Structural plasticity of dendritic secretory compartments during LTP-induced synaptogenesis
Figures
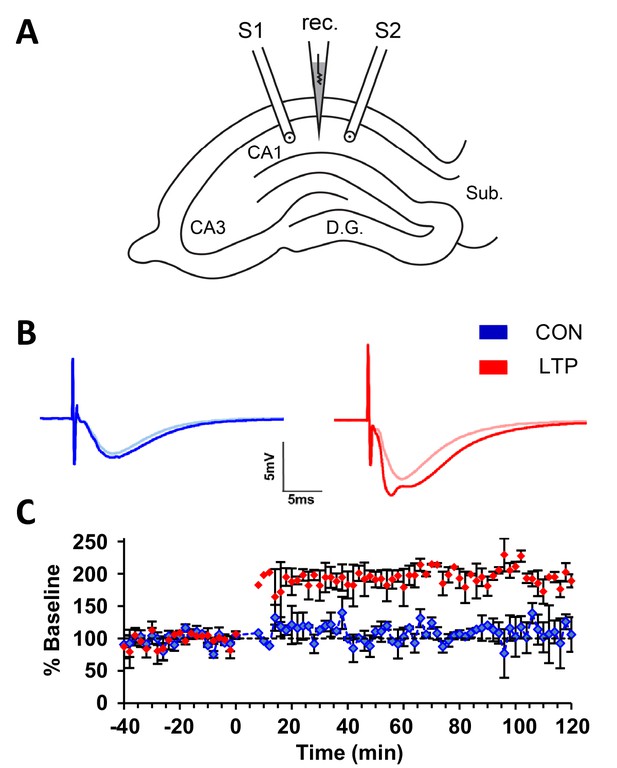
Within-slice experimental design and electrophysiological outcome.
(A) Illustration of an acute slice from a P15 rat hippocampus with a recording electrode (rec.) in the middle of CA1 stratum radiatum between two bipolar stimulating electrodes (S1 and S2). S1 and S2 are separated by 600-800 µm. The two experiments were counterbalanced for which of the two electrodes delivered TBS or control stimulation. Tissue samples collected for 3DEM were located ~120 µm beneath and to the side of the stimulating electrodes. D.G., dentate gyrus; Sub., subiculum. (B) Representative waveforms from control (CON, blue) and TBS (LTP, red) sites. Each waveform is the average of the final 10 responses to each stimulating electrode obtained for the last 20 min before delivery of TBS at time 0 (light color) and for 20 minutes before the end of the experiment at 120 min after TBS (dark color). The stimulus intensity was set at population spike threshold to activate a large fraction of the axons in the field of each stimulating electrode. The positive deflection in the post-TBS waveform at ~3-4 ms reflects synchronous firing of pyramidal cells with LTP. (C) Changes in the slope of the field excitatory postsynaptic potential (fEPSP), expressed as a percentage of the average baseline response to test-pulses, were recorded for 20 min before delivery of TBS at time 0 (red) or control stimulation (blue). Responses were recorded for n=2 slices for 120 min after the first TBS train, then fixed and processed for 3DEM as described in Methods. Error bars are SEM. Adapted from Watson et al. (2016) where it was originally published under a CC BY-NC-ND 4.0 license https://creativecommons.org/licenses/by-nc-nd/4.0/).
-
Figure 1—source data 1
Excel spreadsheet containing the raw numbers that generated the graphs and waveforms for these experiments.
- https://doi.org/10.7554/eLife.46356.003
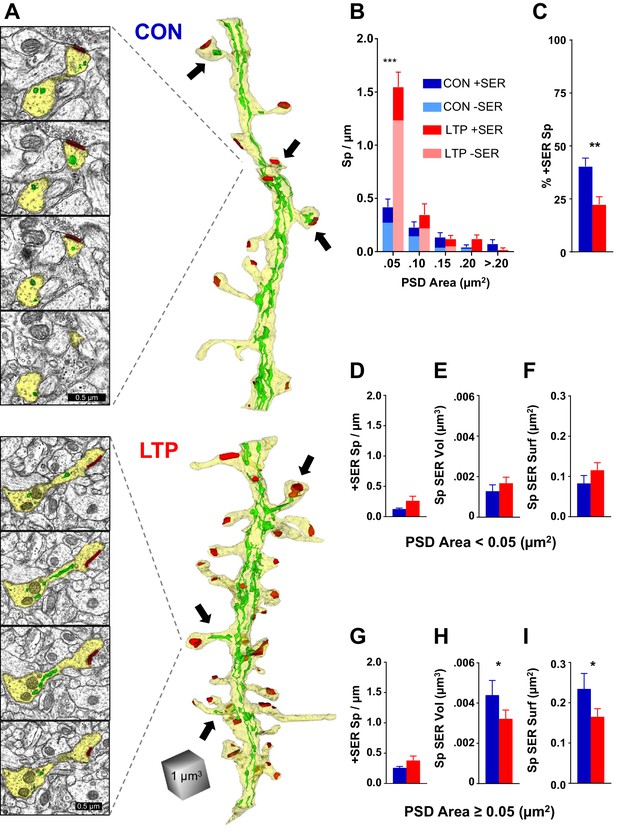
The limited occupancy of spines by SER does not increase during spinogenesis in the LTP condition.
(A) Sample serial section EMs (left) and representative 3D reconstructions of dendrites (right) from control (top) and LTP (bottom) conditions, illustrating dendrites (yellow), SER (green), and synapses (red). Synaptic area was measured as the total surface area of the PSD. Arrows point to SER-containing spines. (B) Spine density (#/µm) binned for PSD area. Significant increase in spines following TBS was carried by spines in the category with the smallest PSD areas (<0.05 µm2; ANOVA F(1,12)=50.707, p=0.00001, η2 = 0.81). No statistically significant changes occurred in the frequency of spines with larger synapses (PSD area 0.05 to 0.1 µm2, ANOVA F(1,12)=1.079, p=0.31941; PSD area 0.1 to 0.15 µm2, ANOVA F(1,12)=0.09638, p=0.76154; PSD area 0.15 to 0.2 µm2, ANOVA F(1,12)=3.5065, p=0.08569; PSD area >0.2 µm2, ANOVA F(1,11)=3.0778, p=0.10484). Control n = 8, LTP n = 8 dendrites. (C) Decrease in percentage of spines containing SER following TBS (ANOVA F(1,12)=10.599, p=0.00688, η2 = 0.87). Control n = 8, LTP n = 8 dendrites. (D–F) SER content for spines with PSD areas less than 0.05 µm2. (D) No statistically significant difference between control and LTP conditions in density of spines with SER (ANOVA F(1,12)=2.59, p=0.13322). Control n = 8, LTP n = 8 dendrites. (E) No statistically significant difference in average SER volume per SER-containing spine between control and LTP conditions (hnANOVA F(1,14)=.73111, p=0.40692). Control n = 12, LTP n = 15 spines. (F) No statistically significant difference in SER surface area per SER-containing spine between control and LTP conditions (hnANOVA F(1,14)=3.3120, p=0.09022). Control n = 12, LTP n = 15 spines. (G–I) SER content for spines with total PSD area equal to or greater than 0.05 µm2. (G) No statistically significant difference in density of spines with SER between control and LTP conditions (ANOVA F(1,12)=2.1641, p=0.16700). Control n = 8, LTP n = 8 dendrites. (H) Reduction in average SER volume per SER-containing spine in the LTP relative to control condition (hnANOVA F(1,38)=5.7205, p=0.02182, η2 = 0.13). Control n = 29, LTP n = 25 spines. (I) Reduction in average SER surface area in SER-containing spines in the LTP relative to control condition (hnANOVA F(1, 38)=4.5873 p=0.03868, η2 = 0.12). Control n = 29, LTP n = 25 spines. Bar graphs show mean ± S.E.M. Control (CON, blue) and TBS (LTP, red).
-
Figure 2—source data 1
Excel spreadsheets containing the raw numbers that generated the graphs in each part of this figure along with the summary of statistics.
- https://doi.org/10.7554/eLife.46356.006
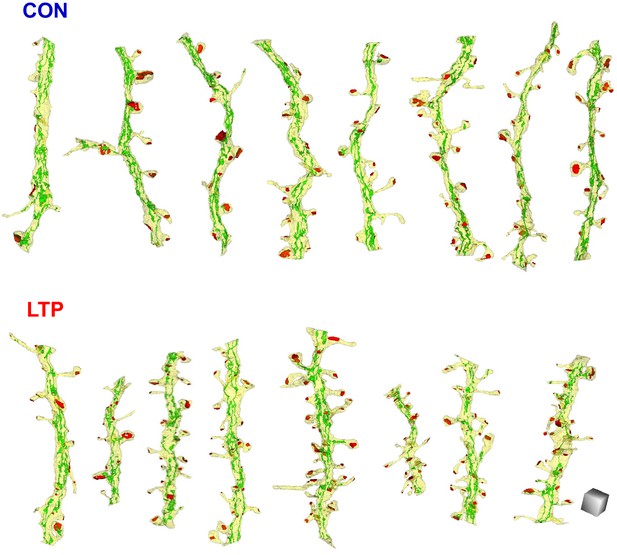
All analyzed dendrites fully reconstructed with SER, aligned left to right from least to greatest spine density.
Scale cube is 0.5 µm on each side.
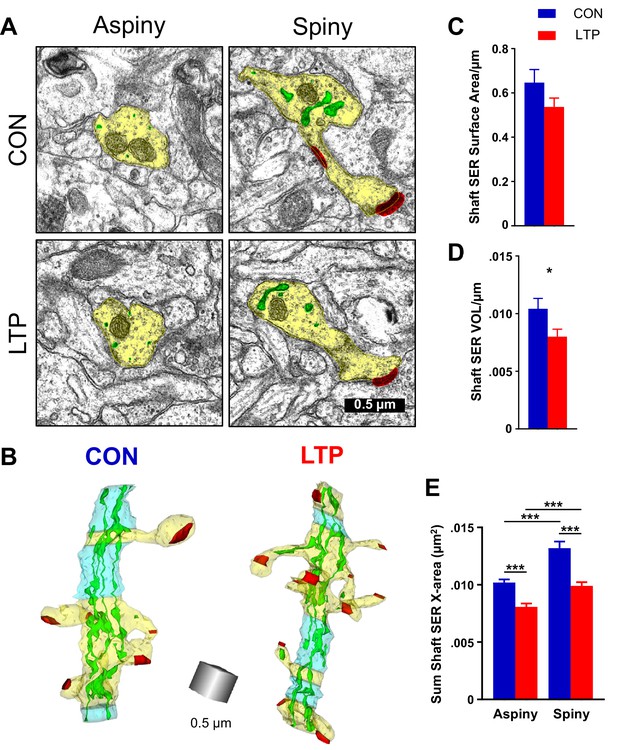
Reduction in shaft SER following LTP.
(A) Electron micrographs showing the dendrite (yellow), SER (green), and synapses (red). For both control and LTP, the SER in the aspiny segments forms small cross-sectioned tubules, whereas in the spiny segments the SER tubules are broadly expanded. (B) Sample 3D reconstructions from serial section electron micrographs of SER-containing dendrites, illustrating spiny segments (yellow) and aspiny segments (blue) while the other colors match Figure 2. Aspiny segments consist of two or more sections (>100 nm) of no spine origins. Spiny segments had at least one spine and were surrounded by aspiny segments. Scale cube is 0.5 µm on each side. (C) No statistically significant differences between control and LTP conditions were found in surface area of SER in the dendritic shaft (ANOVA F(1,12)=3.8833, p=0.07228). Control n = 8, LTP n = 8 dendrites. (D) Volume of dendritic SER network was reduced in the LTP relative to control conditions (ANOVA F(1,12)=6.4397, p=0.02605, η2 = 0.35). Control n = 8, LTP n = 8 dendrites. (E) Summed cross-sectional area of SER tubules and cisterns as a measure of changes in complexity. More SER on spiny than aspiny sections within both control (hnANOVA F(1,1432) = 51.672, p<0.00000, η2 = 0.034; spiny n = 493, aspiny n = 955 sections) and LTP conditions (hnANOVA F(1,324)=17.535, p=0.00003, η2 = 0.013; spiny n = 714, aspiny n = 626 sections). Reduced SER complexity with LTP for both spiny (hnANOVA F(1,1191) = 51.745, p<0.00000, η2 = 0.019; Control n = 493, LTP n = 714 sections) and aspiny sections (hnANOVA F(1,1565) = 29.991, p<0.00000, η2 = 0.042; Control n = 955, LTP n = 626 sections) relative to control. Bar graphs show mean ± S.E.M. Control (CON, blue) and TBS (LTP, red).
-
Figure 3—source data 1
Excel spreadsheets containing the raw numbers that generated the graphs in each part of this figure along with the summary of statistics.
- https://doi.org/10.7554/eLife.46356.008
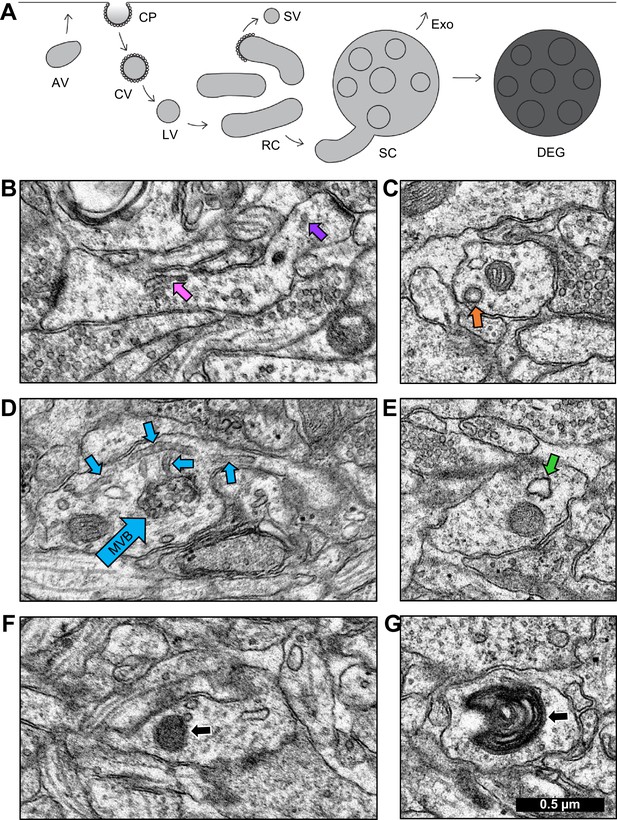
Identification of endosomal compartments.
(A) Model of the dendritic endosomal pathway. Clathrin-coated pits (CPs) invaginate, becoming clathrin-coated vesicles (CVs) and large vesicles (LVs) after coat shedding. Large vesicles fuse to form tubules, recycling complexes (RCs), and sorting complexes (SCs) with a multivesicular body (MVB). From here, the sorted material may be sent to the plasma membrane via small vesicles (SVs) that pinch off coated tips of tubules. MVBs may serve as exosomes (Exo) or primary lysosomes, that are more darkly stained than exosomes due to the acidic cytomatrix of lysosomes (adapted from Cooney et al., 2002). Sample electron micrographs illustrate (B) recycling complex (pink arrow) and small vesicles (purple arrow), (C) clathrin-coated pit (orange arrow), (D) sorting complex (light blue arrows point to multivesicular body (MVB) in the center and tubules around it), (E) amorphous vesicle (green arrow), (F) lysosome (black arrow), and (G) whorl (black arrow). Scale bar in (G) is 0.5 µm for all images.
-
Figure 4—source data 1
Excel spreadsheets containing details of the locations of each object in Figure 4.
- https://doi.org/10.7554/eLife.46356.016
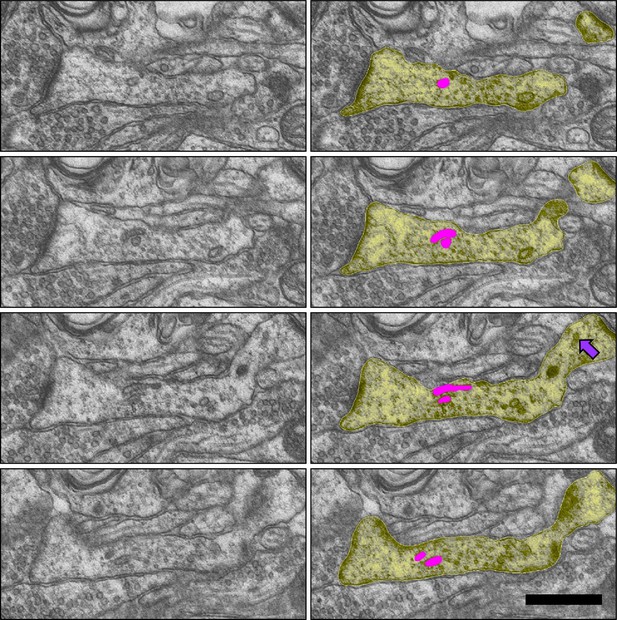
Sample images from the LTP condition of dendritic (yellow) recycling complex with multiple tubules (pink) entering the spine neck, and two small vesicles (purple arrow) in a different dendritic spine.
D28 FZYJV sections 108–111. Scale bar 0.5 µm.
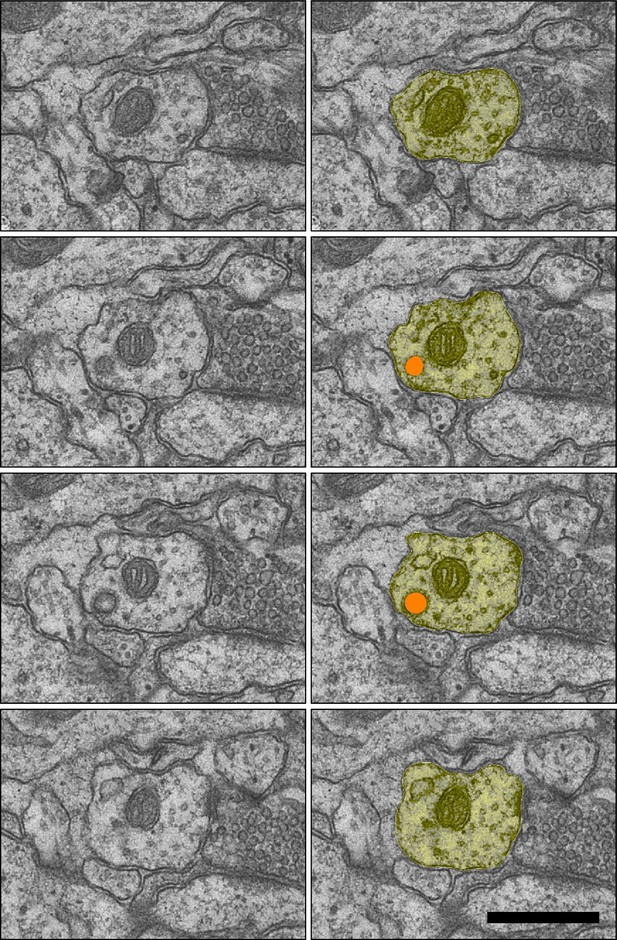
Sample images of coated pit (orange) inside the dendritic shaft (yellow) from a dendrite in the LTP condition, D25 DCPBM sections 121–124.
Scale bar 0.5 µm.
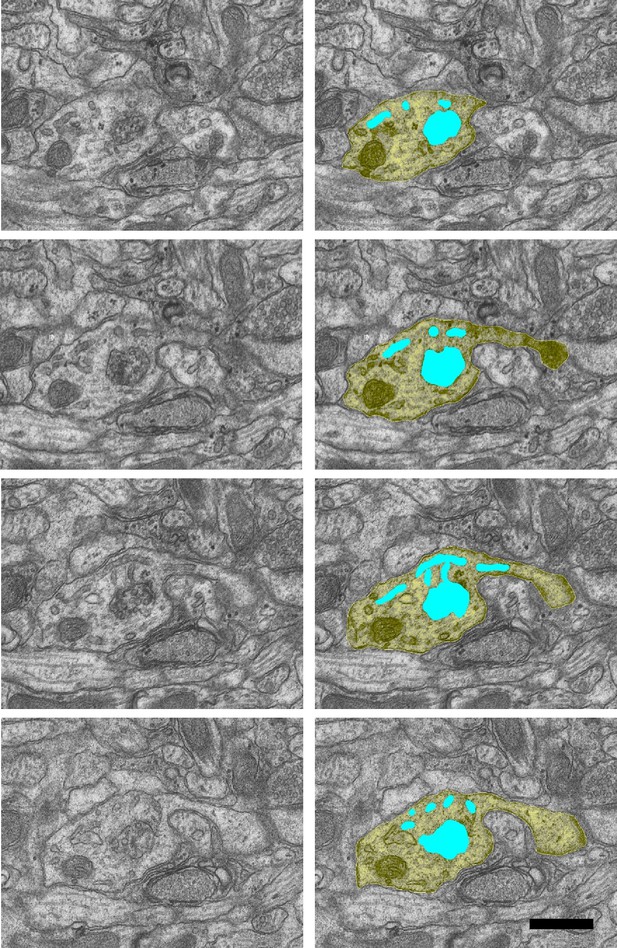
Sample images of sorting complex (turquoise) inside the dendritic shaft from the control condition, with one tubule entering a spine neck (right side row 3).
D26 PWCNZ sections 41–44. Scale bar 0.5 µm.
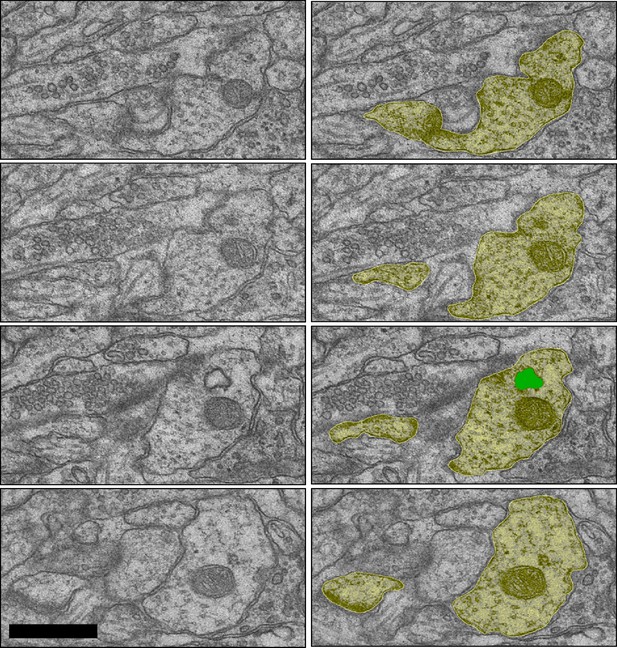
Sample image of an amorphous vesicle in the dendritic shaft of the LTP condition from D35 DCPBM sections 25–28.
Scale bar 0.5 µm.
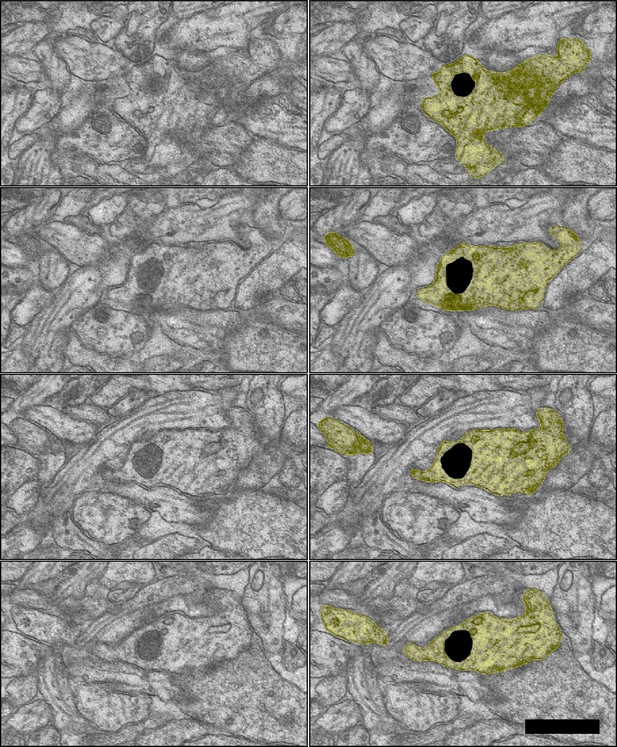
Sample images of degradative lysosome (black) in the dendritic shaft (yellow) of the LTP condition from D17 FZYJV sections 146–149.
Scale bar 0.5 µm.
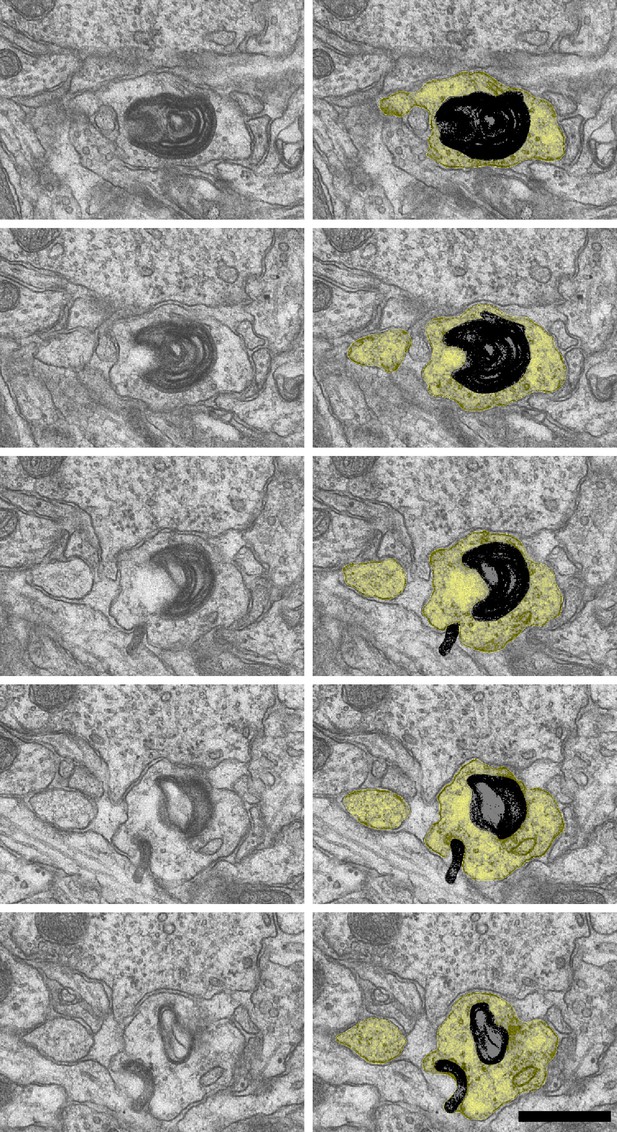
Sample images of degradative whorl (black) in a dendrite (yellow) of the control condition from D69 FXBVK sections 176–180.
Scale bar 0.5 µm.
Video paging through dendritic from the LTP condition including sections 96–121 of D28 FZYJV.
https://doi.org/10.7554/eLife.46356.017A sorting complex (turquoise) in a dendrite (yellow) from the control condition is D26 PWCNZ and includes sections 35–46.
https://doi.org/10.7554/eLife.46356.018A degradative whorl (black) in a dendrite of the control condition from D69 FXBVK sections 170–187.
https://doi.org/10.7554/eLife.46356.019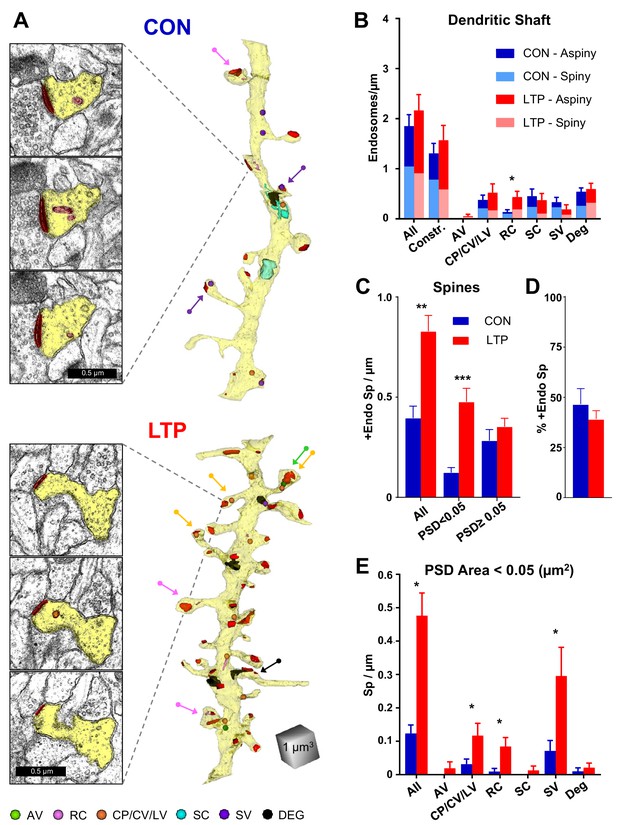
Increased occurrence of endosomes in small spines after LTP.
(A) Sample serial EM sections and representative 3D reconstructed dendrites illustrate the distribution of endosomal compartments from control and LTP conditions. Dendrites are yellow, synapses are red, and color-coded arrows point to endosome-containing spines. The color-coded key in the lower left corner indicates amorphous vesicles (AV), recycling complexes (RC), coated pits (CP), coated vesicles (CV), large vesicles (LV), sorting complexes (SC), small vesicles (SV) and degradative structures (DEG); these abbreviations apply also to the graphs. Vesicles are represented as 100 nm spheres (AV, CP, CV, LV, and SV). The other structures (RC, SC, DEG) are reconstructed in 3D to scale. (B) Endosomal structures in dendritic shafts (#/µm) with relative distributions to aspiny and spiny segments in control (CON) and LTP conditions. Overall, shaft endosomes (hnANOVA F(1,293)=0.93104, p=0.33539), degradative structures (hnANOVA F(1,293)=0.47789, p=0.48993) or constructive endosomal compartments (Constr. = all minus degradative; hnANOVA F(1,293)=0.62167, p=0.43107) did not differ between LTP and control conditions or segment locations. Recycling complexes (RC) were greater in the LTP than control dendritic shafts (hnANOVA F(1,293)=6.4920, p=0.01135, η2 = 0.022), but no significant differences occurred in the other categories: amorphous vesicles (hnANOVA F(1,293)=1.5092, p=0.22025); small vesicles (hnANOVA F(1, 293)=1.1699, p=28031); coated pits, coated vesicles, and large vesicles (hnANOVA F(1,293)=0.89152, p=0.34584); and sorting complexes (hnANOVA F(1,293)=0.45286, p=0.50151). (For control (CON) n = 151 aspiny + spiny segments and for LTP n = 158 aspiny + spiny segments.) (C) More dendritic spines contained endosomes along the dendrites in the LTP than the control condition (ANOVA F(1,12)=18.047, p=0.00113, η2 = 0.60), an effect that was carried by spines with PSD areas less than 0.05 µm2 (ANOVA F(1,12)=23.642, p=0.00039, η2 = 0.66) but not in spines with PSD area >0.05 µm2 (ANOVA F(1,12)=0.84714, p=0.37550). (D) Stability in percentage of spines containing endosomes following TBS (ANOVA F(1,12)=.72158, p=0.41225). (E) Among spines with PSD area less than 0.05 µm2, the increase in occupancy of endosomes was due to more with coated pits, coated vesicles, and large vesicles (ANOVA F(1,12)=4.94433, p=0.046140, η2 = 0.29), recycling complexes (ANOVA F(1,12)=11.009, p=0.00613, η2 = 0.48), and more with small vesicles (ANOVA F(1,12)=5.2575, p=0.04072, η2 = 0.30). No significant changes in spine occupancy occurred for amorphous vesicles (ANOVA F(1,12)=1, p=0.33705), sorting complexes (ANOVA F(1,12)=1, p=0.33705), or degradative structures (ANOVA F(1,12)=0.46689, p=0.5074). Bar graphs show mean ± S.E.M. (For C–E), Control (CON, n = 8 full dendrite reconstructions) and LTP (n = 8 full dendrite reconstructions).
-
Figure 5—source data 1
Excel spreadsheets containing the raw numbers that generated the graphs in each part of this figure along with the summary of statistics.
- https://doi.org/10.7554/eLife.46356.023
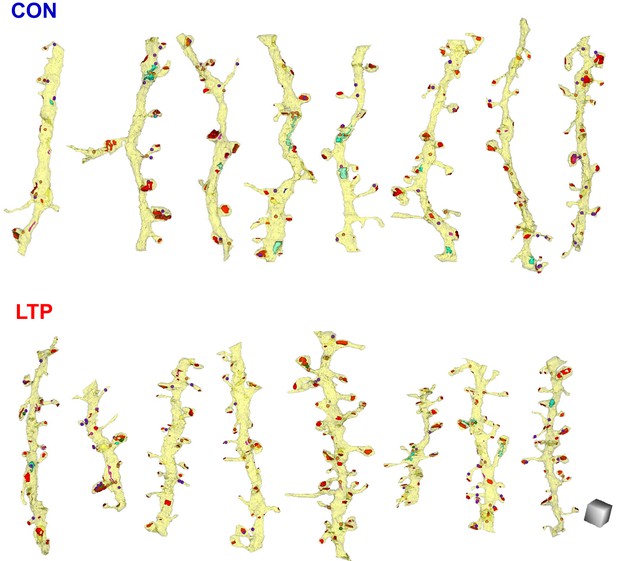
All analyzed dendrites fully reconstructed with constructive endosomes, aligned left to right from least to greatest spine density.
Scale cube is 0.5 µm on each side.
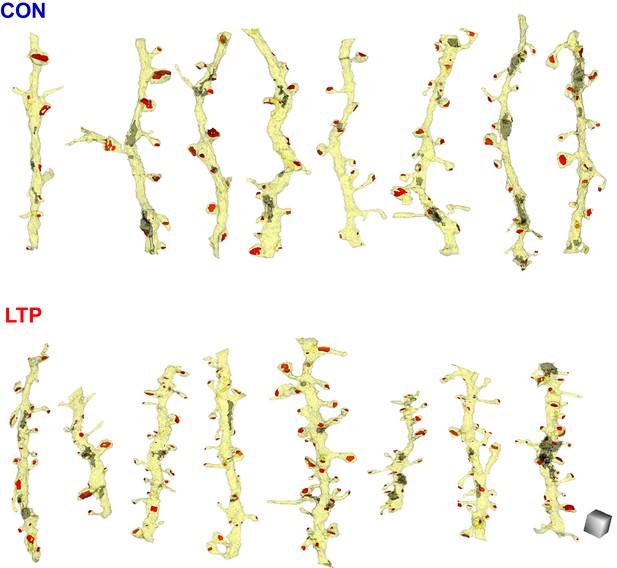
All analyzed dendrites fully reconstructed with intracellular degradative structures, aligned left to right from least to greatest spine density.
Scale cube is 0.5 µm on each side.
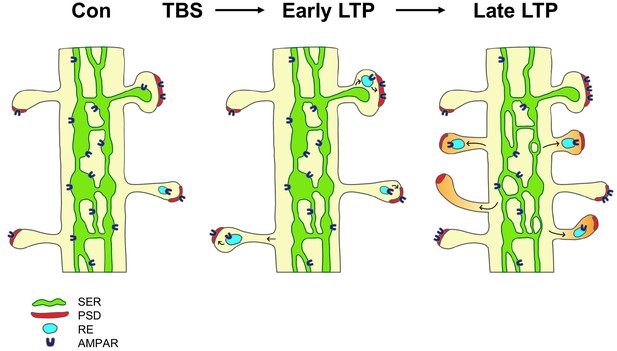
Model of the contribution of dendritic secretory compartments to LTP-induced synaptogenesis.
Smooth endoplasmic reticulum (SER, green), postsynaptic density (PSD, red), small vesicle or recycling endosome (RE, turquoise), new silent spines (orange), control activation (Con), theta-burst stimulation (TBS), long-term potentiation (LTP), AMPA receptors (AMPAR).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Rattus norvegicus, male) | Long-Evans Rat | Charles River | Charles River strain# 006; RRID:RGD_2308852 | |
| Chemical compound, drug | Potassium ferrocyanide | Sigma-Aldrich | Cat# P3289 | |
| Chemical compound, drug | Osmium tetroxide | Electron Microscopy Sciences | Cat# 19190 | |
| Chemical compound, drug | Uranyl acetate | Electron Microscopy Sciences | Cat# 22400 | |
| Chemical compound, drug | LX-112 embedding kit | Ladd Research Industries | Cat# 21210 | |
| Chemical compound, drug | Lead nitrate | Ladd Research Industries | Cat# 23603 | |
| Chemical compound, drug | Pioloform F | Ted Pella | Cat# 19244 | |
| Software, algorithm | Igor Pro 4 | WaveMetrics | https://www.wavemetrics.net/ | |
| Software, algorithm | Reconstruct | Fiala, 2005 | Executable and manual: http://synapseweb.clm.utexas.edu/software-0 | Source at:https://github.com/orgs/SynapseWeb/teams/reconstruct-developers |
| Software, algorithm | STATISTICA 13 Academic | Tibco | https://onthehub.com//statistica/ | |
| Other | Tissue slicer | Stoelting | Cat # 51425 | |
| Other | Vibratome | Leica Biosystems | VT1000S | |
| Other | Ultramicrotome | Leica Biosystems | UC6 | Used with a Diatome Ultra35 knife |
| Other | SynapTek Grids | Ted Pella | Cat# 4514 or 4516 | |
| Other | Diffraction grating replica | Electron Microscopy Sciences | Cat# 80051 | |
| Other | Transmission electron microscope | JEOL | JEM-1230 | |
| Other | Harris Lab wiki | Harris Lab | https://wikis.utexas.edu/display/khlab/ | This wiki site hosts experimental methods used for this paper and updates. |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.46356.025





