Density-dependent resistance protects Legionella pneumophila from its own antimicrobial metabolite, HGA
Figures
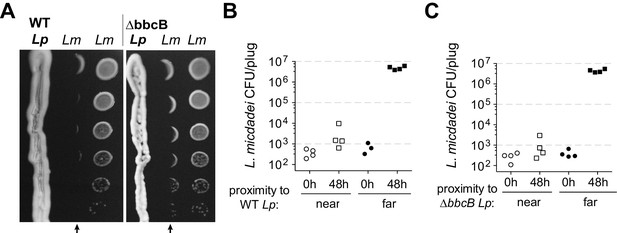
L. pneumophila (Lp) produces a secreted inhibitor independent of surfactant.
(A) When pre-incubated on low-cysteine BCYE charcoal agar plates, Lp produces a zone of inhibition, impacting the growth of nearby L. micdadei (Lm). Arrows mark the edge of inhibition fronts. Droplets of Lm at different dilutions were added to the plate in parallel columns three days after streaking Lp. WT Lp (left panel) generates a similar zone of inhibition as a surfactant-null mutant, ∆bbcB (right panel). (B) Quanitification of Lm growth within (‘near’) or outside of (‘far’) the wild type Lp zone of inhibition. (C) Quanitification of Lm growth within or outside of the ∆bbcB Lp zone of inhibition. In B and C, bacteria were sampled and removed from the plate in a ‘plug’ of fixed area before plating for viable CFUs.
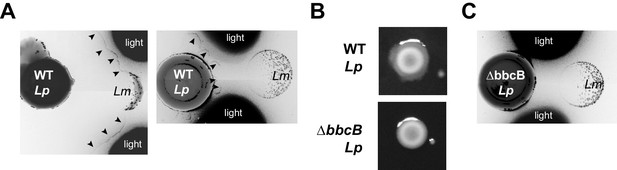
Separation of surfactant and antimicrobial phenotypes.
(A) Lm inhibition sometimes co-occurs with contact with WT Lp’s secreted surfactant (left), but sometimes is observable outside of the surfactant front (right). The leading edge of the surfactant front (arrowheads) is faintly visible when light is reflected off adjacent regions of the plate (‘light’). Each image is a composite of two fields of view, with inverted colors to enable visualization of the surfactant front. (B) Lack of surfactant production from Lp with a deletion of bbcB, evident by a defect in spreading on BCYE plates. (C) The bbcB mutant can inhibit the growth of neighboring Lm, despite lacking surfactant secretion.
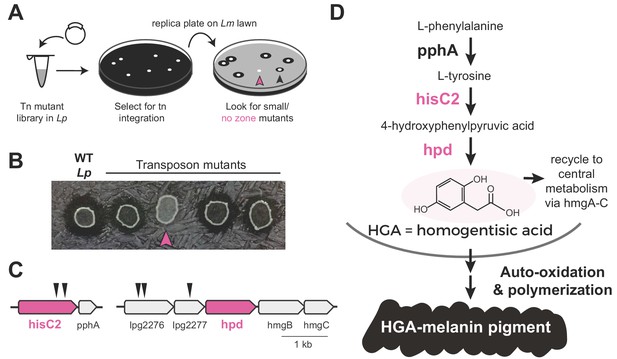
Transposon mutagenesis screen implicates the HGA-melanin pathway in production of the inhibitor.
(A) Screen for mutant L. pneumophila (Lp) that do not inhibit L. micdadei (Lm). Following electroporation of a Mariner-transposon-containing plasmid, Lp mutants were selected for transposon integration. Colonies were patched or replica plated onto a lawn of Lm. Mutants of interest generated a zone of inhibition that was reduced (black arrowhead) or absent (pink arrowhead) compared to WT Lp. (B) Selected transposon mutants produce abnormal zones of inhibition when grown on a lawn of Lm. Pink arrowhead indicates a hisC2::Tn ‘no zone’ mutant. (C) Transposon insertion sites (triangles) identified in the five recovered ‘no zone’ mutants. (D) HGA-melanin synthesis pathway. HGA is exported from the cell where it auto-oxidizes and polymerizes to form HGA-melanin. Genes in pink were validated by complementation to have essential roles in Lm inhibition.
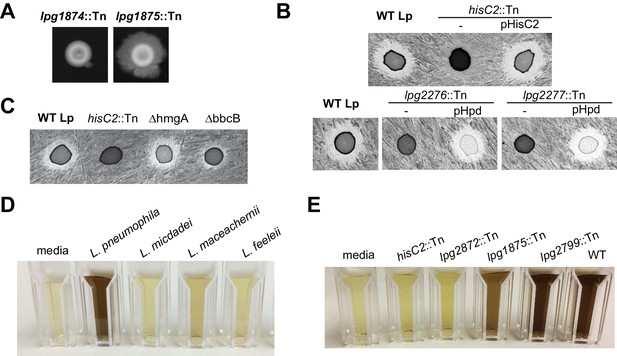
Genetic validation linking inhibition-defective mutants to the HGA-melanin pathway.
(A) Inter-bacterial inhibition does not correlate with surface spreading. For example, two mutants with transposon insertions in lpg1874 (general secretion system protein L) and lpg1875 (general secretion system protein M) share ‘small zone’ inhibition phenotypes, yet show opposite spreading phenotypes on BCYE. (B) Overexpression of either hisC2 or hpd from a plasmid was sufficient to complement the ‘no zone’ phenotype in all recovered mutants. (C) Unlike mutations to hisC2, deletion of the hmgA gene does not disrupt Lm inhibition, showing that the intracellular recycling of HGA is not required for inhibitor production. The bbcB gene is not required for inhibition, but generates a smaller zone of inhibition than wild type. Colors in images from B and C were inverted to facilitate visualization of the zone of inhibition. (D) Pigmentation of AYE conditioned media following 48 hr growth of various Legionella species. While multiple species produce some pigment, many are less pigmented than L. pneumophila. The L. micdadei susceptible strain does not secrete detectable pigment. (E) After 48 hr growth in AYE, none of the ‘no zone’ mutants (represented here by hisC2::Tn) produce pigment. A subset of ‘small zone’ mutants also have pigmentation defects (e.g. lpg2872::Tn and lpg1875::Tn), further implicating the HGA-melanin pathway in inhibition.
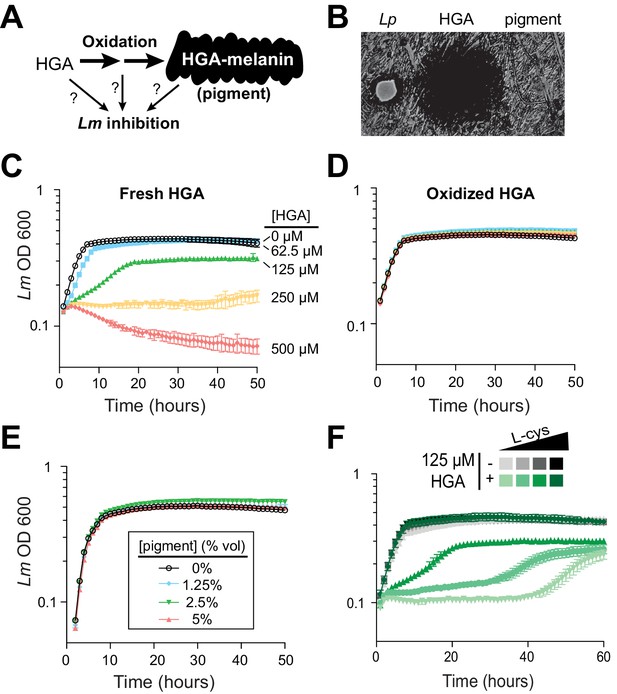
Synthetic HGA inhibits Legionella micdadei growth, depending on its oxidation state.
(A) We tested whether inter-bacterial inhibition is caused by HGA, HGA-melanin, or an oxidative intermediate. (B) Zones of inhibition on a lawn of Lm generated from either live Lp bacteria, synthetic 50 mM HGA, or concentrated pigment extract. HGA prevents Lm growth in a large region (central dark circle) but pigment does not. (C) Growth inhibition of Lm from increasing concentrations of synthetic HGA in rich AYE media. (D) Pre-oxidation of synthetic HGA in AYE media for 24 hr eliminates its inhibitory activity, resulting in normal Lm growth. Concentrations of HGA colored as in panel C. (E) Addition of oxidized HGA-melanin pigment from Lp has little impact on Lm growth in AYE liquid media. (F) In the absence of HGA (gray symbols), titration of L-cysteine (L-cys) from 25–200% of standard AYE media has little impact on Lm growth. In contrast, HGA activity is enhanced in low-cysteine media and decreased in high-cysteine media (green symbols). All error bars show standard deviations among 3–4 replicates.
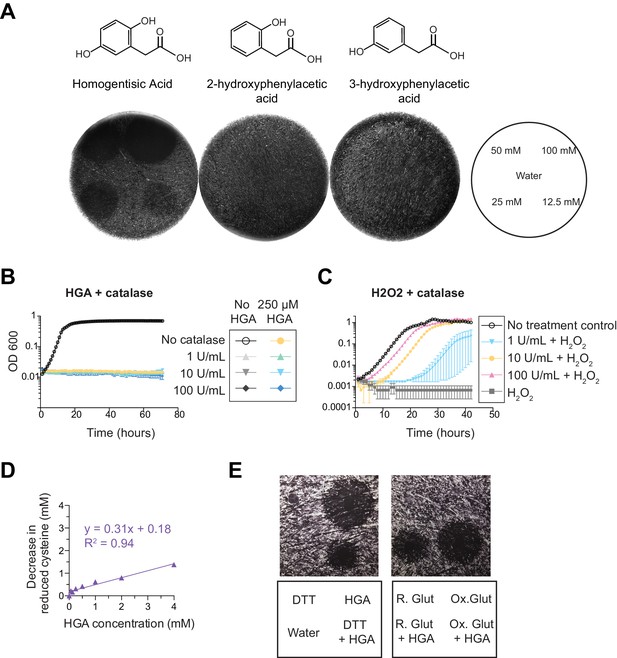
Impacts of chemical compounds on HGA-mediated inhibition of Legionella.
(A) HGA inhibits Lm growth but HGA-related compounds do not. Chemical names and structures are shown above the corresponding plates. On each plate, different concentrations of each compound were spotted onto a lawn of Lm in 10 µL droplets, arranged as indicated at the right. (B) Addition of catalase does not rescue Lm susceptibility to 250 µM HGA. (C) Control experiment showing catalase is active, based on its ability to protect Lp from H2O2. (D) Quantification of HGA’s ability to oxidize cysteine. Data points are in purple, with a linear line of best fit. (E) The potency of HGA is decreased when pre-incubated with reducing agents. 100 mM HGA was mixed with dithiothreitol (DTT), oxidized glutathione (Ox. Glut), or reduced glutathione (R. Glut) for 15 min prior to spotting 10 µL onto a lawn of Lm and allowing to grow for 3 days. Key below each image indicates where each solution was added.
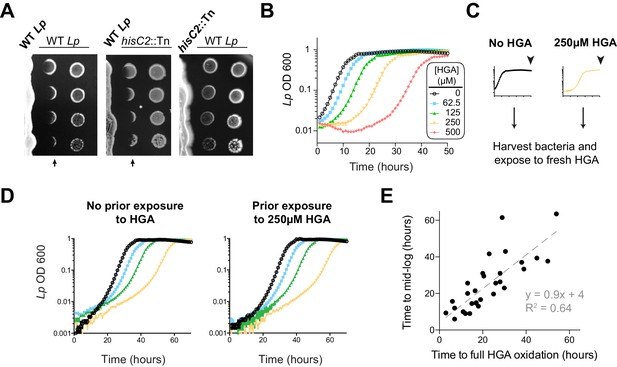
L. pneumophila is susceptible to bacteriostatic inhibition by HGA in rich media.
(A) When pre-incubated on agar plates, Lp produces a zone of inhibition (arrows), preventing the growth of genetically-identical Lp plated 3 days later. The ‘no zone’ mutant hisC2::Tn does not produce a sharp front of inhibition, implicating HGA. (B) Increasing concentrations of synthetic HGA inhibit the growth of Lp, causing a growth delay in rich media. Error bars showing standard deviation are small and mostly obscured by the symbols. (C) To test if Lp population recovery at late time points following HGA exposure is due to the outgrowth of HGA-resistant mutants, we grew Lp with or without HGA and sampled bacteria at the end of the experiment (arrowhead) that were unexposed to HGA or were exposed to 250 µM HGA. These were used to inoculate media ± fresh HGA. (D) Prior HGA exposure did not lead to subsequent resistance. (E) The time for Lp to grow in the presence of HGA is correlated with the time for synthetic HGA to oxidize at each concentration, suggesting that Lp delays growth until HGA has sufficiently oxidized to lose inhibitory activity. Plot shows data combined across eight experiments. Linear fit curve and equation are shown.
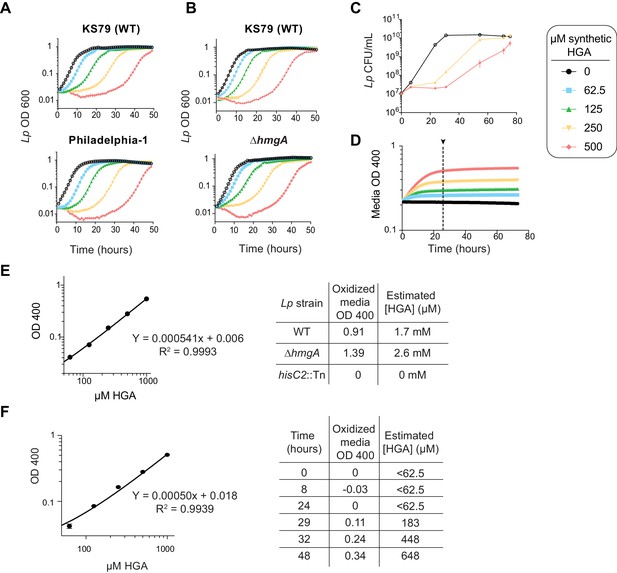
HGA-induced growth delays and quantification of HGA production.
(A–B) Growth curves of Lp strains upon exposure to HGA. For each experiment, we provide a matched wild type (KS79) control for comparison. (A) Clinical isolate Philadelphia-1 and lab strain KS79 exhibited nearly identical growth curves in the presence of HGA. (B) The ∆hmgA deletion strain responded to HGA similarly to wild type KS79, showing that HmgA-C do not play a significant role in HGA susceptibility. (C) Viable CFU counts of KS79 Lp exposed to HGA. Even at high HGA concentrations, in rich media HGA is bacteriostatic at early time points, followed by population recovery. (D–F) Using standard curves of synthetic HGA to estimate the amount of HGA secreted by Lp. (D) OD400 was used to track the oxidation of synthetic HGA in AYE media. The 24 hr timepoint (arrow) was used for the standard curve and all experiments estimating HGA concentrations, as the HGA had completed oxidation by this point. (E) Standard curve showing OD400 of oxidized synthetic HGA used to estimate the amount of HGA secreted into Lp conditioned media after 48 hr growth. The equation for the linear regression is shown along with a table with the OD400 readings and estimated HGA concentrations for each sample. (F) Similar standard curve used for time course experiment of HGA secretion in wild type Lp in Figure 6. Standard curves were generated for each experiment independently, in parallel with experimental samples.
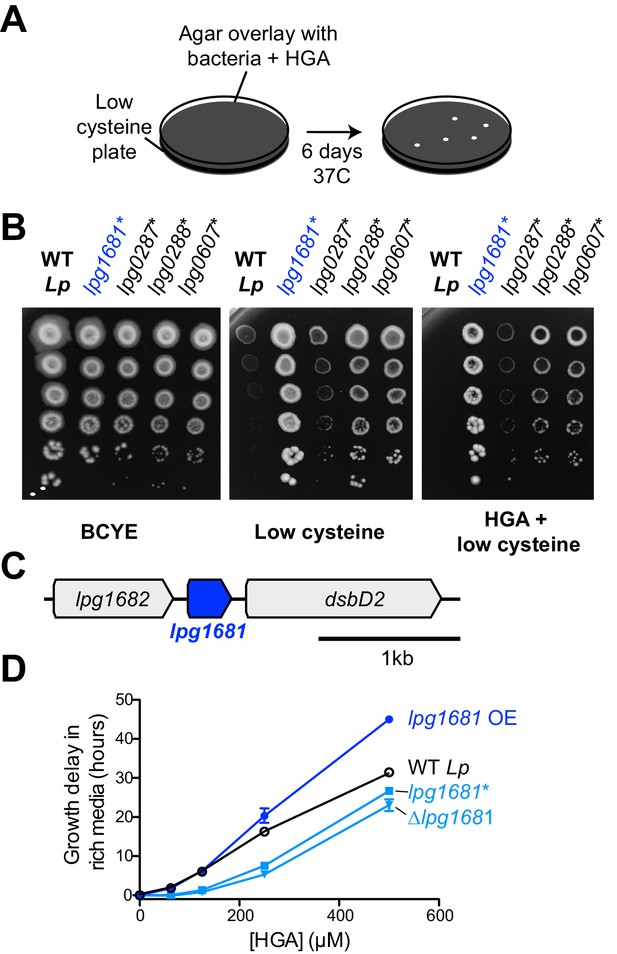
L.pneumophila susceptibility to HGA is modulated by lpg1681.
(A) Scheme to select for Lp spontaneous HGA-resistant mutants. (B) Growth of HGA-selected mutants (*) compared to wild type Lp. All isolates grew better than wild type in selection conditions (HGA + low cysteine), as well as in low cysteine conditions. (C) Syntenic region of lpg1681 in Lp strains. Lpg1681 is a hypothetical gene that lies downstream of lpg1682, a predicted oxidoreductase/dehydrogenase, and upstream of dsbD2, a thiol:disulfide interchange protein. (D) In rich media, a spontaneous lpg1681 mutant (lpg1681*) and the lpg1681 deletion strain (∆lpg1681) are less sensitive to growth inhibition by HGA than wild type Lp, as seen by a shorter growth delay at each concentration of HGA. Overexpression of lpg1681 (OE) heightens sensitivity to HGA (longer growth delay). Graphs here summarize experiments similar to those in Figure 4B. See Figure 5—figure supplement 2 for full data.
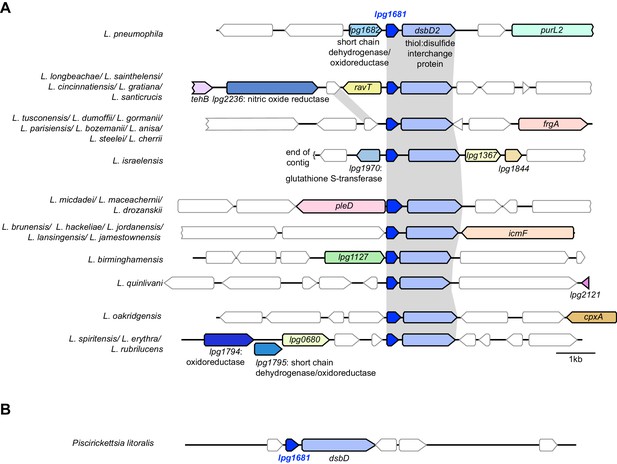
Evolution of genes in the lpg1681 locus among A) Legionella species and B) the fish pathogen Piscirickettsia.
In the species that carry hypothetical gene lpg1681, it always resides upstream of the thiol:disulfide interchange gene dsbD2, despite extensive turnover of neighboring genes. Species separated by a ‘/” have similar syntenic loci. Annotated genes are colored, with redox-related genes in shades of blue. Hypothetical genes are white. Gray shading indicates gene homology among the species.
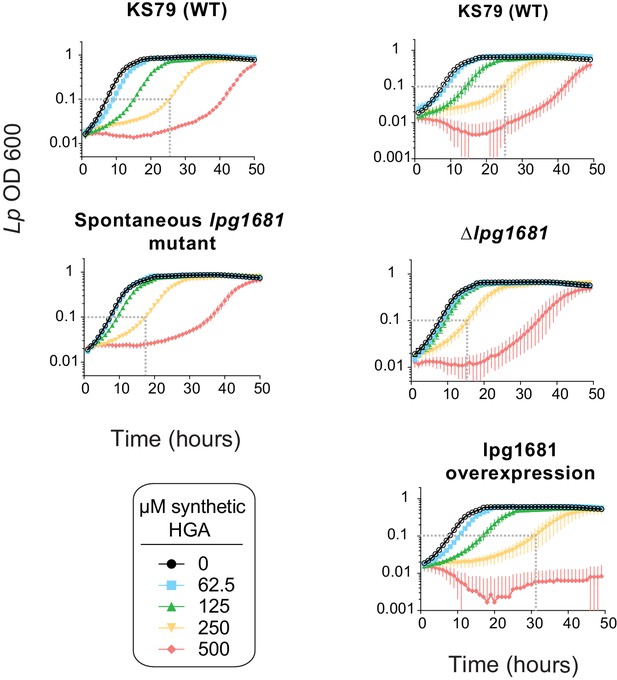
Growth curves of lpg1681 Lp strains upon exposure to HGA.
For each experiment we provide a matched wild type (KS79) control for comparison. The spontaneous lpg1681 mutant and the deletion mutant both had less severe growth delays from HGA than wild type. Conversely, the lpg1681 overexpression strain was sensitized to HGA, with longer growth delays than wild type. Dashed gray line shows time for the 250 µM HGA condition to reach an OD of 0.1, to facilitate comparisons.
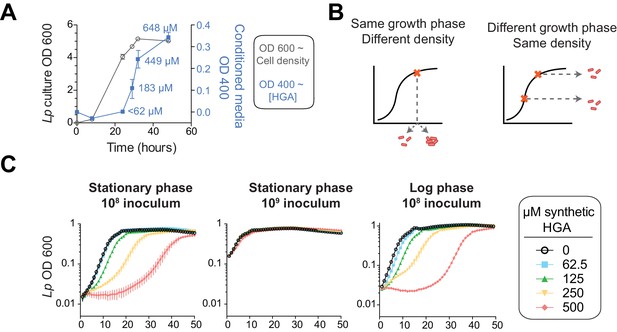
HGA susceptibility in Lp is linked to cell density, independent of inoculum growth phase.
(A) Timing of HGA secretion in Lp in rich media, measured by OD 400 of conditioned media (CM) after allowing for full HGA oxidation (blue boxes, right y-axis). A matched growth curve of Lp is presented for comparison (gray circles, left y-axis). Abundant HGA is secreted during stationary phase. Estimates of secreted HGA concentration (blue) are based on a standard curve of synthetic HGA. Error bars show standard deviations. (B) Schematic showing how experiments controlled for inoculum density vs. growth phase. To test the impact of cell density, a single culture was diluted to multiple densities at the start of the experiment. To test the impact of growth phase, Lp was sampled at multiple stages of growth and diluted to the same CFU/mL at the start of the experiment. (C) Lp growth in AYE rich media is inhibited by HGA when cells are inoculated at relatively low density (10^8 CFU/mL), but HGA is ineffective in inhibiting growth when cells are inoculated at high density (10^9 CFU/mL). Growth phase of Lp inoculum has little impact on HGA susceptibility.
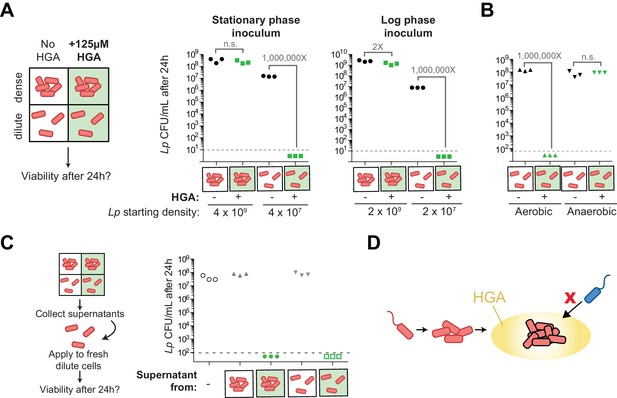
Density-dependent HGA susceptibility is heightened in low-nutrient media and requires HGA’s oxidative intermediates, while high-density resistance is cell-intrinsic.
(A) Viable CFUs following 24 hr incubation of Lp with or without 125 µM HGA in PBS. When incubated at high density, bacteria are almost entirely resistant to HGA, while they are highly sensitive at lower density. Dotted line shows the limit of detection. Brackets indicate the fold change in viable CFUs due to HGA exposure. (B) Viable CFUs following low-density HGA exposure in aerobic or anaerobic conditions. HGA is not toxic to Lp in the absence of oxygen. (C) High-density HGA-resistant cells do not inactivate extracellular HGA. Supernatants from Lp ±HGA at two densities were collected after 24 hr incubation and applied to new dilute cells for 24 hr. Viable CFUs were counted following supernatant exposure. HGA in all supernatants remained active against low-density Lp. (D) Model for HGA activity in extracellular environments. Lp (orange) colonizes a surface and grows to form a microcolony. Once cells are at high density, they secrete abundant HGA (yellow). Through unknown mechanisms, high-density Lp are resistant to HGA’s effects (black outline), while low-density Lp or other Legionella species (blue) are inhibited by HGA and cannot invade the microcolony’s niche.
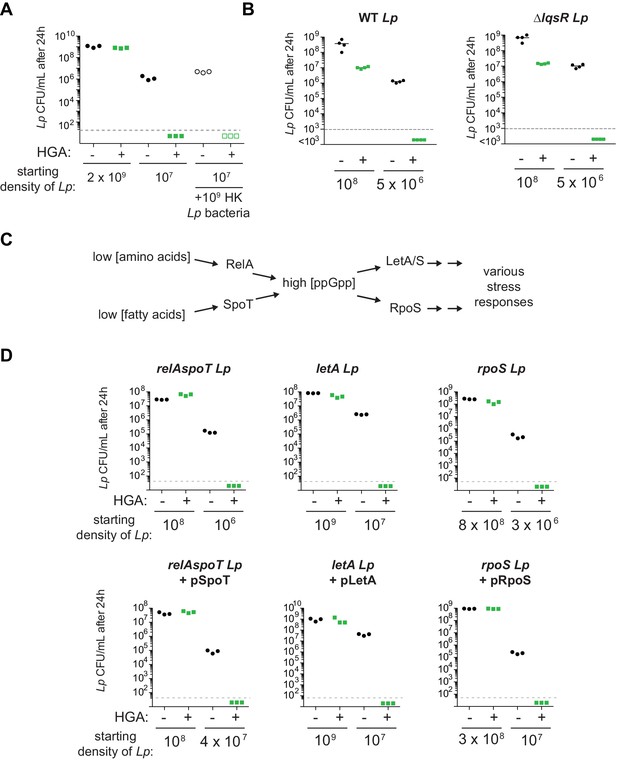
Density-dependent HGA resistance acts independently of bulk cellular material, the lqsR quorum sensing pathway, and the stringent response pathway.
(A) The presence of dense heat-killed bacteria is not sufficient to protect dilute, live Lp cells from HGA inhibition. (B) The susceptibility of wild type Lp is nearly identical to that of Lp with a deletion of lqsR, the proposed quorum sensing response regulator in L. pneumophila. (C) Schematic illustrating part of the stringent response pathway in L. pneumophila. In low nutrient conditions, RelA and SpoT generate the alarmone ppGpp, which activates a variety of downstream stress responses via the LetA/LetS two component system and the RpoS sigma factor. (D) L. pneumophila stringent response mutants show similar, density-dependent susceptibility to HGA as wild type. Complementation of relAspoT, rpoS, and letA mutants by plasmid expression (with pSpoT, pRpoS, and pLetA respectively) similarly had little effect on HGA-mediated inhibition.
Tables
Genes Mutated in HGA-Selected L. pneumophila
https://doi.org/10.7554/eLife.46086.014| Mutated locus | Function or product | # Spontaneous Mutants w/this mutation | Mutations recovered |
|---|---|---|---|
| lpg1681 | Hypothetical Protein | 4 | R49K, R49S, R49G, T50K |
| lpg0288 | YjeK, 2,3-beta-lysine aminomutase | 11 | W8S, Q9*, L26G, K39*, A50P, R101L, D196G, H215R, Q243K, I250F, Q294* |
| lpg0607 | PoxA/YjeA/GenX, Elongation factor P beta-lysine transferrase | 5 | W100*, Q184L, A218V, Q258*, 1 bp deletion in S156 |
| lpg0325 | RpS7 ribosomal 30S protein | 1 | G100D |
| lpg0336 | RplP 50 s ribosomal protein | 1 | G88R |
| lpg0287 | Elongation factor P | 1 | Stop > Q (70 AA Extension) |
| lpg0349 | SecY | 3 | N118K, Q132R, R369L |
| lpg2001 | SecD | 2 | V238F, A277G |
| lpg1504 | AceE pyruvate dehydrogenase | 1 | P272C |
-
* =introduction of a stop codon
Additional files
-
Supplementary file 1
(A) Recovered transposon mutants of L. pneumophila with defects in L. micdadei inhibition (B) Strains and plasmids (C) Primers used for cloning.
- https://doi.org/10.7554/eLife.46086.018
-
Transparent reporting form
- https://doi.org/10.7554/eLife.46086.019





