Pregnancy-associated plasma protein-aa supports hair cell survival by regulating mitochondrial function
Figures
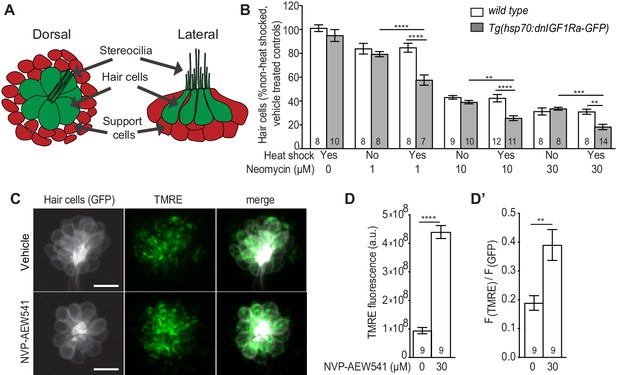
Inhibition of IGF1R activity enhances neomycin-induced hair cell death.
(A) Schematic of lateral line neuromast. (B) Mean percentage of surviving hair cells following induction of dnIGF1Ra-GFP expression. To calculate hair cell survival percentage, hair cell number 4 hr post-neomycin treatment was normalized to mean hair cell number in non-heat-shocked, vehicle-treated larvae of the same genotype. **p<0.01, ***p<0.001, ****p<0.0001 two-way ANOVA, Holm-Sidak post test. N = 7–14 larvae per group (shown at base of bars), three neuromasts perlarva from two experiments. (C) brn3c:GFP labeled hair cells loaded with TMRE in NVP-AEW541 and vehicle treated larvae. (D–D’) Mean TMRE fluorescence (D) and mean TMRE fluorescence normalized to GFP fluorescence (D’) from Z-stack summation projections of brn3c:GFP labeled hair cells. N = 9 larvae per group. Total number of neuromasts included in the analysis = 26 (vehicle treated) and 27 (NVP-AEW541). **p<0.01, ****p<0.0001. Unpaired t test, Welch-corrected.
-
Figure 1—source data 1
Hair cell survival post neomycin in wild type and Tg(hsp70:dnIGF1Ra-GFP) larvae.
- https://doi.org/10.7554/eLife.47061.003
-
Figure 1—source data 2
Mean F(TMRE) and ratio of mean F(TMRE) to mean F(GFP) in wild type and pappaap170 hair cells following treatment with NVP-AEW541.
- https://doi.org/10.7554/eLife.47061.004
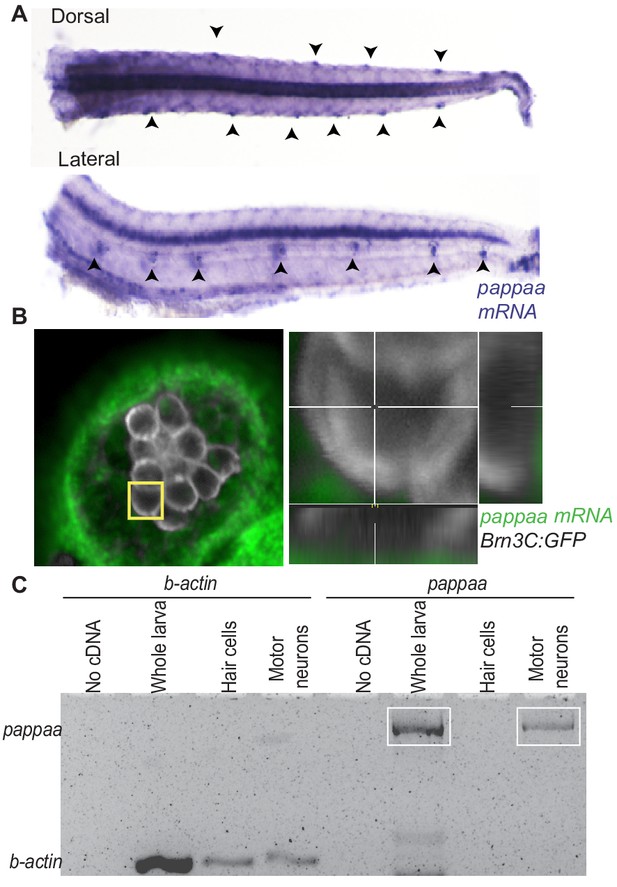
pappaa is expressed by neuromast support cells and motor neurons.
(A) Whole mount in situ hybridization shows pappaa mRNA expression at four dpf by lateral line neuromasts (arrowheads). (B) Fluorescent in situ hybridization of pappaa (green) and brn3c:GFP labeled hair cells (white) shows pappaa mRNA expression by the support cells that surround hair cells. (C) RT-PCR of fluorescently sorted brn3c:GFP labeled hair cells and mnx1:GFP labeled motor neurons shows pappaa expression by motor neurons, but not hair cells. RT-PCR products represent pappaa cDNA fragment.
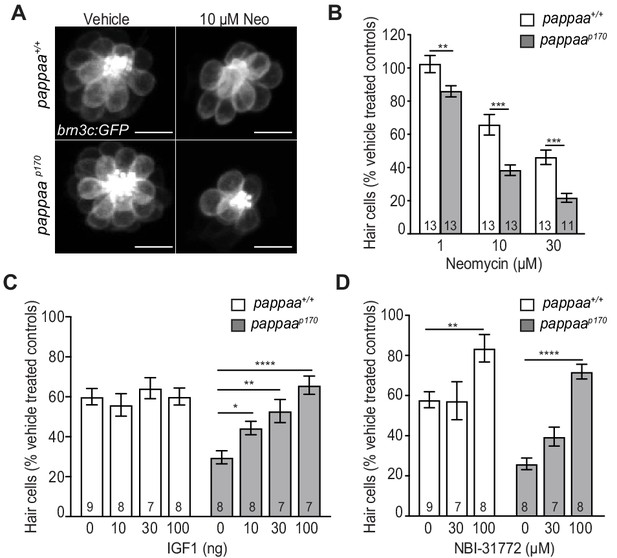
Hair cell survival is reduced in zebrafish pappaap170 larvae.
(A) Representative images of brn3c:GFP labeled hair cells from vehicle or 10 µM neomycin treated larvae. Scale = 10 µm. (B) Mean percentage of surviving hair cells. To calculate hair cell survival percentage, hair cell number 4 hr post-neomycin treatment was normalized to mean hair cell number in vehicle treated larvae of the same genotype. **p<0.01, ***p<0.001, two-way ANOVA, Holm-Sidak post test. N = 11–15 larvae per group (shown at base of bars). Total number of neuromasts included in the analysis = 45 (wild type; vehicle-treated), 39 (wild type; 1 µM neomycin), 39 (wild type; 10 µM neomycin), 39 (wild type; 30 µM neomycin), 45 (pappaap170; vehicle-treated), 39 (pappaap170; 1 µM neomycin), 39 (pappaap170; 10 µM neomycin), and 33 (pappaap170; 30 µM neomycin) from two experiments. (C) Mean percentage of surviving hair cells following co-treatment with IGF1 and 10 µM neomycin. To calculate hair cell survival percentage, hair cell counts after treatment were normalized to hair cell number in vehicle treated larvae of same genotype. *<0.05, **p<0.01 ****p<0.0001. Two-way ANOVA, Holm-Sidak post test. N = 7–11 larvae per group (shown at base of bars). Total number of neuromasts included in the analysis = 33 (wild type; vehicle-treated), 27 (wild type; 10 µM neomycin), 24 (wild type; 10 µM neomycin +10 ng IGF1), 21 (wild type; 10 µM neomycin +30 ng IGF1), 24 (wild type; 10 µM neomycin +100 ng IGF1), 33 (pappaap170; vehicle-treated), 24 (pappaap170; 10 µM neomycin), 24 (pappaap170; 10 µM neomycin +10 ng IGF1), 21 (pappaap170; 10 µM neomycin +30 ng IGF1), and 21 (pappaap170; 10 µM neomycin +100 ng IGF1) (D) Mean percentage of surviving hair cells following co-treatment with NBI-31772 and 10 µM neomycin. To calculate hair cell survival percentage, hair cell counts after treatment were normalized to hair cell number in vehicle treated larvae of same genotype. **p<0.01 ****p<0.0001. Two-way ANOVA, Holm-Sidak post test. N = 7–9 larvae per group (shown at base of bars). Total number of neuromasts included in the analysis = 15 (wild type; vehicle-treated), 27 (wild type; 10 µM neomycin), 21 (wild type; 10 µM neomycin +30 µM NBI-31772), 24 (wild type; 10 µM neomycin +100 µM NBI-31772), 15 (pappaap170; vehicle-treated), 24 (pappaap170; 10 µM neomycin), 21 (pappaap170; 10 µM neomycin +30 µM NBI-31772), and 24 (pappaap170; 10 µM neomycin +100 µM NBI-31772).
-
Figure 3—source data 1
Hair cell survival post neomycin in wild type and pappaap170 larvae.
- https://doi.org/10.7554/eLife.47061.009
-
Figure 3—source data 2
Hair cell survival post co-treatment of IGF1 and neomycin in wild type and pappaap170 larvae.
- https://doi.org/10.7554/eLife.47061.010
-
Figure 3—source data 3
Hair cell survival post co-treatment of NBI-31772 and neomycin in wild type and pappaap170 larvae.
- https://doi.org/10.7554/eLife.47061.011
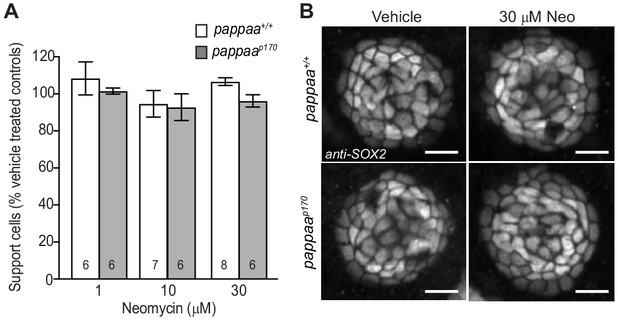
Support cells are not affected by neomycin treatment.
(A) Mean percentage of surviving neuromast support cells at 4 hr post-neomycin treatment. To calculate support cell survival percentage, support cell number 4 hr post-neomycin treatment was normalized to mean support cell number in vehicle treated larvae in the same genotype. Two-way ANOVA, Holm-Sidak post test revealed no significant difference among groups. N = 6–9 larvae per group (shown at base of bars). Total number of neuromasts included in the analysis = 27 (wild type; vehicle-treated), 18 (wild type; 1 µM neomycin), 21 (wild type; 10 µM neomycin), 24 (wild type; 30 µM neomycin), 27 (pappaap170; vehicle-treated), 18 (pappaap170; 1 µM neomycin), 18 (pappaap170; 10 µM neomycin), and 18 (pappaap170; 30 µM neomycin). Error bars = SEM. (B) Representative confocal images of anti-SOX2 labeled support cells that were control or 30 µM neomycin treated. Scale = 10 µm.
-
Figure 3—figure supplement 1—source data 1
Support cell survival post neomycin in wild type and pappaap170larvae.
- https://doi.org/10.7554/eLife.47061.008
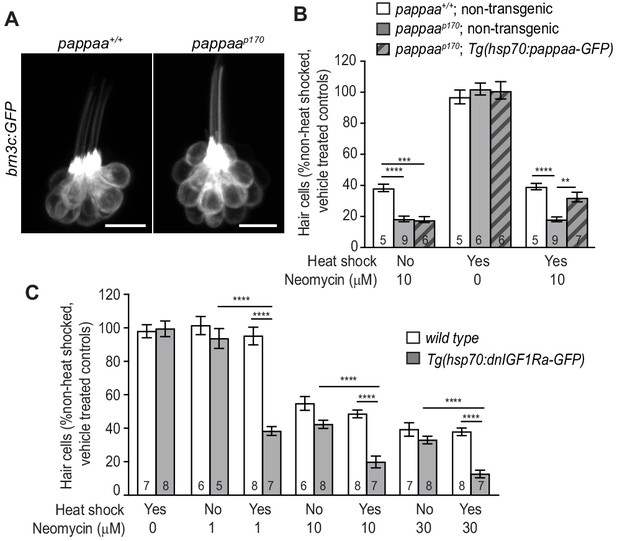
Post-developmental regulation of IGF1 signaling by Pappaa is required for hair cell survival.
(A) Lateral view of brn3c:GFP labeled hair cells in 5dpf wild type and pappaap170 larvae. Scale = 10 µm. (B) Mean percentage of surviving hair cells following post-developmental induction of Pappaa expression in pappaap170 larvae. To calculate hair cell survival percentage, hair cell number 4 hr post-neomycin treatment was normalized to mean hair cell number in non-heat-shocked, vehicle-treated larvae of the same genotype. **p<0.01, ***p<0.001, ****p<0.0001, 2-way ANOVA, Holm-Sidak post test. N = 5–9 larvae per group (shown at base of bars), three neuromasts perlarva. (C) Mean percentage of surviving hair cells following post-developmental induction of dnIGF1Ra-GFP expression. To calculate hair cell survival percentage, hair cell number 4 hr post-neomycin treatment was normalized to mean hair cell number in non-heat-shocked, vehicle-treated larvae of the same genotype. ****p<0.0001, two-way ANOVA, Holm-Sidak post test. N = number of larvae per group (shown at base of bars), three neuromasts per larva.
-
Figure 4—source data 1
Hair cell survival post neomycin in wild type, pappaap170, and Tg(hsp70:pappaa-GFP); pappaap170 larvae.
- https://doi.org/10.7554/eLife.47061.013
-
Figure 4—source data 2
Hair cell survival post neomycin in wild type and Tg(hsp70:dnIGF1Ra-GFP) larvae.
- https://doi.org/10.7554/eLife.47061.014
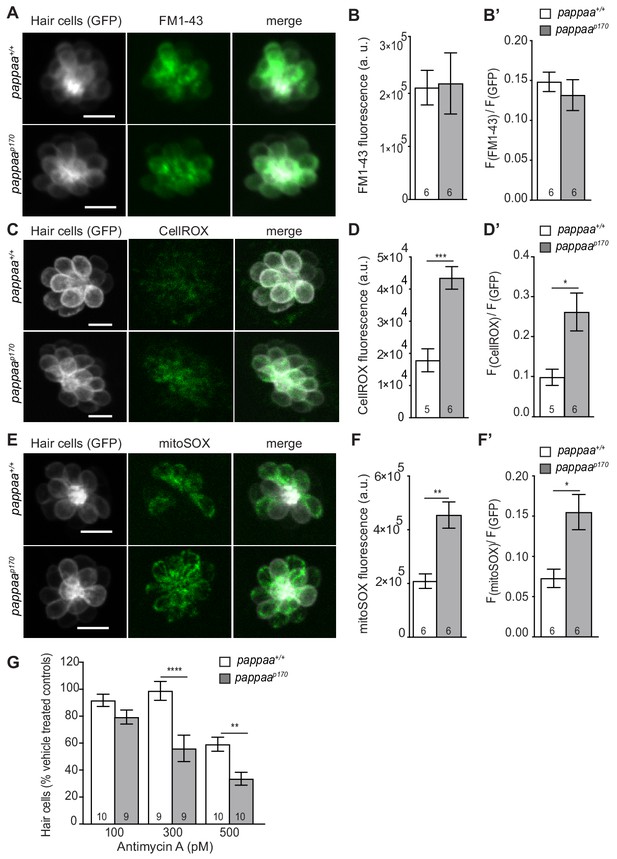
Pappaa regulates mitochondrial ROS generation.
(A, C, E) Still images of live brn3c:GFP hair cells loaded with the amphypathic styryl dye FM1-43 (A) or cytoplasmic or mitochondrial ROS indicators (C: CellROX, E: mitoSOX). Scale = 10 µm. (B, D, F) Mean dye fluorescence (D) and mean dye fluorescence normalized to GFP fluorescence (B’, D’, F’) from Z-stack summation projections of brn3c:GFP labeled hair cells. N = 5–6 larvae per group (shown at base of bars). Total number of neuromasts included in the analysis = 18 (wild type; FM1-43), 18 (pappaap170; FM1-43), 21 (wild type; CellROX), 21 (pappaap170; CellROX), 18 (wild type; mitoSOX), and 22 (pappaap170; mitoSOX). *p<0.05, **<p < 0.01, ***p<0.001. Unpaired t test, Welch-corrected. (G) Mean percentage of surviving hair cells post Antimycin A treatment. To calculate hair cell survival percentage, hair cell counts after treatment were normalized to hair cell number in vehicle treated larvae of same genotype. **p<0.01 ****p<0.0001. Two-way ANOVA, Holm-Sidak post test. N = 9–10 larvae per group (shown at base of bars). Total number of neuromasts included in the analysis = 30 (wild type; vehicle-treated), 30 (wild type; 100pM antimycin a), 27 (wild type; 300pM antimycin a), 30 (wild type; 500pM antimycin a), 27 (pappaap170; vehicle-treated), 27 (pappaap170; 100pM antimycin a), 27 (pappaap170; 300pM antimycin a), and 30 (pappaap170; 500pM antimycin a).
-
Figure 5—source data 1
Mean F(FM1-43) and ratio of mean F(FM1-43) to F(GFP) in wild type and pappaap170 hair cells.
- https://doi.org/10.7554/eLife.47061.019
-
Figure 5—source data 2
Mean F(CellROX) and ratio of mean F(CellROX) to F(GFP) in wild type and pappaap170 hair cells.
- https://doi.org/10.7554/eLife.47061.020
-
Figure 5—source data 3
Mean F(mitoSOX) and ratio of mean F(mitoSOX) to F(GFP) in wild type and pappaap170 hair cells.
- https://doi.org/10.7554/eLife.47061.021
-
Figure 5—source data 4
Hair cell survival post Antimycin A in wild type and pappaap170 larvae.
- https://doi.org/10.7554/eLife.47061.022
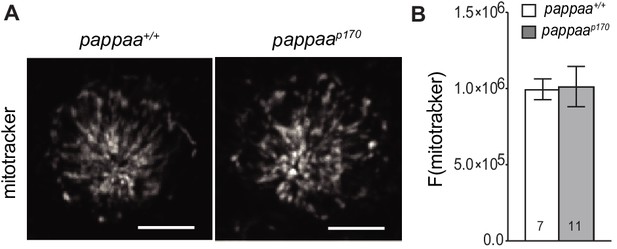
Mitochondrial mass is equivalent in wild type and pappaap170 hair cells.
(A) Still images from live wild type and pappaap170 hair cells loaded with the mitochondrial mass marker, Mitotracker. Scale = 10 µm. (B) Mean mitotracker fluorescence; quantified from Z-stack summation projection. Unpaired t test with Welch correction revealed no significant difference among groups. N = 7–11 larvae per group, (shown at base of bars). Total number of neuromasts included in the analysis = 21 (wild type) and 33 (pappaap170). Error bars = SEM.
-
Figure 5—figure supplement 1—source data 1
Mean F(mitotracker) in wild type and pappaap170 hair cells.
- https://doi.org/10.7554/eLife.47061.017
-
Figure 5—figure supplement 1—source data 2
Mean F(CellROX) and ratio of mean F(CellROX) to mean F(GFP) in wild type and pappaa mutant hair cells.
- https://doi.org/10.7554/eLife.47061.018
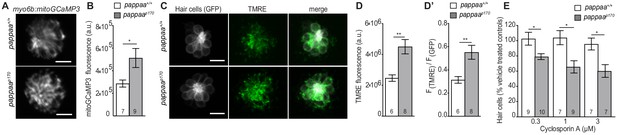
Mitochondrial Ca2+ levels and transmembrane potential are disrupted in pappaap170 hair cells.
(A) Still images from live myo6b:mitoGCaMP3 labeled hair cells. Scale = 10 µm. (B) Mean mitoGCaMP fluorescence; quantified from Z-stack summation projection. *p<0.05. Unpaired t test, Welch-corrected. N = 7–9 larvae per group (shown at base of bars). Total number of neuromasts included in the analysis = 19 (wild type) and 26 (pappaap170). (C) Still images from live brn3c:GFP labeled hair cells loaded with TMRE. Scale = 10 µm. (D) Mean TMRE fluorescence (D) and mean TMRE fluorescence normalized to GFP fluorescence (D’) from Z-stack summation projections of brn3c:GFP labeled hair cells.. N = 6–8 larvae per group (shown at base of bars). Total number of neuromasts included in the analysis = 17 (wild type) and 23 (pappaap170).. **p<0.01. Unpaired t test, Welch-corrected. (E) Mean percentage of surviving hair cells post Cyclosporin A treatment. To calculate hair cell survival percentage, hair cell counts post-treatment were normalized to hair cell numbers in vehicle treated larvae of same genotype. N = 7–10 larvae per group (shown at base of bars). Total number of neuromasts included in the analysis = 45 (wild type; vehicle-treated), 27 (wild type; 0.1 µM CsA), 21 (wild type; 1 µM CsA), 21 (wild type; 3 µM CsA), 42 (pappaap170; vehicle-treated), 30 (pappaap170; 0.1 µM CsA), 27 (pappaap170; 1 µM CsA), and 21 (pappaap170; 3 µM CsA) from two experiments. *p<0.05. Two-way ANOVA, Holm-Sidak post test.
-
Figure 6—source data 1
Mean F(mitoGCaMP) in wild type and pappaap170 hair cells.
- https://doi.org/10.7554/eLife.47061.024
-
Figure 6—source data 2
Mean F(TMRE) and ratio of mean F(TMRE) to F(GFP) in wild type and pappaap170 hair cells.
- https://doi.org/10.7554/eLife.47061.025
-
Figure 6—source data 3
Hair cell survival post Cyclosporin A in wild type and pappaap170 larvae.
- https://doi.org/10.7554/eLife.47061.026
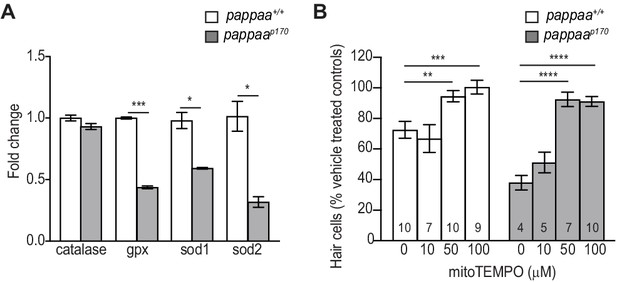
Pappaa regulates expression of mitochondrial antioxidants.
(A) Mean fold change in antioxidants transcript levels in hair cells at five dpf. cDNA was from FACsorted hair cells that were collected from 200 Tg(brn3c:GFP) dissected tails. N = 2–4 technical replicates/gene. *p<0.05, ***p<0.001. Multiple t tests, Holm-Sidak post test. Error bars = SEM. (B) Mean percentage of surviving pappaap170 hair cells following co-treatment with mitoTEMPO and 10 µM neomycin. To calculate hair cell survival percentage, hair cell counts 4 hr post-neomycin treatment were normalized to hair cell counts in vehicle treated pappaap170 larvae. **p<0.01, ***p<0.001, ****p<0.0001. Two-way ANOVA, Holm-Sidak post test. N = 4–10 (shown at base of bars). Total number of neuromasts included in the analysis = 30 (wild type; vehicle-treated), 30 (wild type; 10 µM neomycin), 21 (wild type; 10 µM neomycin +10 µM mitoTEMPO), 30 (wild type; 10 µM neomycin +50 µM mitoTEMPO), 27 (wild type; 10 µM neomycin +100 µM mitoTEMPO), 30 (pappaap170; vehicle-treated), 12 (pappaap170; 10 µM neomycin), 15 (pappaap170; 10 µM neomycin +10 µM mitoTEMPO), 21 (pappaap170; 10 µM neomycin +50 µM mitoTEMPO), and 30 (pappaap170; 100 µM neomycin +10 µM mitoTEMPO). Error bars = SEM.
-
Figure 7—source data 1
Quantification of antioxidant transcript expression in wild type and pappaap170 hair cells.
- https://doi.org/10.7554/eLife.47061.028
-
Figure 7—source data 2
Hair cell survival post co-treatment of mitoTEMPO and neomycin in pappaap170 larvae.
- https://doi.org/10.7554/eLife.47061.029
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Danio rerio) | pappaap170 | Wolman et al., 2015 | RRID:ZFIN_ZDB-GENO-190322-4 | single nucleotide nonsense mutationC > T at position 964 in Exon 3 |
| Strain, strain background (Danio rerio) | Tg(myo6b:mitoGCaMP3) | Esterberg et al., 2014 | RRID:ZFIN_ZDB-GENO-141008-1 | |
| Strain, strain background (Danio rerio) | Tg(brn3c:GFP) | Xiao et al., 2005 | RRID:ZFIN_ZDB-ALT-050728-2 | |
| Strain, strain background (Danio rerio) | Tg(mnx1:GFP) | Rastegar et al., 2008 | RRID:ZFIN_ZDB-GENO-140605-2 | |
| Strain, strain background (Danio rerio) | Tg(hsp70:dnIGF1Ra-GFP) | Kamei et al., 2011 | RRID:ZFIN_ZDB-GENO-110614-4 | |
| Strain, strain background (Danio rerio) | Tg(hsp70:pappaa-GFP) | this paper | Materials and methods subsection maintenance of zebrafish | |
| Strain, strain background (Danio rerio) | TLF | Zebrafish International Resource Center (ZIRC) | RRID:ZFIN_ZDB-GENO-990623-2 | |
| Antibody | anti-myosinVI (rabbit polyclonal) | Proteus biosciences | RRID:AB_10013626 | 1:200 |
| Antibody | anti-SOX2 (rabbit polyclonal) | Abcam | RRID:AB_2341193 | 1:200 |
| Antibody | anti-GFP (rabbit polyclonal) | ThermoFisher Scientific | RRID:AB_221569 | 1:500 |
| Antibody | Alexa 488, secondary (rabbit polyclonal) | ThermoFisher Scientific | RRID:AB_2576217 | 1:500 |
| Peptide, recombinant protein | IGF-1 | Cell sciences | Catalog number: DU100 | |
| Other | FM1-43 | ThermoFisher Scientific | Catalog number: T3136 PubChem CID:6508724 | 3 uM for 30 s |
| Other | CellROX deep red | ThermoFisher Scientific | Catalog number: C10422 | 10 uM for 60 min |
| Other | mitoSOX red | ThermoFisher Scientific | Catalog number: M36008 | 1 uM for 30 min |
| Other | mitotracker greenFM | ThermoFisher Scientific | Catalog number: M7514 PubChem CID:70691021 | 100 nM for 5 min |
| Other | TMRE | ThermoFisher Scientific | Catalog number: T669 PubChem CID:2762682 | 25 nM for 20 min |
| Other | 0.25% trypsin-EDTA | Sigma-Aldrich | Catalog number: T3924 | |
| Other | TRIzol | Invitrogen | Catalog number: 15596026 | |
| Other | Sso fast Eva Green Supermix | BioRad | Catalog number: 1725200 | |
| Chemical compound, drug | Neomycin | Sigma-Aldrich | Catalog number: N1142 PubChem CID:24897464 | |
| Chemical compound, drug | NBI-31772 | Fisher Scientific | Catalog number: 519210 | |
| Chemical compound, drug | antimycin-a | Sigma-Aldrich | Catalog number: A8674 PubChem CID: 24891355 | |
| Chemical compound, drug | CsA | Abcam | Catalog number: ab120114 PubChem ID: 5284373 | |
| Chemical compound, drug | mitoTEMPO | Sigma-Aldrich | Catalog number: SML0737 PubChem CID: 134828258 | |
| Chemical compound, drug | NVP-AEW541 | Selleck | Catalog number: S1034 PubChem CID: 11476171 | |
| Commercial assay or kit | SuperScript II Reverse Transcriptase | Invitrogen | Catalog number: 18064014 | |
| Software, algorithm | Fluoview (FV10-ASW 4.2) | Olympus | RRID:SCR_014215 | |
| Software, algorithm | ImageJ | PMID: 22743772 | RRID:SCR_003070 | |
| Software, algorithm | GraphPad Prism | GraphPad | RRID:SCR_002798 |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.47061.030





