Splicing in a single neuron is coordinately controlled by RNA binding proteins and transcription factors
Figures
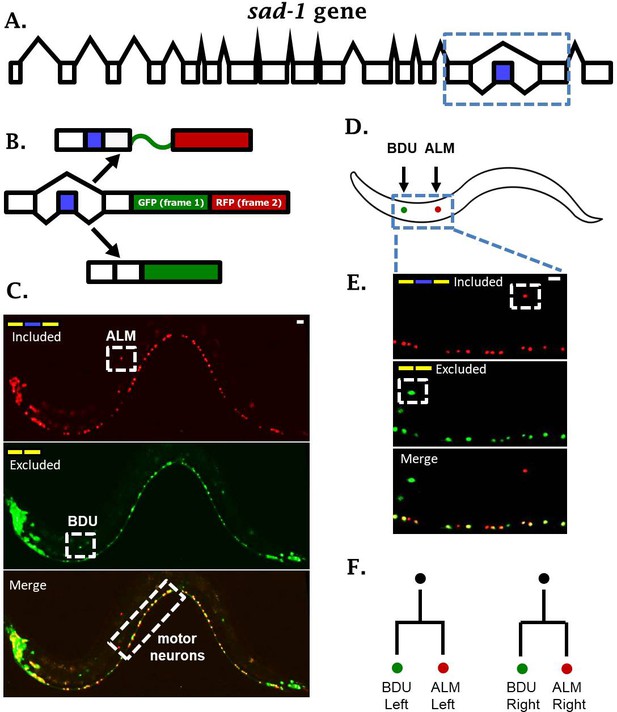
sad-1 is alternatively spliced in single neurons.
(A) The sad-1 gene. Alternative cassette exon in blue. (B) Two-color splicing reporter schematic for sad-1 cassette exon. The cassette exon encodes a + 1 nt frameshift so that when skipped, GFP is produced with an in frame stop codon. When skipped, GFP is read out of frame without stop codons, followed by in-frame translation of RFP. (C) Whole worm fluorescent micrograph demonstrating both exon inclusion (RFP) and skipping (GFP) in many neurons, while certain neurons express only the included (ALM) of skipped (BDU) isoforms. (D–E) Higher magnification focusing on ALM and BDU neurons. (F) BDU and ALM are both paired neurons present on the left and right side of the worm. Each BDU neuron is a sister cell to an ALM neuron, derived from the same neuroblast. Scale bar represents 10 µm.
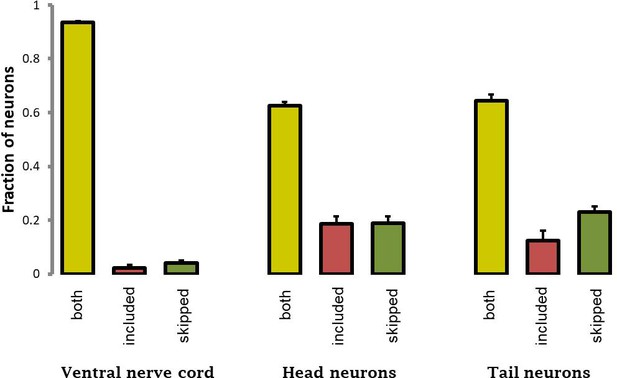
Gross quantification of fraction of neurons expressing included (red), skipped (green), or both (yellow) isoforms of sad-1, in either ventral nerve cord, head, or tail.
https://doi.org/10.7554/eLife.46726.004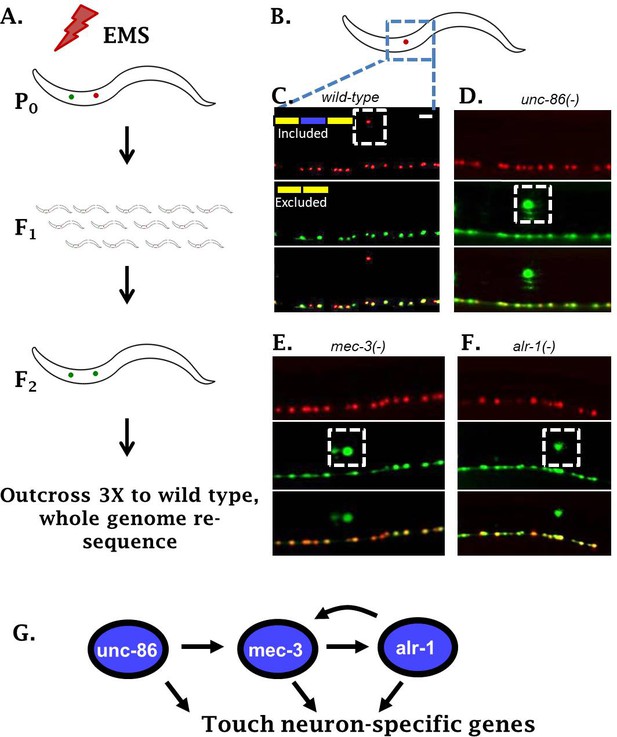
Genetic screen identifies neuronal TFs affecting sad-1 splicing in the ALM neuron.
(A) Schematic of forward genetic screen to identify regulators of sad-1 splicing in the ALM touch neuron. (B–F) ALM neurons (dashed boxes) shift from complete inclusion (RFP) to skipping (GFP) in unc-86(e1416), mec-3(e1338), or alr-1(oy42) TF mutants. Splicing phenotypes fully penetrant (n = 50 animals) (G) Previously-identified roles of the three TFs in a transcriptional cascade to control touch neuron gene expression. Scale bar represents 10 µm.
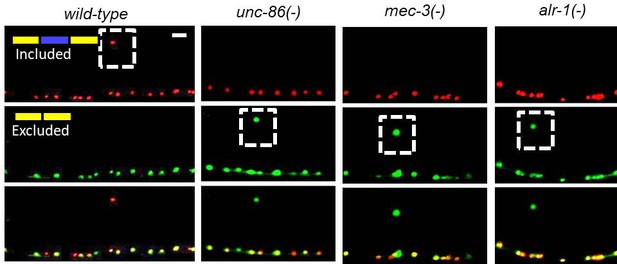
TF alleles identified in genetic screen cause sad-1 splicing defects in ALM: unc-86(csb9), mec-3(csb10), alr-1(csb11).
https://doi.org/10.7554/eLife.46726.006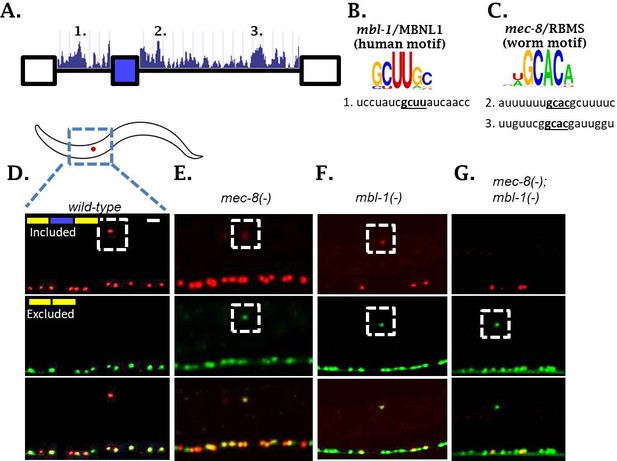
Two neuronal RBPs combinatorially control sad-1 splicing in ALM neurons.
(A) Conservation scores in the introns surrounding sad-1 exon 15, basewise phyloP26way comparison of 26 nematode species (Hubisz et al., 2011). Numbers 1–3 indicate consensus binding motifs for mbl-1 and mec-8 displayed in B-C. (B–C) cis-elements matching consensus binding motifs for mbl-1 and mec-8. (D–F) mec-8 and mbl-1 mutants both cause a partial loss of sad-1 exon inclusion. (G) mec-8; mbl-1 double mutants cause complete loss of exon inclusion, phenocopying the TF mutants. Splicing phenotypes fully penetrant (n = 50 animals) Scale bar represents 10 µm.
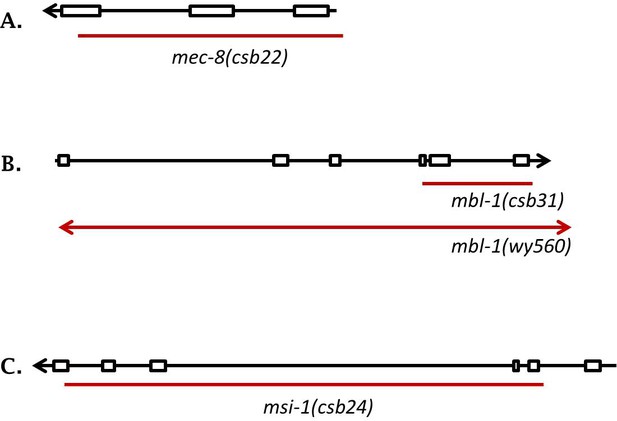
Deletion alleles used in this study, in addition to canonical mutations and mutations identified in our forward genetic screen.
In black is a schematic of the entire gene, in red a representation of the deletion. (A) mec-8(csb22) (B) mbl-1(csb31) and mbl-1(wy560). (C) msi-1(csb24).
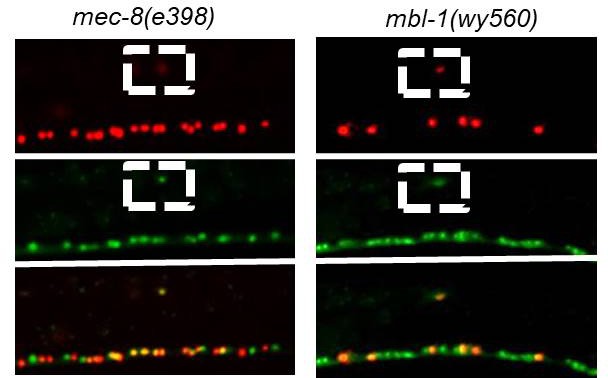
Canonical RBP alleles of mec-8 and mbl-1 affect sad-1 splicing similarly to CRISPR deletions of mec-8 and mbl-1.
mec-8 (e398) premature stop codon mutation. mbl-1(wy560) large deletion. Splicing phenotypes fully penetrant (n = 50 animals).
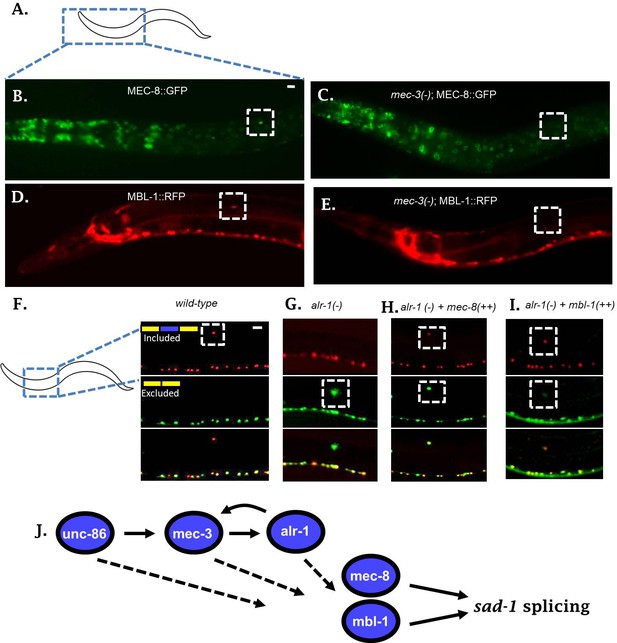
Neuronal TFs establish expression of both mec-8 and mbl-1 to mediate splicing of sad-1 in ALM neurons.
(A–B) A mec-8 translational GFP fosmid reporter reveals strong expression in ALM neuron (strong expression in 28/31 = 90% of animals inspected). (C) In a mec-3 TF mutant, mec-8 expression is absent specifically in ALM (no detectable expression in 43/50 = 86%, dim expression in 7/50 = 14% of animals inspected). (D) mbl-1 translational RFP fosmid reporter is expressed in ALM neuron (strong expression in 19/20 = 95% of animals inspected). (E) In a mec-3 mutant, mbl-1 expression is absent specifically in ALM (no detectable expression in 19/21 = 90%, dim expression in 2/21 = 10% of animals inspected). (F–G) Aberrant splicing of sad-1 in alr-1 TF mutants is partially rescued by over-expression of either mec-8 (6/6 animals examined) or mbl-1 (6/7 animals examined) RBPs (H–I). Scale bar represents 10 µm.
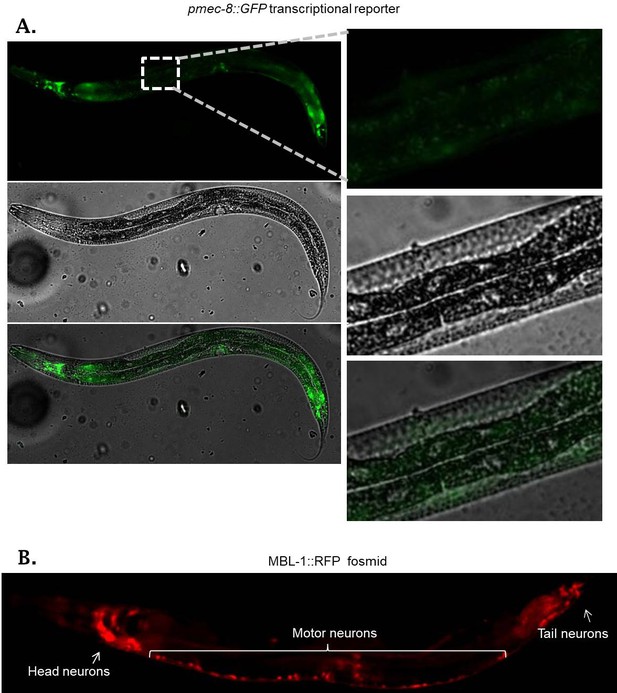
Transcriptional and translational reporters for MEC-8 and MBL-1.
(A) Promoter-GFP fusion for mec-8 exhibits GFP expression in various cells but not in ALM touch neurons. Right panel represents high magnification of ALM region (dotted square in left panel). (B) Micrograph of mbl-1::RFP fosmid as in Figure 4D, demonstrating expression in numerous head neurons, motor neurons, and tail neurons.
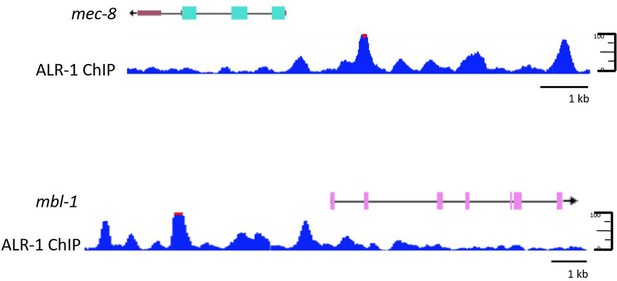
ALR-1 ChIP peaks in mec-8 and mbl-1 promoters.
ALR-1 ChIP peaks from modENCODE data suggest that ALR-1 may bind directly to the promoters of mec-8 (upper panel) and mbl-1 (lower panel), although these peaks do not immediately contain consensus binding motifs for a mouse homologue of alr-1 Arx (Berger et al., 2008). Red bars capping ChIP peaks indicate values greater than the mean +5X the standard deviation.
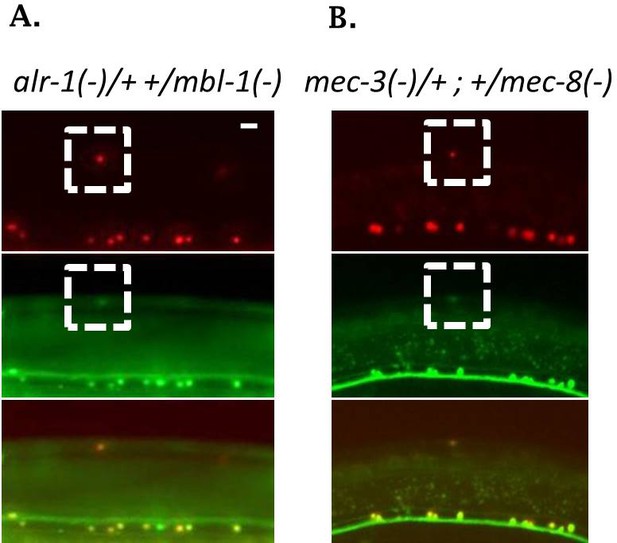
Double heterozygotes for TFs and RBPs display moderate defects in sad-1 splicing in ALM neurons.
(A) alr-1; mbl-1 double heterozygotes. (B) mec-3; mec-8 double heterozygotes. ALM splicing phenotype fully penetrant (n = 10 animals).
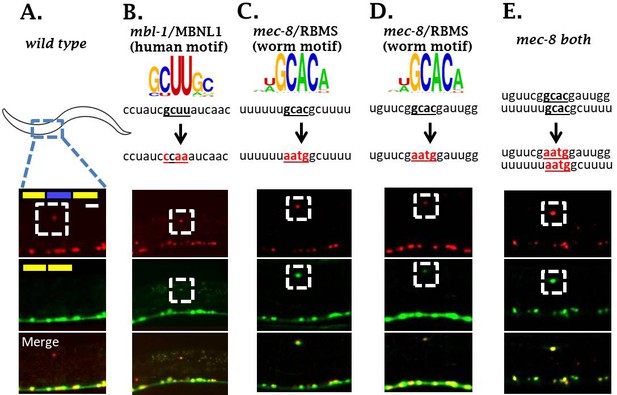
mbl-1 and mec-8 affect sad-1 splicing by direct interaction with sad-1 introns.
(A–B) Mutation of mbl-1 consensus sequence in sad-1 splicing reporter results in aberrant splicing in ALM neurons that phenocopies an mbl-1 mutant. (C–E) Mutation of either mec-8 binding motif, or both simultaneously, likewise results in aberrant sad-1 splicing in ALM neurons. ALM splicing phenotypes fully penetrant (n = 25 animals) Scale bar represents 10 µm.
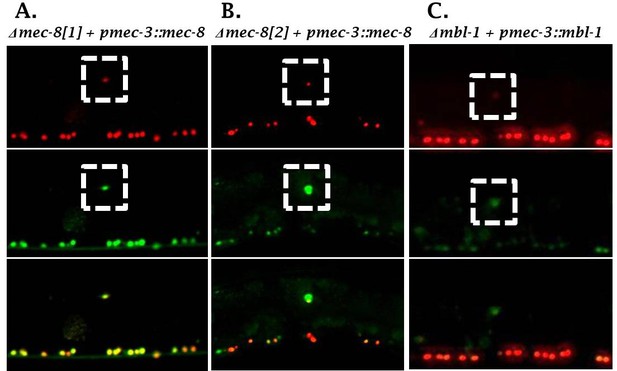
RBP overexpression does not rescue sad-1 splicing defects of cognate cis-element mutant reporters (failure to rescue in n = 20 animals for each condition).
(A) Δmec-8[1] site and (B) Δmec-8[2] site not rescued by pmec-3::mec-8 over-expression. (C) Δmbl-1 site not rescued by pmec-3::mbl-1.
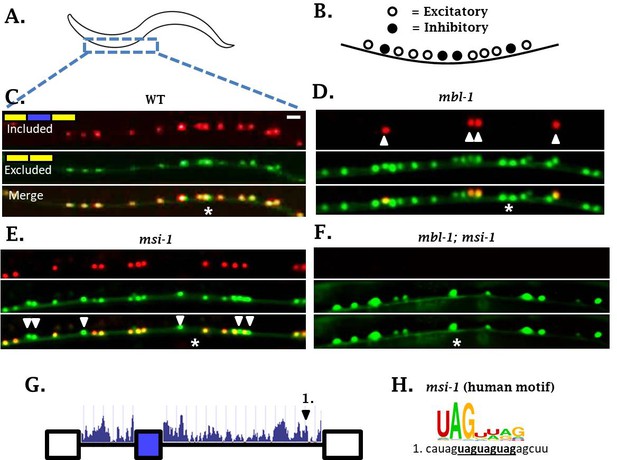
sad-1 splicing in motor neurons of the ventral nerve cord is controlled by mbl-1 and msi-1 RBPs.
(A–C) In wild-type worms, sad-1 is partially included in both excitatory and inhibitory motor neurons. (D) In mbl-1 mutants, exon inclusion is lost in excitatory motor neurons, but remains in inhibitory motor neurons (arrowheads). (E) msi-1 mutants lose exon inclusion in inhibitory motor neurons (arrowheads) but not in excitatory motor neurons. (F) mbl-1; msi-1 double mutants lose exon inclusion in all motor neurons in the ventral nerve cord. Splicing phenotypes in ventral nerve cord invariant (n = 15 animals) (G) Conservation scores (determined as in Figure 3A) in the introns surrounding sad-1 exon 15. Number one indicates consensus binding motifs for msi-1. (H) cis-elements matching consensus binding motifs for msi-1. Asterisk indicates anterior-posterior position of ALM neuron as anatomical reference. Splicing phenotypes fully penetrant (n = 50 animals). Scale bar represents 10 µm.
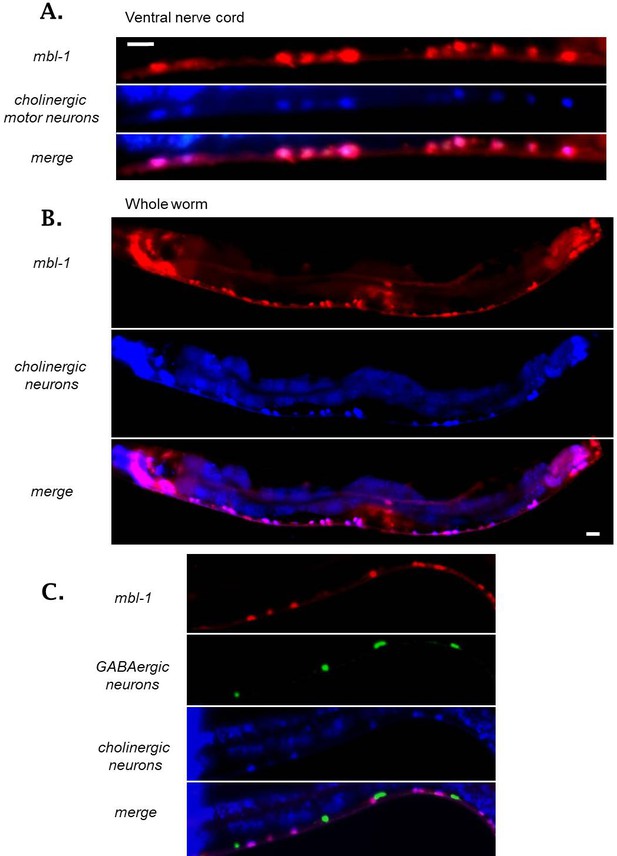
MBL-1, visualized by a translational RFP fusion is expressed specifically in the excitatory cholinergic neurons of the ventral nerve cord, visualized by an unc-17::BFP promoter reporter.
(A) High-magnification micrograph of portion of the ventral nerve cord, (B) Micrograph of whole worm to demonstrate expression of MBL-1 in Cholinergic motor neurons throughout the extent of the nerve cord. (C) As in A, GABAergic neurons labeled with GFP, cholinergic neurons with BFP, demonstrating MBL-1 expression in cholinergic motor neurons but not in GABAergic motor neurons.
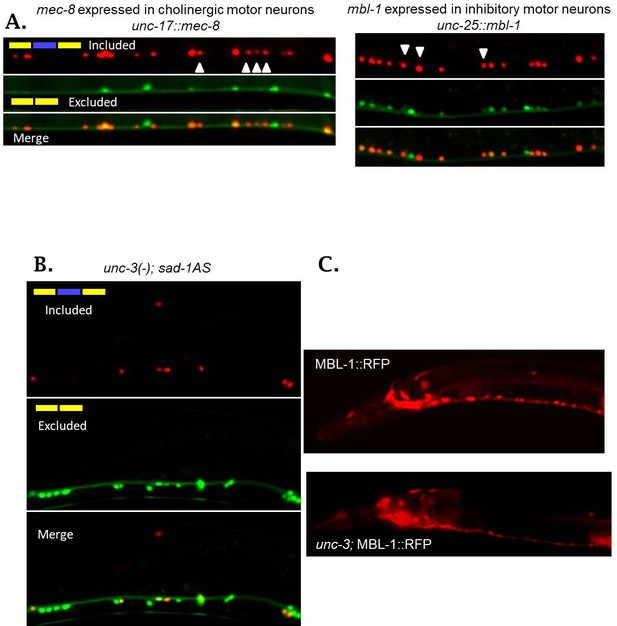
sad-1 splicing is controlled by distinct RBPs and TFs in ventral nerve cord motor neurons.
(A) Ectopic expression of mec-8 in excitatory motor neurons (via unc-17 promoter) changes sad-1 splicing in excitatory motor neurons from partial exon inclusion to complete exon inclusion (n = 7/11 animals examined). (B) Ectopic expression of mbl-1 in inhibitory motor neurons (via unc-25 promoter) has similar effects in inhibitory motor neurons (n = 9/13 animals examined). (C) unc-3 mutants exhibit partial (12/25 worms examined) or complete (13/25 worms) loss of sad-1 exon inclusion in excitatory motor neurons. (D) unc-3 mutants exhibit partially-penetrant loss of mbl-1 expression in excitatory motor neurons (25/65 animals exhibit complete loss, 28/65 partial (example displayed), 12/65 no detectable change).
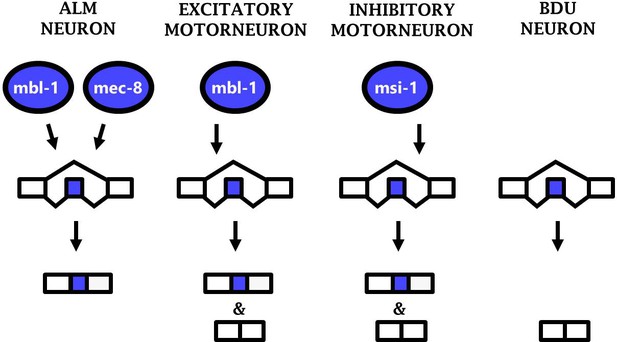
Phenotypic convergence at the level of splicing regulation.
Different RBPs act in different neuron types to carry out the common function of mediating sad-1 exon inclusion.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain | unc-86(csb9) | This study | JAC401 | Norris Lab. SMU. Dallas, TX. |
| Strain | mec-3(csb10) | This study | JAC402 | Norris Lab. SMU. Dallas, TX. |
| Strain | alr-1(csb11) | This study | JAC403 | Norris Lab. SMU. Dallas, TX. |
| Strain | unc-86(e1416) | CGC, University of Minnesota | CB1416 | |
| Strain | mec-3(e1338) | CGC, University of Minnesota | CB1338 | |
| Strain | alr-1(oy42) | CGC, University of Minnesota | PY1598 | |
| Strain | mec-8(e398) | CGC, University of Minnesota | CB398 | |
| Strain | mec-8(csb22) | This studdy | JAC626 | Norris Lab. SMU. Dallas, TX. |
| Strain | mbl-1(csb31) | This study | JAC635 | Norris Lab. SMU. Dallas, TX. |
| Strain | mbl-1(wy560) | CGC, University of Minnesota | JAC002 | |
| Strain | msi-1(csb24) | This study | JAC628 | Norris Lab. SMU. Dallas, TX. |
| Strain | mec-8(csb22); mbl-1(wy560) | This study | ADN342 | Norris Lab. SMU. Dallas, TX. |
| Strain | mec-8(csb22); mbl-1(csb31) | This study | JAC670 | Norris Lab. SMU. Dallas, TX. |
| Strain | msi-1(csb24); mbl-1(csb31) | This study | ADN257 | Norris Lab. SMU. Dallas, TX. |
| Strain | pmec-3::mec-8 | This study | ADN431 | Norris Lab. SMU. Dallas, TX. |
| Strain | pmec-3::mbl-1 | This study | ADN514 | Norris Lab. SMU. Dallas, TX. |
| Strain | punc-25::mbl-1 | This study | ADN515 | Norris Lab. SMU. Dallas, TX. |
| Strain | punc-17::mec-8 | This study | ADN505 | Norris Lab. SMU. Dallas, TX. |
| Strain | Δmbl-1 cis-element sad-1 splicing reporter | This study | ADN319 | Norris Lab. SMU. Dallas, TX. |
| Strain | Δmec-8[1] cis-element sad-1 splicing reporter | This study | ADN364 | Norris Lab. SMU. Dallas, TX. |
| Strain | Δmec-8[2] cis-element sad-1 splicing reporter | This study | ADN377 | Norris Lab. SMU. Dallas, TX. |
| Strain | Δmec-8[both] cis-element sad-1 splicing reporter | This study | ADN333 | Norris Lab. SMU. Dallas, TX. |
| Strain | sad-1 splicing reporter | This study | JAC017 | Norris Lab. SMU. Dallas, TX. |
| Strain | MEC-8::GFP reporter fosmid | This study | JAC447 | Norris Lab. SMU. Dallas, TX. |
| Strain | MBL-1::RFP reporter fosmid | This study | JAC576 | Norris Lab. SMU. Dallas, TX. |
| Strain | pmec-8::GFP | CGC, University of Minnesota | BC11068 | |
| Strain | unc-3(e151) | CGC, University of Minnesota | CB151 |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.46726.020





