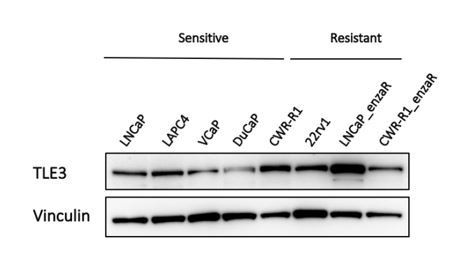
TLE3 loss confers AR inhibitor resistance by facilitating GR-mediated human prostate cancer cell growth
Figures
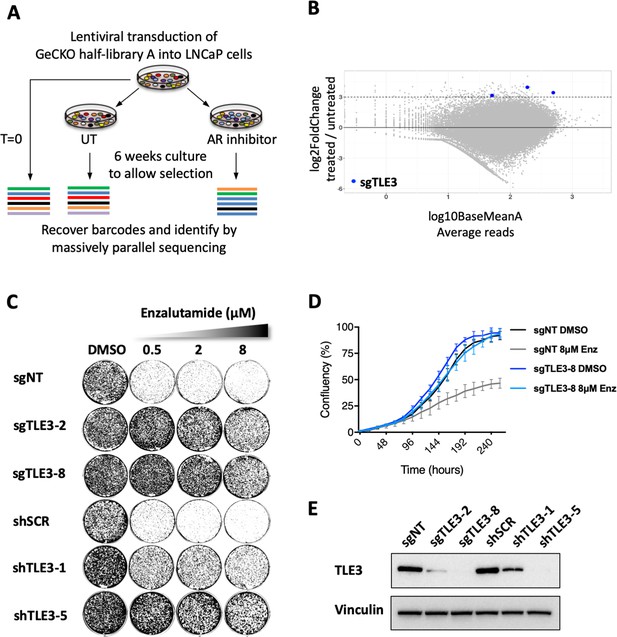
Genome-wide screen identifies TLE3 as a modulator of AR inhibitor sensitivity.
(A) Overview of the genome-wide CRISPR-Cas9 resistance screen. (B) Representation of the relative abundance of the gRNA barcode sequences of the CRISPR-Cas9 resistance screen. The y-axis shows the enrichment (relative abundance of apalutamide treated/untreated) and the x-axis shows the average sequence reads of the untreated samples. (C) Long-term growth assay (14 days) showing the functional phenotype of LNCaP cells harboring TLE3 knockout or knockdown vectors, cultured in the presence of vehicle or enzalutamide. Cells harboring a non-targeting sgRNA (sgNT) or scrambled shRNA (shSCR) were used as a control. (D) Quantitative analysis of live cell proliferation in real-time for control cells and TLE3KO cells in the absence or presence of enzalutamide. (E) Western blot showing TLE3 protein levels for control cells and TLE3KO cells shown in C and D. Vinculin was used as a loading control.
-
Figure 1—source data 1
Normalized readcounts CRISPR screen.
Normalized readcounts obtained with massively parallel sequencing are shown for each condition of the CRISPR screen (with three replicates per condition): timepoint 0, untreated and ARN-treated cells.
- https://cdn.elifesciences.org/articles/47430/elife-47430-fig1-data1-v2.xlsx
-
Figure 1—source data 2
DESeq2 analysis of the CRISPR screen.
Output file showing the raw data obtained with DESeq2 for the CRISPR resistance screen. For each gRNA the results, as indicated at the top of each column, are shown.
- https://cdn.elifesciences.org/articles/47430/elife-47430-fig1-data2-v2.xlsx
-
Figure 1—source data 3
MAGeCK analysis of the CRISPR screen.
Output file showing the raw data obtained with MAGeCK for the CRISPR resistance screen. For each gRNA the results, as indicated at the top of each column, are shown.
- https://cdn.elifesciences.org/articles/47430/elife-47430-fig1-data3-v2.xls
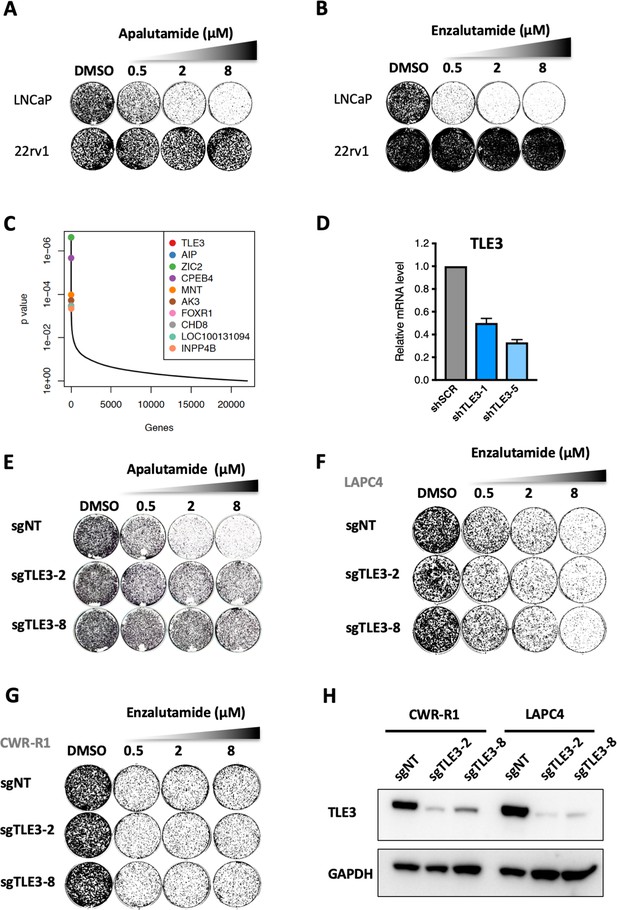
Genome-wide CRISPR screen identifies TLE3 as a modulator of AR inhibitor sensitivity.
(A–B) Long-term growth assay of LNCaP cells treated with apalutamide or enzalutamide for 14 days. Enzalutamide-resistant 22rv1 cells were used as control. (C) MAGeCK analysis showing hits obtained from the CRISPR-Cas9 resistance screen. (D) TLE3 mRNA expression levels in LNCaP cells harboring shRNAs targeting TLE3. Cells with scrambled shRNA (shSCR) were used as a control. (E) Long-term growth assays showing the functional phenotype for control and TLE3KO LNCaP cells cultured in the presence of vehicle or apalutamide at indicated concentrations for 14 days. (F–G) Long-term growth assays showing the functional phenotypes for indicated cell lines carrying control or TLE3-targeting gRNAs , cultured in the presence of vehicle or enzalutamide at indicated concentrations. (H) Western blot analysis showing TLE3 expression levels for the cells shown in F and G. GAPDH was used as a loading control.
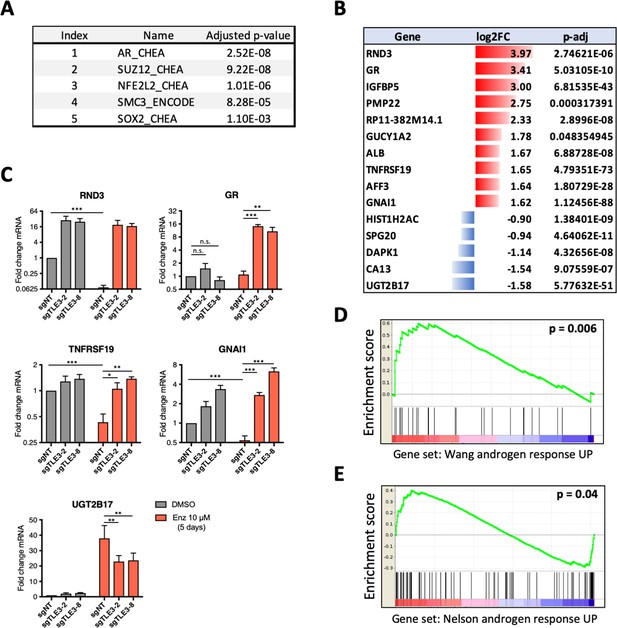
Transcriptomics analyses comparing control and TLE3KO cells cultured in the presence of vehicle or 10 μM enzalutamide for 5 days.
(A) Enrichment analysis for transcription factors associated with genes differentially expressed in enzalutamide-treated control cells compared to TLE3KO cells. (B) Overview of the fold changes in gene expression of the most differentially expressed genes in control cells versus TLE3KO cells treated with enzalutamide. (C) Validation (qPCR) of mRNA expression levels for several genes shown in B. Bars represent average data from at least three independent experiments ± SEM. P-values are indicated with ***p<0.001, **p<0.01 and *p<0.05 (two-tailed t-test). (D–E) GSEA for genes differentially expressed in control cells compared to TLE3KO cells, treated with 10 μM enzalutamide using indicated gene sets.
-
Figure 2—source data 1
Readcounts RNA-seq experiment comparing control and TLE3KO cells.
The normalized readcounts (transcripts) for each gene are shown for control and TLE3KO cells treated with vehicle or 10 μM enzalutamide for 5 days.
- https://cdn.elifesciences.org/articles/47430/elife-47430-fig2-data1-v2.xlsx
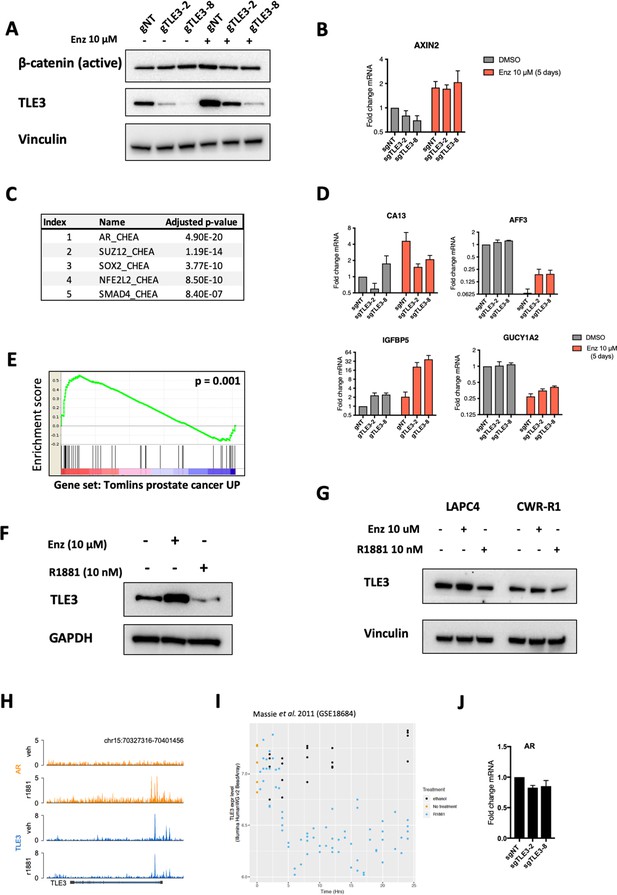
Transcriptomics analyses comparing control and TLE3KO cells.
(A) Western blot results showing expression levels of active ß-catenin, TLE3 and Vinculin (loading control) in indicated cell lines cultured with and without enzalutamide. (B) Expression levels of AXIN2 as determined by qPCR in control and TLE3KO cells cultured in the presence of absence enzalutamide. (C) Top five transcription factors associated with genes differentially expressed in control versus TLE3KO cells in the untreated condition. (D) Validation of several genes most differentially expressed in TLE3KO cells compared to control cells cultured in the presence of enzalutamide. (E) GSEA for genes differentially expressed in control versus TLE3KO cells treated with enzalutamide, using the indicated geneset. (F) Western blot analysis for TLE3 expression levels in LNCaP cells cultured as indicated for 5 days. GAPDH was used as a loading control. (G) Western blot showing TLE3 protein expression levels in LAPC4 and CWR-R1 cells cultured with indicated drugs for 5 days. (H) Snapshot of coverage profiles for TLE3 and AR binding at the TLE3 locus under indicated conditions. (I) RNA-seq data showing TLE3 mRNA levels of LNCaP cells treated with vehicle or R1881 for 24 hr (time course). (J) RT-qPCR analysis showing AR expression in untreated control and TLE3KO cells.
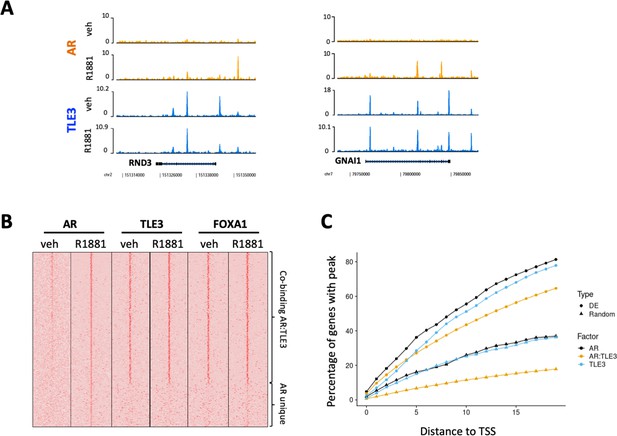
ChIP-seq analyses for transcription factor binding at differentially expressed genes in control cells compared to TLE3KO cells cultured in the presence of 10 μM enzalutamide.
(A) Coverage profiles for TLE3 and AR at the loci of two genes (RND3 and GNAI1). (B) Heatmap of AR, TLE3 and FOXA1. (co-)binding at genes differentially expressed in TLE3KO compared to control cells treated with enzalutamide are shown. The binding of AR, TLE3 and FOXA1 at these sites is shown for androgen-depleted or R1881-stimulated (4 hr) conditions in parental LNCaP cells. (C) ChIP-seq peak enrichment near the Transcription Start Sites (TSS) of differentially expressed (DE) genes and a random set of genes. The fraction of genes with a peak for TLE3, AR or both transcription factors at indicated distance from the TSS is shown for both genesets.
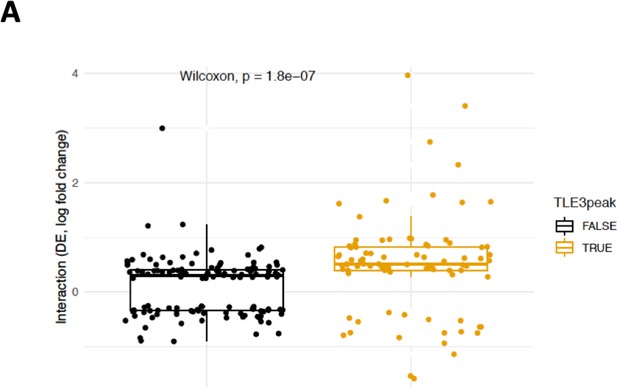
TLE3 binding status proximal to genes differentially expressed control cells compared to TLE3KO cells.
(A) Boxplot showing the Log2 fold change expression of genes associated with TLE3KO and enzalutamide treatment. Genes have been grouped based on TLE3 binding status at the loci of these genes; no TLE3 binding (black) and TLE3 binding (orange).
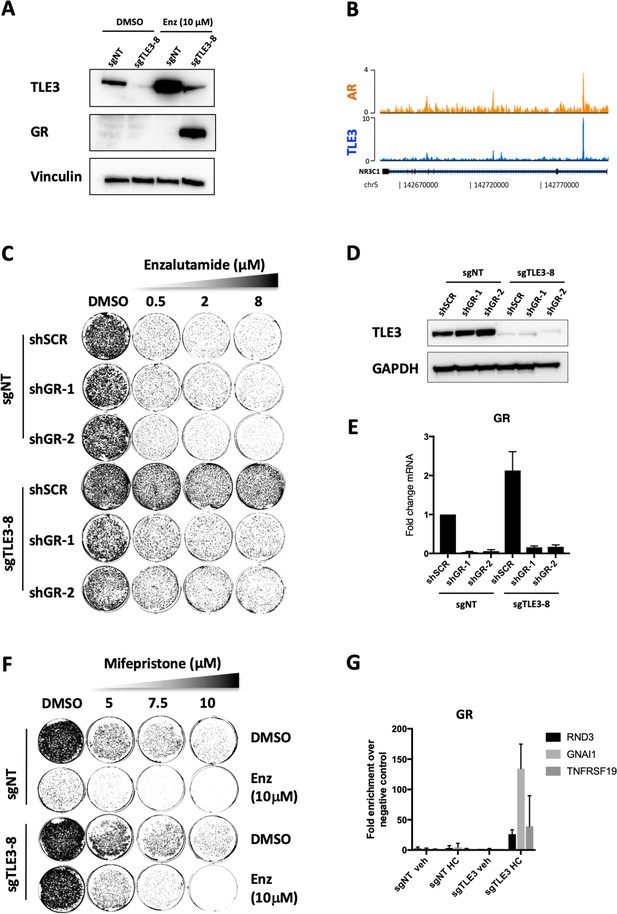
GR inhibition resensitizes TLE3KO cells to enzalutamide treatment.
(A) Western blot showing protein expression levels of TLE3 and GR in control and TLE3KO cells cultured vehicle or enzalutamide (B) Coverage profiles for TLE3 and AR binding at the GR locus. (C) Long-term growth assay (14 days) showing the drug resistance phenotype of control and TLE3KO cells with and without GR knockdown in the presence of vehicle or enzalutamide. (D) Western blot analysis for TLE3 protein levels in control and TLE3KO cells shown in C, using GAPDH as a loading control. (E) mRNA levels for GR in control and TLE3KO cells carrying shSCR or shGR constructs, shown in C. (F) Long-term growth assay (14 days) for cells harboring a control sgRNA or TLE3-targeting sgRNA cultured in the presence of vehicle, enzalutamide, mifepristone or the combination at indicated concentrations. (G) ChIP-qPCR showing GR occupancy at enhancers proximal to indicated genes. All samples were cultured in the presence of 10 μM enzalutamide with or without 1 μg/ml hydrocortisone (HC) as indicated.
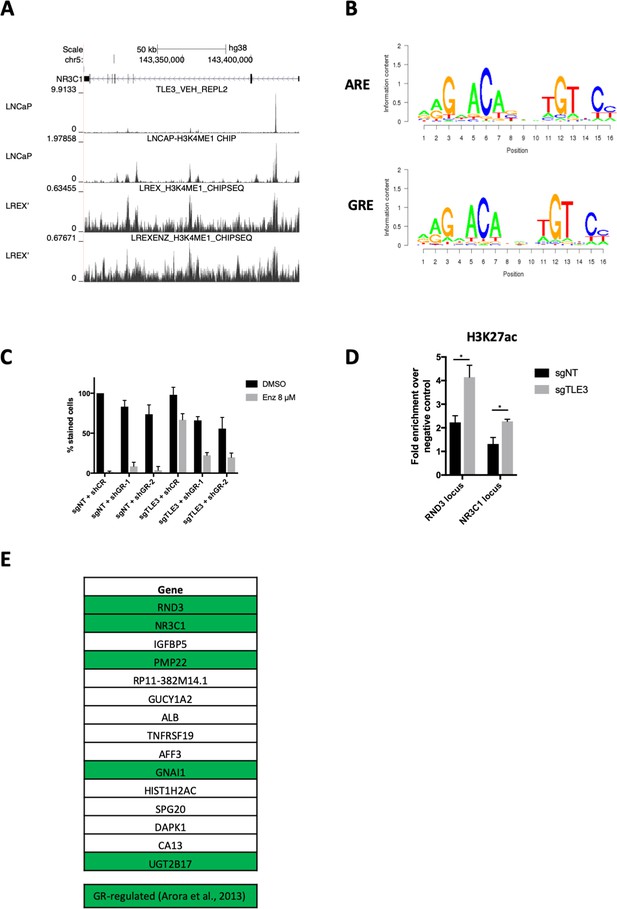
GR-mediated gene regulation in TLE3KO cells.
(A) Coverage profiles for TLE3 or H3K4me1 at the GR locus in LNCaP and LREX’ cells. (B) Core consensus sequences for AR and GR. (C) Quantification of long-term colony formation assays showing the functional phenotype of control and TLE3KO, with or without GR knockdown, in cells treated with vehicle or enzalutamide. (D) ChIP-qPCR for H3K27 acetylation in control and TLE3KO cells at indicated loci. (E) Overview of the genes listed in Figure 2B with GR-regulated genes (as shown by Arora et al., 2013) highlighted in green.
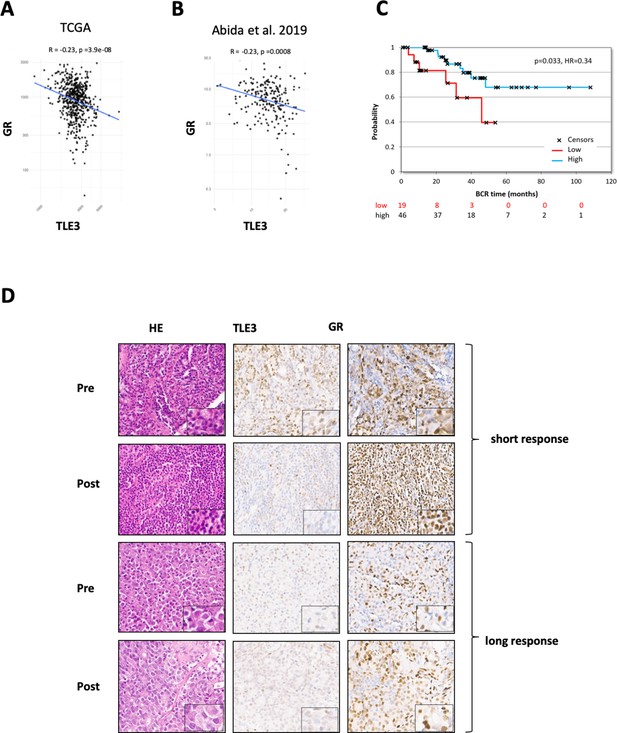
TLE3 and GR expression in tumors of prostate cancer patients.
(A–B) RNA-seq analysis showing the correlation between TLE3 and GR expression in tumor samples from prostate cancer patients. (C) Kaplan-Meier curve showing the biochemical recurrence of prostate cancer patients from the TCGA dataset, only patients receiving anti-hormonal therapy were included (65 patients) using an optimal cut-off for high versus low TLE3 expression. (D) Immunohistochemistry for H and E, TLE3 and GR in tumor biopsy samples collected from two CRPC patients pre- and post-enzalutamide treatment.
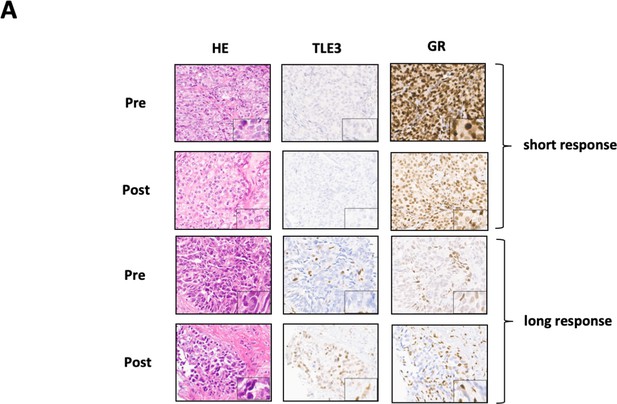
TLE3 and GR protein expression in CRPC patient samples pre- and post enzalutamide treatment.
(A) Immunohistochemistry for H and E, TLE3 and GR in tumor biopsy samples collected from two CRPC patients pre- and post-enzalutamide treatment.
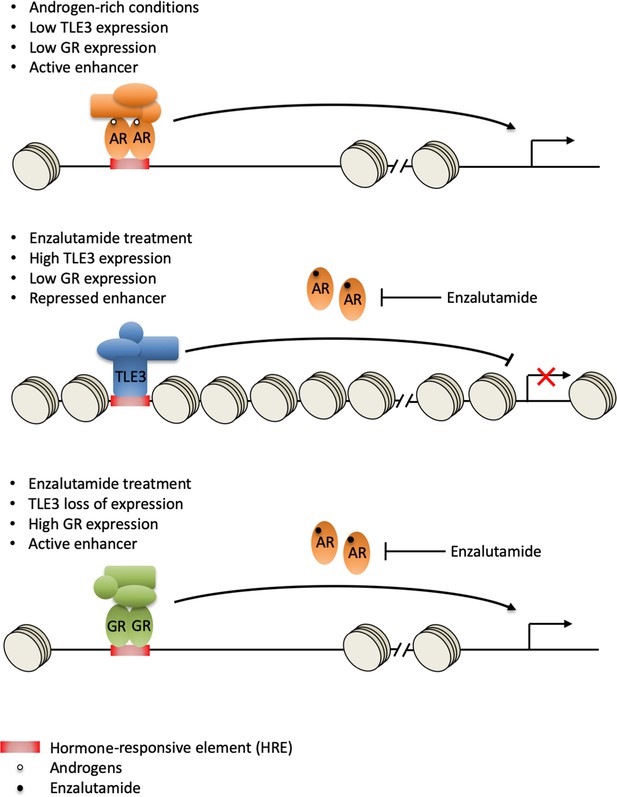
Model for GR-mediated enzalutamide resistance in TLE3KO prostate cancer cells.
In the presence of androgens, TLE3 expression is repressed and enhancers are active. AR regulates target gene expression, including repression of the GR locus (top panel). Upon enzalutamide treatment, TLE3 is upregulated and enhancers are inactive. TLE3 represses expression of AR target genes including GR (middle panel). Enzalutamide treatment in the context of TLE3 loss leads to upregulation of GR which is able to substitute for AR at active enhancers, leading to drug resistance (bottom panel).
Additional files
-
Supplementary file 1
Motif enrichment analysis.
Motif enrichment analysis showing sequence motifs that are enriched at genes differentially expressed in enzalutamide-treated control cells compared to TLE3KO cells.
- https://cdn.elifesciences.org/articles/47430/elife-47430-supp1-v2.pdf
-
Supplementary file 2
Key Resources Table.
- https://cdn.elifesciences.org/articles/47430/elife-47430-supp2-v2.docx
-
Supplementary file 3
TCGA prostate cancer dataset (from https://portal.gdc.cancer.gov/) used for Figure 5C.
- https://cdn.elifesciences.org/articles/47430/elife-47430-supp3-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/47430/elife-47430-transrepform-v2.docx