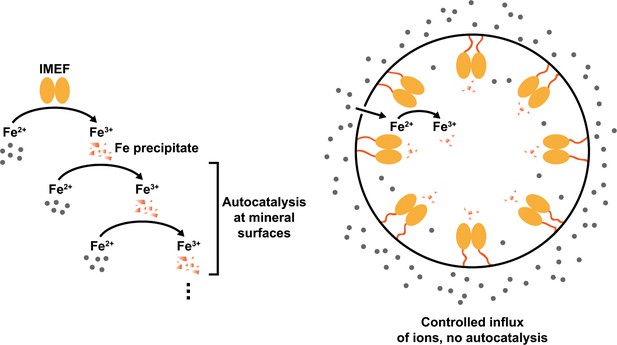
Model explaining the observed ferroxidase activities of free IMEF cargo (left) and the cargo-loaded IMEF encapsulin (right).
Without a protein shell creating a barrier between the compartment interior and outside, autocatalytic oxidation of ferrous iron on enzymatically formed ferric iron precipitates is observed (left) leading to a sigmoidal time-course curve as shown in Figure 4g in the main text. However, in the presence of a protein shell, a characteristic hyperbolic enzyme catalysis curve is observed (Figure 4h). This suggests that the encapsulin shell strictly controls the influx of iron to the compartment interior and thus the internal concentration of iron substrate available to encapsulated IMEF cargo proteins. The overall effect of this arrangement is that mineralization inside the IMEF encapsulin is controlled and autocatalytic runaway iron precipitation prevented. This is likely of key importance for the functioning of this novel iron storage system in the bacterial cytoplasm resulting in an iron storage system able to safely store essential but also toxic iron in a soluble and bioavailable form.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (E. coli) | MegaX DH10B T1R | Thermo Fischer Scientific | C640003 | Cloning strain |
| Strain, strain background (E. coli) | One Shot BL21 Star (DE3) | Thermo Fischer Scientific | C601003 | Expression strain |
| Strain, strain background (Geobacillus stearothermophilus) | ATCC 7953 | ATCC | ATCC 7953 | |
| Recombinant DNA reagent | pETDuet1 | EMD Millipore | 71146-3 | Expression vector |
| Sequence-based reagent | Codon-optimized IMEF cargo protein gene + encapsulin capsid protein gene containing overhangs for Gibson Assembly (oligonucleotide gBlock) | Integrated DNA Technologies (IDT) | based on accessions: WP_039238473.1; WP_03923847 | gttaagtataagaaggagatatacaATGAAGGAAGAACTGGATGCTTTCCATCAGATTTTCACTACGACCAA AGAGGCAATCGAACGTTTTATGGCGATGCTGACCCCGGTCATTGAGAACGCGGAGGACGATCATGAGCGCCTGTATTATCATCATATCTACGAAGAGGAGGAGCAACGTCTGTCGCGCCTGGACGTTCTGATCCCACTGATCGAAAAGTTTCAAGATGAAACCGACGAAGGCCTCTTCTCCCCCTCCAACAACGCCTTTAACCGTCTGCTTCAGGAGCTGAATCTGGAAAAATTCGGTTTGCATAACTTTATCGAGCATGTTGACCTGGCCCTTTTTAGTTTCACCGACGAGGAACGCCAGACATTGCTTAAAGAACTGCGTAAAGATGCCTATGAAGGCTATCAGTATGTTAAAGAAAAACTGGCAGAAATTAACGCTCGTTTTGATCACGATTACGCAGACCCGCATGCGCACCATGATGAACACCGTGACCATCTTGCGGATATGCCCTCAGCGGGTTCATCGCACGAAGAAGTGCAGCCTGTTGCACATAAAAAGAAAGGTTTCACGGTGGGTTCATTAATCCAGTAAATTTCGCTTAAATATTACCGCTAGCTCAAAAAGGAGGAAAAGTGAATGAACAAAAGCCAACTTTATCCGGATTCACCACTGACGGATCAGGACTTCAACCAATTAGACCAAACCGTGATTGAGGCTGCTCGTCGTCAGCTGGTGGGTCGTCGCTTCATTGAGTTATATGGCCCATTGGGGCGTGG CATGCAGAGTGTCTTCAACGATATCTTCATGGAGTCTCATG AAGCGAAAATGGACTTCCAGGGCAGCTTTG ACACGGAGGTAGAGTCCTCCCGTCGTGTAAACTATACCATTCCGATGTTATATAAAGACTTCGTGCTTTACTGGCGCGATCTGGAACAGAGCAAGGCACTCGATATTCCGATCGACTTTTCAGTGGCAGCGAACGCTGCCCGCGACGTTGCGTTCCTGGAAGATCAGATGATTTTCCATGGAAGCAAAGAATTTGATATCCCGGGTCTGATGAACGTGAAAGGTCGCCTGACCCATCTGATTGGCAATTGGTATGAGTCGGGTAACGCCTTTCAGGATATTGTGGAGGCCCGCAATAAATTACTCGAAATGAACCACAATGGCCCATATGCTCTCGTGCTGTCCCCGGAGCTGTACTCACTCTTA CATCGTGTGCATAAAGACACGAATGTGCTGGAGATCGAACACGTGCGCGAGTTGATTACTGCTGGGGTTTTTCAGTCGCCTGTCCTCAAAGGGAAAAGTGGTGTGATCGTAAACACCGGTCGCAACAATCTGGATTTGGCTATCTCGGAAGATTTTGAGACTGCATACCTGGG CGAGGAAGGTATGAACCATCCCTTTCGCGTGTACGAGACAGTTGTTCTGCGCATCAAACGCCCGGCGGCCATTTGTACTTTAATCGATCCGGAAGAATAAattaacctaggctgctgccaccgct |
| Sequence- based reagent | Codon-optimized IMEF cargo protein gene w/o TP + encapsulin capsid protein gene containing overhangs for Gibson Assembly (oligonucleotide gBlock) | Integrated DNA Technologies (IDT) | based on accessions: WP_039238473.1; WP_039238471 | gttaagtataagaaggagatatacaATGAAGGAAGAACTGGATGCTTTCCATCAGATTTTCACTACGACCAAAGAGGCAATCGAACGTTTTATGGCGATGCTGACCCCGGTCATTGAGAACGCGGAGGACGATCATGAGCGCCTGTATTATCATCATATCTACGAAGAGGAGGAGCAACGTCTGTCGCGCCTGGACGTTCTGATCCCACTGATCGAAAAGTTTCAAGATGAAACCGACGAAGGCCTCTTCTCCCCCTCCAACAACGCCTTTAACCGTCTGCTTCAGGAGCTGAATCTGGAAAAATTCGGTTTGCATAACTTTATCGAGCATGT TGACCTGGCCCTTTTTAGTTTCACCGACGAGGAACGCCAGACATTGCTTAAAGAACTGCGTAAAGATGCCTATGAAGGCTATCAGTATGTTAAAGAAAAACTGGCAGAAATTAACGCTCGTTTTGATCACGATTACGCAGACCCGCATGCGCACCATGATGAACACCGTGACCATCTTGCGGATATGCCCTCAGCGGGTTCATCGCACGAAGAAGTGCAGCCTGTTGCATAAATTTCGCTTAAATATTACCGCTAGCTCAAAAAGGAGGAAAAGTGAATGAACAAAAGCCAACTTTATCCGGATTCACCACTGACGGATCAGGACTTCAACCAATTAGACCAAACCGTGATTG AGGCTGCTCGTCGTCAGCTGGTGGGT CGTCGCTTCATTGAGTTATATGGCCCA TTGGGGCGTGGCATGCAGAGTGTCTTCAACGATATCTTCATGGAGTCTCATGAAGCGAAAATGGACTTCCAGGGC AGCTTTGACACGGAGGTAGAGTCCTCCCGTCGTGTAAACTATACCATTCCGATGTTATATAAAGACTTCGTGCTTTACTGGCGCGATCTGGAACAGAGCAAGGCACTCGATATTCCGATCGACTTTTCAGTGGCAGCGAACGCTGCCCGCGACGTTGCGTTCCTGGAAGATCAGATGATTTTCCATGGAAGCAAAGAATTTGATATCCCGGGTCT GATGAACGTGAAAGGTCGCCTGACCCATCTGATTGGCAATTGGTATGAG TCGGGTAACGCCTTTCAGGATATTG TGGAGGCCCGCAATAAATTACTCGAAATGAACCACAATGGCCCATATGCTCTCGTGCTGTCCCCGGAGCTGTACTCACT CTTACATCGTGTGCATAAAGACACGAATGTGCTGGAGATCGAACACGTGCGCGAGTTGATTACTGCTGGGGTTTTTCAGTCGCCTGTCCTCAAAGGGAAAAGTGGTGTGATCGTAAACACCGGTCGCAACAATCTGGATTTGGCTATCTCGGAAGATTTTGAGACTGCATACCTGGGCGAGGAAGGTATGAACCATCCCTTTCGCGTGTACGAGACAGTTGTTCTGCGCATCAAACGCCCGGCGGCCATTTGTACTTTAATCGATCCGGAAGAATAAattaacctaggctgctgccaccgct |
| Commercial assay or kit | Gibson Assembly Master Mix | New England Biolabs | E2611L | |
| Commercial assay or kit | 14% Novex Tris-Glycine Gel | Thermo Fischer Scientific | XP00140BOX | |
| Commercial assay or kit | MIDAS screen | Molecular Dimensions | MD1–59 | Crystallization screen |
| Commercial assay or kit | Pierce Coomassie Plus (Bradford) Assay | Thermo Fischer Scientific | 23236 | Protein concentration determination4 |
| Chemical compound, drug | Isopropyl-β-D-thiogalactoside | Millipore Sigma | 10724815001 | |
| Chemical compound, drug | Lysozyme | Millipore Sigma | L6876 | |
| Chemical compound, drug | DNAse I | Millipore Sigma | 11284932001 | |
| Chemical compound, drug | Ni-NTA agarose resin | Qiagen | 30210 | |
| Chemical compound, drug | Polyethylene glycol 8000 | Millipore Sigma | 1546605 | |
| Chemical compound, drug | Uranyl formate | EMS | 22450 | |
| Chemical compound, drug | Formaldehyde 37% in water | Millipore Sigma | 252549 | |
| Chemical compound, drug | Glutaraldehyde 25% in water | Millipore Sigma | G5882 | |
| Chemical compound, drug | Picric acid | Millipore Sigma | 197378 | |
| Chemical compound, drug | Sodium cacodylate | Millipore Sigma | C0250 | |
| Chemical compound, drug | Uranyl acetate | EMS | 22400 | |
| Chemical compound, drug | Propylene oxide | Millipore Sigma | 82320 | |
| Chemical compound, drug | Epon | EMS | 14910 | |
| Chemical compound, drug | Glycolic acid | Millipore Sigma | 798053 | |
| Chemical compound, drug | Trisodium citrate | Millipore Sigma | S1804 | |
| Chemical compound, drug | Ammonium iron (II) sulfate | Millipore Sigma | F1543 | |
| Chemical compound, drug | ortho-phenylenediamine | Millipore Sigma | P9029 | |
| Chemical compound, drug | Hydrogen peroxide 30 % in water | Millipore Sigma | 216763 | |
| Software, algorithm | Genome Neighborhood Network Tool (GNT) | Gerlt et al., 2015 | https://efi.igb.illinois.edu/efi-gnt/ | |
| Software, algorithm | blastp | NIH NCBI | https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins | |
| Software, algorithm | Clustal Omega | McWilliam et al., 2013 | https://www.ebi.ac.uk/Tools/msa/clustalo/ | |
| Software, algorithm | Simply Phylogeny | Madeira et al., 2019 | https://www.ebi.ac.uk/Tools/phylogeny/simple_phylogeny/ | |
| Software, algorithm | Geneious 9.14 | Biomatters Ltd | https://www.geneious.com/ | |
| Software, algorithm | UCSF Chimera 1.13 | Pettersen et al., 2004 | https://www.cgl.ucsf.edu/chimera/ | |
| Software, algorithm | Open Source PyMOL | Schroedinger LLC | https://github.com/schrodinger/pymol-open-source | |
| Software, algorithm | I-TASSER | Roy et al., 2010 | https://zhanglab.ccmb.med.umich.edu/I-TASSER/ | |
| Software, algorithm | IDT Codon Optimization Tool | Integrated DNA Technologies (IDT) | https://www.idtdna.com | |
| Software, algorithm | MotionCor2 | Zheng et al., 2017 | https://omictools.com/motioncor2-tool | |
| Software, algorithm | CTFFIND4 | Rohou and Grigorieff, 2015 | http://grigoriefflab.janelia.org/ctffind4 | |
| Software, algorithm | SAMUEL | Liao Lab | https://liao.hms.harvard.edu/samuel | |
| Software, algorithm | Sam Viewer | Liao Lab | https://liao.hms.harvard.edu/samviewer | |
| Software, algorithm | Relion 3.0 | Scheres, 2012 | https://www3.mrc-lmb.cam.ac.uk/relion/index.php?title=Main_Page | |
| Software, algorithm | SPIDER | Frank et al., 1996 | https://spider.wadsworth.org/spider_doc/spider/docs/spider.html | |
| Software, algorithm | ResMap | Swint-Kruse and Brown, 2005 | http://resmap.sourceforge.net/ | |
| Software, algorithm | Coot 0.8.9.1 | Emsley et al., 2010 | https://www2.mrc-lmb.cam.ac.uk/personal/pemsley/coot/ | |
| Software, algorithm | Phenix 1.14 | Adams et al., 2010 | http://www.phenix-online.org/ | |
| Software, algorithm | XDS | Kabsch, 2010 | http://xds.mpimf-heidelberg.mpg.de/ | |
| Software, algorithm | ACRIMBOLDO_LITE | Sammito et al., 2015 | http://chango.ibmb.csic.es/arcimboldo_lite | |
| Software, algorithm | Phaser | McCoy et al., 2007 | https://www.phaser.cimr.cam.ac.uk/index.php/Phaser_Crystallographic_Software | |
| Software, algorithm | SHELX | Thorn and Sheldrick, 2013 | http://shelx.uni-goettingen.de/ | |
| Software, algorithm | CCP4 | Winn et al., 2011 | http://www.ccp4.ac.uk/ | |
| Software, algorithm | REFMAC5 | Murshudov et al., 1997 | http://www.ccp4.ac.uk/html/refmac5.html | |
| Software, algorithm | Fiji-ImageJ 1.52h | Schindelin et al., 2012 | https://fiji.sc/ | |
| Software, algorithm | UCSFImage4 | omicX | https://omictools.com/ucsfimage-tool | |
| Other | 200 Mesh Gold Grids | EMS | FCF-200-Au | |
| Other | 400 Mesh Cu Holy Carbon Grids | EMS | Q410CR1.3 |
Additional files
-
Supplementary file 1
Table of ferritin-like proteins (Flps) identified in IMEF operon-containing Firmicutes.
All identified ferritins (Ftn), bacterioferritins (Btf) and DNA-binding proteins from starved cells (Dps) found in IMEF operon strains. IMEF and encapsulin capsid protein IDs are shown as well. 97% of IMEF operon-containing strains do not encode Ftn, 93% do not encode Bfr and 92% do not encode either. However, 93% of IMEF operon-encoding strains encode Dps systems. This likely indicates that the encapsulin based IMEF system represents the major iron storage system in 92% of the listed strains. It also indicates that IMEF systems do likely not function as unusual Dps system given that the vast majority of strains encode standard Dps system. Blast searches were carried out using the NCBI Blastp server with the following sequences as queries: Ftn: OTY20392, Bfr: EEK74551, Dps: WP_039234032, IMEF: WP_039238473, Encapsulin: WP_039238471.
- https://doi.org/10.7554/eLife.46070.018
-
Supplementary file 2
Table of cryo-EM data collection statistics for IMEF-loaded encapsulin.
- https://doi.org/10.7554/eLife.46070.019
-
Supplementary file 3
List of ferritin-like protein and IMEF protein IDs used to construct the phylogenetic tree shown in Figure 3A.
IMEF: Iron-mineralizing encapsulin-associated Firmicute cargo, EncFlp: ferritin-like proteins (Flps) found within encapsulin operons containing targeting peptides, noEncFlp: Flps found outside encapsulin operons not containing a targeting peptide, Bfr: bacterioferritin, Rr: rubrerythrin, Mam-Ftn: mammalian ferritin, Bac-Ftn: bacterial ferritin, Dps: DNA-binding proteins from starved cells.
- https://doi.org/10.7554/eLife.46070.020
-
Supplementary file 4
X-ray structure determination and refinement statistics for the IMEF cargo protein.
- https://doi.org/10.7554/eLife.46070.021
-
Supplementary file 5
EELS data of electron dense cores of purified IMEF encapsulins produced in E. coli under high iron conditions.
#Fe from EELS analysis values were extracted using Gatan software. Errors combine estimated statistical error in this measurement with known error for cross section. For density calculations, particles were approximate as spheres.
- https://doi.org/10.7554/eLife.46070.022
-
Transparent reporting form
- https://doi.org/10.7554/eLife.46070.023





