Synergy between serum amyloid A and secretory phospholipase A2
Figures
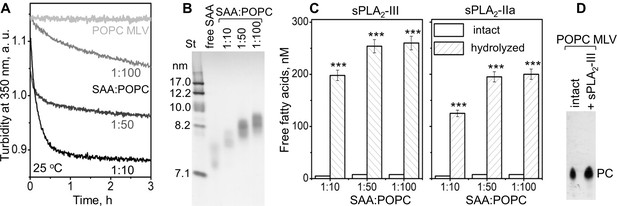
SAA remodels phospholipid bilayers into small particles that form substrates for sPLA2.
(A) POPC MLV (~200 nm) were incubated with SAA at 25°C, and the time course of MLV remodeling into smaller particles was monitored by turbidty at 350 nm. Protein:lipid molar ratios were 1:10, 1:50 and 1:100, as indicated. Protein-free POPC MLV were used as a control. The data for POPC MLV alone (light gray) closely superimposed similar data using POPC MLV with either apoA-I or apoA-II at a 1:100 protein:lipid weight ratio (not shown to avoid overlap). (B) Non-denaturing PAGE of the POPC mixtures with SAA, performed after 6 hr of incubation, shows the formation of SAA-POPC complexes. SAA:POPC molar ratios are indicated; lipid-free SAA is shown for comparison. Gels in this and other figures were stained with Denville Blue protein stain. (C) SAA-POPC particles shown in panel (B) were isolated by size-exclusion chromatography (SEC) and used as substrates for sPLA2-III (left) or sPLA2-IIa (right). Free fatty acids produced per nm of lipid are shown as average values of three independent measurements with standard errors of mean. The protein:lipid molar ratios in the initial incubation mixtures are indicated; sPLA2-free particles were used as controls. FFA produced in the presence and in the absence of sPLA2 were compared using the t-test; ***p<0.001. In SAA-POPC particles, approximately 70% of the total lipid was hydrolyzed by sPLA2-III and 60% by sPLA2-IIa. (D) Thin-layer chromatography analysis of POPC MLV before (intact) or after their incubation with sPLA2-III (+ sPLA2-III). The PC band is indicated; the absence of the lysoPC band underneath the PC band indicates the absence of significant hydrolysis.
-
Figure 1—source data 1
Free fatty acid analysis of SAA:POPC complexes hydrolysed by sPLA2.
- https://doi.org/10.7554/eLife.46630.004
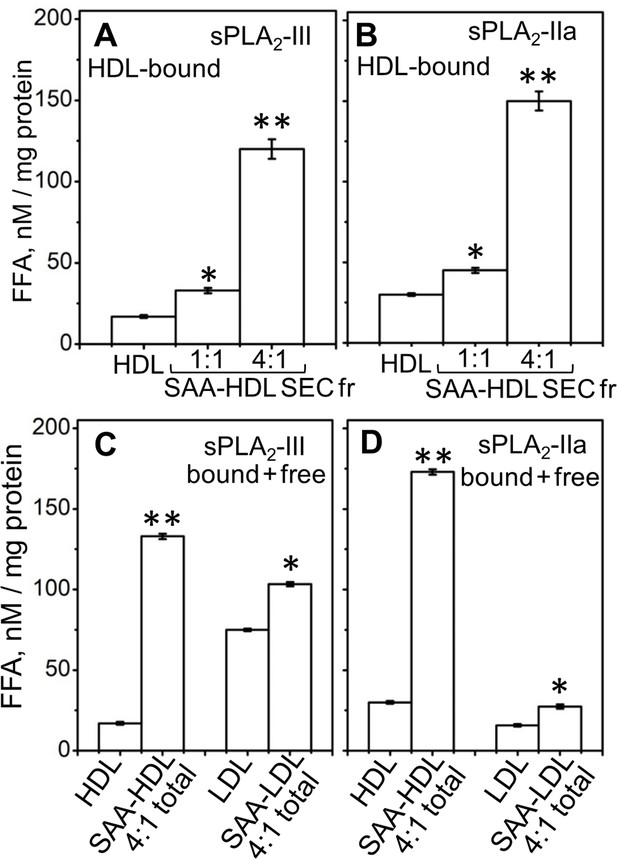
Effect of SAA on the lipolysis of HDL and LDL by sPLA2-III or sPLA2-IIa.
(A, B) FFA were generated upon lipolysis of either native HDL or SAA-HDL. For SAA-HDL, the SEC fraction containing HDL-bound proteins (Sec Fr) was isolated from the total incubation mixtures, which contained 1:1 or 4:1 SAA:apoA-I molar ratios as indicated. (C, D) FFA were generated upon lipolysis of HDL, LDL and their total incubation mixtures with SAA. Protein weight ratios were 1:4 SAA to apoA-I in HDL or 1:4 SAA to apoB in LDL. The amounts of FFA produced per mg of apoA-I (in HDL) or apoB (in LDL) are shown as the means of three independent measurements with the standard deviations of these means. FFA generated in the presence and in the absence of SAA were compared using one-way ANOVA; *, p<0.05; **, p<0.01. A characterization of protein-containing complexes formed upon incubation of SAA with human plasma lipoproteins is shown in Figure 2—figure supplement 1.
-
Figure 2—source data 1
Free fatty analysis of HDL and LDL hydrolysed by sPLA2.
- https://doi.org/10.7554/eLife.46630.007
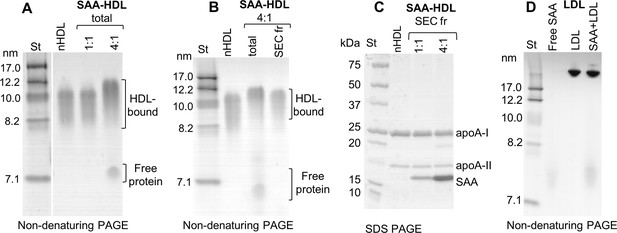
Characterization of protein-containing complexes formed upon incubation of SAA with human plasma lipoproteins.
(A) Non-denaturing PAGE of incubation mixtures of SAA and single-donor human HDL (SAA-HDL). Molar ratios of SAA to apoA-I, the main HDL protein, are 1:1 or 4:1 (as indicated on the lanes). Fractions corresponding to HDL-bound and HDL-unbound (free) protein are indicated. Non-modified HDL (nHDL) is shown for comparison. Molecular size standards (St) are shown. These and all other gels were stained with Denville Blue protein stain. (B) Non-denaturing PAGE of SAA-HDL samples containing 4:1 molar ratio of SAA to apoA-I. The total incubation mixture (total) and the HDL-containing fraction isolated by SEC (SEC fr) are shown. (C) SDS PAGE of the HDL fraction isolated by SEC (SEC fr) from the total SAA-HDL incubation mixture. The total SAA-HDL contained a molar ratio of 1:1 or 4:1 SAA:apoA-I (as indicated on the lanes). (D) Non-denaturing PAGE of single-donor LDL incubated with SAA (SAA + LDL) at a 1:1 protein weight ratio of SAA to apoB. Lipid-free SAA and LDL alone are included for comparison.
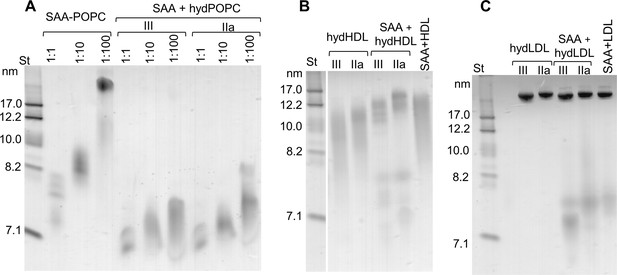
SAA forms complexes with hydrolyzed model or plasma lipids.
(A) SAA was incubated with either unmodified POPC to form SAA-POPC complexes or with hydrolyzed POPC to form SAA + hydPOPC complexes (see 'Materials and methods' for details). Protein:POPC molar ratios were 1:1, 1:10 or 1:100 as indicated; sPLA2-III or sPLA2-IIa was used as indicated. (B, C) Human plasma lipoproteins including HDL (B) and LDL (C) were hydrolyzed with sPLA2 group-III or -IIa to form hydHDL or hydLDL, respectively. In samples marked SAA + hydLDL (B) or hydHDL (C), the hydrolyzed lipoproteins were incubated with SAA using protein weight ratios of 1:1 SAA:apoA-I (for HDL) or 1:1 SAA:apoB (for LDL) as described in 'Materials and methods'. Similar incubation mixtures of SAA with non-hydrolyzed HDL (SAA+HDL) or LDL (SAA+LDL) are shown for comparison. Figure 3—figure supplement 1 shows non-denaturing PAGE that monitors the remodeling of SAA-containing model and plasma lipoproteins upon their hydrolysis by sPLA2.
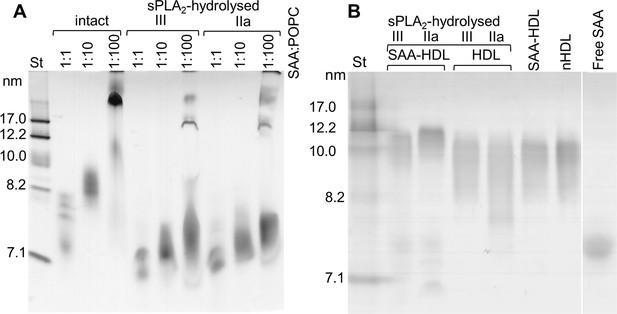
Non-denaturing PAGE monitors remodeling of SAA-containing model and plasma lipoproteins upon their hydrolysis by sPLA2.
(A) SAA-POPC complexes were prepared by incubating SAA with POPC SUV using SAA:POPC molar ratios of 1:1, 1:10 or 1:100 (indicated in the lanes). Lanes for intact SAA-POPC particles are shown, as well as those for similar particles that have been hydrolyzed with sPLA2-III or sPLA2-IIa. (B) Native HDL and HDL that have been enriched with exogenous SAA using an SAA:apoA-I molar ratio of 1:4 (SAA-HDL) hydrolyzed by sPLA2-III or sPLA2-Ila (as indicated on the lanes). Non-hydrolyzed native HDL (nHDL), SAA-HDL and lipid-free SAA were used as controls.
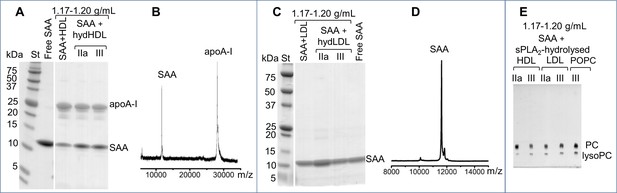
Biochemical analysis of the ~7 nm complexes formed by SAA and lipolytic products.
SAA-containing complexes, which were obtained upon lipolysis of model (SAA-POPC) or plasma lipoproteins (HDL and LDL) by sPLA2, were isolated at 1.17–1.20 g/mL density (Figure 4—figure supplement 1). These isolated complexes, marked SAA + hydHDL (A), SAA + hydLDL (B) or SAA + sPLA2-hydrolyzed HDL, LDL or POPC (E), were analyzed for protein (A–D) and lipid composition (E). SDS PAGE (A) and matrix-assisted laser desorption ionization – time of flight mass spectrometry (B) of SAA + hydHDL revealed SAA and apoA-I; the protein mass detected by mass spectrometry was 11,606 Da for SAA and 28,086 for apoA-I (B). Similar analyses of SAA + hydLDL complexes showed only SAA (C, D). (E) Thin-layer chromatography showed the presence of PC and lysoPC in the SAA-containing ~7 nm complexes that were obtained from all hydrolyzed lipoproteins (HDL, LDL) or model lipids (POPC). Lipid-free SAA and the 1.17–1.20 g/mL density fraction isolated from SAA mixtures with non-hydrolyzed HDL or LDL (SAA + HDL in panel (A) and SAA + LDL in (C)) are shown for comparison.
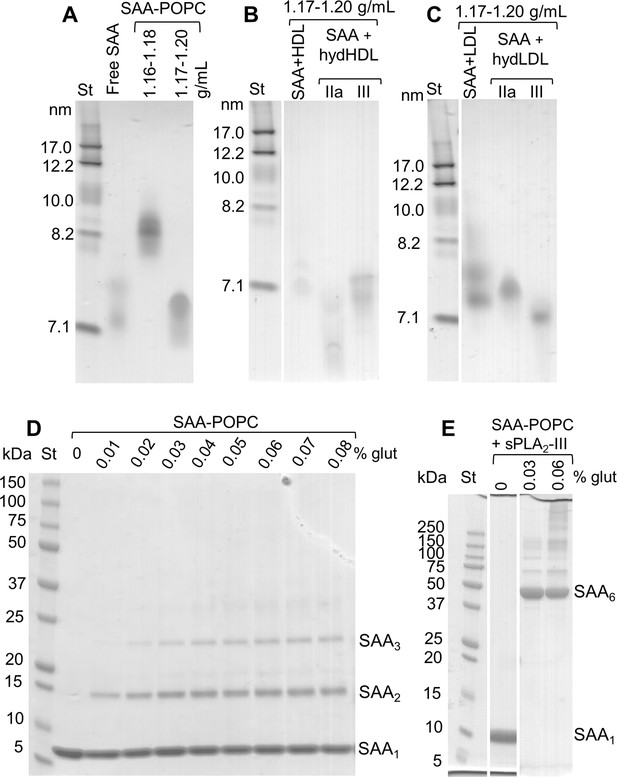
Gel electrophoresis of isolated SAA complexes with the products of phospholipid hydrolysis.
All gels were stained with Denville Blue protein stain. (A) Non-denaturing PAGE of model SAA-POPC complexes, which were prepared as described in 'Materials and methods' using 1:10 SAA:POPC mol:mol and hydrolyzed by sPLA2-III. The intact complexes were isolated in the density range 1.16–1.18 g/mL, and the hydrolyzed complexes in the density range 1.17–1.20 g/mL (as indicated on the lanes). Lipid-free SAA is shown for comparison. Non-denaturing PAGE of (B) HDL and (C) LDL from human plasma that were hydrolyzed by sPLA2-III or sPLA2-IIa (as indicated) and incubated with SAA to form 7–7.5 nm complexes (as described in the Figure 3 legend). These complexes were isolated by density gradient centrifugation in the density range 1.17–1.20 g/mL (as indicated on the lanes). Similar density fractions isolated from SAA mixtures with intact HDL or LDL (labeled SAA + HDL in panel (B) and SAA + LDL in panel (C)) are shown for comparison. SDS PAGE of model SAA-POPC complexes that were either (D) intact or (E) hydrolyzed by sPLA2-III. The complexes (containing 0.5 mg/ml protein in 50 mM PBS at pH 7.5) have been cross-linked with glutaraldehyde (as described in 'Materials and methods); the cross-linker concentrations are indicated. Gel bands corresponding to SAA monomers and oligomers are indicated.
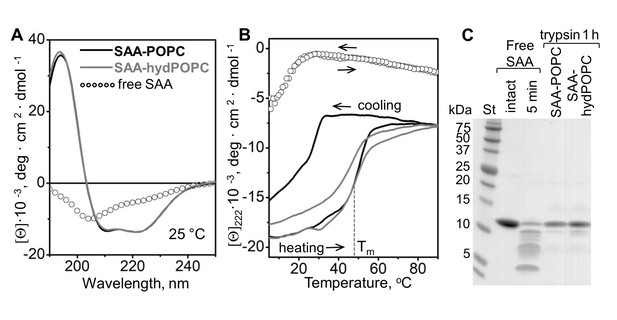
Structure and stability of SAA complexes with the products of POPC hydrolysis.
Intact SAA-POPC complexes (prepared using 1:10 protein:lipid molar ratio) were hydrolyzed with sPLA2-III, and the 1.17–1.20 g/ml density fraction containing ~7 nm particles was isolated (for details, see main text, Figure 3 and 'Materials and methods'). Far-UV CD spectra at 25°C (A) and the melting data (B) are shown for these isolated ~7 nm complexes (SAA-hydPOPC), the parent particles (SAA-POPC) and lipid-free SAA. The melting data show CD signal at 222 nm, [Θ]222(T), that monitors α-helical structure during heating and cooling from 10°C to 90°C at a rate of 60 °C/h. Arrows show directions of the temperature changes; the dotted line indicates the melting temperature for SAA-hydPOPC, Tm = 48 ± 2°C. (C) Limited tryptic digestion of free SAA, SAA-POPC, and SAA-hydPOPC complexes monitored by SDS PAGE. Trypsin was incubated at room temperature for 5 min with free SAA or for 1 hr with SAA-lipid complexes (see 'Materials and methods' for details). Figure 5—figure supplement 1 shows the structure and stability of the 7–7.5 nm complexes formed by SAA and the products of LDL hydrolysis. Figure 5—figure supplement 2 shows the structure and stability of SAA complexes with lysoPC and POPC.
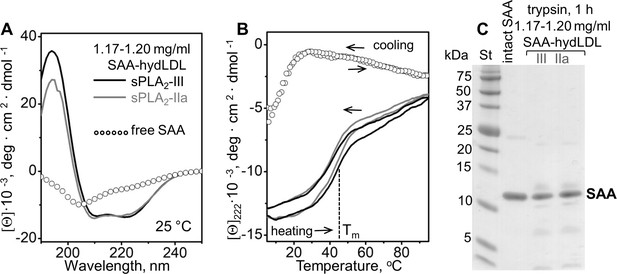
Structure and stability of the 7–7.5 nm complexes formed by SAA and the products of LDL hydrolysis.
SAA was incubated with human plasma LDL (SAA-hydLDL) that had been hydrolyzed either with sPLA2-III or with sPLA2-IIa. The ~7–7.5 nm complexes, which were isolated from the incubation mixtures in the density range 1.17–1.20 g/ml, are shown in Figure 4—figure supplement 1. (A) Far-UV CD spectra of the 7–7.5 nm complexes, which were obtained from LDL hydrolyzed with either sPLA2-III (dark solid line) or sPLA2-IIa (light solid line), were recorded at 25°C. A similar spectrum for lipid-free SAA is shown for comparison (open circles). (B) The melting data of the complexes were recorded by CD at 222 nm during heating and cooling from 10°C to 90°C at a rate of 60 °C/h. Arrows show the directions of temperature changes; the dotted line indicates the melting temperature, Tm. Similar data for lipid-free SAA are shown for comparison. Line coding is the same as in panel (A). (C) SDS PAGE showing the products of limited tryptic digestion of these complexes after 1 hr of incubation with trypsin at room temperature. Intact SAA in the absence of trypsin is shown for comparison.
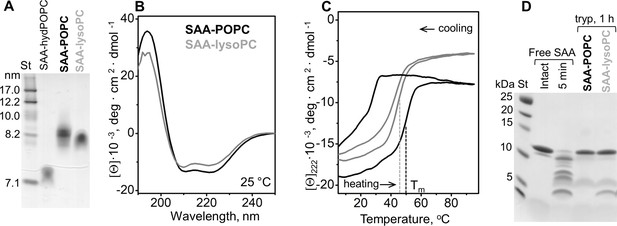
Structure and stability of SAA complexes with lysoPC and POPC.
Complexes of SAA with either lysoPC (SAA-lysoPC) or POPC (SAA-POPC) were prepared using 1:10 SAA:PC molar ratio and analyzed as follows. (A) Non-denaturing PAGE of these SAA-POPC and SAA-lysoPC complexes. SAA complexes with hydrolyzed POPC (SAA-hydPOPC), which were prepared at a 1:10 protein:lipid molar ratio, are shown for comparison. (B) Far-UV CD spectra of SAA-POPC (black) and SAA-lysoPC (gray) complexes at 25°C were used to assess the protein secondary structure. (C) Thermal stability of the complexes was assessed by CD signal at 222 nm recorded during heating and cooling from 10°C to 90°C at a rate of 60 °C/h. Arrows show the directions of temperature changes; dotted lines indicate melting temperatures, Tm. Line coding is the same as in panel (B). (D) Limited tryptic digestion of free SAA and of SAA-POPC and SAA-lysoPC complexes monitored by SDS PAGE. The complexes were incubated with trypsin at room temperature for 1 hr (see 'Materials and methods' for details). Free SAA that was intact or has been incubated with trypsin for 5 min is shown for comparison (left lanes).
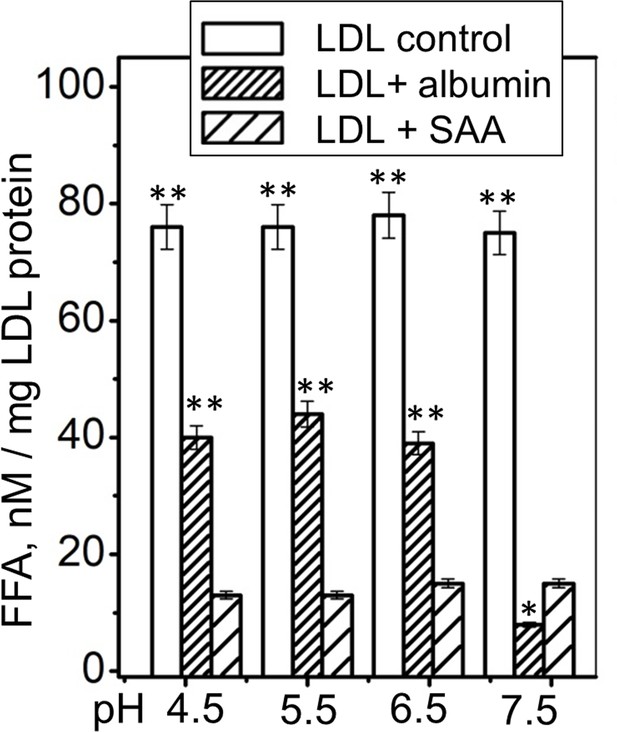
FFA removal by SAA and by albumin as a function of pH.
Single-donor LDL were hydrolyzed with sPLA2-III and incubated at pH 4.5–7.5 either alone (control), with albumin (LDL + albumin) or with SAA (LDL + SAA). Next, LDL were re-isolated by density gradient centrifugation and their residual FFA content was determined. The results are shown as the mean of three independent measurements with their standard errors. FFA levels in controls and in albumin-treated LDL were compared with those in SAA-treated LDL using one-way ANOVA; *, p<0.5; **, p<0.01. Figure 6—figure supplement 1 shows that SAA and albumin sequester naturally occurring FFA from human plasma LDL at pH 7.5. Figure 6—figure supplement 2 shows that human SAA 1.1 sequesters the products of phospholipid hydrolysis from model and plasma lipoproteins.
-
Figure 6—source data 1
Free fatty acid analysis of LDL at different pH.
- https://doi.org/10.7554/eLife.46630.020
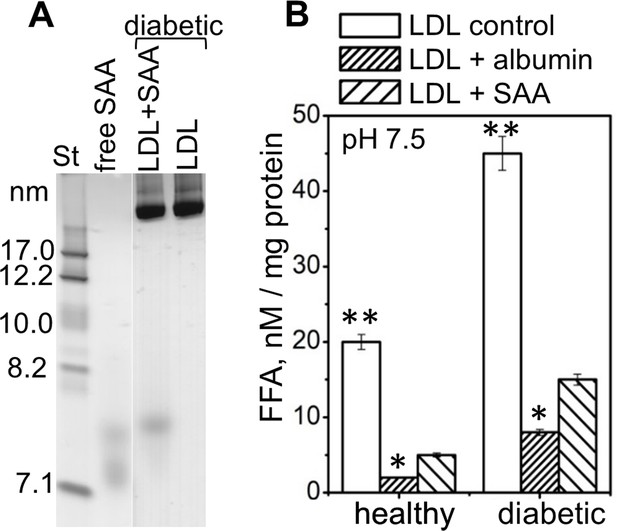
SAA and albumin sequester naturally occurring FFA from human plasma LDL at pH 7.5.
LDL were obtained from pooled plasma of diabetic and from healthy normolipidemic patients as previously described (Jayaraman et al., 2017b). Diabetic LDL had 20% higher content of endogenous FFA compared to normolipidemic LDL. (A) Non-denaturing PAGE shows ~7.5 nm species in the incubation of SAA with diabetic LDL (LDL + SAA). Diabetic LDL alone and lipid-free SAA alone are shown for comparison. (B) Quantification of FAA removal by SAA and by human serum albumin from healthy and diabetic LDL at pH 7.5. LDLs (0.2 mg/ml apoB) were incubated either alone, with SAA (0.2 mg/ml), or with albumin (0.2 mg/ml). LDL re-isolation was then carried out by density centrifugation and analysis of the residual FFA content in these LDL. Mean values with the standard error of means are shown. FFA levels in control and in albumin-treated LDL (LDL control and LDL + albumin) were compared with those in SAA-treated LDL (LDL + SAA) using one-way ANOVA; *, p<0.05, **, p<0.01.
-
Figure 6—figure supplement 1—source data 1
Free fatty analysis of normal and diabetic LDL.
- https://doi.org/10.7554/eLife.46630.017
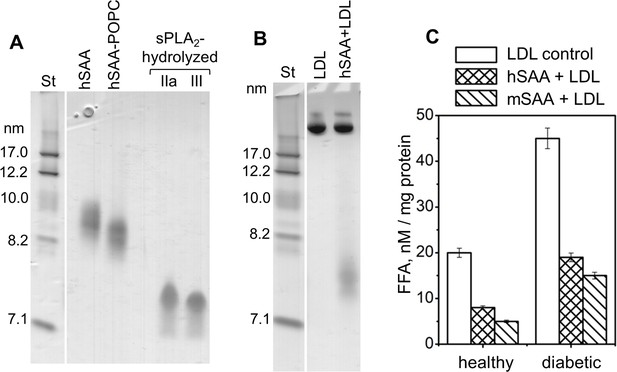
Non-denaturing PAGE shows that human SAA 1.1 (hSAA1.1) sequesters the products of phospholipid hydrolysis from model and plasma lipoproteins.
(A) Complexes of hSAA1.1 with POPC, which were prepared using 1:10 protein-lipid molar ratio, were hydrolyzed with sPLA2-IIa or sPLA2-III (as indicated on the lanes). Intact non-hydrolyzed complexes (hSAA-POPC) and hSAA1.1 alone (hSAA) are shown for comparison. (B) Non-denaturing PAGE of normal human plasma LDL alone (LDL) or in mixtures with SAA (hSAA + LDL) at 1:1 weight ratio of SAA to apoB. SAA in these samples is not bound to LDL. (C) FFA analysis in LDL alone (LDL control) or in the presence of human or murine SAA (hSAA + LDL or mSAA + LDL). Human LDL were obtained from the pooled plasma of healthy (left) or diabetic patients (right) as described in the 'Materials and methods'. FFA levels in these LDL were compared with those in the control LDL using one-way ANOVA; **, p<0.01. The data represent the mean of three independent measurements and the standard errors.
-
Figure 6—figure supplement 2—source data 1
Free fatty acid analysis of LDL incubated with human and mouse SAA.
- https://doi.org/10.7554/eLife.46630.019
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.46630.021





