Silencing cryptic specialized metabolism in Streptomyces by the nucleoid-associated protein Lsr2
Figures
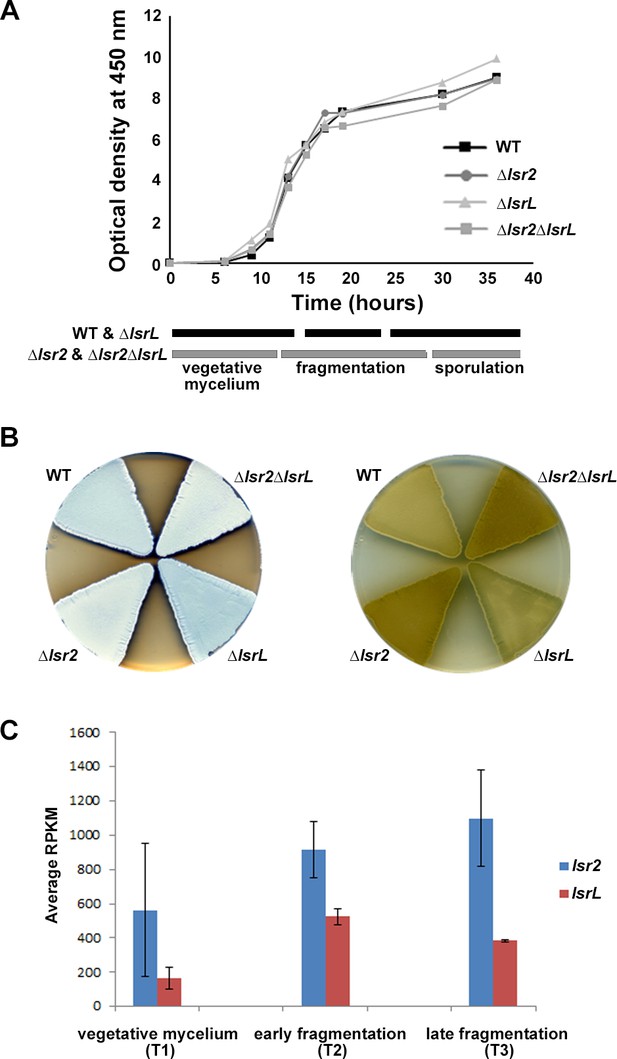
Expression and phenotypic analyses of lsr2 and lsrL in S. venezuelae.
(A) Growth curves and developmental stages (as determined using light microscopy) of wild type (WT), and the three different lsr2/lsrL mutants, over a 40 hr time course in MYM liquid medium. (B) Comparing development (left) and melanin/brown pigment production (right – underside of plate) of wild type (WT), versus single (Δlsr2 and ΔlsrL) and double (Δlsr2ΔlsrL) mutant strains after 2 d growth on MYM agar medium. The white color of the lsr2-containing mutants reflects aerial hyphae formation, while the darker color associated with the wild type and lsrL mutant strains indicates culture sporulation. (C) Comparative transcript levels (RPKM = reads per kilobase per million) for lsr2 and lsrL at three time points [T1, T2, T3, representing the three developmental stages indicated in (A)] in liquid MYM medium, as assessed using RNA-sequencing data. Data are presented as mean ± standard deviation (n = 2).
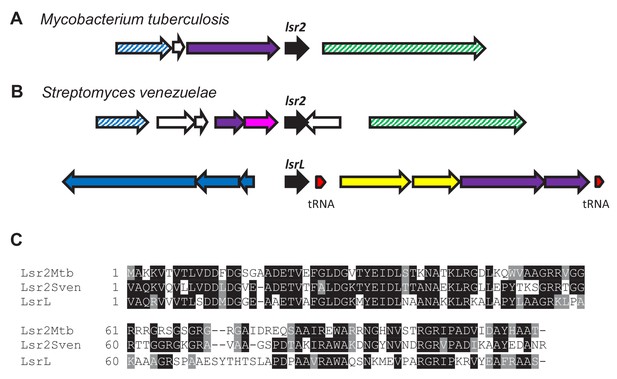
Sequence similarity and genetic organization of lsr2 and lsrL (lsr2-like) genes and their products in Streptomyces and Mycobacterium.
(A) Location of lsr2 in Mycobacterium tuberculosis. (B) Location of lsr2 and lsrL in Streptomyces venezuelae. lsr2/lsrL are shaded in black; tRNAs in red; transcriptional regulators in purple; transport and amino acid metabolism in pink; transport and metabolism of nucleotides and coenzymes in blue; protein turnover in green; ABC transporters in yellow; and hypothetical genes in white. Hatched colors indicate homologs. (C) Amino acid sequence alignment of Lsr2 from M. tuberculosis (Mtb), S. venezuelae (Sven) and LsrL. Identical residues are shaded in black, while gray shading indicates similar residues.
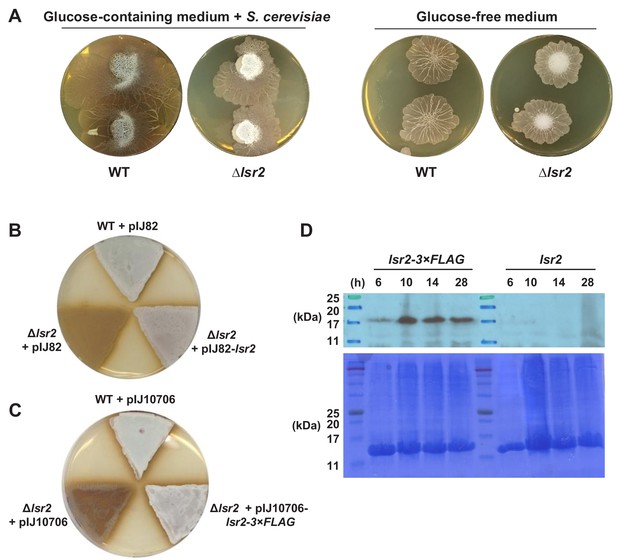
Phenotypic comparison of lsr2 deletion mutants with wild-type S. venezuelae.
(A) Wild type and Δlsr2 strains exploring on glucose-containing medium (YPD) in association with yeast (left), or on glucose-free (YP) medium without yeast (right). (B) Phenotype of wild-type S. venezuelae carrying the empty vector pIJ82 (top), the lsr2 null mutant carrying the empty vector pIJ82 (bottom left), or complemented with the native lsr2 (bottom right). Equivalent numbers of spores were grown on MYM solid agar and images were taken after 24 hr. (C) Phenotype of wild-type S. venezuelae carrying the empty plasmid pIJ10706 (top), the lsr2 null mutant carrying the same plasmid (bottom left), or complemented with a C-terminally triple FLAG-tagged lsr2 fusion (bottom right). As for (A), images were taken after 24 hr growth on MYM solid agar. (D) Top panel: western blot analysis of the S. venezuelae lsr2 mutant strain complemented with either the wild type lsr2 gene (right) or the lsr2−3 × FLAG fusion (left). Protein extracts were prepared after 6, 10, 14 and 28 hr growth in MYM broth. Bottom panel: Coomassie staining of equivalent samples to those used for western blotting, was used to control for differential sample loading. Protein size markers (BLUeye, Sigma) are shown with relevant molecular masses annotated in kDa.
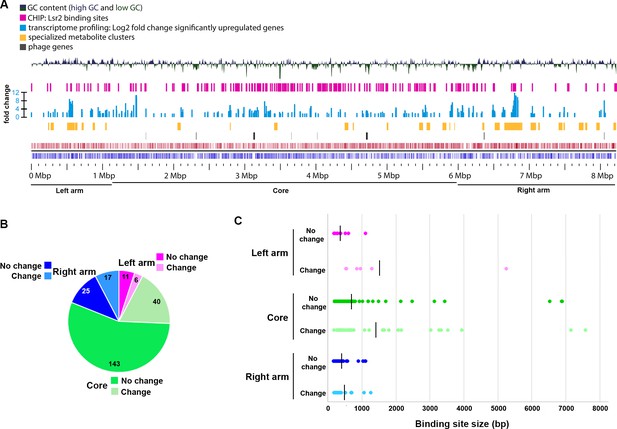
Composition of S. venezuelae chromosome in relation to Lsr2 binding sites and differentially affected genes.
(A) Panels are described from the bottom up. Bottom panels: coding sequences and relative strand organization (forward orientation shown in red; reverse orientation shown in blue) across the S. venezuelae chromosome, with left arm, core and right arm regions indicated. Above that, the regions shown with black bars indicate the relative position of predicted phage genes, while those in yellow indicate the location of specialized metabolic clusters. The light blue bars represent genes whose expression is significantly upregulated in the lsr2 mutant (fold change indicated), while the pink bars indicate Lsr2 binding sites, as determined by ChIP-seq. The top panel depicts the GC content of the chromosome, relative to the average percentage (72.4%). The peaks above the middle line indicate a GC content above 72.4%, while those below indicate a GC content below 72.4%. Image was generated using GView (Petkau et al., 2010). (B) Binding sites within the left arm (pink), core (green) or right arm (blue), and relative proportion of sites associated with transcriptional changes (no change in transcription = darker color; change in transcription = lighter color). Shown within each segment is the number of binding sites associated with that region. (C) Size of binding sites (in base pairs, bp) in each of the chromosome regions, separated into those associated with altered transcription (change), versus not (no change). Color scheme is as described in (B). The average binding site size for each group is indicated by the vertical black line.
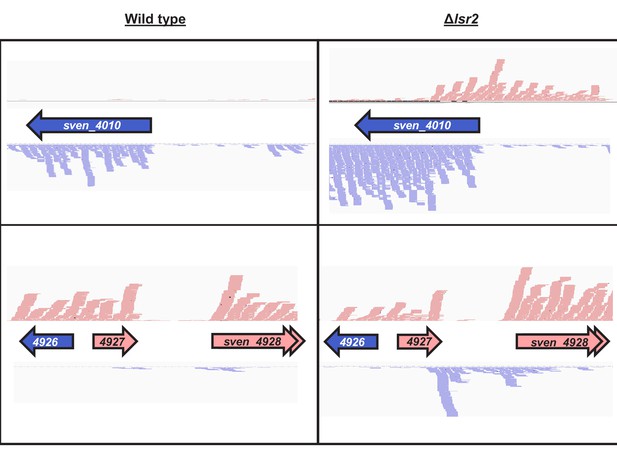
Increased antisense RNA expression in an lsr2 mutant strain compared with wild type.
Gene names/numbers are as indicated. Blue arrows/RNA-seq reads indicate expression in the reverse direction, while pink arrows/RNA-seq reads indicate expression in the forward direction. Note there is little forward expression for sven_4010 in the wild type strain, and little reverse expression for sven_4927/28 in the wild type strain.
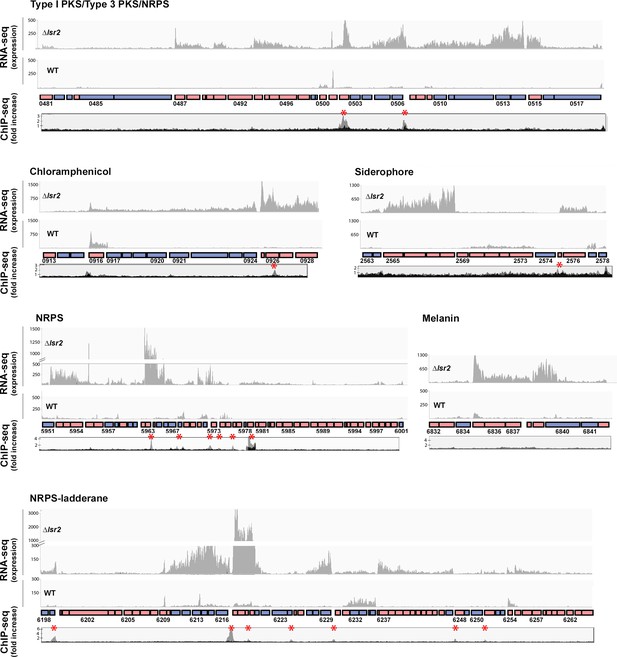
Lsr2 binding sites and expression analysis of select Lsr2-regulated specialized metabolic clusters.
For each of the six specialized metabolic clusters shown, genes oriented in the forward direction are shown as pink boxes, while those in the reverse direction are shown as blue boxes. Select genes are labeled with their corresponding sven numbers (e.g. 0481). RNA-sequencing results are shown above each cluster, with graphs depicting expression levels. For each, the lsr2 read profile is shown on the top, while the wild type profile is shown on the bottom. The ChIP-seq profiles (below the gene cluster) are shown as ‘fold increase’, with the gray profiles indicating regions associated with 3× FLAG tagged Lsr2 (where an anti-FLAG antibody was used for the immunoprecipitation), and the black profile representing the negative control (strain expressing a non-FLAG-tagged Lsr2 variant). Lsr2−3× FLAG binding sites are indicated with a red asterisk.
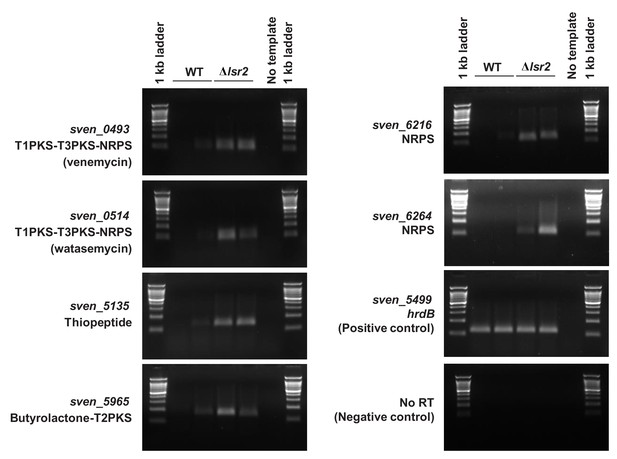
Semi-quantitative RT-PCR using RNA (biological replicates) isolated from S. venezuelae wild type (WT) or ∆lsr2 in early stationary phase.
sven_0493, sven_0514, sven_5135, sven_5965, sven_6216 and sven_6264 are genes predicted to be located within different specialized metabolic gene clusters by antiSMASH 4.0 (Blin et al., 2017). Primers annealing within the constitutively expressed vegetative sigma factor hrdB were used as a positive control for RNA abundance and integrity. No-RT reactions (RNA template) and no-template reactions served as controls for DNA contamination.
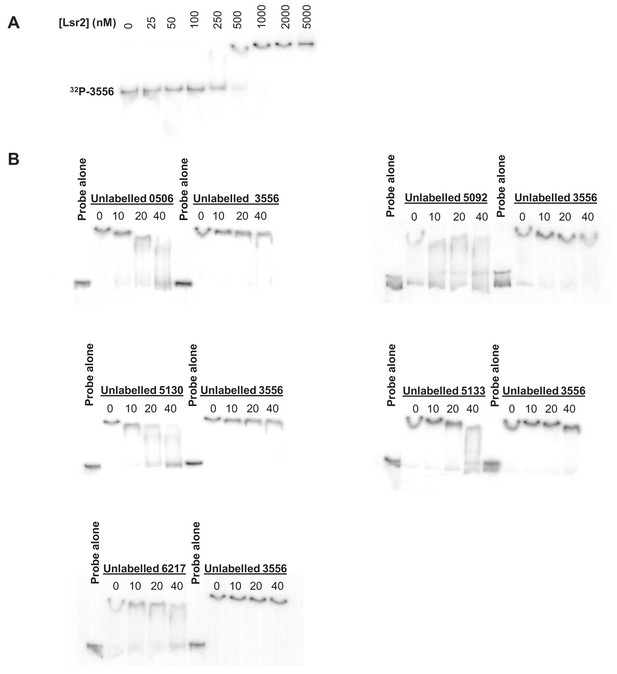
Electrophoretic mobility shift assays with Lsr2.
(A) Lsr2 directly binds labeled DNA probes in vitro, with 1 μM Lsr2 being sufficient to shift 10 nM labeled non-specific probe (where non-specific refers to a sequence not bound by Lsr2 in vivo). (B) Competition EMSAs, in which different unlabeled probes were used to compete for Lsr2 binding (1 μM) with a labeled, non-specific probe (SVEN_3556). Five sequences identified as Lsr2-binding sites in the ChIP-seq experiment (SVEN_5092, SVEN_6217, SVEN_0506, SVEN_5130 and SVEN_5133), alongside a negative control sequence (SVEN_3556 – not an Lsr2 ChIP target), were added at 10, 20 or 40 × excess (40–160 nM, with test probe concentrations being 4 nM). All EMSAs were conducted in triplicate, with representative results shown.

Schematic of the Lsr2-binding site between sven_5106–5107.
(A) DNA content, and relevant transcriptional details for the flanking genes are as detailed below the sequence. (B) Shown in red are the specific mutations made in the wild type sequence shown in (A), to raise the GC-content, without disrupting the promoter regions for either gene.
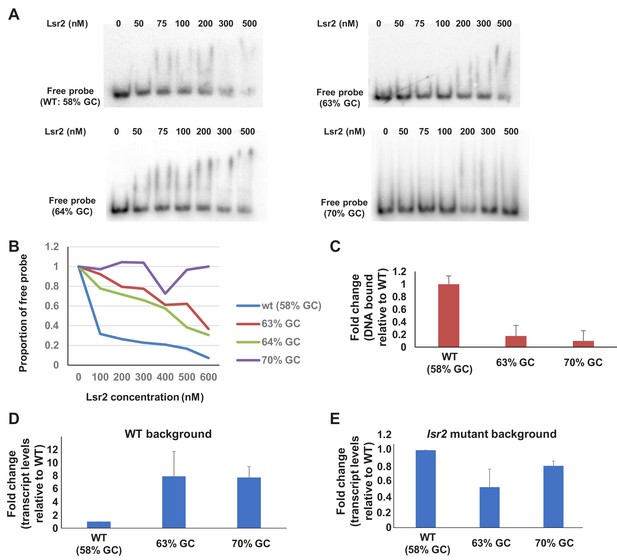
Effect of increasing GC content on Lsr2 binding and activity.
(A) EMSAs performed using probes (1 nM) having increasing GC-content, together with increasing concentrations of Lsr2. Probes corresponded to a 124 bp region between the Lsr2 targeted binding site between sven_5106 and sven_5107, where AT nucleotides (outside of the promoters for the flanking genes) were mutated to increase the GC content. (B) Quantification of free probe remaining following incubation with increasing concentrations of Lsr2, from EMSAs shown in (A). The different probes are represented by the colors indicated on the right. (C) Quantitative PCR of a region corresponding to the beginning of sven_5105, after ChIP (using anti-FLAG-antibodies specific for Lsr2−3× FLAG) to follow Lsr2 binding to a wild type or mutated (increasing GC content) sequence between sven_5106–5107, relative to binding to the wild type sequence. (D) RT-qPCR assessing transcript levels of sven_5106 in a wild type Lsr2-expressing strain (D) bearing wild type or mutant Lsr2 binding sequences between sven_5106–5107. (E) Same as (D), only conducted in an lsr2 mutant background, to ensure that the mutations in the Lsr2 binding site only affected Lsr2 binding, and did not generally enhance promoter activity for sven_5106. All EMSA experiments were performed in duplicate or triplicate, with representative results shown. ChIP-qPCR and RT-qPCR experiments were performed using biological duplicates and technical triplicates.
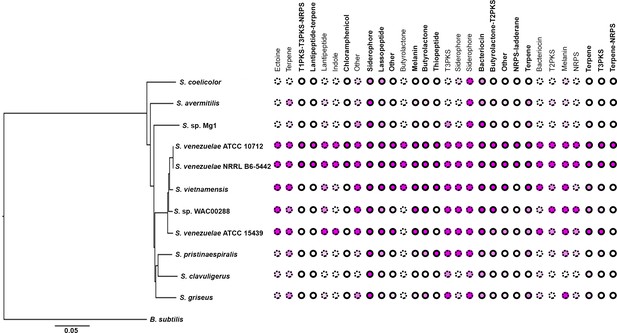
Conservation of specialized metabolic clusters in diverse streptomycetes.
Phylogenetic tree of diverse Streptomyces species, with the relative conservation of each specialized metabolic cluster from S. venezuelae shown in the right. Conservation is based on BLAST analyses, with <20% (white), 20% (light pink), 40% (medium pink) and 60% (dark pink) indicating query coverage and overall degree of cluster conservation. Clusters bound by Lsr2 are indicated with bolded names and solid circles, while those not bound by Lsr2 are depicted with dotted circles.
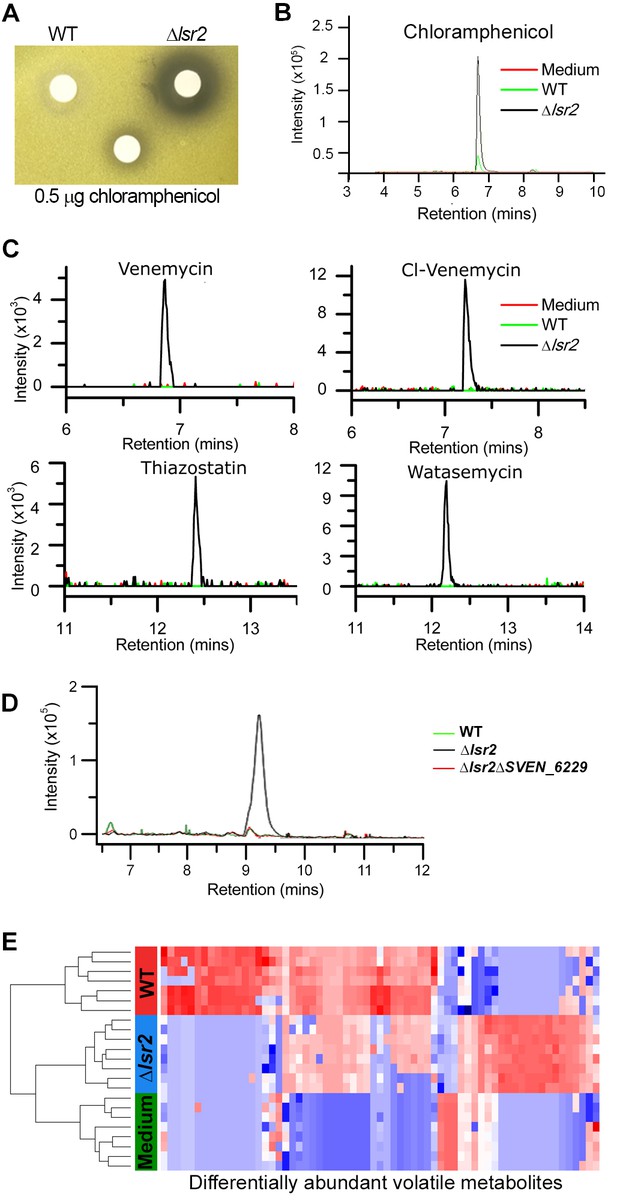
Specialized and volatile metabolite comparisons between wild type and lsr2 mutant strains.
(A) Bioactivity of S. venezuelae extracts against Micrococcus luteus. Wild type and lsr2 mutant strains were cultured for 18 hr prior to extraction in methanol and reconstitution in DMSO. Extracts were applied to Whatman filter discs, alongside a chloramphenicol positive control. (B) Extracted ion chromatogram for chloramphenicol (m/z 321.005), from LC/MS analysis of methanol extracts from wild type and lsr2 mutant cultures grown in MYM liquid medium for 3 d, alongside a medium (MYM) control. (C) Extracted ion chromatograms of [M-H]−=219.040 (venemycin); [M-H]−=252.992 (chlorinated venemycin); [M + H]+ = 353.099 (watasemycin); and [M + H]+ = 339.083 (thiazostatin), from LC/MS analyses of methanol extracts of wild type and lsr2 mutant strains grown for 3 d in MYM liquid medium. (D) Extracted ion chromatogram of m/z 281.068, from LC/MS analysis of methanol extracts of wild type, lsr2 mutant and double lsr2 sven_6229 mutant strains, grown in MYM liquid medium for 3 d. (E) Heat map depicting the 65 volatile compounds (columns) that were significantly different in relative abundance (p<0.05 after BH correction) between wild type (WT, red) and lsr2 mutant (Δlsr2, blue) strains. Sterile media (Media, green) is included for comparison. Cell color corresponds to relative compound abundance after log10-transformation, mean-centering, and unit-scaling, ranging from low abundance (blue) to high abundance (red). Dendrogram (left) was constructed using Euclidean distance as the distance metric.
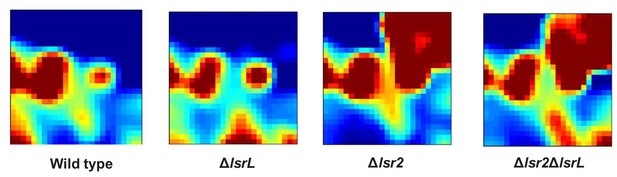
Differential metabolic phenotype heat map for wild type, lsr2, lsrL and lsr2/lsrL mutant strains.
Methanol extracts from triplicate cultures were analyzed by LC/MS. The resulting peaks were averaged and the medium control was subtracted. All metabolites were then compiled into the self-organizing metabolic maps shown here. Red: most abundant; blue: least abundant.
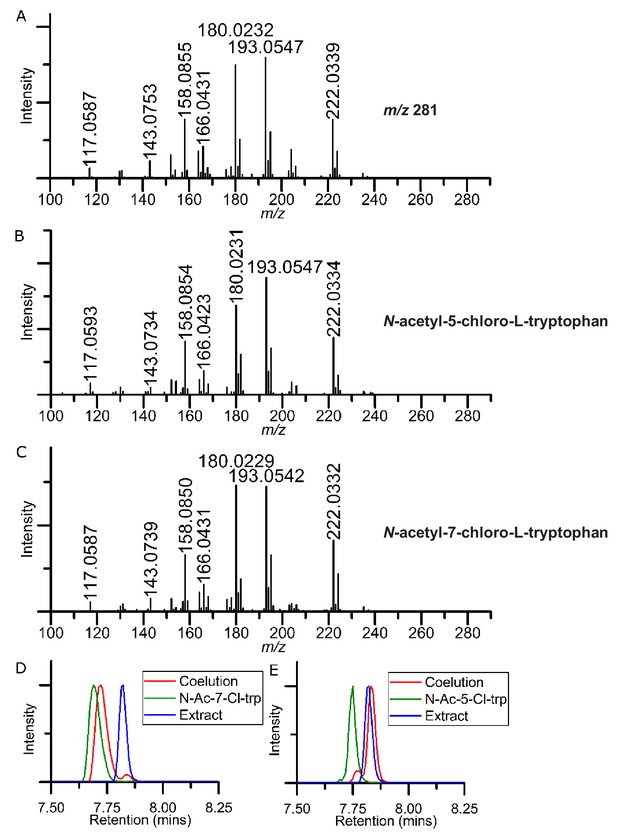
Comparison of unknown m/z 281 species to synthetic N-acetyl-5-chloro-L-tryptophan and N-acetyl-7-chloro-L-tryptophan.
(A) Collision-induced (CID) fragmentation spectrum of m/z 281 from lsr2 mutant strain. (B) CID fragmentation spectrum of N-acetyl-7-chloro-L-tryptophan. (C) CID fragmentation spectrum of N-acetyl-5-chloro-L-tryptophan. (D) Coelution of m/z 281 extracted ion chromatograms of synthetic standard of N-acetyl-7-chloro-L-tryptophan, lsr2 extract, and coelution. (E) Coelution of m/z 281 extracted ion chromatograms of synthetic standard of N-acetyl-5-chloro-L-tryptophan, lsr2 extract, and coelution.
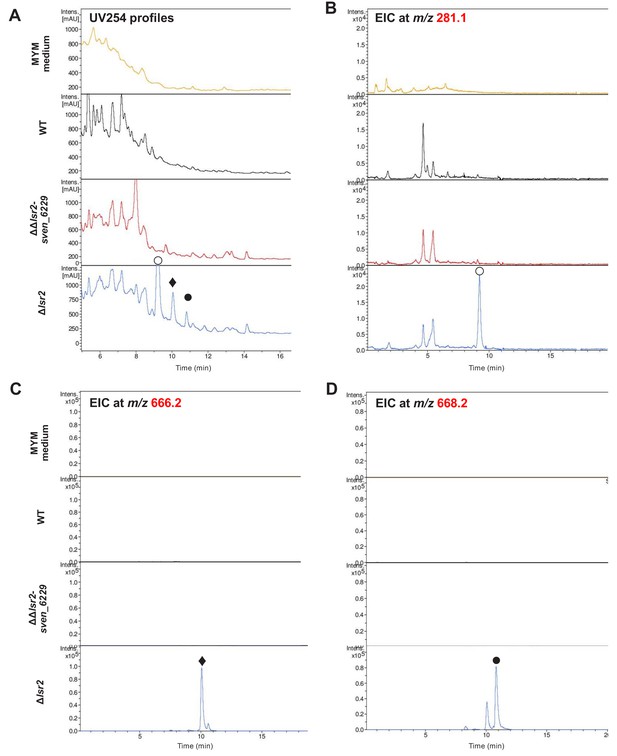
Deletion of a halogenase-encoding gene leads to loss of multiple molecules.
(A) UV254 profiles for medium control (MYM), wild type S. venezuelae (WT), the double lsr2 sven_6229 mutant, and the single lsr2 mutant. In the lsr2 mutant, three new peaks are marked with a filled diamond, and open and filled circles. (B) Positive ion mode xtracted ion chromatogram (EIC) for the open circle peak at m/z 281.1. (C) EIC for the filled diamond peak at m/z 666.2. (D) EIC for the filled circle peak at m/z 668.2.
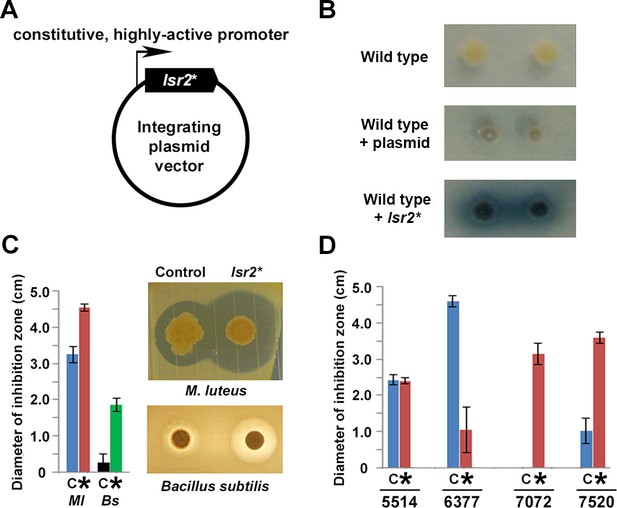
Manipulating Lsr2 activity can stimulate new antibiotic production in diverse Streptomyces.
(A) Lsr2 activity ‘knockdown’ construct, where a DNA-binding defective variant of Lsr2 (lsr2*) is under the control of a constitutive (ermE*) promoter, and is on a plasmid vector bearing an apramycin resistance marker, that is capable of integrating into the chromosomes of most, if not all, streptomycetes. (B) Growth of single Streptomyces coelicolor colonies on Difco nutrient agar. Top: wild type; middle: wild type carrying the empty plasmid; bottom: wild type carrying the plasmid with dominant negative lsr2* variant. (C) Antibiotic bioassay using the wild Streptomyces strain WAC4718, bearing either the control (empty) plasmid (C), or the Lsr2 knockdown construct (*), tested against the indicator strains M. luteus (Ml) or Bacillus subtilis (Bs). Bars indicate standard error (n = 4). (D) Antibiotic bioassay using four different wild Streptomyces strains (WAC5514, WAC6377, WAC7072, and WAC7520), carrying either empty plasmid (C) or the Lsr2 knockdown construct (*), tested against M. luteus as an indicator strain. Bars indicate standard error (n = 4).
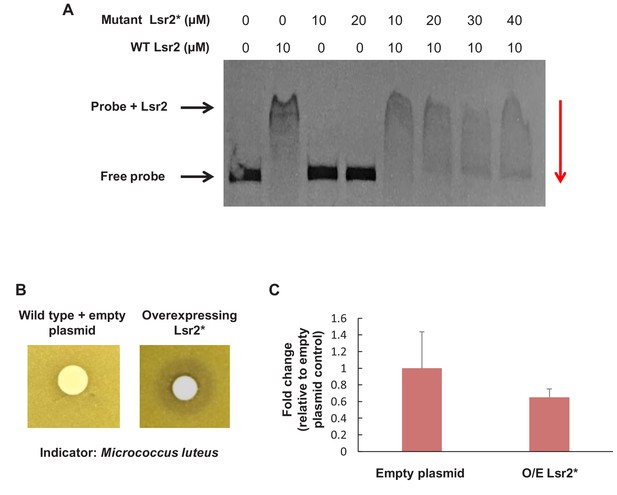
Dominant negative Lsr2 variant (Lsr2*) inhibits DNA binding by wild type Lsr2 and promotes antibiotic production in S. venezuelae.
(A) Electrophoretic mobility shift assay illustrating DNA binding by wild type Lsr2, no DNA binding by the R82A mutant allele (mutant Lsr2*), and effective inhibition of Lsr2 binding by the mutant variant, based on the increasing abundance of ‘free probe’ as mutant concentrations rise. Probe: 1.5 kb region corresponding to the Lsr2 binding site in the chloramphenicol biosynthetic cluster. Reactions were separated on a 1% agarose gel stained with ethidium bromide. (B) Bioactivity of S. venezuelae extracts against Micrococcus luteus. Wild type S. venezuelae carrying either a control (empty) plasmid (left), or one overexpressing the mutant 9R82A) Lsr2 variant (right; Lsr2*) were cultured for 18 hr prior to extraction in methanol and reconstitution in DMSO. Extracts were applied to Whatman filter discs, and cultures were grown overnight. (C) Quantitative PCR analysis of Lsr2-3xFLAG-binding to the validated target site sven_0926, following ChIP, for a strain carrying an empty plasmid (control) or overexpressing Lsr2* (O/E Lsr2*). Overexpressing Lsr2* led to a 40% drop in binding by Lsr2. ChIP experiments were conducted in duplicate, and qPCR was done for each replicate in technical triplicate.
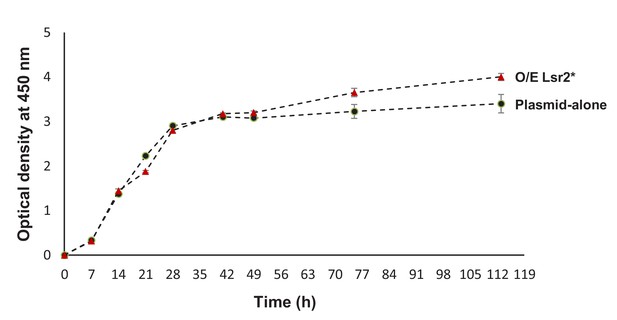
Growth curves of WAC04718 containing either an empty plasmid or one expressing Lsr2*.
Strains were grown in Bennet’s medium for up to 120 hr, with optical density (at 450 nm) readings being taken at regular intervals. Experiments were conducted in duplicate, with standard deviation shown.
Tables
Specialized metabolic clusters and their control by Lsr2.
https://doi.org/10.7554/eLife.47691.012| Predicted specialized metabolic gene cluster# (characterized product) | 1st gene | Last gene | Number of upregulated genes (q-value < 0.01; >4 fold change) | % of upregulated genes in each cluster | Number of Lsr2- associated sites (q-value < 0.01) |
|---|---|---|---|---|---|
| Ectoine | SVEN_0223 | SVEN_0234 | 1 | 8.33 | 0 |
| Terpene | SVEN_0261 | SVEN_0306 | 1 | 0 | 0 |
| T1PKS-T3PKS-NRPS (venemycin/watasemycin/thiazostatin) | SVEN_0463 | SVEN_0531 | 28 | 42.65 | 2 (SVEN_0502*, 506) |
| Lantipeptide – terpene | SVEN_0540 | SVEN_0561 | 0 | 0 | 1 (SVEN_0557) |
| Lantipeptide (venezuelin) | SVEN_0612 | SVEN_0630 | 3 (1 repressed) | 16.67 | 0 |
| Indole (acryriaflavin) | SVEN_0755 | SVEN_0772 | 0 | 0 | 0 |
| Chloramphenicol | SVEN_0913 | SVEN_0928 | 14 | 93.33 | 1 (SVEN_0926) |
| Other | SVEN_1844 | SVEN_1884 | 0 | 0 | 0 |
| Siderophore (desferrioxamine-like) | SVEN_2566 | SVEN_2577 | 5 | 41.67 | 1 (SVEN_2576) |
| Lassopeptide | SVEN_3103 | SVEN_3132 | 2 | 6.90 | 1 (SVEN_3116*−7) |
| Other | SVEN_4061 | SVEN_4110 | 1 | 2.04 | 1 (SVEN_4069–70) |
| Butyrolactone (gaburedin) | SVEN_4179 | SVEN_4189 | 0 | 0 | 0 |
| Melanin | SVEN_4620 | SVEN_4662 | 0 | 0 | 4 (SVEN_4629–30, 4632, 4634–5, 4651) |
| Butyrolactone | SVEN_5076 | SVEN_5111 | 3 | 11.54 | 2 (SVEN_5091–92, 5106*−07) |
| Thiopeptide | SVEN_5119 | SVEN_5145 | 3 | 11.54 | 3 (SVEN_5127–8, 5129–31, 5132–3) |
| T3pks | SVEN_5351 | SVEN_5383 | 0 | 0 | 0 |
| Siderophore | SVEN_5413 | SVEN_5426 | 0 | 0 | 0 |
| Siderophore | SVEN_5471 | SVEN_5482 | (1 repressed) | 9.09 | 0 |
| Bacteriocin | SVEN_5817 | SVEN_5840 | 3 | 15.15 | 1 (SVEN_5817*) |
| Butyrolactone – T2PKS | SVEN_5951 | SVEN_6002 | 19 | 45.10 | 6 (SVEN_5963*−4, 5968*−9, 5972–3, 5974, 5975–6, 5979) |
| Other | SVEN_6112 | SVEN_6204 | 3 (1 repressed) | 3.26 | 1 (SVEN_6199) |
| NRPS-ladderane | SVEN_6134 | SVEN_6282 | 50 | 36.49 | 7 (SVEN_6199, 6216–7, 6219–20, 6225*, 6230, 6247–8, 6251) |
| Terpene | SVEN_6436 | SVEN_6490 | 6 | 16.67 | 1 (SVEN_6458) |
| Bacteriocin | SVEN_6527 | SVEN_6535 | (1 repressed) | 0 | 0 |
| T2PKS | SVEN_6767 | SVEN_6814 | 2 | 4.26 | 0 |
| Melanin | SVEN_6833 | SVEN_6842 | 4 | 44.44 | 0 |
| NRPS | SVEN_7032 | SVEN_7080 | 0 | 0 | 0 |
| Terpene | SVEN_7101 | SVEN_7119 | (two repressed) | 0 | 1 (SVEN_7109–10*) |
| T3PKS | SVEN_7223 | SVEN_7259 | 1 | 2.78 | 2 (SVEN_7235, 7237*−8) |
| Terpene-NRPS | SVEN_7417 | SVEN_7452 | 0 | 0 | 4 (SVEN_7427–8, 7440,7447–9, 7449–50) |
-
#:Cluster prediction by antiSMASH; * Asterisks indicate regulatory genes bound by Lsr2
Bold: clusters containing an Lsr2 binding site
-
Gray shading: clusters containing differentially expressed genes
Additional files
-
Supplementary file 1
Differentially expressed genes in an lsr2 mutant, relative to wild type (qvalue <0.01;>4 fold change).
- https://doi.org/10.7554/eLife.47691.021
-
Supplementary file 2
Transcript levels of known global antibiotic regulatory genes.
- https://doi.org/10.7554/eLife.47691.022
-
Supplementary file 3
ChIP-seq identified Lsr2-FLAG-bound sequences.
- https://doi.org/10.7554/eLife.47691.023
-
Supplementary file 4
(A) Number of core binding sites and AT-rich 20 nt stretches within Lsr2 binding sequences. (B) Start and end positions within the chromosome, for AT-rich 'core' sequences (5 of 6 A/T residues). (C) Position and sequence of AT-rich 20 nt stretches. (D) Number of AT-rich core and 20-nt stretches in random 500 bp and 1000 bp sequences
- https://doi.org/10.7554/eLife.47691.024
-
Supplementary file 5
Highly abundant (intensities > 105) unique or significantly upregulated (>5 fold) compounds.
- https://doi.org/10.7554/eLife.47691.025
-
Supplementary file 6
Putative identification of the most abundant, differentially expressed volatile compounds by wild type and Δlsr2 strains of S. venezuelae.
- https://doi.org/10.7554/eLife.47691.026
-
Supplementary file 7
Bacterial strains, plasmid and cosmids used in this work.
- https://doi.org/10.7554/eLife.47691.027
-
Supplementary file 8
Oligonucleotides and synthetic DNA used in this study.
- https://doi.org/10.7554/eLife.47691.028
-
Transparent reporting form
- https://doi.org/10.7554/eLife.47691.029