Tight nuclear tethering of cGAS is essential for preventing autoreactivity
Figures
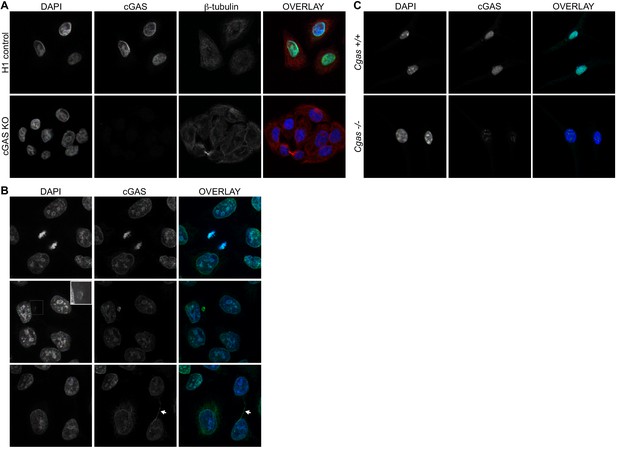
Endogenous cGAS is predominantly a nuclear protein.
(A) Clonal lines of HeLa cells were generated using lentiCRISPR encoding either a non-targeting H1 control gRNA (top row) or a cGAS-targeted gRNA (cGAS KO). Cells were fixed with methanol, stained with antibodies to human cGAS and beta-tubulin, counter-stained with DAPI, and visualized by confocal microscopy. (B) We noted three reproducible patterns of cGAS localization in addition to the nucleus: condensed mitotic chromatin (top row), structures resembling micronuclei (middle row), and tendril-like bridges between cells. (C) Mouse Cgas+/+ and Cgas-/- primary bone marrow-derived macrophages were stained using a mouse antibody to cGAS and processed as in (A).
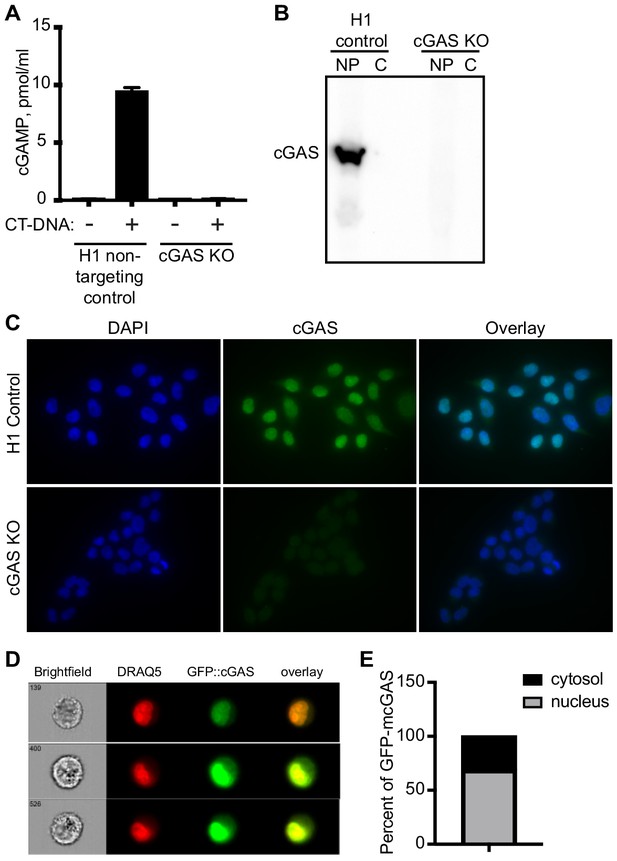
Characterization of clonal cGAS KO HeLa cells and microscopy conditions for cGAS visualization.
(A) HeLa cells were transduced with LentiCRISPR encoding H1 non-targeting control gRNA or cGAS-targeted gRNA, cloned by limiting dilution, and tested for production of cGAMP in cell lysates four hours after transfection of calf thymus DNA (CT-DNA). (B) Lysates from H1 control and cGAS KO clonal HeLa cells were separated into cytosol (C) and nuclear pellet (NP), and then blotted for endogenous cGAS. (C) H1 control and cGAS KO HeLa cells were fixed with 4% paraformaldehyde, permeabilized with 0.1% Triton X-100, and stained for endogenous cGAS. (D) cGAS KO HeLa cells were transduced with pSLIK lentivirus encoding doxycyline-inducible GFP-mouse cGAS (mcGAS), and then treated with doxycyline to induce GFP-mcGAS expression. Cells were then stained with DRAQ5 and visualized with an imaging flow cytometer (Amnis ImageStream). Representative images shown in (D). (E) Analysis of ImageStream data for thousands of individual cells showing percent nuclear and cytosolic localization of GFP-mcGAS.
Compiled z-series confocal images of the HeLa cells visualized in Figure 1A.
Tubulin is red, cGAS is green, DAPI is blue.
Rendered z-series of HeLa cells rotated around the y-axis.
Tubulin is red, cGAS is green, DAPI is blue.
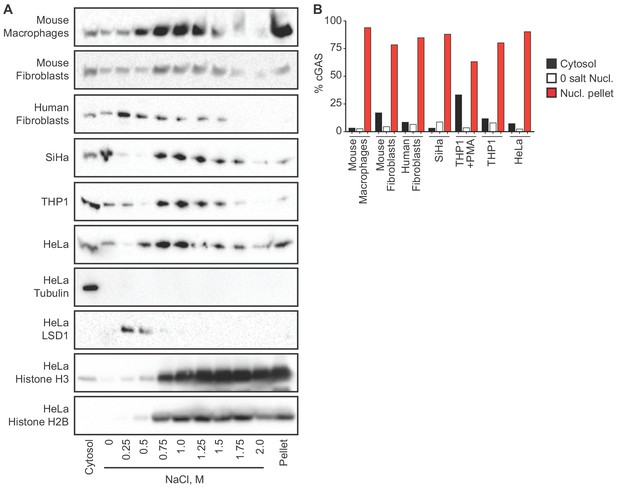
cGAS is tightly tethered in the nucleus.
(A) Mouse and human cell lines were separated into cytosolic and nuclear fractions, followed by sequential stepwise elutions of nuclear pellets with the indicated concentrations of NaCl. cGAS and the indicated control proteins (shown for HeLa cells) were monitored throughout the elution by western blot. (B) Densitometry measurements quantitating the relative amounts of endogenous cGAS protein in the cytosol, the 0 salt nuclear lysis, and the cumulative nuclear pellet.
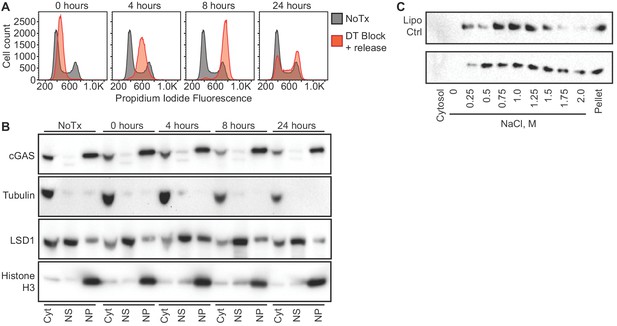
cGAS nuclear localization is independent of cell cycle phase or activation status.
(A) HeLa cells were arrested at the G1/S border using double thymidine (DT) block, followed by release and harvest at the indicated time points for measurement of DNA content. (B) Cells from (A) were fractionated and cGAS localization was determined by western blot. (C) HeLa cells were transfected with Lipofectamine alone (Lipo) or with CT-DNA for 4 hr, followed by extraction, salt elution, and western blot for endogenous cGAS.
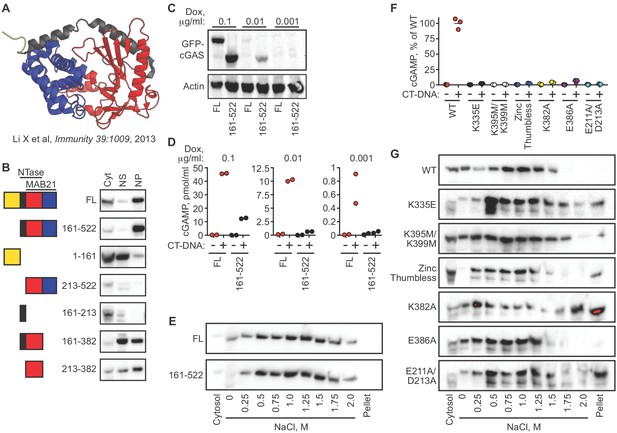
cGAS nuclear localization and tethering are independent of robust DNA binding, dimerization, condensation, and catalytic activity.
(A) Structure of human cGAS with domains colorized. (B) TERT-immortalized human foreskin fibroblasts were reconstituted with the indicated Dox-inducible GFP-cGAS lentivirus constructs, treated with 0.1 μg/ml Dox for 24 hr, and then separated into cytosolic (Cyt), nuclear supernatant (NS), and nuclear pellet (NP) fractions. FL: Full-Length. (C) cGAS-deficient HeLa cells reconstituted with the indicated GFP-cGAS constructs were induced for 24 hr with three doses of Dox. Whole cell lysates that recover all cGAS were prepared and blotted with anti-GFP antibody. (D) Cells from (C) were transfected with CT-DNA for four hours, followed by cGAMP measurement in lysates by modified ELISA. (E) Cells described in C-D were treated with 0.1 μg/ml Dox for 24 hr to induce GFP-cGAS expression, then harvested and used for sequential fractionation and salt elution as in Figure 2. (F) Dox-inducible, full-length mouse cGAS constructs were introduced into hTERT-immortalized human fibroblasts. Cells were treated with 0.1 μg/ml Dox for 24 hr followed by stimulation for 4 hr and measurement of cGAMP in cell lysates. (G) Unstimulated cells from (F) were fractionated and blotted for cGAS.
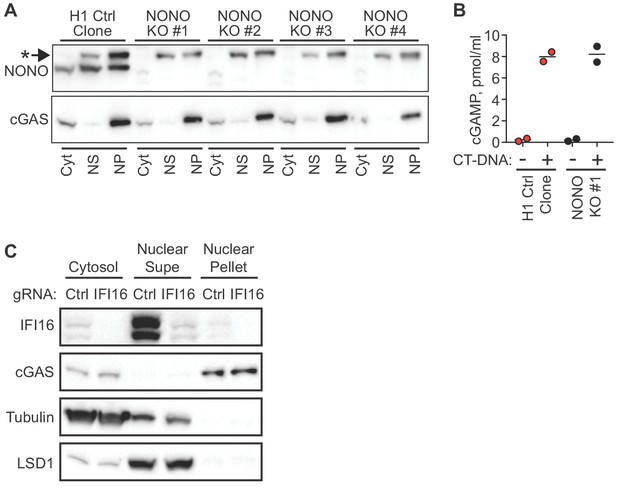
NONO and IFI16 are dispensable for cGAS nuclear localization and tethering.
(A) H1-targeted control HeLa clone and four independent NONO KO HeLa clones were lysed and separated into Cytosol (Cyt), Nuclear supe (NS), and nuclear pellet (NP) fractions, and blotted for NONO and endogenous cGAS. The asterisk indicates a non-specific band detected by the NONO antibody. (B) Cells from (A) were transfected with CT-DNA for four hours, followed by cGAMP measurement in lysates by modified ELISA. (C) TERT-immortalized human fibroblasts were transduced with lentiCRISPR constructs encoding either control gRNA or gRNA targeting IFI16, selected with puromycin to enrich for transduced cells, and then fractionated and blotted with the indicated antibodies.
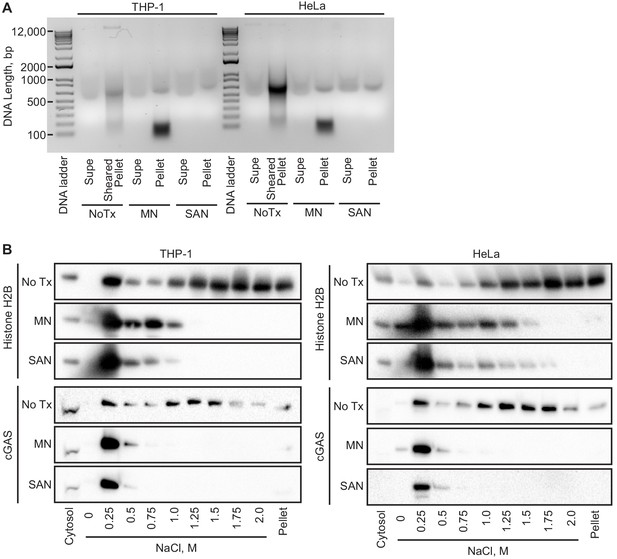
Intact chromatin is required for cGAS tethering.
(A) THP-1 or HeLa cell nuclear extracts were left untreated (NoTx), treated after the 0 salt wash step with micrococcal nuclease (MN), or treated with at the 0.25 M NaCl elution step with Salt Active Nuclease (SAN). Supernatants (supe) and pellets were collected, and the untreated pellet was sonicated to shear large genomic DNA. DNA was extracted, run on an agarose gel, and visualized with SYBR Safe. (B) Extracts treated as described above were used for sequential salt elution, followed by blotting for Histone H2B or cGAS.
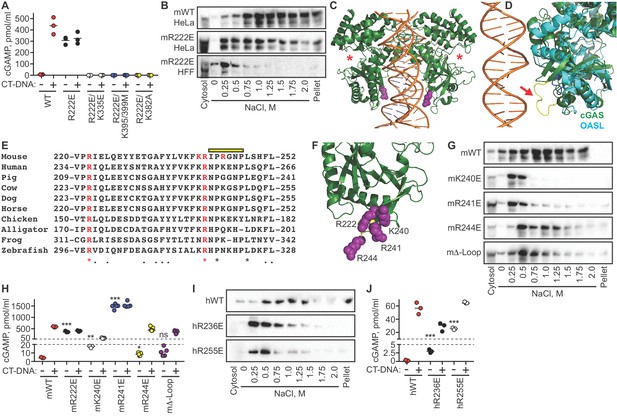
The cGAS tethering surface prevents autoreactivity.
(A) cGAS KO HeLa cells were reconstituted with the indicated Dox-inducible murine cGAS lentivirus constructs, treated with 0.1 μg /ml Dox for 24 hr, then transfected for four hours with lipofectamine alone or lipofectamine + CT DNA, followed by measurement of cGAMP in cell lysates. (B) Salt elution profiles of WT and R222E mouse cGAS; the top two panels are in HeLa cells and the bottom panel is in TERT-immortalized human fibroblasts. (C) Crystal structures of mouse cGAS (green) assembled on DNA, modeled from PDBID 5N6I. DNA is orange, R222 is highlighted in purple spheres, and the locations of the active sites are noted with red asterisks. (D) Overlay of mouse cGAS (green) and human OASL (cyan; PDBID 4XQ7). The unique loop in cGAS is colored yellow and indicated with the red arrow. (E) Alignments of cGAS across vertebrate phylogeny, with conserved, positively charged residues highlighted in red and the central loop amino acids indicated by the yellow bar. (F) The positively charged residues near R222 are highlighted in purple spheres. (G) Salt elution profiles of cGAS KO HeLa cells transduced with the indicated murine cGAS constructs after induction with 0.1 μg/ml Dox for 24 hr. (H) Cells from (G) were treated with 0.1 μg/ml Dox for 24 hr, then transfected for four hours with lipofectamine alone or lipofectamine + CT-DNA, followed by measurement of cGAMP in cell lysates. (I) Salt elution profiles of cGAS KO HeLa cells transduced with the indicated human cGAS constructs after induction with 0.1 μg/ml Dox for 24 hr. (J) Cells from (I) were treated as in (H), followed by measurement of cGAMP in cell lysates. Statistical comparisons were made between resting WT cGAS and each mutant, using one-way ANOVA of log-transformed biological replicates. *p=0.0155; **p=0.0018; ***p<0.0001; ns: not significant.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (human) | HeLa | ATCC | CCL-2 | |
| Cell line (human) | SiHa | ATCC | HTB-35 | |
| Cell line (human) | THP1 | ATCC | TIB-202 | |
| Cell line (human) | HEK 293T | ATCC | CRL-3216 | |
| Cell line (human) | Primary human foreskin fibroblasts | Millipore | Cat # SCC-058 | |
| Cell line (mouse) | mouse bone marrow macrophages, C57BL/6J | Gray EE et al, J Immunol 2015 195:1939 PMID: 27496731 | ||
| Cell line (human) | HeLa H1 LentiCRISPR control clonal lines | This Paper | ||
| Cell line (human) | HeLa cGAS LentiCRISPR clonal lines | This Paper | ||
| Cell line (human) | Hela cGAS KO pSLIK GFP | This Paper | ||
| Cell line (human) | Hela cGAS KO pSLIK GFP FL mcGAS | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK GFP mcGAS 161–522 | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK mcGAS WT | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK mcGAS R222E | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK mcGAS R222E/K335E | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK mcGAS R222E/K335E | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK mcGAS R222E/K395M/K399M | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK mcGAS R222E/K382A | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK mcGAS K240E | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK mcGAS R241E | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK mcGAS R244E | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK mcGAS ΔLoop | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK hcGAS WT | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK hcGAS R236E | This Paper | ||
| Cell line (human) | HeLa cGAS KO pSLIK hcGAS R255E | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK GFP-mcGAS full length (FL) | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK GFP-mcGAS (161-522) | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK GFP-mcGAS (1-161) | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK GFP-mcGAS (213-522) | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK GFP-mcGAS (161-213) | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK GFP-mcGAS (161-382) | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK GFP-mcGAS (213-382) | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK mcGAS K335E | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK mcGAS K395M/K399M | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK mcGAS Zinc thumbless | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK mcGAS K382A | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK mcGAS E386A | This Paper | ||
| Cell line (human) | hTERT human fibroblast pSLIK mcGAS E211A/D213A | This Paper | ||
| Cell line (human) | hTERT human fibroblast H1 lentiCRISPR control | Gray EE et al, Immunity 2016 45:255 PMID: 27496731 | ||
| Cell line (human) | hTERT human fibroblast IFI16 lentiCRISPR | Gray EE et al, Immunity 2016 45:255 PMID: 27496731 | ||
| Cell line (mouse) | mouse bone marrow macrophages, Cgas-/- | Gray EE et al, J Immunol 2015 195:1939 PMID: 26223655 | ||
| Cell line (mouse) | WT MEF (mouse embryonic fibroblasts) | Gray EE et al, J Immunol 2015 195:1939 PMID: 26223655 | ||
| Transfected construct | pSLIK-Neo | Addgene | Plasmid #25735 | |
| Transfected construct | pSLIK-Blasticidin | This Paper | ||
| Antibody | rabbit monoclonal anti-cgas D1D3G | Cell Signaling | cat # 15102S | |
| Antibody | rabbit monoclonal anti-cgas, mouse specific | Cell Signaling | cat # 31659 | |
| Antibody | rabbit monoclonal anti-LSD1 C69G12 | Cell Signaling | cat # 2184S | |
| Antibody | rabbit polyclonal anti-histone H3 | abcam | cat # ab1791 | |
| Antibody | rabbit polyclonal anti-histone H2B | abcam | cat # ab1790 | |
| Antibody | rabbit polyclonal anti-GFP | abcam | cat # 6556 | |
| Antibody | mouse monoclonal anti-Ifi16 | abcam | cat # 55328 | |
| Antibody | rabbit polyclonal anti-NONO | Sigma Aldrich | cat # N8789-200UL | |
| Antibody | rabbit polyclonal anti- α/β tubulin | Cell Signaling | cat # 2148 | |
| Antibody | mouse monoclonal anti-β actin | SIGMA | cat # A5441 | |
| Antibody | rabbit IgG control | Fisher | cat # 10500C | |
| Antibody | rabbit polyclonal anti-cgas | SIGMA | cat # HPA031700 | |
| Antibody | mouse monoclonal anti-β-tubulin | Cell Signaling | cat # 86298 | |
| Commercial assay or kit | NE-PER nuclear and cytoplasmic extraction reagents | Thermo Fisher Scientific | cat # PI78835 | |
| Commercial assay or kit | propidium iodide | SIGMA | cat # P4170 | |
| Commercial assay or kit | Luciferase Assay System | Promega | cat # E4550 | |
| Commercial assay or kit | 2',3'-Cyclic GAMP Direct EIA Kit | Arbor Assays | cat # K067-H1 | |
| Chemical compound, drug | thymidine | VWR | cat # 80058–750 | |
| Chemical compound, drug | SAN (salt active nuclease) | SIGMA | cat # SRE0015 | |
| Chemical compound, drug | calf thymus DNA | SIGMA | cat # D4764 | |
| Chemical compound, drug | micrococcal nuclease | New England Biolabs | cat # M0247S | |
| Chemical compound, drug | DAPI | SIGMA | cat # D9542 | |
| Chemical compound, drug | SYBR safe | Apex Bio | cat # A8743 | |
| Chemical compound, drug | DRAQ5 | Thermo Fisher | cat # 62251 | |
| Chemical compound, drug | NP-40 substitute | SIGMA | cat # 74385 | |
| Chemical compound, drug | protease inhibitor tablet | Pierce | cat # PIA32955 | |
| Chemical compound, drug | Prolong Gold Antifade mountant | Thermo Fisher | cat # P36930 | |
| Chemical compound, drug | Roche Block | Sigma | cat # 11921673001 | |
| Other | Immobilon-FL PVDF, 0.45 µm western blot membrane | SIGMA | cat # IPFL00010 | |
| Other | Immobilon-PSQ PVDF,0.2 µm western blot membrane | SIGMA | cat # ISEQ00010 |





