Self-sperm induce resistance to the detrimental effects of sexual encounters with males in hermaphroditic nematodes
Figures
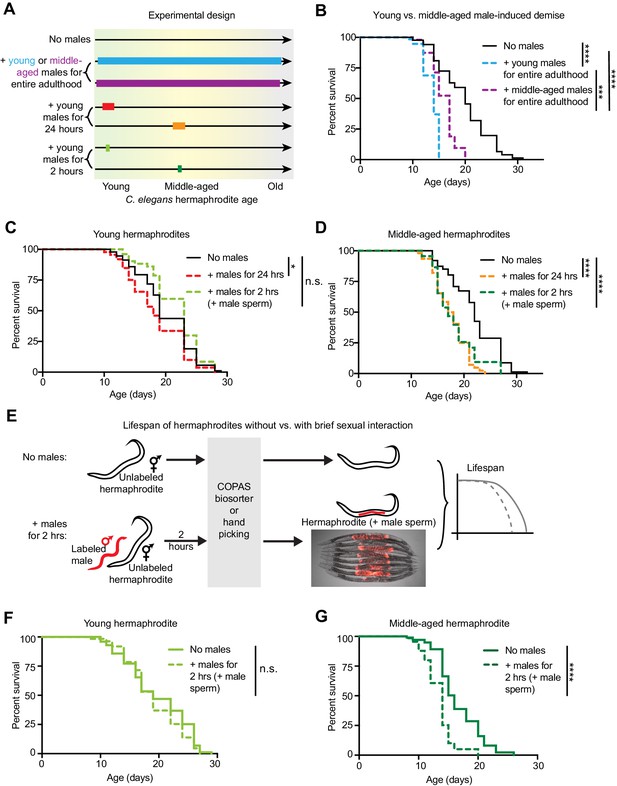
The length of sexual encounters and age of the sexual partners influences the detrimental effect of males on hermaphrodite lifespan.
(A) Scheme describing the lengths of sexual interactions between C. elegans males and hermaphrodites and the ages of the sexual partners used in this study. Young was defined as the first day of adulthood (day 3 of life), and middle-aged as the 5th day of adulthood (day 7 of life). (B) Long and frequent sexual interactions with young males (blue dashed line) reduced hermaphrodite lifespan (p<0.0001 vs. no males). Long and frequent sexual interactions with middle-aged males (purple dashed line) also reduced hermaphrodite lifespan compared to hermaphrodites without males (p<0.0001), but middle-aged males shortened hermaphrodite lifespan less than young males (p=0.0003). (C-D) Hermaphrodites that interacted with males for 24 hr when young from day 3 to 4 of life (C, red dashed line) or middle-aged from day 6 to 7 of life (D, orange dashed line) lived shorter than hermaphrodites that never interact with males (p=0.029 and p<0.0001, respectively). Hermaphrodites that mated within a brief (2 hr) interaction with males when young on day 3 of life (C, lime green dashed line) did not have a shortened lifespan (n.s. vs. no males) but hermaphrodites that mated within a brief period (2 hr) when middle-aged on day 7 of life (D, green dashed line) did have a significantly shortened lifespan (p<0.0001 vs. no males). (E) To control for the mating efficiency differences between ages and genotypes (see Supplementary file 3), we measured the lifespans of only C. elegans individuals that had successfully mated by identifying and isolating hermaphrodites that have fluorescent male sperm (represented by the red tilde in the scheme) from those that are unmated and lack fluorescent male sperm using either hand picking or a large particle COPAS large particle biosorter. The presence of fluorescent male sperm is indicative of fertilization (Figure 1—figure supplement 1B, C), though this was not specifically measured for the lifespan assays. Hermaphrodites that received male sperm following two hours with males were compared to hermaphrodites that never interacted with males but that were hand-picked or run through the COPAS large particle biosorter. (F-G) Using a different method of isolating mated hermaphrodites, we also found that young hermaphrodites (F) were resistant to mating-induced demise if they received male sperm and seminal fluid during a brief, 2 hr interaction with males (n.s. vs. no males) but that older hermaphrodites (G) were sensitive and lived shorter following a brief, 2 hr interaction with males (p<0.0001 vs. no males). In panels C and D, hermaphrodites that received fluorescent male sperm were isolated by hand and the males were him-5(e1467) mutants. In panels F and G, males were him-8(e1489) mutants with a male-specific GFP reporter (Ppkd-2::GFP) and hermaphrodites with fluorescent male sperm were isolated with the COPAS large particle biosorter. For each condition, 67–114 animals were used to quantify lifespan. Lifespan data are plotted as Kaplan-Meier survival curves and p-values were determined using Mantel-Cox log ranking. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, n.s. = not significant. See also Supplementary file 2 for extended statistics and replicates.
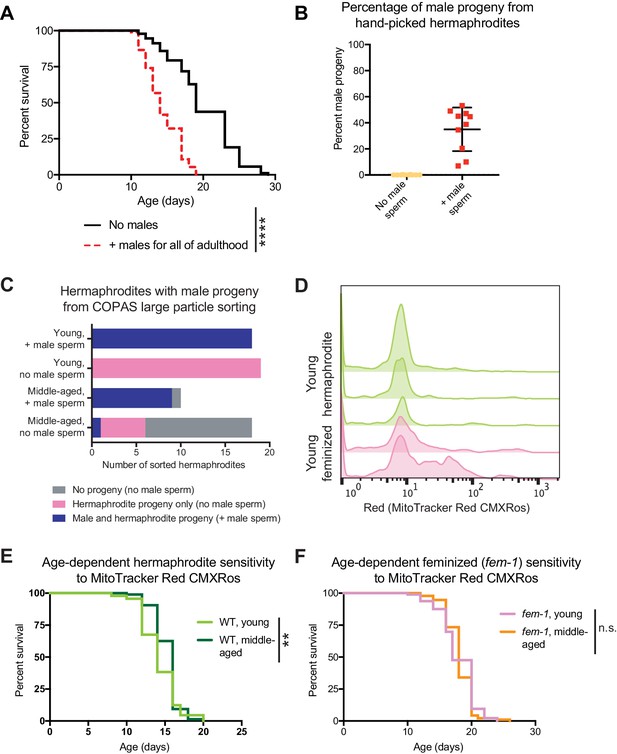
Effect of long-term exposure to young males and development of the sperm tracking method.
(A) The continuous presence of young males beginning at the onset of adulthood (day 3 of life) shortens hermaphrodite lifespan (p<0.0001). (B-C) Accuracy tests for the presence of fluorescence in the uterus and spermatheca of hermaphrodites as an indication of successful mating following 2 hr with fluorescently labeled males. The production of male progeny is used as an orthogonal indication of successful mating. In panel B, the percentage of male progeny produced by hermaphrodites that were identified by eye using a fluorescent dissecting microscope as containing fluorescent male sperm (red points) compared to hermaphrodites that never encountered a male (yellow points). In panel C, the number of hermaphrodites that produced hermaphrodite only progeny, male and hermaphroditic progeny or no progeny. Hermaphrodites that contained fluorescent male sperm (identified by the COPAS large particle biosorter) are compared to hermaphrodites that never encountered a male. (D) Histograms of the amount of male sperm (red fluorescence, quantified by the COPAS large particle biosorter) received by young hermaphrodites (WT, green) or young feminized individuals (fog-2[q71], pink) following a brief interaction with males. The individual replicates are displayed. On the far left of the plot are individuals that did not receive sperm (unmated) and centered around 10 on the x-axis are the individuals that received male sperm. Note that the majority of worms that received male sperm received similar amounts but that a smaller portion of the hermaphrodites or feminized individuals may have consumed some of the MitoTracker Red CMXRos labeled bacteria, leading to higher levels of fluorescence. Young hermaphrodites and feminized individuals do not have different levels of multiple mating during a two-hour period (Figure 2—figure supplement 1E). (E-F) Young and middle-aged hermaphrodites (E) and feminized worms (fem-1[hc17], F) do not have different sensitivities to 24 hr of exposure to 5 ng/mL MitoTracker Red CMXRos in OP50-1 bacteria (p=0.0091 for E and n.s. for F). For the lifespan assays, 94–107 animals were used for each experimental condition. Lifespan data are plotted as Kaplan-Meier survival curves and p-values were determined using Mantel-Cox log-ranking. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, n.s. = not significant. See Supplementary file 2 for additional replicates and statistics.
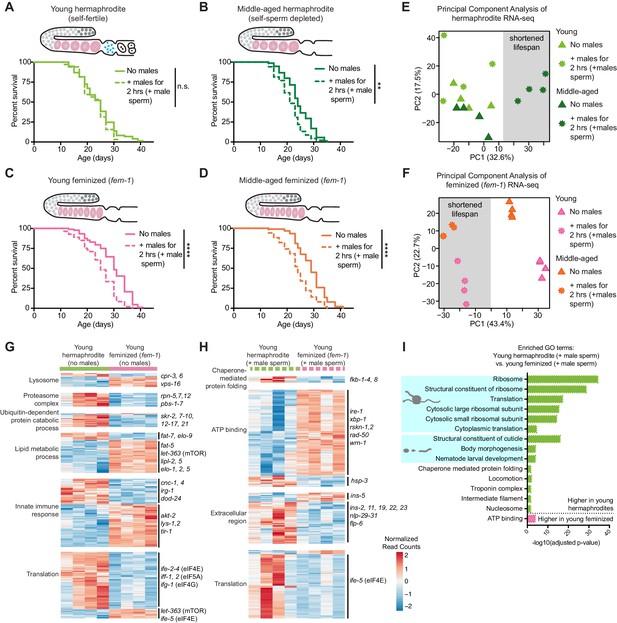
The presence of self-sperm is necessary for the resistance of young hermaphrodites to a brief encounter with males.
(A-B) Young, self-fertile, wild-type hermaphrodites with self-sperm (A, day 3 of life) that received male sperm after a brief interaction with males had a normal lifespan (n.s. vs. no males) whereas middle-aged, wild-type hermaphrodites that are self-sperm depleted (B, day 7 of life) had a shortened lifespan (p=0.0013 vs. no males). (C-D) The lifespan of feminized C. elegans that lack self-sperm at all ages (fem-1[hc17]) was reduced if they received male sperm during a brief, 2 hr interaction with males either when young (C, day 3 of life, p<0.0001 vs. no males) or middle-aged (D, day 7 of life, p<0.0001 vs. no males). Images above the lifespan curves in panels A-D show the state of the germline with oocytes in pink and self-sperm in blue. (E-F) Principal Component Analysis (PCA) of the normalized read counts from the entire transcriptomes of hermaphrodite (E) and feminized (F) C. elegans that never interacted with males and that received male sperm during a two-hour interaction with males. (G-H) Heatmaps of the normalized read counts for the differentially expressed genes that comprise select GO terms that were enriched when comparing young hermaphrodite and young feminized (fem-1) individuals without male exposure (G) and that received male sperm following a brief interaction with males (H). The four replicates for each condition are shown. (I) Selected, enriched GO terms from the differentially expressed genes between young hermaphrodite versus young feminized that received male sperm. GO terms that were enriched in the genes expressed more highly in young hermaphrodites are shown in green and GO terms enriched in the genes more highly expressed in young feminized individuals are in pink. P-values were calculated with the Fisher’s exact test and corrected for multiple hypothesis testing with Benjamini-Hochberg. A complete list of all significantly enriched GO terms can be found in Figure 2—source data 2. All individuals were raised at the restrictive temperature (25°C) until the onset of adulthood (day 3 of life) and then moved to 20°C for the remainder of their lifespan. Worms that received male sperm from him-5(e1467) males were isolated by hand picking individuals with fluorescent male sperm in their uterus or spermatheca. For all lifespan assays 52–109 animals were tested per condition. Lifespan data are plotted as Kaplan-Meier survival curves and p-values were determined using Mantel-Cox log ranking. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, n.s. = not significant. See also Supplementary file 2 for extended statistics and replicates.
-
Figure 2—source data 1
The DESeq2 output (differential expression) from the RNA-seq analysis.
- https://doi.org/10.7554/eLife.46418.008
-
Figure 2—source data 2
The complete list of GO terms whose enrichment was determined using the significantly differentially expressed genes when comparing young hermaphrodites vs. young feminized individuals (selected, enriched GO results are displayed in Figure 2G–I and in Figure 2—figure supplement 2C).
- https://doi.org/10.7554/eLife.46418.009
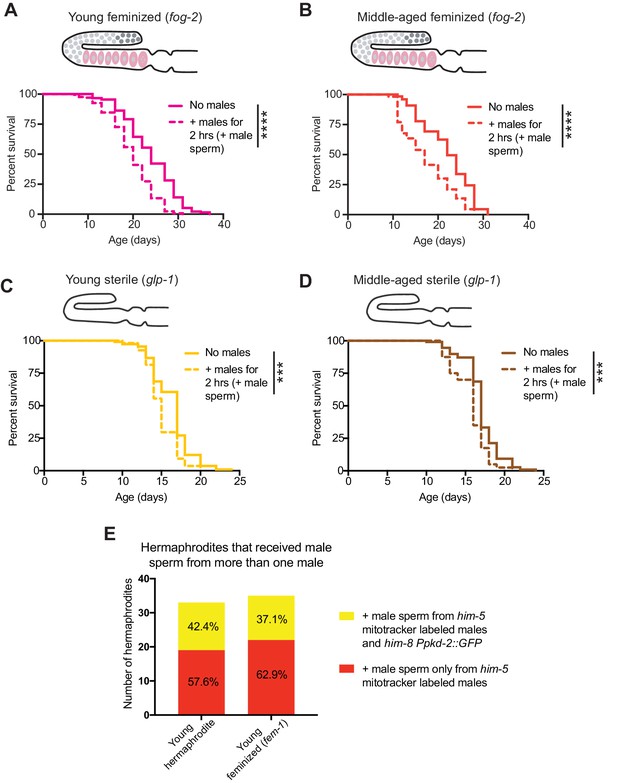
The effect of a brief encounter with males on feminized and sterile individuals.
(A-B) Receiving male sperm (dashed lines) when young (day 3 of life) or middle-aged (day 7 of life) reduced the lifespan of feminized (fog-2[q71]) individuals (p<0.0001 for both young and middle-aged). (C-D) Receiving male sperm (dashed lines) when young (C, day 3 of life) or middle-aged (D, day 6 of life) shortened the lifespan of sterile (glp-1[e2144]) individuals (p=0.0004 and p=0.0003, young and middle-aged, respectively). E) Despite a higher mating efficiency (Supplementary file 3), young feminized (fem-1[hc17]) individuals and young hermaphrodites did not have major differences in their frequency of receiving male sperm from two different males in the two-hour sexual interaction period. For the lifespan assays, 40–219 animals were used. Mated worms (dashed lines) were selected by hand-picking based on the presence of fluorescent male sperm in their uterus or spermatheca following a two-hour interaction with him-5(e1467) males. Lifespan data are plotted as Kaplan-Meier survival curves and p-values were determined using Mantel-Cox log ranking. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, n.s. = not significant. See also Supplementary file 2 for extended statistics and replicates. The experiment using fog-2 (and the WT control, see Supplementary file 2) was performed at 20°C and the experiment using glp-1 (and the WT control, see Supplementary file 2) was performed at the restrictive temperature, 25°C.
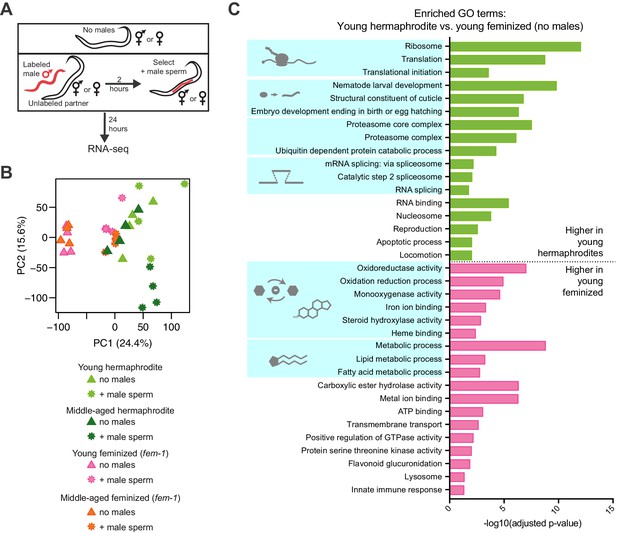
RNA-seq of young and middle-aged hermaphrodites and feminized individuals with and without receiving male sperm.
(A) Scheme describing the experimental set up for the RNA-seq experiment. (B) Principal Component Analysis (PCA) of the normalized read counts from the complete transcriptomes of young and middle-aged hermaphrodites and feminized individuals that received male sperm and those that did not interact with males. (C) Selected, enriched GO terms from the young hermaphrodite versus young feminized differentially expressed genes that never interacted with a male. GO terms that were enriched in the genes expressed more highly in young hermaphrodites are shown in green and GO terms enriched in the genes more highly expressed in young feminized individuals are in pink. P-values were calculated with the Fisher’s exact test and were corrected for multiple hypothesis testing with Benjamini-Hochberg. A complete list of all significantly enriched GO terms can be found in Figure 2—source data 2.
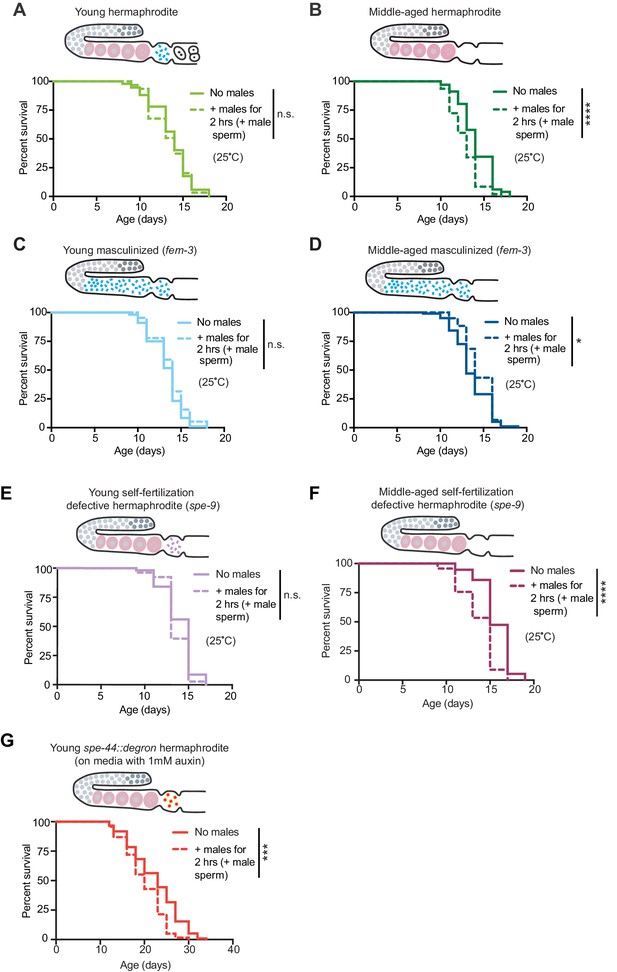
The presence of self-sperm is sufficient for the resistance to mating-induced demise.
(A-D) Hermaphrodites with a masculinized germline (fem-3[q20]) that have self-sperm when young (C, day 3 of life) and middle-aged (D, day 6 of life), were resistant to a brief, 2 hr interaction with males and lived a normal lifespan. As a control, wild-type hermaphrodites (A and B) were also tested and showed the expected resistance to a brief interaction with males when young and shortened lifespan following a brief interaction with males when middle-aged. (E-F) Young hermaphrodites that have fertilization-defective self-sperm (spe-9[hc88], panel E, day 3 of life) were protected from a brief, 2 hr interaction with males when young (n.s. vs. no males) but when self-sperm are depleted with age (F, day 7 of life), their lifespan was shortened if they received male sperm during brief interaction with males (p<0.0001 vs. no males). (G) Young hermaphrodites that have defective sperm due to the absence of the SPE-44 transcription factor (fxIs1[pie-1p::TIR1::mRuby]; spe-44(fx110[spe-44::degron]) grown on 1 mM auxin until adulthood), are not fully protected from a brief, 2 hr interaction with males (p=0.0001 vs. no males). Images above the lifespan curves show the state of the germline. Mated worms (dashed lines) were selected by hand-picking based on the presence of fluorescent male sperm in their uterus or spermatheca following a two-hour interaction with him-5(e1467) males. Lifespans were performed with 31–144 animals per condition. Masculinized (fem-3[q20]) versus WT and fertilization-defective self-sperm (spe-9[hc88]) experiments (A-F) were performed at the restrictive temperature, 25°C. The SPE-44 auxin-inducible degradation experiment (G) was performed at 20°C and controls for this experiment are found in Supplementary file 2 and Figure 3—figure supplement 1C–D. Lifespan data are plotted as Kaplan-Meier survival curves and p-values were determined using Mantel-Cox log ranking. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, n.s. = not significant. See also Supplementary file 2 for extended statistics and replicates.
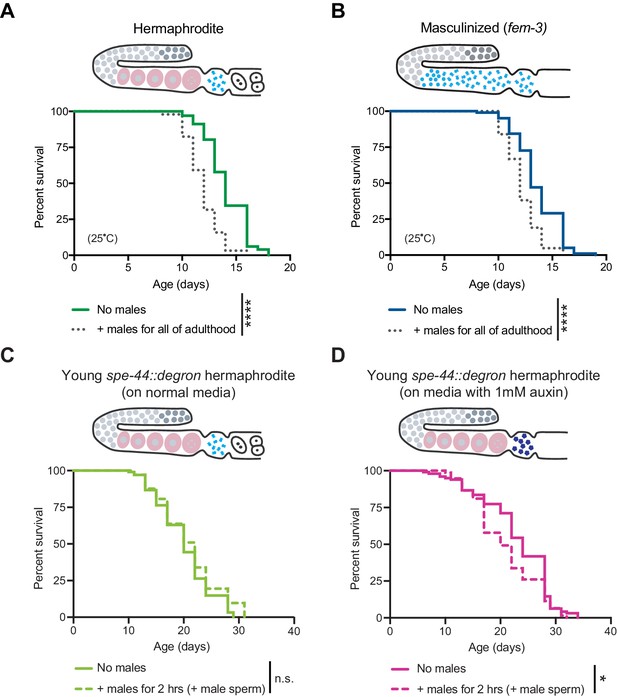
The effect of males on hermaphrodites and masculinized individuals.
(A-B) Middle-aged hermaphrodites but not middle-aged masculinized individuals succumbed to brief mating-induced demise (Figure 3B,D). However, both hermaphrodites and masculinized individuals were susceptible to long-term male-induced demise and lived shorter when young males were present throughout all of adulthood (dotted lines, p<0.0001 vs. no males for A and B). (C-D) Young hermaphrodites that have defective self-sperm due to the absence of the SPE-44 transcription factor (panel D, fxIs1[pie-1p::TIR1::mRuby]; spe-44(fx110[spe-44::degron]) grown on 1 mM auxin), are not fully protected from a brief, 2 hr interaction with males (p=0.0303 vs. no males). The same strain cultured without auxin and thus with normal levels of SPE-44 and normal sperm (panel C) lives a normal lifespan after a brief 2 hr interaction with males (n.s. vs. no males). Masculinized (fem-3[q20]) versus WT experiments were performed at the restrictive temperature, 25°C. SPE-44 auxin-inducible degradation experiments (C and D) were performed at 20°C and the worms in which SPE-44 degradation was induced (D) were grown on NGM with 1 mM auxin until adult day 1. For panels C and D, mated worms (dashed lines) were selected by hand-picking based on the presence of fluorescent male sperm in their uterus or spermatheca following a two-hour interaction with him-5(e1467) males. For the lifespan assays, 46–113 animals were used. Lifespan data are plotted as Kaplan-Meier survival curves and p-values were determined using Mantel-Cox log ranking. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, n.s. = not significant. See also Supplementary file 2 for extended statistics and replicates.
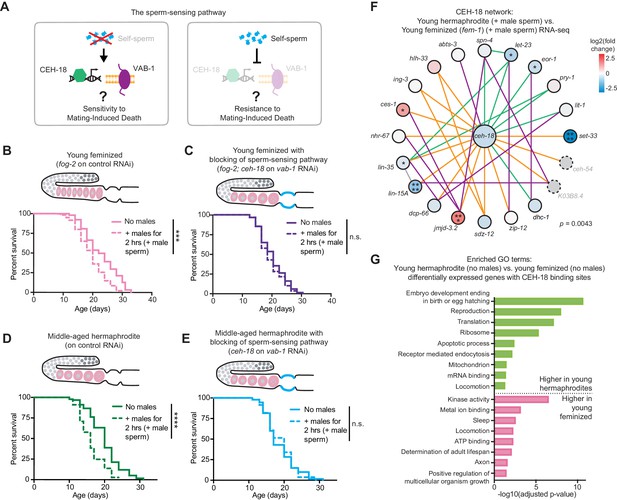
Self-sperm act via repression of a somatic sperm-sensing pathway to mediate resistance to mating-induced demise.
(A) A model for the role of self-sperm in mating-induced demise resistance. The absence of self-sperm (right panel) activates the CEH-18 and VAB-1 sensing pathway in the somatic gonad. (B-C) Young, feminized (fog-2[q71]) worms (B, day 4 of life) were sensitive to a brief interaction with males when young (B, p=0.0001 vs. no males) but when the sperm-sensing pathway was blocked by loss of CEH-18 and VAB-1 (fog-2[q71]; ceh-18[mg57] grown on vab-1 targeting RNAi bacteria), these worms (C) were resistant to a brief, 2 hr interaction with males and had a normal lifespan (n.s. vs. no males). (D-E) Middle-aged, self-sperm depleted hermaphrodites (D, day 7 of life) were sensitive to a brief mating with males (D, p<0.0001 vs. no males) but when the sperm-sensing pathway was blocked by loss of CEH-18 and VAB-1 (ceh-18[mg57] grown on vab-1 targeting RNAi bacteria), these middle-aged worms (E) were resistant to a brief, 2 hr interaction with males and had a normal lifespan (E, n.s. vs. no males). For all experiments, worms that received male sperm (dashed lines) were selected by hand-picking based on the presence of fluorescent male sperm in their uterus or spermatheca following a two-hour interaction with him-5(e1467) males. Images above the lifespan curves in panels B-E show the state of the germline with oocytes in pink and the blocking of sperm-sensing pathway by a blue line. Lifespans were performed with 86–115 animals per condition. Lifespan assays were performed at 20°C. Lifespan data are plotted as Kaplan-Meier survival curves and p-values were determined using Mantel-Cox log ranking. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, n.s. = not significant. See also Supplementary file 2 for extended statistics and replicates. (F) The CEH-18 interaction network calculated by GeneMANIA (Warde-Farley et al., 2010). Lines indicate genetic interactions (green), co-expression (purple), physical interactions (orange), and ‘other’ (gray). The differential expression of the genes in the network is shown with a blue-red gradient. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, no stars = not significant. Network members not detected by the RNA-seq are in gray with dotted outlines. The number of differentially expressed genes in this network was statistically enriched (p=0.0043) as measured using the hypergeometric distribution test. See Figure 2—source data 1 for exact differential expression values. (G) Selected, enriched GO terms from the genes that are differentially expressed between young hermaphrodite versus young feminized that never interacted with a male and contain a CEH-18 binding site as defined by CEH-18 ChIP-seq (Kudron et al., 2018), see Material and methods. GO terms that were enriched in the genes expressed more highly in young hermaphrodites are shown in green and GO terms enriched in the genes more highly expressed in young feminized individuals are in pink. P-values were calculated with the Fisher’s exact test and were corrected for multiple hypothesis testing with Benjamini-Hochberg. A complete list of all significantly enriched GO terms can be found in Figure 4—source data 2.
-
Figure 4—source data 1
The intersection of the DESeq2 output (differential expression) and the CEH-18 binding sites (Kudron et al., 2018).
- https://doi.org/10.7554/eLife.46418.014
-
Figure 4—source data 2
The complete list of GO terms whose enrichment was determined using the significantly differentially expressed genes associated with CEH-18 binding peaks when comparing young hermaphrodites vs. young feminized individuals (selected, enriched GO results are displayed in Figure 4G and Figure 4—figure supplement 1C).
- https://doi.org/10.7554/eLife.46418.015
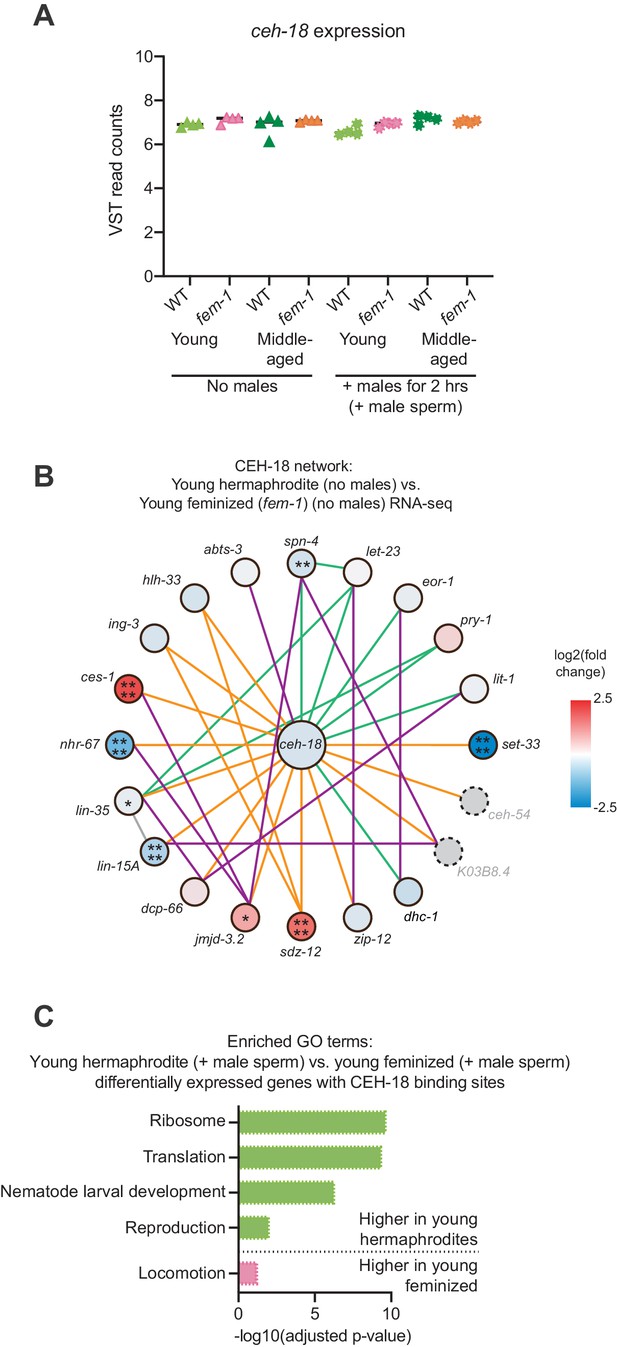
CEH-18 expression and network analysis.
(A) The sperm-sensing gene ceh-18 was not differentially expressed between young and middle-aged hermaphrodites and feminized individuals. Shown here are the VST-normalized read counts for ceh-18. DESeq2 calculated differentially expression values can be found in Figure 2—source data 1. (B) Network analysis for the sperm-sensing transcription factor CEH-18 determined by GeneMANIA (Warde-Farley et al., 2010). Lines indicating genetic interactions are in green, co-expression in purple, physical interactions in orange, and ‘other’ in gray. The differential expression of the genes in the network is shown with a blue-red gradient. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, no stars = not significant. Network members not detected by the RNA-seq are in gray with dotted outlines. The number of differentially expressed genes in these networks was not statistically enriched as measured using the hypergeometric distribution test. See Figure 2—source data 1 for exact differential expression values. (C) Selected, enriched GO terms from the genes that are differentially expressed between young hermaphrodite versus young feminized that mated with a male and contain a CEH-18 binding site as defined by CEH-18 ChIP-seq (Kudron et al., 2018), see Material and methods. GO terms that were enriched in the genes expressed more highly in young hermaphrodites are shown in green and GO terms enriched in the genes more highly expressed in young feminized individuals are in pink. P-values were calculated with the Fisher’s exact test and were corrected for multiple hypothesis testing with Benjamini-Hochberg. A complete list of all significantly enriched GO terms can be found in Figure 4—source data 2.
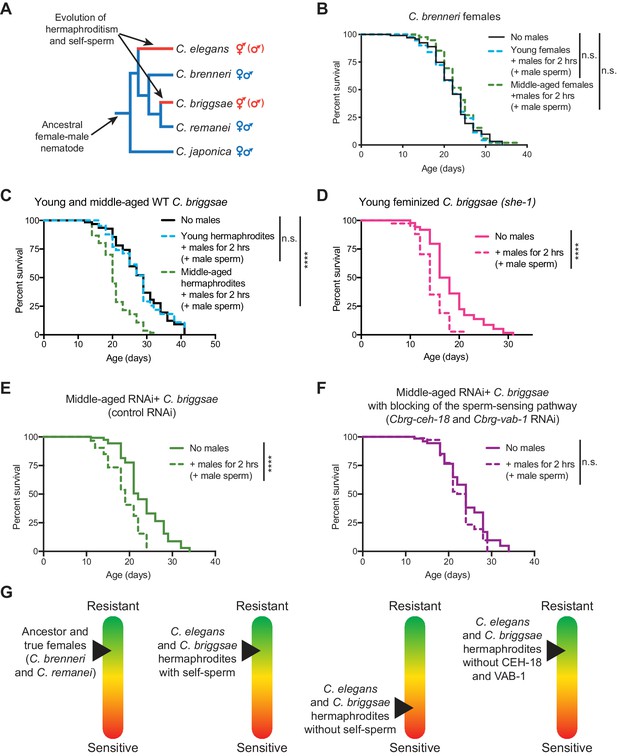
The importance of self-sperm in protecting young hermaphrodites against males independently evolved in a distantly-related nematode species.
(A) The phylogeny of Caenorhabditis nematodes with the hermaphroditic lineages shown in red. (B) C. brenneri females lived a normal lifespan if they mated with a male during brief, two-hour interaction when young or middle-aged (n.s. vs. no males). For C. brenneri, we defined young as day 4 of life and middle-aged as day 10 of life. (C) C. briggsae hermaphrodites had a normal lifespan if they received male sperm during a brief interaction with males when they were young (day 3 of life, blue dashed line, n.s vs. no males) but had a shortened lifespan if they received male sperm when middle-aged (day 8 of life, green dashed line, p<0.0001 vs. no males). (D) Feminized C. briggsae (she-1[v35]) had a shortened lifespan if they received male sperm following a brief interaction with males when young (day 3 of life, p<0.0001 vs. no males). (E-F) Middle-aged, self-sperm depleted C. briggsae have a shortened lifespan following mating with a male (E, p<0.0001 vs. no males) but when the orthologs of the sperm-sensing pathway (Cbr-ceh-18 and Cbr-vab-1) was blocked by RNAi knock-down, these worms lived a normal lifespan (F, n.s. vs. no males). Experiments were performed using a transgenic C. briggsae strain that is capable of RNAi knock-down by dsRNA ingestion (mfIs42[Cel-sid-2; Cel-myo-2::DsRed]). (G) A scheme summarizing the resistance of sensitivity of different species, aged individuals, and mutants to mating-induced demise. Mated worms (dashed lines) were selected by hand-picking based on the presence of fluorescent male sperm in their uterus or spermatheca. C. briggsae males are him-8(v186) mutants (Wei et al., 2014). Lifespans were performed with 82–151 animals. Lifespan data are plotted as Kaplan-Meier survival curves and p-values were determined using Mantel-Cox log ranking. *p<0.05, **p<0.01, p < *** 0.001, ****p<0.0001, n.s. = not significant. See also Supplementary file 2 for extended statistics and replicates.
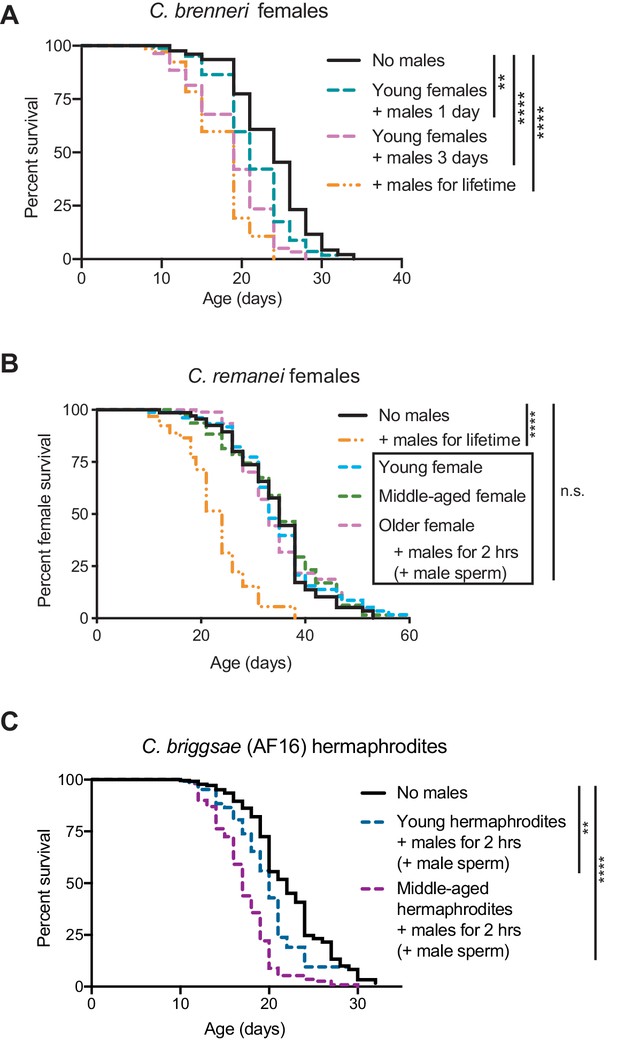
The evolution of self-sperm mediated protection from mating-induced demise.
(A) C.brenneri female lifespan was shortened if they interacted with males for a longer period of time. Interacting with males for their entire lifetime significantly shortened lifespan (orange dashed line, p<0.0001 vs. no males). C. brenneri female lifespan was also reduced when interacting with males for one day when young (blue dashed line, p=0.0011 vs. no males) or for three days when young (pink dashed line, p<0.0001 vs. no males). (B) C. remanei females lived a normal lifespan after a brief, 2 hr mating with males at all the ages tested (n.s. vs. no males) but succumbed when they interacted with young males for their entire lifetime (p<0.0001 vs. no males). The ages when the females mated with males are day 4 of life (young), day 11 of life (middle-aged), and day 18 of life (older). (C) The wild-type C. briggsae strain AF16 was more resistant to a brief interaction with males when young (day 3 or 4 of life, p=0.0093 vs. no males and p<0.0001 vs. middle-aged interaction) compared to when middle-aged (day 7 or 8 of life, p<0.0001 vs. no males). We note that this strain had an extremely high rate of leaving the agar plate and censored at a rate of approximately 60–90%. Because of this high rate of censorship, the results from three independent experiments have been combined and are displayed together. For the results from each independent experiment, see Supplementary file 2. Lifespan assays were performed with 79–425 animals. For the conditions in which sexual interactions were very brief (2 hr), mated worms (dotted lines in panels B and C) were selected by hand-picking based on the presence of fluorescent male sperm in their spermatheca or uterus. Lifespan data are plotted as Kaplan-Meier survival curves and p-values were determined using Mantel-Cox log ranking. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, n.s. = not significant.
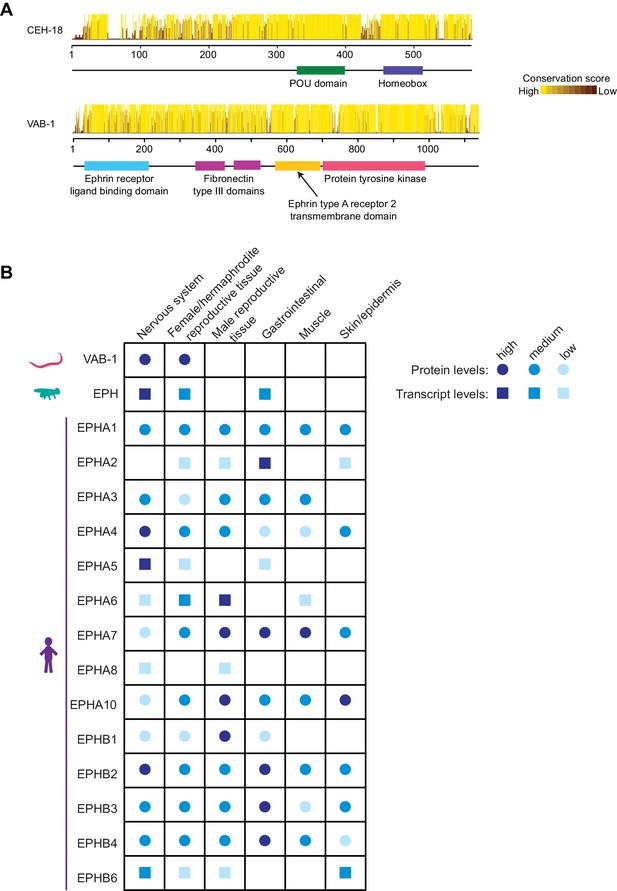

Conservation of CEH-18 and VAB-1.
(A) Conservation scores for the Caenorhabditis orthologs of CEH-18 and VAB-1. (B) Tissue-specific protein (circles) or RNA (squares) expression levels for Ephrin receptors in C. elegans (www.wormbase.org), Drosophila (www.flyatlas.gla.ac.uk/FlyAtlas2/index.html and Leader et al., 2018), and humans (www.proteinatlas.com and Uhlén et al., 2015; Uhlen et al., 2010).

RNAi knock-down of the sperm-sensing pathway in C.briggsae.
(A-B) Representative images of the germlines of feminized (mfIs42[Cel-sid-2; Cel-myo-2::DsRed]; she-1[v35]) C. briggsae individuals cultured on control RNAi (empty vector, A) or Cbr-ceh-18 and Cbr-vab-1 RNAi (B). The proximal gonads (PG) are indicated. (C) Quantification of the number of oocytes present in the gonads of feminized (mfIs42[Cel-sid-2; Cel-myo-2::DsRed]; she-1[v35]) C. briggsae individuals cultured on control RNAi (empty vector, green data points) or Cbr-ceh-18 and Cbr-vab-1 RNAi (purple data points). Each data point represents one gonad arm. 12–18 animals were counted, and the p-value was calculated using the Mann-Whitney test. (D) The presence of unfertilized oocytes in the uterus of feminized (mfIs42[Cel-sid-2; Cel-myo-2::DsRed]; she-1[v35]) C. briggsae individuals cultured on control RNAi (empty vector) or Cbr-ceh-18 and Cbr-vab-1 RNAi. 12–18 animals were quantified. (E) Knock-down in the C. briggsae pharynx bulb of the transgenic marker Cele-myo-2::DsRed by ingested dsRNA was measured on day 8 of life. 30–34 animals were measured and the p-value was calculated using the Mann-Whitney test.
Additional files
-
Supplementary file 1
Nematode strains used in this study.
The complete list of all strains used in this study, with their genotype and source listed.
- https://doi.org/10.7554/eLife.46418.020
-
Supplementary file 2
Lifespan assay results.
The data for the lifespan assays displayed in the figures as well as lifespan assays whose plots are not shown in the manuscript. Each set of lifespan assays performed together is separated from the other sets of assays by a blank row in the table. ‘Temp.’ describes the temperature at which the worms were grown. Note that 25→20 indicates that the hermaphrodites and feminized individuals were grown during development at the restrictive temperature and then shifted to a lower temperature on day 3 of life as described in the methods. All p-values were determined using Mantel-Cox log-ranking.
- https://doi.org/10.7554/eLife.46418.021
-
Supplementary file 3
Mating efficiencies.
The mating efficiencies performed. The individuals whose mating efficiencies were measured are underlined in each set of mating partners. Successful mating was determined by the presence of fluorescent male sperm in the spermatheca or uterus. The presence of fluorescent male sperm is indicative of fertilization (see Figure 1—figure supplement 1B,C) though this was not specifically measured in this assay. See Source data 1 for a complete list of the mating efficiencies for each plate of 20 hermaphrodites (or germline mutants) with 40 males that were used to calculate the median and p-values.
- https://doi.org/10.7554/eLife.46418.022
-
Source data 1
The raw data that comprise Supplementary file 3.
- https://doi.org/10.7554/eLife.46418.023
-
Transparent reporting form
- https://doi.org/10.7554/eLife.46418.024





