Telomere dysfunction cooperates with epigenetic alterations to impair murine embryonic stem cell fate commitment
Figures
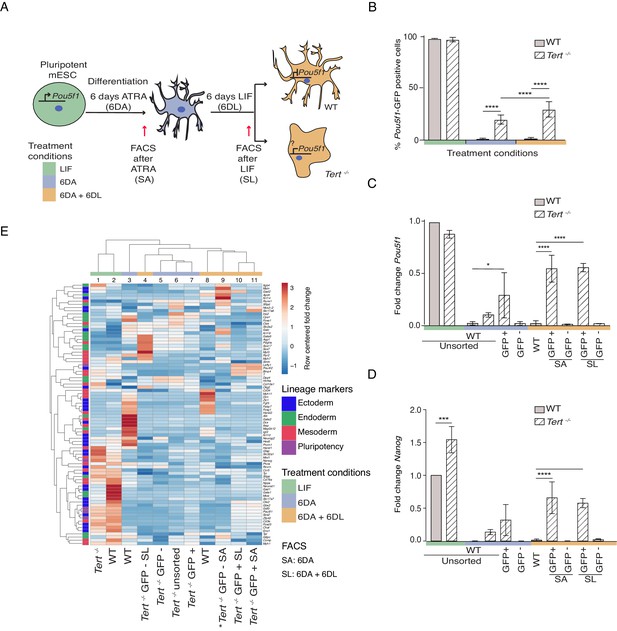
Murine ESCs with short telomeres fail to complete differentiation commitment and to suppress pluripotency gene expression.
(A) Schematic representation of the experimental design. Wild-type (WT) and Tert-/- mESCs transduced with Pou5f1 -GFP were maintained in a pluripotent state in the presence of leukemia inhibitory factor (LIF). Murine ESCs were next induced to differentiate by all-trans retinoic acid (ATRA) treatment for 6 days, followed by re-seeding in LIF-containing media for an additional 6 days to assess their ability to remain stably differentiated (GFP-). The GFP status of mESCs was monitored by flow cytometry (FACS) and sorted samples were collected at each experimental step: (i) LIF (pluripotent), (ii) 6 days ATRA (6 DA) and (iii) 6 days ATRA followed by 6 days LIF (6 DA + 6 DL). SA: mESCs sorted after 6 DA. SL: mESCs sorted after 6 DA + 6 DL (B) FACS analysis of Pou5f1-GFP reporter expression in mESCs at each experimental step. Data are represented as mean ± SD. Data represent n = 3 biological replicates: for n = 1 and n = 2 biological replicates, there were three technical replicates; for the n = 3 biological replicate, there were two technical replicates, for a total of 8 samples. Statistical analysis was performed using two-way ANOVA, * (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001). Color bars underneath the x-axis represent the same treatment groups as indicated in (A). (C, D) Assessment of relative fold change in Pou5f1 and Nanog expression (WT pluripotent set as 1) by RT-qPCR, normalized to five housekeeping genes: Actb, B2m, Gapdh, Gusb and Hsp90ab1. Data are represented as mean ± SD (n = 3). Statistical analysis was performed using ordinary one-way ANOVA, * (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001). Color bars underneath the x-axis represent the same treatment groups as indicated in (A). (E) Heatmap representation of relative fold change in gene expression of 77 lineage-specific markers (endoderm, mesoderm and ectoderm) and five pluripotency genes from the Qiagen Mouse Cell Lineage Identification qPCR Array (n = 3 except for one sample marked * where n = 2, see legend D). Relative fold change in gene expression was calculated using WT pluripotent mESC (2 days in LIF) as control (fold change = 1) and normalized to five housekeeping genes: Actb, B2m, Gapdh, Gusb and Hsp90ab1. The color scale represents the row centered fold change values (blue = lower, white = intermediate, red = higher). Euclidean distance clustering and complete linkage was applied to visualize similarity between samples. See Materials and methods for details.
-
Figure 1—source data 1
Murine ESCs with short telomeres fail to complete differentiation commitment and to suppress pluripotency gene expression.
Source data of (i) the GFP percentage values represented in the histogram in Figure 1B and Figure 1—figure supplement 1F–G; (ii) Raw Ct values and information relative to the Qiagen qPCR array relative to Figure 1C,D,E and Figure 1—figure supplement 1C,D; (iii) Raw telomere signal intensities presented in Figure A-figure supplement 1A-B.
- https://cdn.elifesciences.org/articles/47333/elife-47333-fig1-data1-v2.xlsx
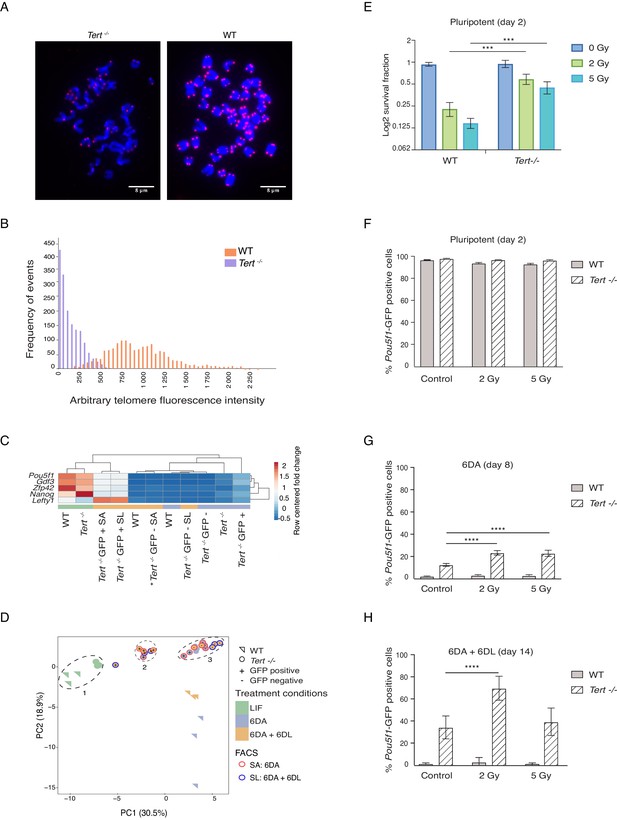
Telomere status, gene expression, and gamma-irradiation response in WT and Tert-/- mESCs.
(A) Relative telomere length measurement by Quantitative Fluorescence In-situ Hybridization (Q-FISH) analysis of WT and Tert-/- mESCs at passage 50. Representative photo of metaphase preparations. DNA/chromosomes are stained with DAPI (blue) and telomeres can be visualized in red (Cy3). The scale is indicated with a microbar within each image. See Materials and methods for further detail regarding sample preparation and image acquisition. (B) Histogram of relative telomere length measured as frequency of events vs. arbitrary telomere fluorescence intensity, extracted from data as in (A). The data represent at least 1587 telomeres analysed per sample. (C) Heatmap representation of pluripotency gene expression assessed by RT-qPCR (n = 3 biological replicates, except for one sample marked with an asterisk * Tert -/- GFP SA, where n = 2 biological replicates). Relative fold change in gene expression was calculated using WT pluripotent mESCs set as control (fold change = 1) and normalized to five housekeeping genes: Actb, B2m, Gapdh, Gusb and Hsp90ab1. The color scale represents the row centered fold change values (blue = lower, white = intermediate, red = higher). Euclidean distance clustering and complete linkage was applied to visualize similarity between samples. Refer to A for sample labelling. (D) Principal component analysis (PCA) of RT-qPCR data, see also heatmap in Figure 1E. Percentage values indicate the principal components with the greatest variance, in the first and second dimensions, respectively. Circles indicate groups referred to in the text (1, 2, 3). Refer to Figure 1A for sample labeling. See also Figure 1—source data 1. (E) Survival fraction (logarithmic scale) of mESCs grown in LIF-containing media, measured by propidium iodide staining, 2 days after gamma-irradiation at the indicated doses. Data are represented as mean + SD (n = 3 biological replicates). Statistical analysis was performed using two-way ANOVA, (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001). (F, G, H) Differentiation analysis of irradiated mESCs. GFP+ status was measured by flow cytometry (F) after 2 days in LIF-containing media (pluripotent) (G) followed by 6 days in ATRA (6 DA, day 8) (H) after 6 days of LIF re-addition (6 DA + 6 DL, day 14). Data are represented as mean + SD (n = 3 biological replicates). Statistical analysis was performed using two-way ANOVA, (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001).
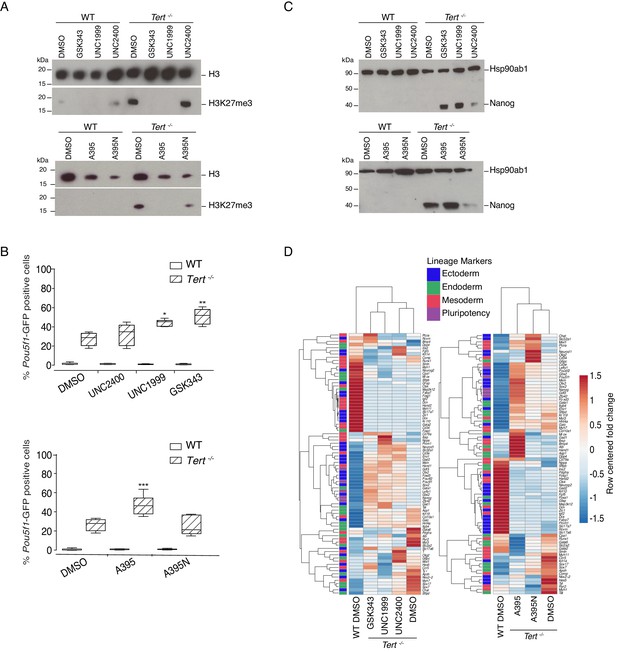
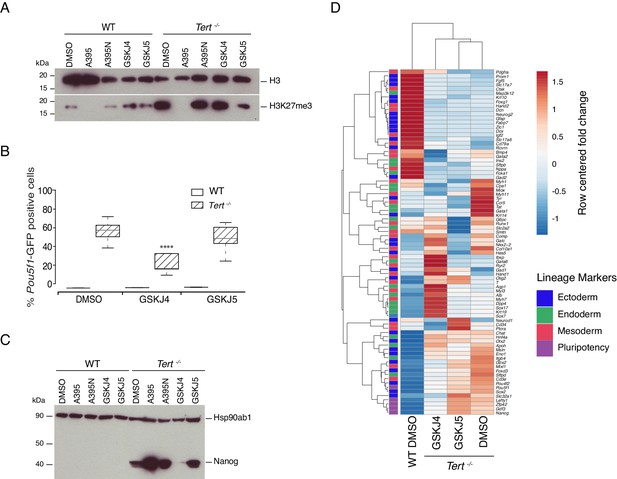
Murine ESCs with short telomeres exhibit altered H3K27me3 levels and incomplete differentiation that is exacerbated by PRC2 inhibition.
(A) Western blot analysis of H3K27me3 levels in WT and Tert-/- mESCs, in the presence of active PRC2 inhibitors (GSK343, UNC1999 and A395), their corresponding controls (UNC2400 and A395N), or the vehicle-only control DMSO. H3 was used as a loading control. Representative blot from n = 3 biological replicates (see Figure 2—figure supplement 1D for quantification of additional representative blots). (B) FACS analysis of Pou5f1-GFP reporter expression in mESCs. Data are represented as mean ± SD. Data represent n = 3 biological replicates: for n = 1 and n = 2 biological replicates, there were three technical replicates; for the n = 3 biological replicate, there were two technical replicates, for a total of 8 samples. Statistical analysis was performed using two-way ANOVA, * (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001). Graphs were generated using the online tool BoxPlotR (http://shiny.chemgrid.org/boxplotr/). (C) Western blot analysis of Nanog protein expression. Hsp90ab1 was used as a loading control. Representative blot from n = 3 biological replicates (see Figure 2—figure supplement 1F). (D) Heatmap representation of relative fold change in gene expression of 77 lineage-specific markers (endoderm, mesoderm and ectoderm) and five pluripotency genes from the Qiagen Mouse Cell Lineage Identification qPCR Array (n = 3). Gene expression analysis was performed on 6 DA + 6 DL mESCs treated with active PRC2 inhibitors (GSK343, UNC1999 and A395), the inactive control compounds (UNC2400 and A395N) or DMSO. Relative fold change in gene expression was calculated using Tert-/- DMSO as control (fold change = 1) and normalized to five housekeeping genes: Actb, B2m, Gapdh, Gusb and Hsp90ab1. The color scale represents the row centered fold change values (blue = lower, white = intermediate, red = higher). Euclidean distance clustering and complete linkage was applied to visualize similarities between samples.
-
Figure 2—source data 1
Murine ESCs with short telomeres exhibit altered H3K27me3 levels and incomplete differentiation that is exacerbated by PRC2 inhibition.
Source data of (i) the GFP percentage values represented in the histogram in Figure 2B and Figure 2—figure supplement 1E; (ii) Fold change values presented in Figure 2—figure supplement 1B; (iii) Raw data from the western blot quantification presented in Figure 2—figure supplement 1D,F; (iv) Raw Ct values and information relative to the Qiagen qPCR array relative to Figure 2D and Figure 2—figure supplement 1G,H.
- https://cdn.elifesciences.org/articles/47333/elife-47333-fig2-data1-v2.xlsx
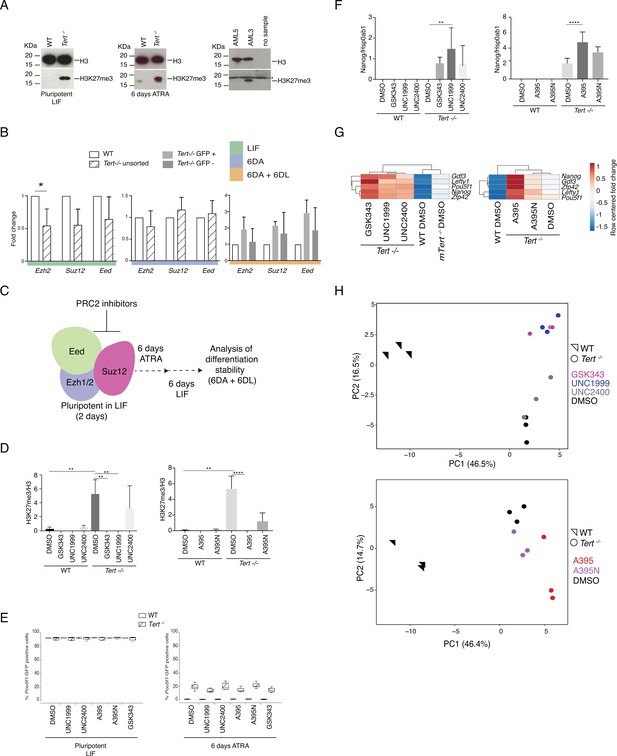
The impact of PRC2 inhibitors on H3K27me3, gene expression, and differentiation of WT and Tert-/- mESCs.
(A) Western blot of H3K27me3 in pluripotent (maintained in LIF) or ATRA treated (6 DA) WT and Tert-/- mESCs. H3 was probed as a loading control. Representative blot from n = 3. A western blot of the OCI-AML3 and EZH2R690H-mutant27 OCI-AML5 (showing a drastic decrease of PRC2 activity) lines was also performed as a control for the antibody (Lehnertz et al., 2017). A non-specific signal, which is also evident in the lane in which no sample was added (at right) is marked with an asterisk (*). (B) Assessment of relative fold change in Ezh2, Suz12 and Eed expression (relative to WT) by RT-qPCR, normalized to two housekeeping genes: B2m and Gapdh, Data are represented as mean + SD (n = 3 biological replicates). Statistical analysis was performed using ordinary one-way ANOVA, * (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001). See Figure 1A for color coding. (C) Schematic representation of the experimental design. PRC2 inhibitors were added to the mESC culture media prior to differentiation induction and maintained during differentiation (6 DA) and after LIF re-addition (6 DA + 6 DL). See Materials and methods for details. (D, F) Quantification of 3 biological replicates of the western blot data in Figure 2A,C using Image J. The y-axis indicates the relative level of the protein species of interest normalized to the level of the control protein, as indicated. Statistical analysis was performed using ordinary one-way ANOVA, * (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001). (E) FACS analysis of Pou5f1-GFP reporter expression in mESCs. Data are represented as mean ± SD. Data represent n = 3 biological replicates: for n = 1 and n = 2 biological replicates, there were three technical replicates; for the n = 3 biological replicate, there were two technical replicates, for a total of 8 samples. Statistical analysis was performed using two-way ANOVA, * (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001). Graphs were generated using the online tool BoxPlotR (http://shiny.chemgrid.org/boxplotr/). The left graph represents pluripotent cells after 2 days of treatment with the inhibitor. The right graph represents the cells treated with the inhibitors for 2 days in LIF followed by 6 days in ATRA (6 DA) (G) Heatmap representation of 5 pluripotency gene expression assessed by RT-qPCR (n = 3 biological replicates). Relative fold change in gene expression was calculated using Tert-/- DMSO as the control (fold change = 1) and normalized to five housekeeping genes: Actb, B2m, Gapdh, Gusb and Hsp90ab1. The color scale represents the row centered fold change values (blue = lower, white = intermediate, red = higher). Sample clustering by Euclidean distance and complete linkage was applied to visualize similarity between samples. (H) Principal component analysis (PCA) of RT-qPCR data, also shown in the heatmap in Figure 2D. WT mESCs, triangles; Tert-/- mESCs, circles. Color legend for the indicated treatment groups is shown on the right. Note that not all biological replicates treated with the Eed inhibitor A395 exhibited an identical profile (lower panel; red circles at right).
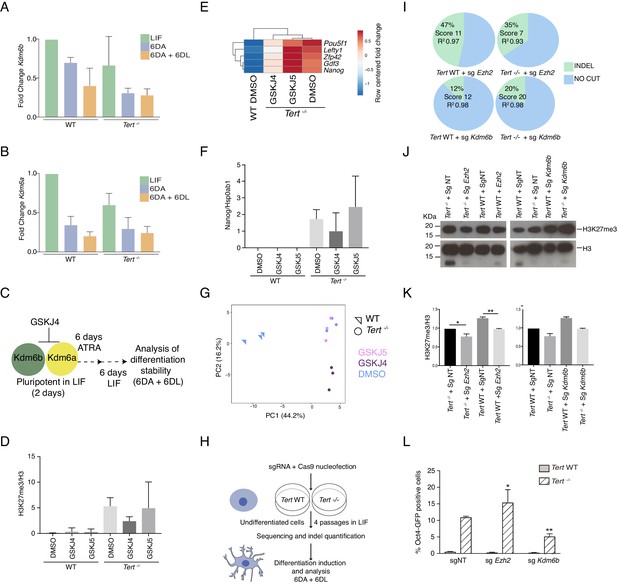
Inhibition of Kdm6a/b histone demethyltransferase activity partially rescues cell fate commitment.
(A, C) Western blot analysis of H3K27me3 or Nanog protein levels in WT and Tert-/- mESCs, in the presence of DMSO only, GSKJ4 or GSKJ5. For clarity, no image cropping was performed between lanes and the blot includes an independent replicate of the A395/A395N treatments shown in Figure 2. Hsp90ab1 and H3 are used as loading control for Nanog and H3K27me3 respectively. Representative blots from n = 3 biological replicates. Quantification of western blots is shown in Figure 3—figure supplement 1D,G. (B) FACS analysis of Pou5f1-GFP reporter expression in mESCs. Data are represented as mean ± SD (n = 3 biological replicates each with three technical replicates). Statistical analysis was performed using two-way ANOVA, * (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001). (D) Heatmap representation of relative fold change in gene expression of 77 lineage-specific markers (endoderm, mesoderm and ectoderm) and five pluripotency genes from the Qiagen Mouse Cell Lineage Identification qPCR Array (n = 3 biological replicates). Gene expression analysis was performed on 6 DA + 6 DL mESCs treated with the Kdm6a/b inhibitor (GSKJ4), the inactive molecule (GSKJ5) or DMSO control. Relative fold change in gene expression was calculated using Tert-/- DMSO as control (fold change = 1) and normalized to five housekeeping genes: Actb, B2m, Gapdh, Gusb and Hsp90ab1. The color scale represents the row centered fold change values (blue = lower, white = intermediate, red = higher). Euclidean distance clustering and complete linkage was applied to visualize similarity between samples.
-
Figure 3—source data 1
Inhibition of Kdm6a/b demethylase activity partially rescues cell fate commitment.
Source data of (i) the GFP percentage values represented in the histogram in Figure 3C and Figure 3—figure supplement 1L; (ii) Raw Ct values and information relative to the Qiagen qPCR array relative to Figure 3E,F and Figure 3—figure supplement 1B,D,E; (iii) Raw data from the western blot quantification presented in Figure 3—figure supplement 1A,C,J,K; (iv) Fold change values presented in Figure 3—figure supplement 1A,B; (v) Indel frequency as showed in Figure 3—figure supplement 1I–L.
- https://cdn.elifesciences.org/articles/47333/elife-47333-fig3-data1-v2.xlsx
The impact of PRC2 or Kdm6a/b inhibition on H3K27me3, gene expression, and differentiation in WT and Tert-/- mESCs.
(A, B) Assessment of relative fold change in Kdm6b and Kdm6a expression (relative to WT 2 DL) by RT-qPCR, normalized to two housekeeping genes: B2m and Gapdh, Data are represented as mean + SD (n = 3 biological replicates). Statistical analysis was performed using ordinary one-way ANOVA, * (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001). See Figure 1A for the color coding. (C) Schematic representation of the experimental design. The Kdm6a/b histone demethyltransferase inhibitor GSKJ4, or its inactive analogue GSKJ5, was added to the cell culture media prior to differentiation induction and maintained during differentiation (6 DA) and after LIF re-addition (6 DA + 6 DL). See Materials and methods for details. (D, F) Quantification of western blot data in Figure 3A,C using image J (n = 3). The y-axis indicates the relative level of the protein species of interest normalized to the level of the control protein, as indicated. Statistical analysis was performed using ordinary one-way ANOVA, * (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001). (E) Heatmap representation of pluripotency gene expression assessed by RT-qPCR (n = 3 biological replicates). Relative fold change in gene expression was calculated using Tert -/- DMSO as the control (fold change = 1) and normalized to five housekeeping genes: Actb, B2m, Gapdh, Gusb and Hsp90ab1. The color scale represents the row centered fold change values (blue = lower, white = intermediate, red = higher). Euclidean distance clustering and complete linkage was applied to visualize similarity between samples. (G) Principal component analysis (PCA) of RT-qPCR data, see also heatmap data in Figure D. WT ESCs, triangles; Tert-/- ESCs, circles. Color legend for treatment group as at right. All samples were analysed after 6D ATRA + 6D LIF, as in (C). (H) Schematic representation of the experimental design. The synthetic guide RNA (sgRNA) and Cas9 were electroporated into cells and propagated for four passages. At this time, Indel percentages within each population were quantified with the Synthego Ice Algorithm, and the differentiation analysis was commenced as described in Figure 1. (I) Indel percentages. Knockout scores represent the proportion of indels that indicate a frameshift or are greater than 21 bp in length. R2 values indicate the confidence level of indel determination based on the sequencing quality. For more details, visit the Synthego website (https://ice.synthego.com/#/). See also Figure 3—source data 1. Three biological replicates were analysed for each sgRNA. (J) Western blot analysis of H3K27me3 levels in WT and Tert-/- mESCs after electroporation of single gRNA (Non-targeting, NT or as indicated). H3 was used as a loading control. Representative blot from n = 2 biological replicates. (K) Quantification of western blot data in Figure 3—figure supplement 1 (J), using image J (n = 2). The y-axis indicates the relative level of the protein species of interest normalized to the level of the control protein, as indicated. Statistical analysis was performed using ordinary one-way ANOVA, * (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001). (L) FACS analysis of Pou5f1-GFP reporter expression in mESCs. Data are represented as mean ± SD (n = 3 biological replicates) sgNT: non-targeting guide RNA. Statistical analysis was performed using two-way ANOVA, * (p<0.0332), ** (p<0.0021), *** (p<0.0002), **** (p<0.0001).
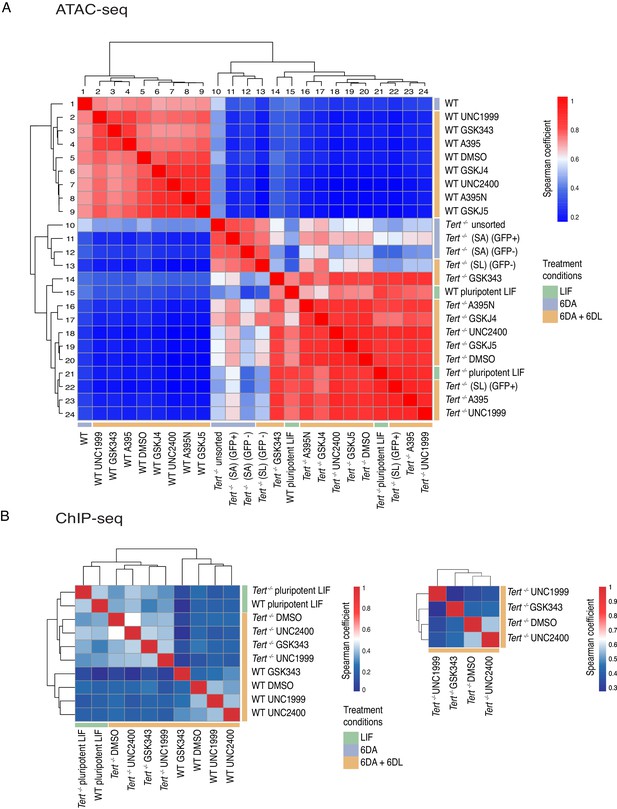
Telomere dysfunction affects the chromatin accessibility landscape during mESC differentiation.
(A) Spearman correlation analysis of called peaks following ATAC-seq analysis of Tert-/- and WT mESCs assessed throughout differentiation, with or without the addition of PRC2 or Kdm6a/b inhibitors, as indicated (a minimum of 2 biological replicates were analysed per sample; see Materials and methods and Source data files for Figure 4). (B) Spearman correlation analysis of called peaks following H3K27me3 ChiP-seq analysis of the indicated samples. At left, the data are shown for one replicate, and at right, for a subset of samples in which two biological replicates were analysed.
-
Figure 4—source data 1
Supplemental information for high throughput sequencing metadata related to ATAC-seq.
- https://cdn.elifesciences.org/articles/47333/elife-47333-fig4-data1-v2.xls
-
Figure 4—source data 2
Supplemental Table 1 related to ATAC-seq data.
- https://cdn.elifesciences.org/articles/47333/elife-47333-fig4-data2-v2.xlsx
-
Figure 4—source data 3
Supplemental Table 1 related to ChIP-seq data.
- https://cdn.elifesciences.org/articles/47333/elife-47333-fig4-data3-v2.xlsx
-
Figure 4—source data 4
Supplemental information for high-throughput sequencing metadata related to ChIP-seq.
- https://cdn.elifesciences.org/articles/47333/elife-47333-fig4-data4-v2.xls
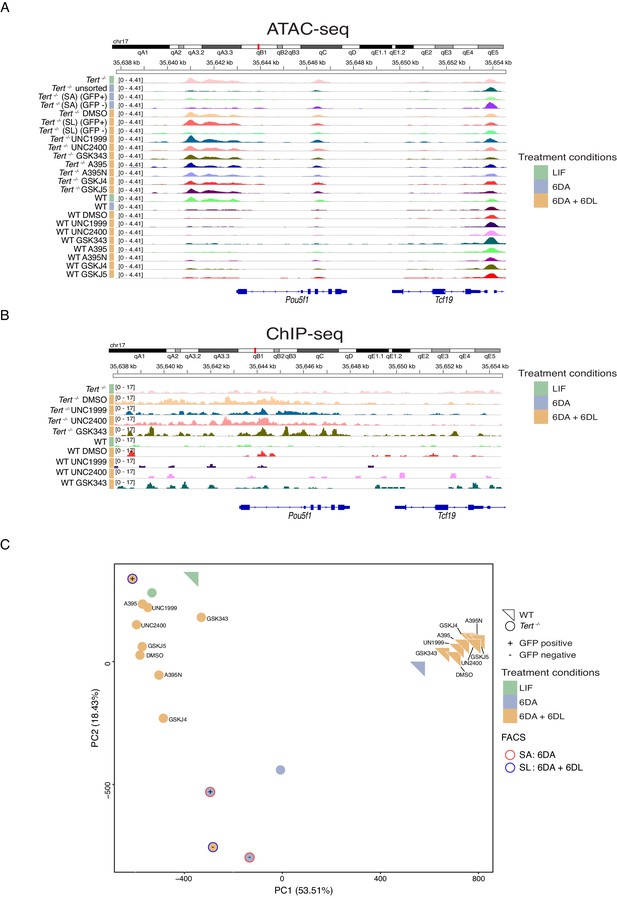
Chromatin accessibility, including at the Pou5f1 promoter, in WT and Tert-/- mESCs during differentiation.
(A, B) IGV screenshot of the genomic region surrounding the Pou5f1 promoter, derived from either ATAC-seq data (A) or H3K27me3 ChIP-seq data (B) as shown in Figure 4. One representative replicate was arbitrarily chosen for each sample. Track Height was defined to fit to full screen mode on IGV and was scaled equivalently for all samples within the same panel, using the auto scale mode. (C) Principal component analysis of called peaks following ATAC-seq analysis, of the same samples as shown in (A). WT mESCs, triangles; Tert-/- mESCs, circles. The symbol color indicates the treatment stage, as indicated in the legend at right, and in Figure 1A schematic. Circle outline colors indicate samples sorted for GFP status after 6D ATRA (SA, red) or a further 6D ATRA + 6D LIF (SL, blue). See also Figure 4—source data 2 .
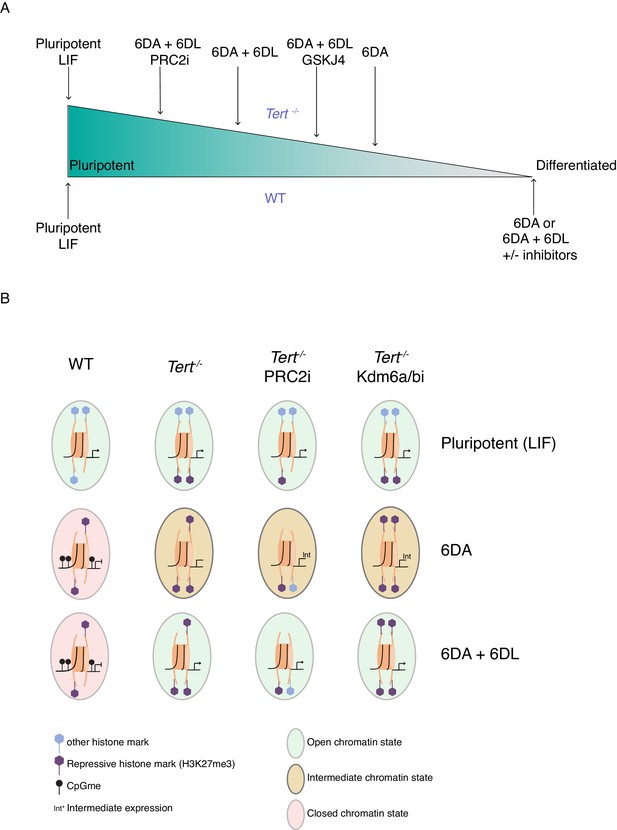
The unstable differentiation of mESC with short telomeres is exacerbated by epigenetic remodelling.
(A) Schematic of the cell state of WT and Tert-/- ESCs upon differentiation induction, based on our data. The slope is intended only to illustrate hypothetical relative differences between cell states. (B) Schematic representation of the potential epigenetic landscape(s) at pluripotency gene promoters in WT, Tert-/-, or Tert-/- mESCs treated with PRC2 or Kdm6a/b inhibitors.
Additional files
-
Supplementary file 1
Key resources table.
Supplemental information about sequence-based reagents, cells lines, antibodies, chemical compounds, software, algorithms and commercial kits used in this study.
- https://cdn.elifesciences.org/articles/47333/elife-47333-supp1-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/47333/elife-47333-transrepform-v2.docx





