Fog signaling has diverse roles in epithelial morphogenesis in insects
Figures
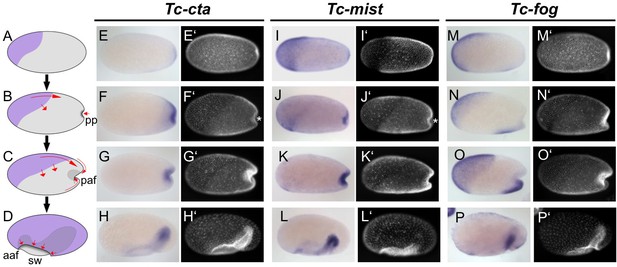
Expression of Fog signaling components during early embryogenesis.
(A–D) Schematics showing embryo condensation as described in the text. Serosa is shown in purple, germ rudiment tissue is shown in gray, arrows display tissue movements. aaf: anterior amniotic fold, paf: posterior amniotic fold, pp: primitive pit, sw: serosal window. (E–P’) Whole mount ISH and DNA staining for Tc-cta (E–H), Tc-mist (I–L) and Tc-fog (M–P). (E’–P’) nuclear (DAPI) staining of respective embryos. Anterior is left, ventral is down (where possible to discern).
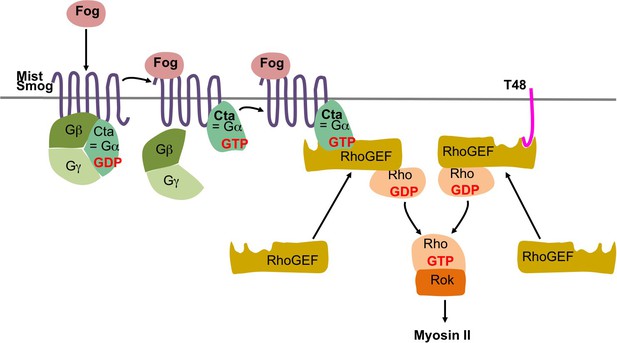
Fog and T48 pathway in Drosophila.
In Drosophila the secreted protein Fog activates the two G protein coupled receptors (GPCRs) Mist (Mesoderm-invagination signal transducer, also known as Mthl1 (Methuselah-like1)) (Manning et al., 2013) and Smog (Jha et al., 2018; Kerridge et al., 2016). The active receptors act through a heterotrimeric G protein composed of Gα12/13, Gβ13F, and Gγ1 (Kerridge et al., 2016) by converting the Gα12/13 subunit, known as Concertina (Cta) from its GDP bound inactive to its GTP bound active form. The complex then dissociates into free Gα12/13 and a Gβ13F/Gγ1 dimer, which regulate their respective intracellular effectors. In particular, active Gα12/13 recruits RhoGEF2 to the apical plasma membrane where it acts as a guanine nucleotide exchanges factor for Rho1. Rho1-GTP then activates Rok (Rho kinase). Rok phosphorylates the regulatory light chain of non-muscle myosin II promoting apical myosin contractility. RhoGEF2 can also be recruited to the apical membrane through the transmembrane protein T48 which interacts with RhoGEF2 via its PDZ-binding domain (Kölsch et al., 2007).
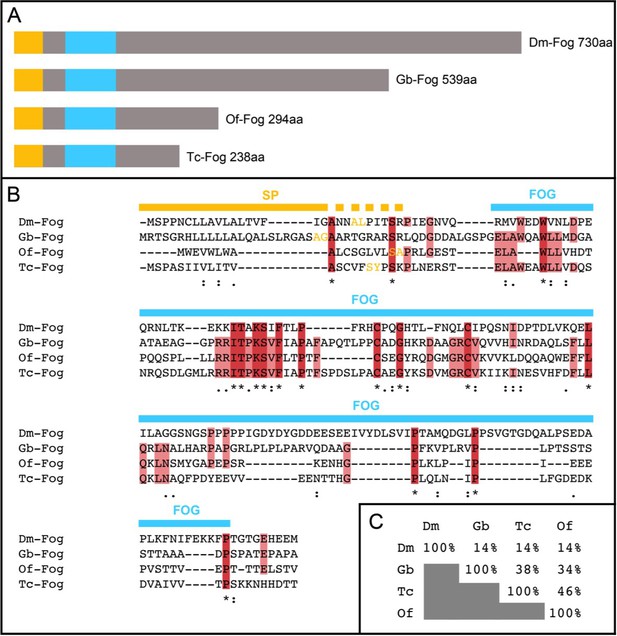
Insect Fog proteins.
(A) Schematic representation of Fog proteins from Drosophila melanogaster (Dm-Fog), Gryllus bimaculatus (Gb-Fog), Oncopeltus fasciatus (Of-Fog) and Tribolium castaneum (Tc-Fog). The proteins contain an N-terminal signal peptide (yellow) followed by a conserved Fog domain (blue) and a highly variable C-terminal domain. (B) Sequences of the N-terminal regions including the signal peptide (SP, yellow) and the Fog domain (FOG, blue). Amino acids marked in yellow indicate the predicted cleavage side of the SP (identified via SignalP 4.1 (http://www.cbs.dtu.dk/services/SignalP/)). (C) Sequence conservation within the Fog domain as indicated as percentage of identical amino acids.
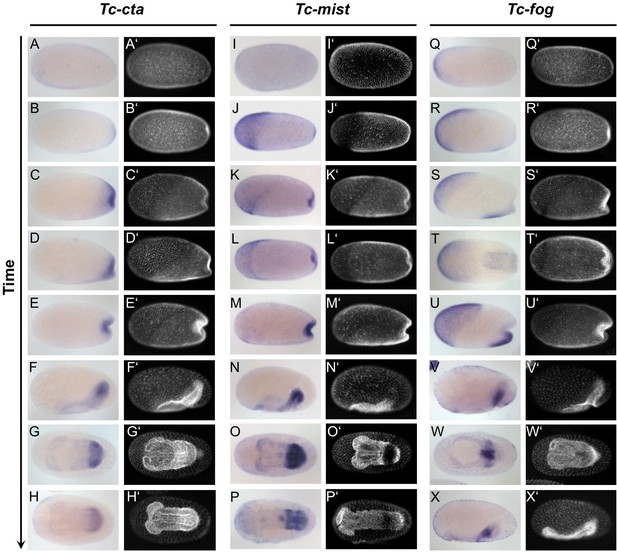
Expression of Tc-cta, Tc-mist and Tc-fog during early embryogenesis.
(A–X) Whole mount ISH for Tc-cta (A–H), Tc-mist (I–P) and Tc-fog (Q–X). (A’–X’) DAPI staining of respective embryos. Anterior is left, all panels show optical sagittal sections, except G, H, L, O, P, T and W, which show ventral surface views.
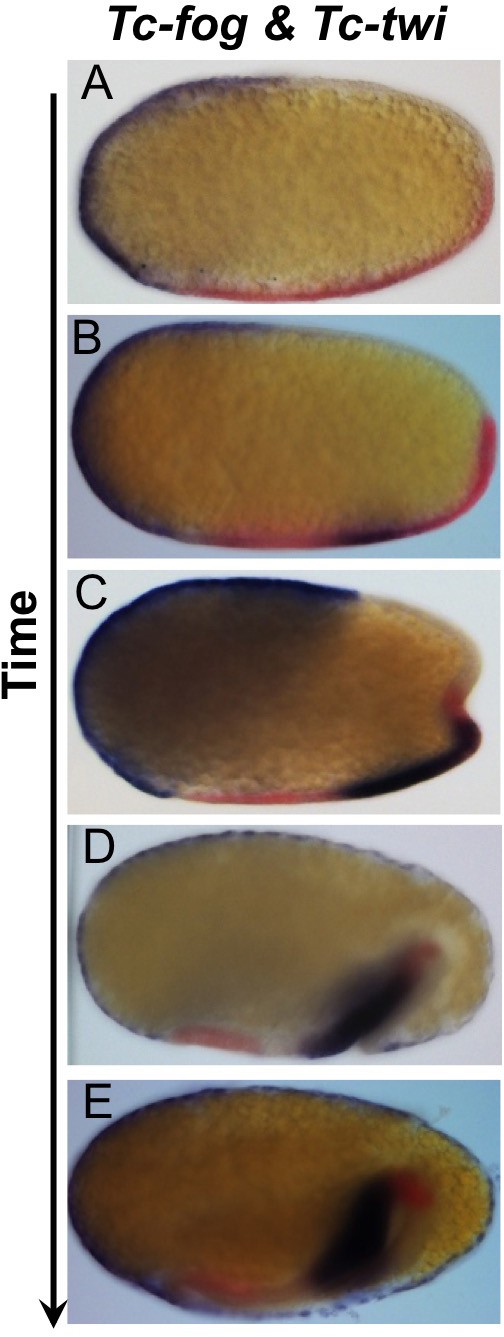
Tc-fog and Tc-twi are co-expressed only within the posterior presumptive mesoderm.
Double whole mount ISH for Tc-twi (red) and Tc-fog (blue). Anterior is left, all panels show optical sagittal sections.
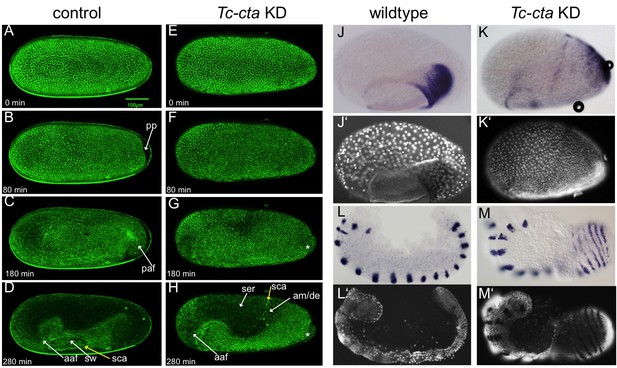
Fog signaling is required for posterior amniotic fold formation.
(A–H) Stills from live fluorescent imaging of LifeAct-eGFP transgenic embryos, ranging from late blastoderm to germband extension stages. (A–D) wildtype control. (E–H) Tc-cta KD. The asterisk marks a cluster of cells that becomes visible at the posterior pole. (J, K) Tc-pnr is expressed in a broad dorsal domain. (J’, K’) nuclear (DAPI) staining of respective embryos. (L, M) Tc-gsb expression marks forming and differentiating segments in elongating germ band embryos. (L’, M’) nuclear (DAPI) staining of respective embryos. (J, L) Wildtype. (K, M) Tc-cta KD. aaf: anterior amniotic fold, am/de: amnion dorsal ectoderm, paf: posterior amniotic fold, pp: primitive pit, sca: supracellular actin cable, sw: serosal window. Anterior is left, ventral is down.
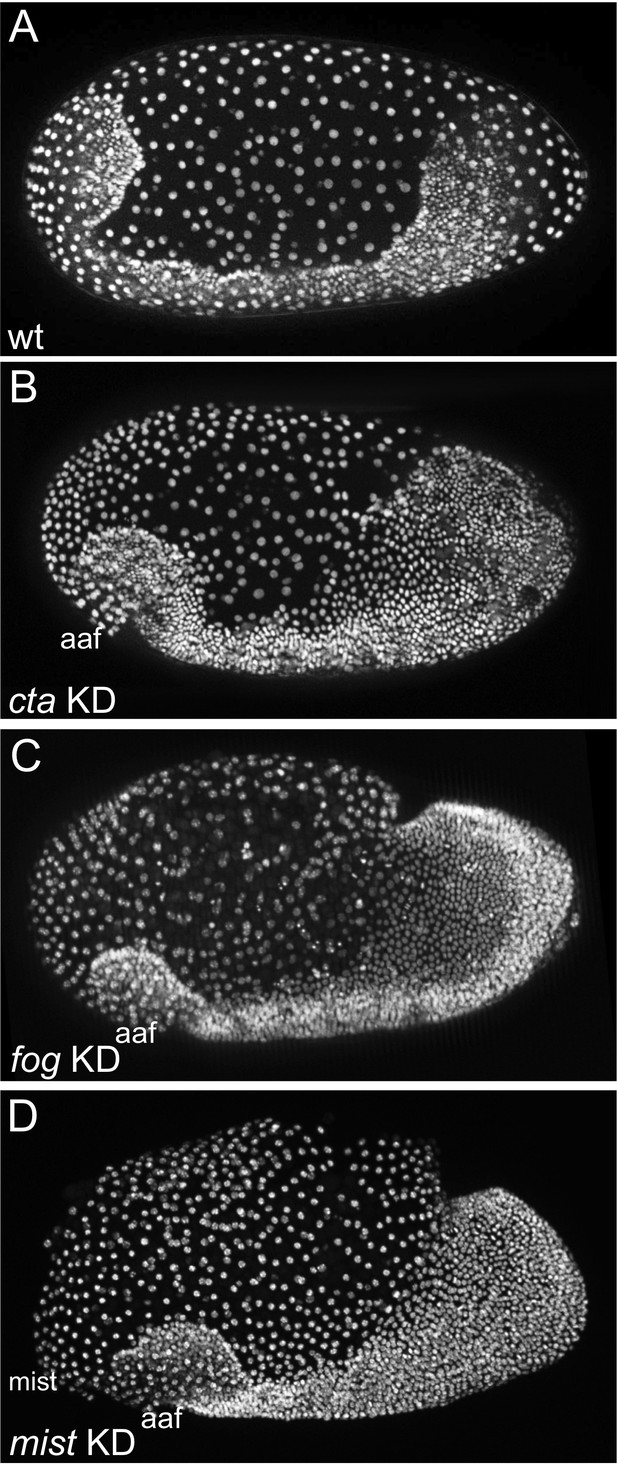
KD of Tc-fog, Tc-mist and Tc-cta by RNAi results in similar phenotypes.
(A–D) Nuclear (DAPI in A, C and D; nGFP transgene in B) staining of embryos of similar age. (A) Wildtype embryo at early elongating germ band stage. (B–D) Tc-cta, Tc-fog or Tc-mist KD embryos. aaf: anterior amniotic fold. Anterior is left, ventral is down.
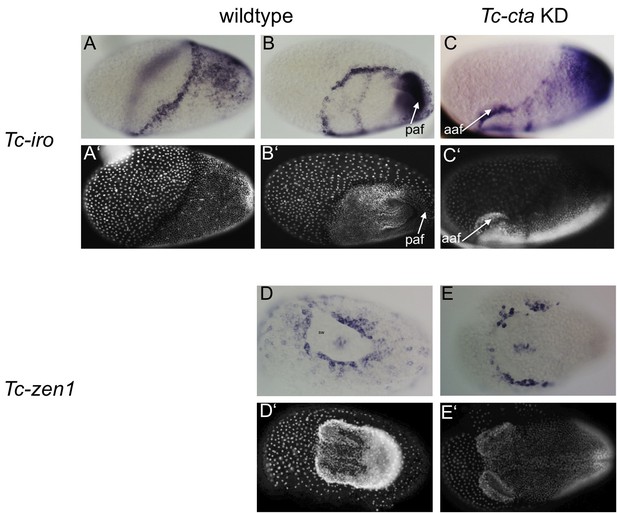
Fog signaling is required for posterior amniotic fold formation.
(A– C) Tc-iro expression (ISH). (A’– C’) DAPI staining of respective embryos. In wildtype embryos, Tc-iro is expressed at the serosa/germ rudiment border and in a dorsal germ rudiment domain. (C) Upon Tc-cta KD, the dorsal germ rudiment domain remains on the dorsal side of the egg. (D, E) Tc-zen1 expression marks the rim of the serosal window (sw). (D’, E’) DAPI staining of respective embryos. (D) Wildtype embryo forms a serosal window while (E) Tc-cta KD embryo lacks a circumferential serosal window. Anterior is to the left for all embryos. (A) is dorsolateral, (B) is ventrolateral, (C) is lateral, (D, E) are ventral views. aaf: anterior amniotic fold, paf: posterior amniotic fold.
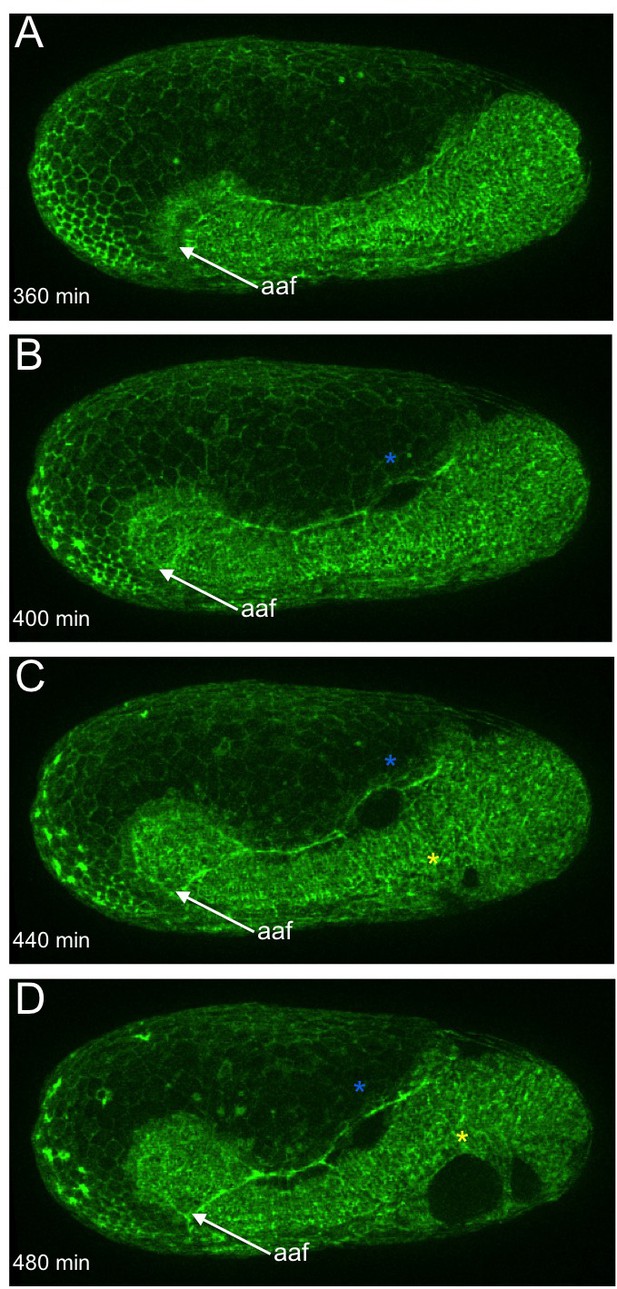
Morphogenetic defects in late Tc-cta KD embryos.
(A–D) Stills from fluorescent live imaging of a Tc-cta KD embryo carrying a LifeAct-eGFP transgene. aaf: anterior amniotic fold, blue asterisk: lateral rupture, yellow asterisk: ventral rupture. Anterior is left, ventral is down.
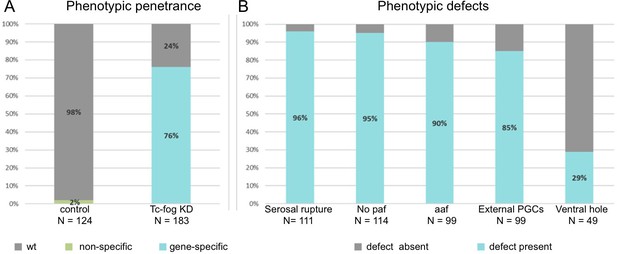
Frequencies of phenotypic defects upon Tc-fog KD.
(A) 98% of offspring of beetles injected with H2O show normal development (N = 124). 76% (N = 183) of offspring of beetles injected with Tc-fog dsRNA showed phenotypic defects. (B) The 139 embryos with phenotypic defects were sub-scored for the defects described in Figure 2, Figure 3, Figure 4 and Figure 2—figure supplement 3. Depending on the position and stage of the embryo not all defect types could be evaluated for each embryo. Therefore, the total number of scorable embryos changes for each phenotypic type. paf: posterior amniotic fold, aaf: anterior amniotic fold, PGCs, primordial germ cells.
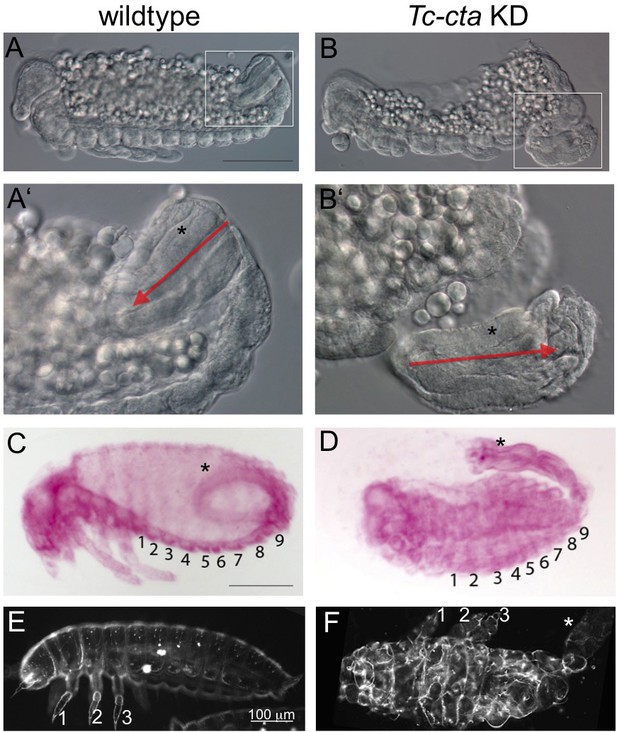
Late development of Tc-cta KD embryos.
(A, B) Optical cross sections (DIC) of fixed embryos after germband retraction. The elongation of the hindgut has started. (A’, B’) Enlarged views of the areas enframed in A, B. Red arrows indicate the polarity of the hindgut (asterisk). (C, D) Fuchsin staining of embryos after dorsal closure. Abdominal segments are numbered. (C) The hindgut (asterisk) forms a curved structure inside the posterior abdomen. (D) The hindgut has not been internalized and develops outside the embryo. This phenotype was observed in 84% of the KD embryos (N = 19). (E, F) Darkfield micrographs of larval cuticle. Legs are numbered. (F) Segments are highly distorted, but formation of structured appendages indicates normal segment polarity. The hindgut is external (white asterisk). An external hindgut was observed in 56% (N = 34) of cuticle preparation of KD embryos.
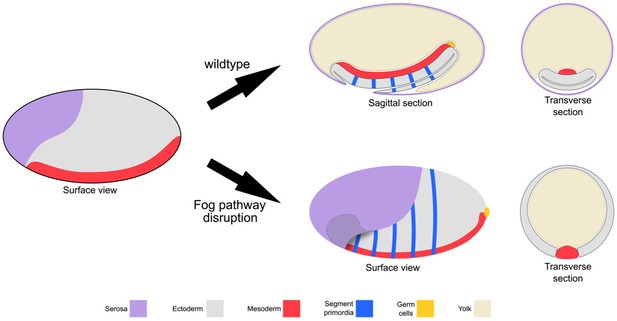
Schematic representation of the embryonic phenotype produced by Fog pathway disruption.
Schematics showing wildtype development and the effects on embryo formation of RNAi disruption of Tc-fog, Tc-mist or Tc-cta. Anterior is left, ventral is down.
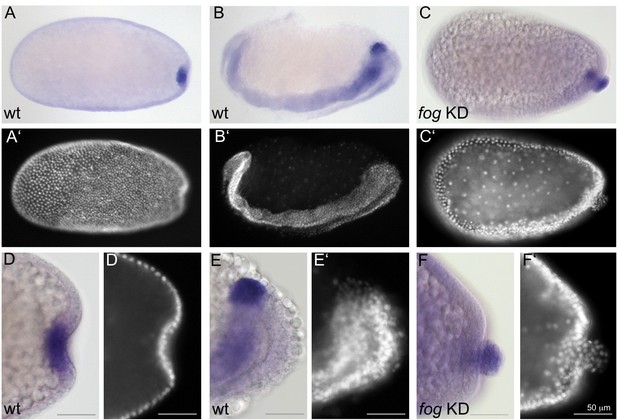
Fog signaling affects the positioning of the primordial germ cells.
Whole mount ISH for the germ cell marker Tc-tapas. (A, B, D, E) Wildtype. (C, F) Tc-fog KD. (A–C) Optical sagittal sections of whole embryos. (D–F) Optical sagittal sections of posterior regions. (A’–F’) DAPI staining of the respective embryos. (A, D) Wildtype embryo at primitive pit stage. (B, E) Wildtype embryo at early germ band extension stage. (C, F) Tc-fog KD embryos at stage corresponding to primitive pit stage in wildtype. Anterior is left, ventral is down.
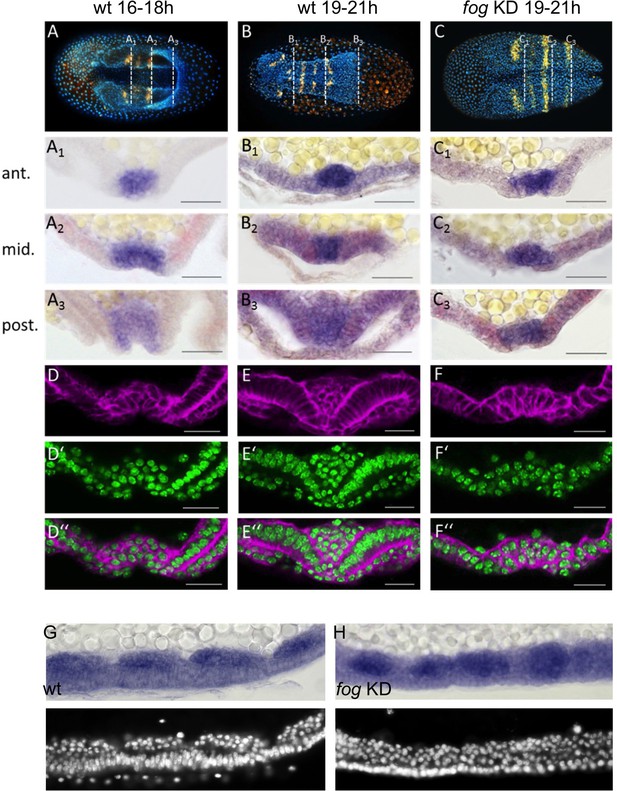
Tc-fog RNAi delays mesoderm internalization.
(A–C) Ventral views of whole mount embryos (anterior left) stained for the segmental marker Tc-gsb (yellow), nuclei (DAPI; blue). Embryos are also stained for Tc-twi expression but this is only visible in A (dark blue ventral domain). (A–B) Wildtype at horseshoe or early germband extension stage with two or four trunk Tc-gsb stripes, respectively. (C) Tc-fog KD at a stage corresponding to (B) with four trunk Tc-gsb stripes (the forth stripe is visible in lateral views). (A1–C3) transverse cryosections of the embryos shown in (A–C) with Tc-twi expression (blue) and Tc-gsb expression (red). The position of each section is indicated by a white dashed line in (A–C). (D–F’’) Transverse cryosections at posterior positions of embryos corresponding in age to those in (A–C) showing F-actin (phalloidin; magenta) and nuclei (sytox; green). (G, H) Sagittal cryosections of embryos during germband elongation (anterior left) showing Tc-twi expression (dark blue). (G’, H’) nuclear (DAPI) staining of respective embryos. Only a portion of the germband comprising four segments is shown. For corresponding sections showing the entire embryo see Figure 5—figure supplement 4. The different appearance of the mesoderm upon Tc-fog KD compared to wildtype is likely due to the delayed maturation of the mesoderm and the overall aberrant morphogenetic movements of KD embryos. For all embryos the serosa was removed during preparation. The cell sheet covering the ventral side of control embryos is the amnion (am) or amnion/dorsal ectoderm (am/de). Staging was done at 25°C.
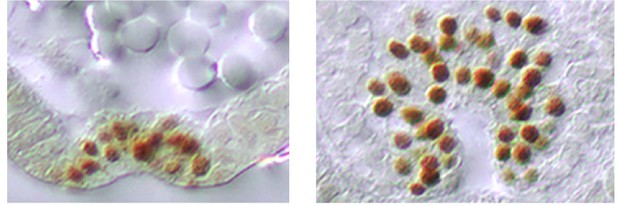
Mesoderm internalization varies along the AP axis in Tribolium.
Twi protein expression within the ventral furrow of a gastrulating Tribolium embryo. Cross section through anterior (A) and posterior (B) region of the same embryo. The sections are modified from Handel et al. (2005).
© 2005 Springer Nature. Reprinted with permission from Springer Nature Customer Service Centre GmbH: Springer Dev. Genes Evol., ‘Tribolium castaneum twist: gastrulation and mesoderm formation in a short-germ beetle’, Handel et al. (Copyright 2005). This figure is not available under CC-BY and is exempt from the CC-BY 4.0 license.
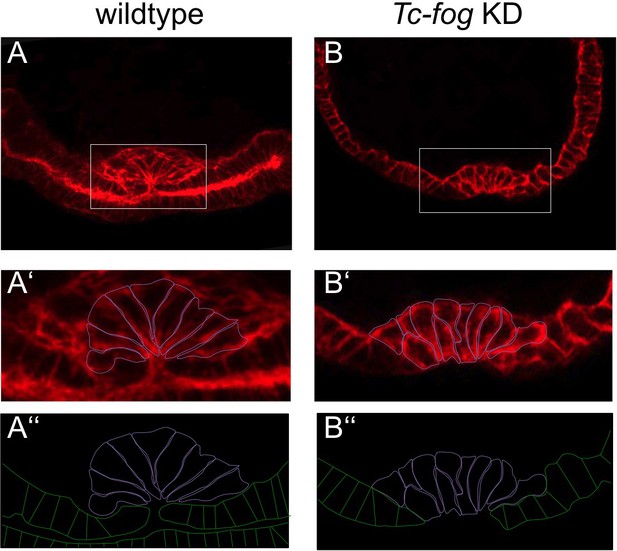
Apical constrictions during mesoderm internalization.
Transverse cryosections at posterior positions of wildtype (A) and Tc-fog KD (B) embryos of similar age stained for F-actin. (A’, B’) Magnified insert with mesoderm cells outlined in magenta. (A’’, B’’) Cell outlines of mesoderm (magenta) and ectoderm (green). (A) High number of apically constricting cells in wildtype. The mesoderm has been largely internalized. (B) Few apically constricting cells upon Tc-fog KD. The mesoderm has not been fully internalized.
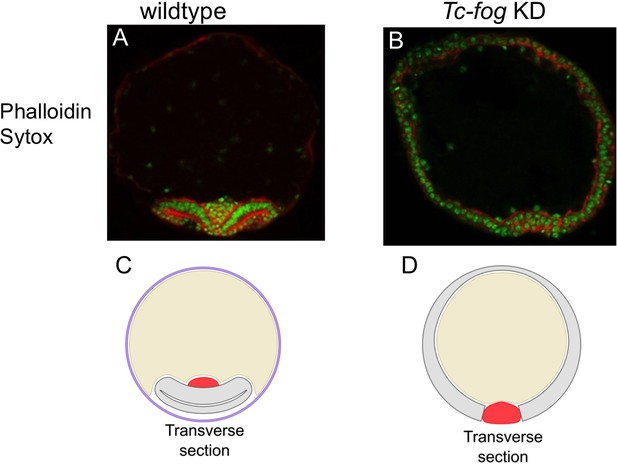
Delayed mesoderm internalization upon Tc-fog KD.
(A, B) Transverse section of staged embryos stained for F-actin (Phalloidin; red) and nuclei (Sytox; green). (A) Wildtype. The amnion/dorsal ectoderm covers the ventral side of the embryo. The mesoderm is internalized. The serosa was lost in the course of the fixation and staining procedure. (B) Tc-fog KD by pRNAi. The amnion/dorsal ectoderm remains at the dorsal side. Mesoderm internalization is delayed. (C, D) Schematic representation of transverse sections of wildtype (C) and Tc-fog KD (D) embryos. Purple: serosa; Yellow: yolk, Gray: Amnion plus embryonic ectoderm; Red: Mesoderm.
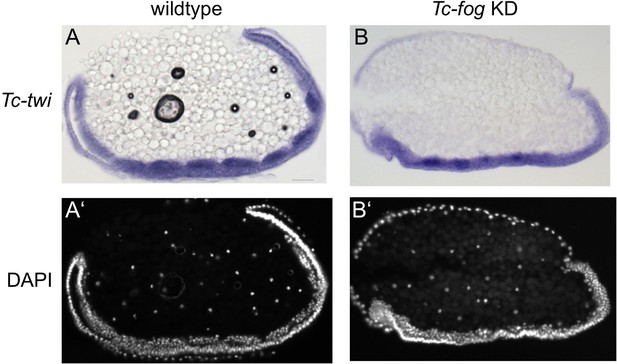
The mesoderm is internalized upon Tc-fog KD.
(A, B) Sagittal cryosections of embryos at the early elongating germ band stage showing Tc-twi expression (ISH, dark blue). (A’, B’) DAPI staining of respective embryos. (A) Wildtpype. Tc-twi is expressed in each segment in regular cell clusters. The amnion covers the embryo at the ventral side. The serosa has been removed. (B) Tc-fog KD by pRNAi. Tc-twi is also expressed in segmental, albeit less regular cell clusters. The embryo is not covered ventrally since the amnion remains at the dorsal side.
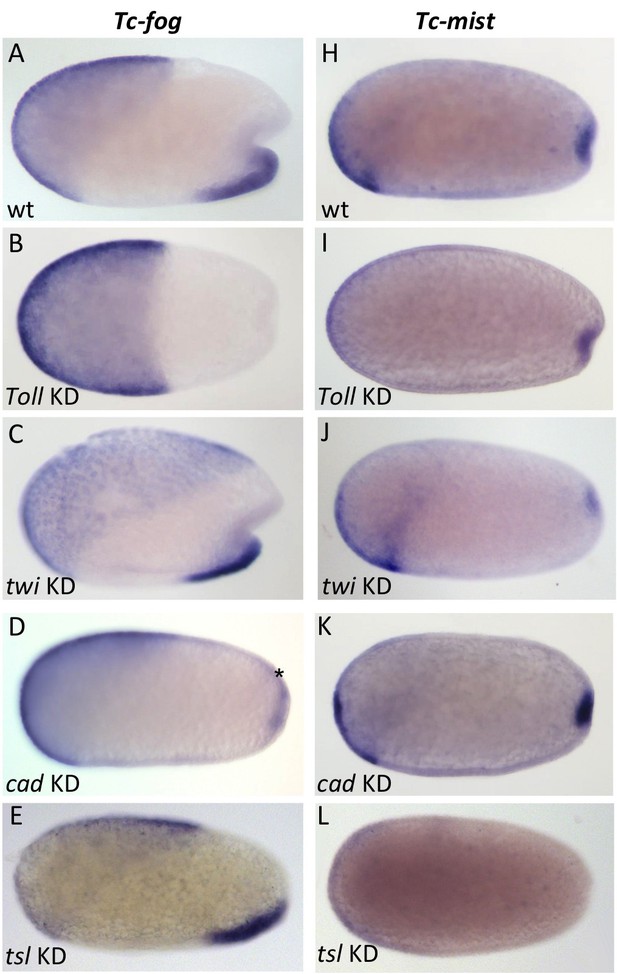
Regulation of Tc-fog and Tc-mist expression.
Whole mount ISH for Tc-fog (A–E) and Tc-mist (H–L) expression in wildtype embryos (A, H) and embryos in which DV and AP genes have been knocked down (B-E, I-L; specific KD shown in panels). All embryos are at primitive pit stage except A and C, which are at the early posterior amniotic fold stage. The asterisk in (D) indicates the appearance of weak Tc-fog expression within a posterior-dorsal domain.
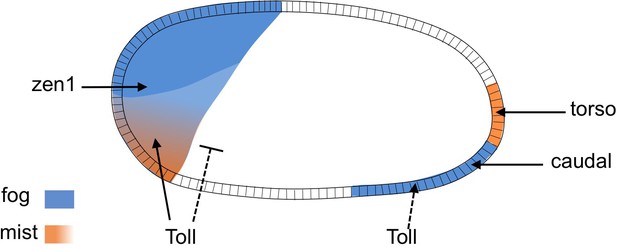
Regulation of Tc-fog and Tc-mist by DV and AP patterning genes.
Arrowheads indicate activation, the bar-headed line indicates inhibition. Dashed lines indicate that the regulation is indirect.
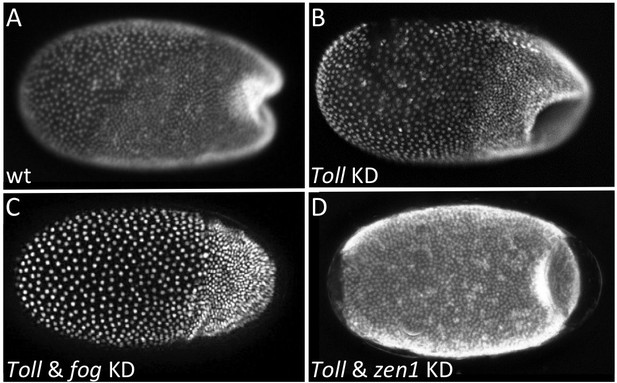
Local Tc-fog expression and posterior folding.
Nuclear (DAPI) staining of wildtype (A) and KD (B, C, D; specific KD shown in panels) embryos at the early posterior amniotic fold stage. Anterior is left, ventral is down (where possible to discern).
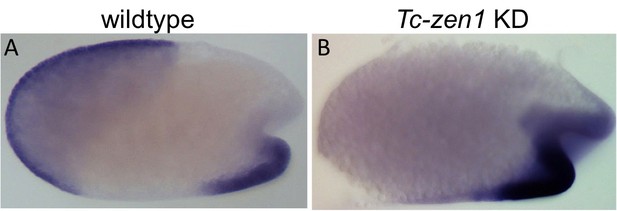
Tc-fog expression after Tc-zen1 KD.
Whole mount ISH for Tc-fog. Optical sagittal sections. (A, B) anterior Tc-fog expression is lost upon KD of Tc-zen1. Anterior is left, ventral is down.
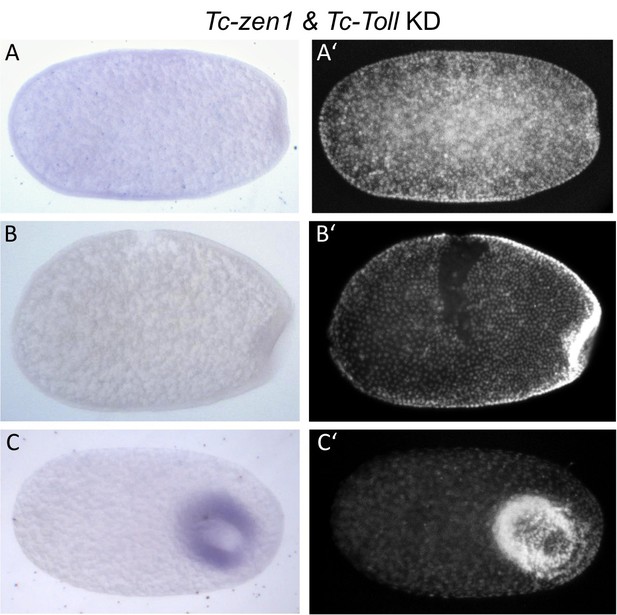
Tc-fog expression after Tc-Toll and Tc-zen1 KD.
Whole mount ISH for Tc-fog. Optical sagittal sections. (A–C) Tc-fog expression is lost (or greatly reduced) upon Tc-Toll and Tc-zen1 double KD by pRNAi. (A’–C’) nuclear (DAPI) staining of respective embryos. Anterior is left, ventral is down (where possible to discern).
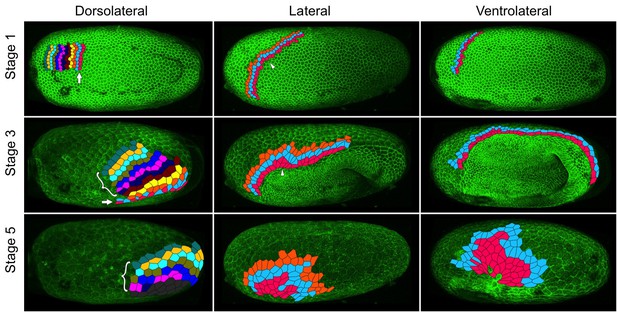
Serosal cells undergo intercalation during serosal expansion.
Stills from confocal live imaging of wildtype embryos with cell membranes marked via transient expression of GAP43-YFP. The tracked cells are colored as rows parallel to the serosa/germ rudiment boundary (pink closest to the boundary). The arrow indicates the region where cells intercalate to increase the number of cells at the serosa/germ rudiment boundary during stages 1–3. The arrowhead indicates one example region where cells intercalate to leave the serosa/germ rudiment boundary during stages 1–3. The bracket indicates cells located away from the serosa/germ rudiment boundary that undergo intercalation during stages 1–5. The dark gray cells in the lower left panel went out of the frame of view and could not be tracked for the full movie. In the right panels, new cells were tracked from when they became visible halfway through embryo formation (middle panel). Cell outlines were manually drawn using projection views of individual timepoints and individual z-sections. All panels show maximum intensity projections of one egg hemisphere. Anterior is left, ventral is down.
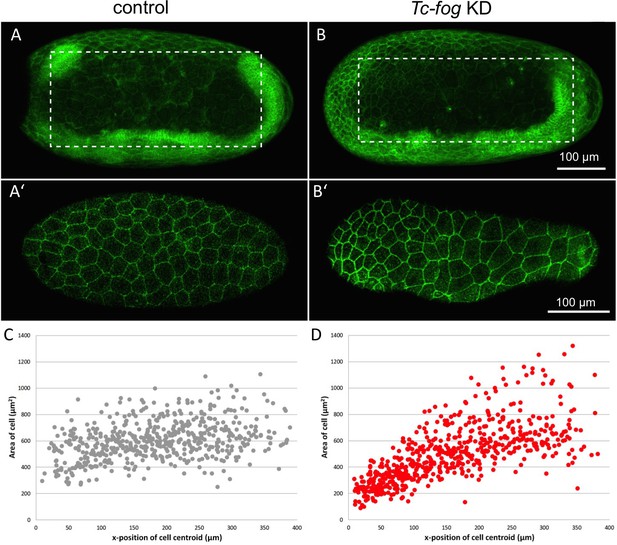
Fog signaling affects serosal expansion.
(A, B) Stills from confocal live imaging of wildtype (A) and Tc-fog eRNAi (B) embryos with cell membranes marked via transient expression of GAP43-YFP. Embryos are undergoing germband elongation. (A’, B’) Single optical section of the serosa from the dashed box region in (A, B) showing cells whose areas were measured. Quantification was performed on single optical section to avoid artefacts caused by curvature of the egg. (C, D) scatter plots showing serosa cell areas (wildtype (gray): 535 cells across eight embryos, Tc-fog eRNAi (red): 578 cells across seven embryos) together with AP position of cell centroids. Measurements were performed manually. (A, B) are average intensity projections of one egg hemisphere. Anterior is left, ventral is down.
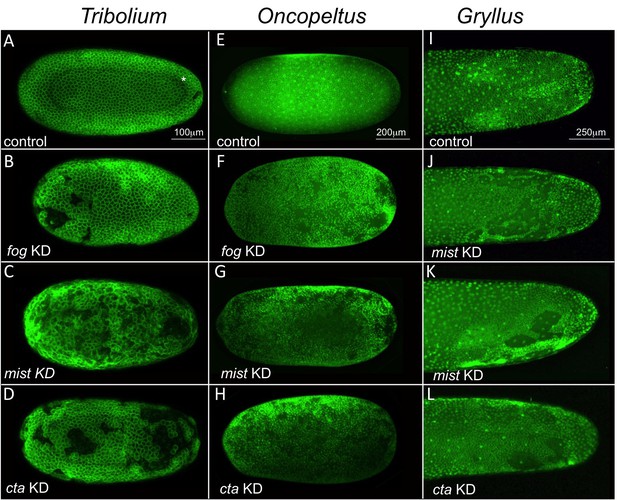
Fog signaling is required for blastoderm formation in Tribolium, Oncopeltus and Gryllus.
(A–D) Stills from confocal live imaging of wildtype (A), and Tc-fog, Tc-mist, Tc-cta eRNAi (B–D) Tribolium embryos with cell membranes marked via transient expression of GAP43-YFP. Embryos are at late blastoderm stage. White asterisk in (A) indicated hole within the blastoderm which later closes. (E–H) O. fasciatus blastoderm stage wildtype (E), and Of-fog, Of-mist, Of-cta pRNAi (F–G) embryos with nuclei stained (Sytox) to reveal their distribution. (I–L) Stills from fluorescent live imaging of control (I), and Gb-mist, Gb-cta pRNAi (J–L) G. bimaculatus embryos with nuclei labeled via a histone2B-eGFP transgene. The phenotype of Gb-mist pRNAi is stronger in (J) than in (K). The latter embryo recovered during later development. (A–D) are average intensity projections of one egg hemisphere, (I–L) are maximum focus projections of one egg hemisphere. Anterior is left for all embryos. (I, J, L) are ventral views, (K) is a ventrolateral view with ventral down.
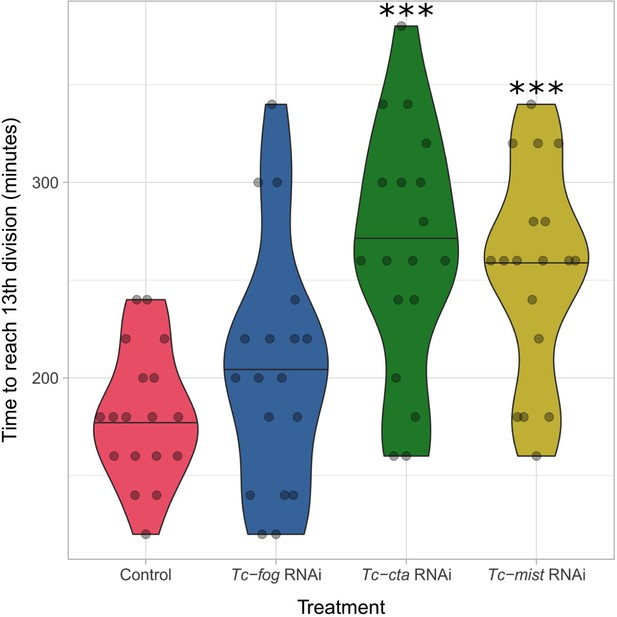
Developmental delay upon KD of Fog pathway components.
Violin plot showing effect of Tc-fog, Tc-cta or Tc-mist KD on timing of early development. All data points are shown in a quasi-random offset. ***p<0.001 (unpaired t-test) comparing control and KD.
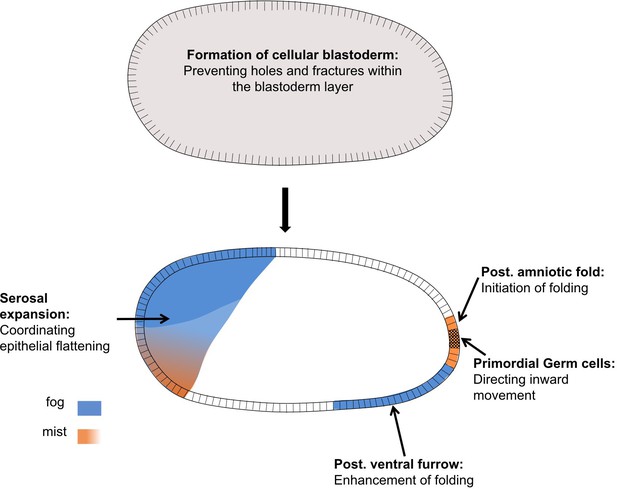
Distinct functions of Fog signaling in Tribolium.
Summary schematic showing the different roles of fog signaling during early Tribolium embryogenesis.
Videos
Fluorescent live imaging of wildtype and Tc-fog RNAi nGFP transgenic embryos.
Maximum intensity projections of one egg hemisphere are shown with anterior to the left and ventral to the bottom.
Fluorescent live imaging of wildtype and Tc-cta RNAi LifeAct-GFP transgenic embryos.
Maximum intensity projections of one egg hemisphere are shown with anterior to the left and ventral to the bottom.
Fluorescent live imaging of the posterior pole of a wildtype embryo transiently expressing GAP43-YFP.
Apical constrictions are visible at the center of the forming fold. Embryo was mounted with the posterior pole towards the objective and the resulting movie was digitally rotated. Maximum intensity projection of posterior view is shown as well as a transverse section along the dorsal/ventral midline. Ventral is to the bottom.
Single section through the subapical region of the cells at the posterior pole of a wildtype embryo transiently expressing GAP43-YFP.
Cells constrict over time and this occurs in a pulsatile manner, and cell intercalation is also visible. Ventral is to the bottom.
Fluorescent live imaging of wildtype embryos transiently expressing GAP43-YFP.
Embryos were mounted with their posterior poles towards the objective. Maximum intensity projection of posterior view is shown. Ventral is to the bottom.
Fluorescent live imaging of the presumptive serosa and germ rudiment epithelium of a wildtype embryo transiently expressing GAP43-YFP.
The cuboidal-to-squamous transition of the serosa cells during germband formation can be seen. Maximum intensity projection of the epithelium as well as transverse and sagittal sections along midpoints of the frame are shown. Anterior is to the left, ventral to the bottom.
Fluorescent live imaging of aTc-cta RNAi LifeAct-GFP transgenic embryo.
Maximum intensity projection of one egg hemisphere is shown with anterior to the left and ventral to the bottom.
Fluorescent live imaging of Tc-Toll1 and Tc-fog double RNAi nGFP transgenic embryo.
Maximum intensity projection of one egg hemisphere is shown with anterior to the left.
Fluorescent live imaging of the dorsolateral side of a wildtype embryo transiently expressing GAP43-YFP.
Serosa cells at/near the serosa/germ rudiment boundary were tracked and colored as rows (pink cells closest to the boundary). Only cells that were visible from the beginning of the timelapse are shown. Maximum intensity projection of one egg hemisphere is shown with anterior to the left and ventral to the bottom.
Fluorescent live imaging of the lateral side of a wildtype embryo transiently expressing GAP43-YFP.
Serosa cells at/near the serosa/germ rudiment boundary were tracked and colored as rows (pink cells closest to the boundary). Only cells that were visible from the beginning of the timelapse are shown. Maximum intensity projection of one egg hemisphere is shown with anterior to the left and ventral to the bottom.
Fluorescent live imaging of the ventrolateral side of a wildtype embryo transiently expressing GAP43-YFP.
Serosa cells at/near the serosa/germ rudiment boundary were tracked and colored as rows (pink cells closest to the boundary). Roughly half the cells (at the anterior) are tracked from the beginning of the timelapse, while the rest are tracked from halfway through embryo formation (at 135 min) when they became visible. Maximum intensity projection of one egg hemisphere is shown with anterior to the left and ventral to the bottom.
Fluorescent live imaging of part of the forming serosa in a wildtype embryo transiently expressing GAP43-YFP.
A group of cells intercalating via rosette formation are tracked. The field of view was manually stabilized to follow this group of cells. Anterior is to the left.
Fluorescent live imaging of the wildtype and Tc-fog weak KD embryos transiently expressing GAP43-YFP.
Serosa cells at/near the serosa/germ rudiment boundary were tracked. The top panels show cells coloured as rows (pink cells closest to the boundary). The bottom panels show cells colored randomly and intercalation events between tracked cells are shown with white triangles. Maximum intensity projection of one egg hemisphere is shown with anterior to the left and ventral to the bottom.
Fluorescent live imaging of a Tc-fog weak KD embryo transiently expressing GAP43-YFP.
Maximum intensity projection of one egg hemisphere with anterior to the left and ventral to the bottom is shown in the center. A transverse section near the anterior pole is to the left, a transverse section near the posterior pole is to the right, a sagittal section along the middle of the egg is shown to the bottom.
Fluorescent live imaging of wildtype and Tc-fog, Tc-T48, and Tc-fog and Tc-T48 double.
RNAi embryos transiently expressing GAP43-YFP. Average intensity projections of one egg hemisphere are shown with anterior to the left and ventral to the bottom.
Fluorescent live imaging of wildtype and Tc-fog, Tc-cta, and Tc-mist RNAi embryos transiently expressing GAP43-YFP.
Average intensity projections of one egg hemisphere are shown with anterior to the left and ventral to the bottom (where possible to discern).
Fluorescent live imaging of additional Tc-fog, Tc-cta, and Tc-mist RNAi embryos transiently expressing GAP43-YFP.
Further examples of blastoderm formation defects to demonstrate the variability in the phenotypes. Average intensity projections of one egg hemisphere are shown with anterior to the left and ventral to the bottom (where possible to discern).
Fluorescent live imaging of wildtype and Gb-mist RNAi histone2B-EGFP transgenic embryos.
Maximum focus projections of one egg hemisphere are shown as ventral views with anterior to left.
Additional files
-
Supplementary file 1
Primer List.
Primers used to produce anti-sense RNA for in-situ hybridization and dsRNA for RNAi-mediated gene knockdown.
- https://doi.org/10.7554/eLife.47346.048
-
Transparent reporting form
- https://doi.org/10.7554/eLife.47346.049





