Tailored tetravalent antibodies potently and specifically activate Wnt/Frizzled pathways in cells, organoids and mice
Figures
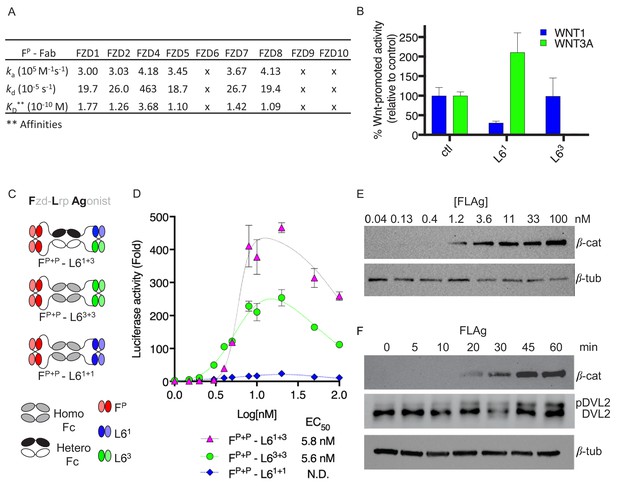
Design and validation of FLAgs as activators of the Wnt-βcatenin pathway.
(A) Surface plasmon resonance (SPR) binding kinetics of FP Fab. Kinetics were derived from curves for soluble FP Fab interacting with immobilized FZD CRD, and ‘x’ indicates no detectable binding. (B) Anti-LRP6 Ab inhibitory activity. Inhibition of WNT1 or WNT3A signaling by indicated LRP6 Abs in the diabody-Fc format was assessed using stimulation with purified WNT3A or upon WNT1 cDNA transfection. (C) Molecular architecture of tetravalent FLAgs. (D) Activation of βcatenin signaling by FLAgs. Dose response curves are shown for the activation of a LEF/TCF reporter gene (y-axis) in HEK293T cells by serial dilutions of pan-specific FLAg proteins (FP+P-L61+1, FP+P-L63+3 and FP+P-L61+3) (x-axis). Error bars indicate SEM, n = 3. (E) Levels of βcatenin protein in RKO cells after 30 min treatment with the indicated concentrations of pan-FLAg (FP+P-L61+3). Representative blot of three replicates. (F) Time course of βcatenin and phosphorylated Disheveled-2 (p-DVL2) protein levels in RKO cells treated with 10 nM pan-FLAg (FP+P-L61+3). Representative blot of three replicates.
-
Figure 1—source data 1
Source data for Figure 1B.
- https://doi.org/10.7554/eLife.46134.004
-
Figure 1—source data 2
Source data for Figure 1B.
- https://doi.org/10.7554/eLife.46134.005
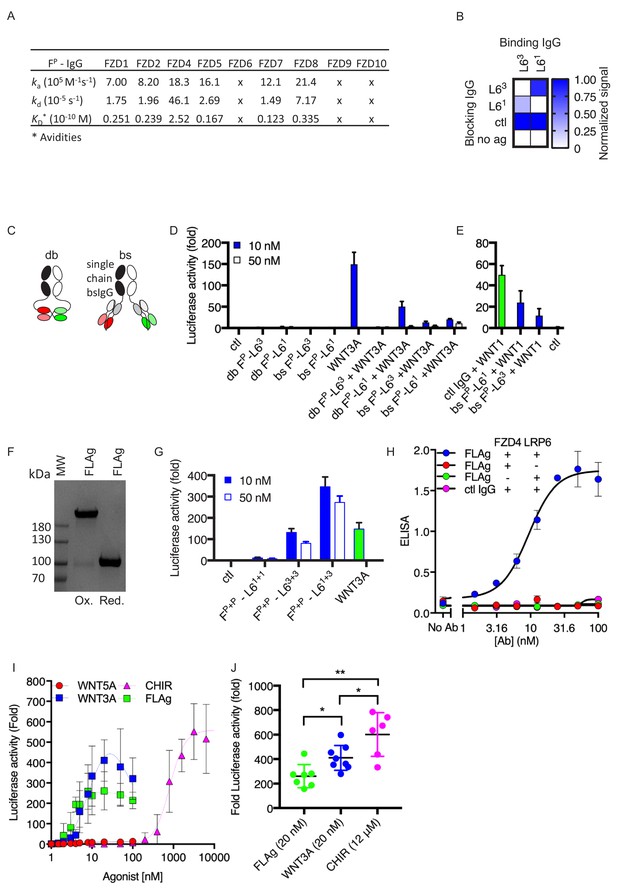
Development of Synthetic Modular Tetravalent FZD.
(A) SPR binding kinetics of FP IgG. 'x’ indicates no detectable binding. (B) Epitope binning for L61 and L63 IgG by competitive blocking measured using BLI. The normalized signal is shown for the binding of L61 or L63 (x-axis) to immobilized LRP6-Fc pre-blocked with the indicated Ab (y-axis). (C) Schematic representation of bispecific diabody-Fc (db) and single-chain bispecific IgG (bs) modalities recognizing FZD and LRP6 receptors. (D–E) Effects of bispecific IgGs and diabody-Fcs on Wnt-βcatenin signaling, as measured using HEK29T cells stably expressing a LEF/TCF reporter and with or without WNT1 transient transfection or treatment with WNT3A (0.5 μg/ml). (F) Coomassie-stained SDS-PAGE analysis of purified FLAg FP+P-L61+3 under oxidizing (Ox) or reducing (red) conditions. (G) βcatenin signaling stimulated by FLAg or WNT3A conditioned media, measured using LEF/TCF reporter assay in HEK293T cells. (H) Co-binding of FLAg FP+P-L61+3 to absorbed LRP6-Fc (LRP6) and soluble concentrations of FZD4-His (FZD4) by ELISA with anti-His-HRP detection. (I) Dose response analysis of βcatenin signaling activity stimulated with mouse WNT3A (R and D systems 1324-WN/CF), human/mouse WNT5A (R and D systems, 645-WN/CF) purified proteins, FLAg FP+P-L61+3 or CHIR99021. HEK293T cells stably expressing a LEF/TCF reporter were stimulated with the indicated doses for 16 hr. (J) The fold change of luciferase activity at maximal stimulation of Wnt reporter cells treated with 20 nM FLAg FP+P-L61+3 , 20 nM purified WNT3A, or 12 μM CHIR99021.
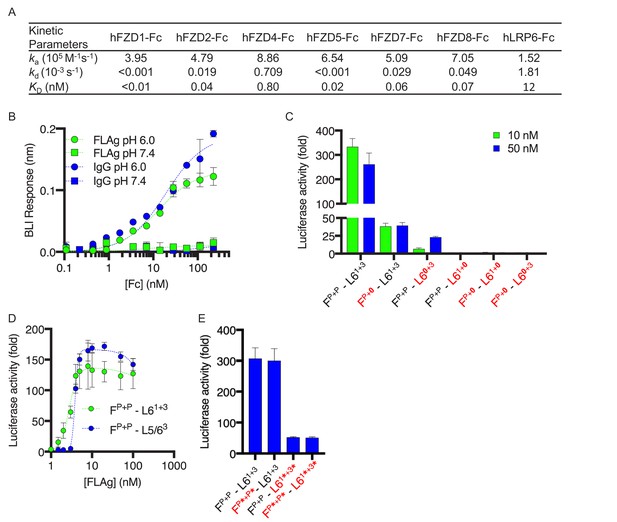
Characterization and structure-function activity relationships for FLAg FP+P-L61+3.
(A) BLI measurements of kinetic parameters for FP+P-L61+3 binding to immobilized, Fc-tagged FZD CRDs and LRP6 ECD (B) Dose response curves for BLI measurements of FP+P-L63+3 and IgG binding to immobilized FcRn at pH 6 or 7.4. Error bars indicate SD, n = 2. (C) Agonist activities of FP+P-L61+3 variants with disabled paratopes. ‘0’ indicates substitution with an anti-MBP paratope. HEK293T cells expressing a LEF/TCF reporter gene were stimulated with 10 or 50 nM of the indicated FLAg, and the fold increase in luciferase activity was determined relative to an unstimulated control. Error bars indicate SEM, n = 3. (D) Activation of βcatenin signaling by FLAgs. Dose response curves are shown for the activation of a LEF/TCF reporter gene (y-axis) by serial dilutions of indicated FLAgs (x-axis). Error bars indicate SEM, n = 3. (E) Agonist activities of FP+P-L61+3 variants with scFvs in place of diabodies. Asterisks (*) indicate scFvs. Assays were performed as in (C). Error bars indicate SEM, n = 3.
-
Figure 2—source data 1
Source data for Figure 2B.
- https://doi.org/10.7554/eLife.46134.008
-
Figure 2—source data 2
Source data for Figure 2C.
- https://doi.org/10.7554/eLife.46134.009
-
Figure 2—source data 3
Source data for Figure 2D.
- https://doi.org/10.7554/eLife.46134.010
-
Figure 2—source data 4
Source data for Figure 2E.
- https://doi.org/10.7554/eLife.46134.011
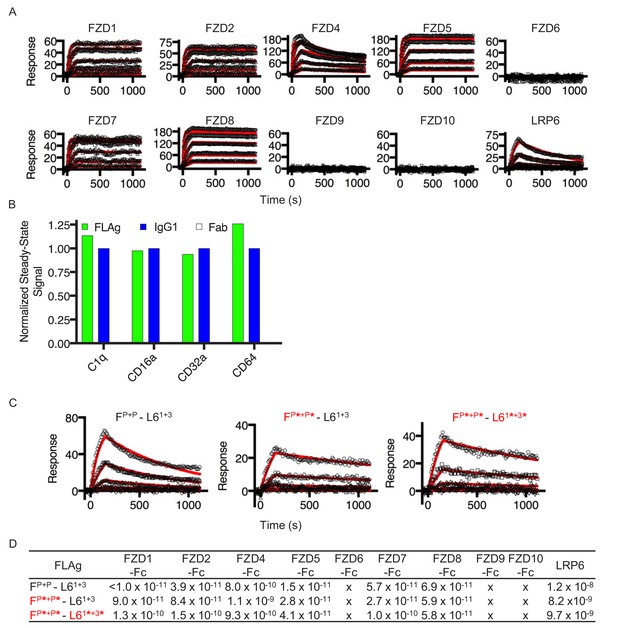
Biophysical properties of FLAgs.
(A) Surface plasmon resonance sensograms of FZD recognition by FP+P - L61+3 at concentrations ranging from 100nM- 1.23pM. (B) BLI steady-state measurements for FLAg, IgG and a negative control Fab binding to immobilized mediators of immune effector functions. Signals for FLAg were normalized to the IgG and Fab binding controls, n = 1. (C) Surface plasmon resonance sensograms of LRP6 recognition for the indicated FLAg at concentrations ranging from 100 nM to 1.23 nM. (D) Binding kinetics table of the FLAg constrained as scFv or diabodies to the various FZD isoforms.
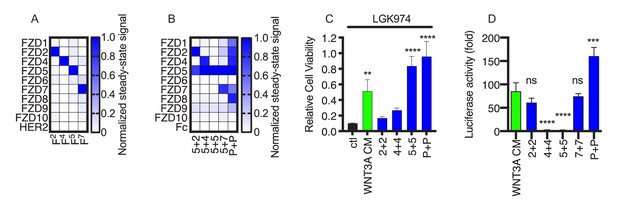
FLAgs with tailored specificities and activities.
(A) Anti-FZD Fab specificity. The heat map represents the steady state BLI signal for 100 nM Fab (x-axis) binding to immobilized, Fc-tagged FZD CRD or negative control HER2 ECD (y-axis), normalized to the highest signal. (B) Specificity of FLAgs for FZD CRDs. For each FLAg (x-axis) of the form Fx+y-L61+3, where ‘x’ and ‘y’ are paratopes from monospecific Fabs shown in (A), the heat map represents the normalized steady state BLI signal, as described in (A). (C) Effects of FLAgs on the viability of HPAF-II cells treated with the Porcupine inhibitor LGK974. Cells were treated with 100 nM LGK974 alone (ctl), or along with WNT3A conditioned media (WNT3A) or 100 nM FLAg of the form Fx+x-L61+3 (‘x + x’ indicated on the x-axis) with specificities shown in (C). Endpoint proliferation was measured by quantification of crystal violet staining and cell viability was normalized to the viability of cells that were not treated with LGK974. N = 3 independent experiments. One-way ANOVA with Dunnett’s multiple comparison test with control treated. **p<0.01, ****p<0.0001. (D) Delineation of FLAg specificity for βcatenin mediated signaling in HEK293 cells. Cells stably expressing a LEF/TCF luciferase reporter were stimulated with various FLAgs specific for individual Frizzled receptors. FZD2 (F2+2-L61+3) and FZD7 (F7+7-L61+3) specific FLAgs as well as the pan-FLAg (FP+P-L61+3) robustly stimulate βcatenin signaling whereas FZD4 and FZD5 specific FLAgs are inactive in this context. Error bars indicate SEM, n = 5. One-way ANOVA with Dunnett’s multiple comparison test with control treated. ****p<0.0001, ***p=0.0003, ns = not significant.
-
Figure 3—source data 1
Source data for Figure 3A.
- https://doi.org/10.7554/eLife.46134.014
-
Figure 3—source data 2
Source data for Figure 3B.
- https://doi.org/10.7554/eLife.46134.015
-
Figure 3—source data 3
Source data for Figure 3C.
- https://doi.org/10.7554/eLife.46134.016
-
Figure 3—source data 4
Source data for Figure 3D.
- https://doi.org/10.7554/eLife.46134.017
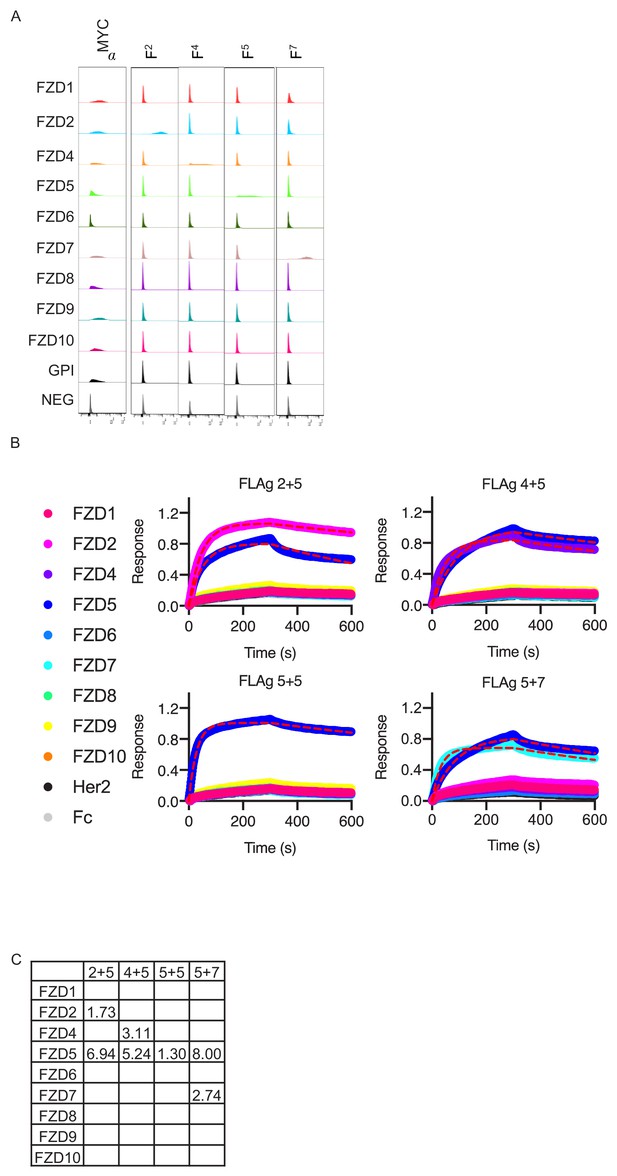
Selectivity of Fabs for FZD CRD expressed on cells.
(A) Flow cytometry specificity analysis of selective FZD Fabs (Steinhart et al., 2017). The specificity of each Fab was determined by measuring binding to a panel of recombinant CHO lines displaying each FZD ECD at the cell surface using a GPI anchor (Myc-FZD ECD-GPI). Cells were stained with 10 nM of each Fab and binding evaluated using flow cytometry. (B) Raw chromatograms of FLAg specificity by BLI. (C) Table of extrapolated KD (nM) from locally fit kinetics of each FLAg to human FZD ECDs using a 1:1 Langmuir model.
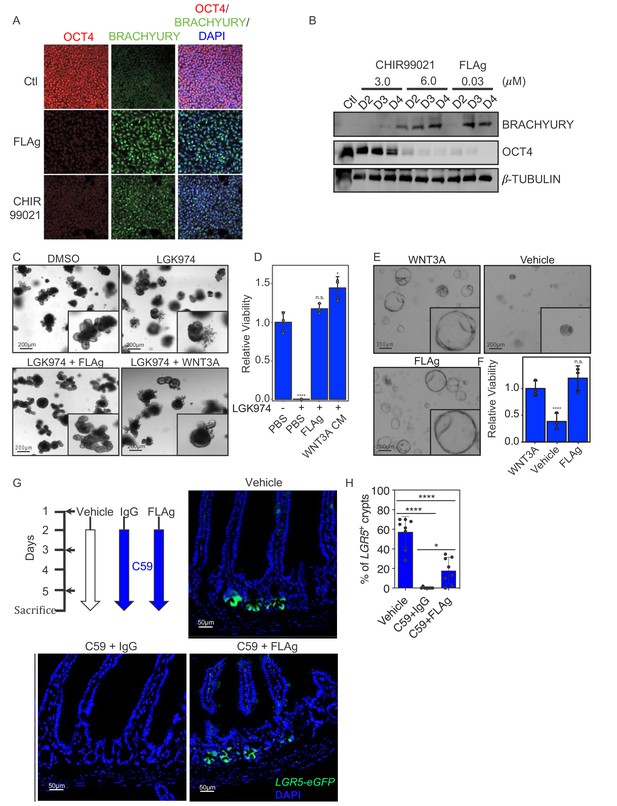
FLAg activity in cells, organoids and animals.
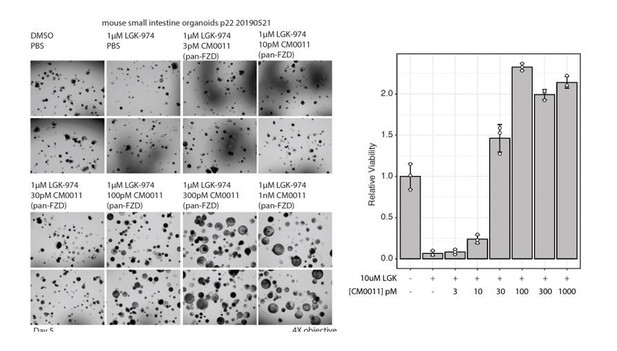
(A,B) Mesoderm differentiation of hPSCs induced by FLAg or the GSK3 inhibitor CHIR99021. Human PSCs were treated with 30 nM pan-FLAg (FP+P-L61+3) or with 3 or 6 μM CHIR99021 for the indicated times. Cells were fixed and visualized by immunofluorescence (A) or lyzed and analyzed by western blotting (B) to assess the levels of the pluripotency marker OCT4 and the early mesodermal lineage marker BRACHYURY. Immunofluorescence staining and western blots are representative of three and two independent experiments respectively. (C) Representative images of mouse small intestinal organoids with indicated treatments (1 μM LGK974, 1 μM LGK974 +30 nM pan-FLAg, 1 μM LGK974 +50% WNT3A conditioned media) (D) Viability of mouse small intestinal organoids following indicated treatments. Bars represent mean fold change ±s.d., representative of n = 3 independent experiments. Statistical analysis was performed by one-way ANOVA Dunnett’s test *p≤0.05, ****p≤0.0001. (E) Culture of human colon organoids with indicated treatments (10 nM pan-FLAg, control conditioned media or 50% WNT3A conditioned media). (F) Viability of human colon organoids following indicated treatments. Bars represent mean fold change ±s.d., representative of n = 3 independent experiments. Statistical analysis was performed by one-way ANOVA Dunnett’s test ****p≤0.0001. (G) Schematic of the in vivo workflow to evaluate FLAg efficacy. Vehicle (PBS), control IgG (10 mg/kg i.p) or FLAg (10 mg/kg i.p) groups were injected on day 1,3 and 5. The IgG and FLAg groups were further administered C59 (50 mg/kg, oral gavage) twice daily starting on day2. Representative fluorescence images of small intestinal sections from LGR5-GFP mice treated with Vehicle, IgG +C59 or pan-FLAg +C59. LGR5-GFP (green) is expressed in the stem cells at the bottom of crypts. Cell nuclei were counterstained with DAPI. (H) Quantification of LGR5 +crypts. Percentage of LGR5 +crypts was evaluated by counting at least 100 crypts per mouse in four different mice in two independent experiments (eight mice total per group). Statistics, one way ANOVA followed by Tukey test compared to vehicle treated group. *p<0.05, ****p<0.0001.
-
Figure 4—source data 1
Source data for Figure 4D.
- https://doi.org/10.7554/eLife.46134.019
-
Figure 4—source data 2
Source data for Figure 4E.
- https://doi.org/10.7554/eLife.46134.020
-
Figure 4—source data 3
Source data for Figure 4H.
- https://doi.org/10.7554/eLife.46134.021
Additional files
-
Supplementary file 1
FLAg amino acid sequences.
- https://doi.org/10.7554/eLife.46134.022
-
Supplementary file 2
Key Resources Table.
- https://doi.org/10.7554/eLife.46134.023
-
Supplementary file 3
FLAg nucleotide sequences.
- https://doi.org/10.7554/eLife.46134.024
-
Transparent reporting form
- https://doi.org/10.7554/eLife.46134.025