Neural stem cell temporal patterning and brain tumour growth rely on oxidative phosphorylation
Figures
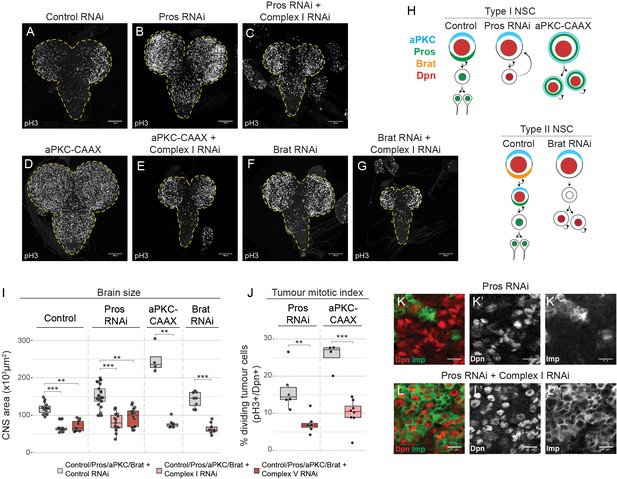
Brain tumours require OxPhos for growth.
(a–g) phospho Histone H3 (pH3) staining in the CNS of third instar larvae (L3) with NSC-specific expression (Wor-GAL4;Tub-GAL80ts) of control RNAi (a), Pros-RNAi (b,c), aPKC-CAAX (d,e) or Brat-RNAi (f, g), either without (b,d,f) or with (c,e,g) RNAi against a complex I subunit (NDUFS1). Maximum intensity projections through the entire CNS; dashed lines outline the CNS. (h) NSC lineages before and after tumourigenic transformation. (i,j) Brain size (i) and mitotic index of Dpn+ tumour cells (j) from L3 larvae expressing the indicated transgenes in NSCs. Datapoints indicate individual brains from one to four biological replicates. (k,l) Dpn (red, k’,l’) and Imp (green, k’’,l’’) immunostaining in Pros-RNAi tumours, without (k) or with (l) a complex I RNAi. Scale bars are 100 µm (a–g) or 10 µm (k,l).
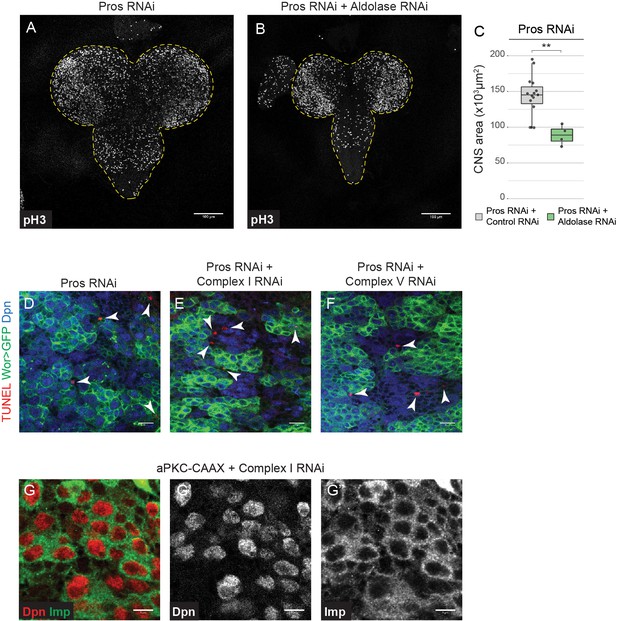
Brain tumours require OxPhos and glycolysis for their growth.
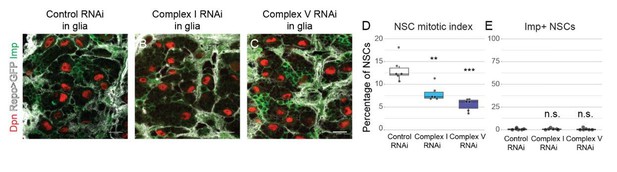
(a,b) pH3 in the CNS of L3 larvae. Maximum intensity projections through the CNS; dashed lines outline the CNS. (c) Brain size (area of CNS, maximum intensity projections) from L3 larvae. Datapoints indicate individual brains from one biological replicate. (d–f) TUNEL, GFP (NSCs, Wor-GAL4 >mCD8 GFP) and Dpn (NSCs) staining in the VNC of L3 larvae with NSC-specific expression of the indicated transgenes; arrowheads indicate TUNEL-positive cells. (g) Dpn and Imp in aPKC-CAAX tumours with NSC-specific expression of complex I RNAi. Scale bars are 100 µm (a,b), 10 µm (d–f) or 5 µm (g).
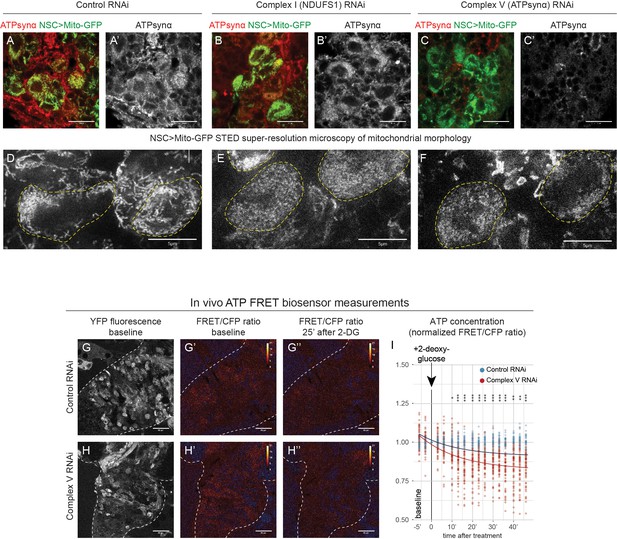
OxPhos RNAi in NSCs affects mitochondrial function.
(a–c) ATPsynα (mitochondria, Complex V subunit) and GFP (mitochondria in NSCs and their progeny; Wor-GAL4 >Mito GFP) staining in the VNC of L3 larvae with NSC-specific expression of the indicated RNAi. Complex V RNAi (c) targets the Drosophila ATPsynα. (d–f) Representative STED super-resolution images of NSCs expressing mitochondria targeted GFP (Mito-GFP) and the indicated RNAi; dashed lines outline NSCs, the other cells are neuronal progeny with perdurance of Mito-GFP. (g–h) Measurement of ATP specifically in NSCs expressing a genetically encoded ATP FRET sensor together with the indicated RNAi. Baseline YFP fluorescence (g,h); pseudocoloured FRET/CFP ratio images are shown at baseline (g’,h’) and 25 min after application of 200 mM 2-deoxyglucose (2-DG) to inhibit glycolysis (g’’,h’’); dashed lines outline the CNS. (i) Quantification of FRET/CFP ratio over time; datapoints indicate individual NSCs from four brains in one biological replicate; the trendline indicates an exponential model fitted to the values. Findings were confirmed in two additional biological replicates (data not shown). Scale bars are 5 µm (d–f), 10 µm (a–c) or 50 µm (g,h).
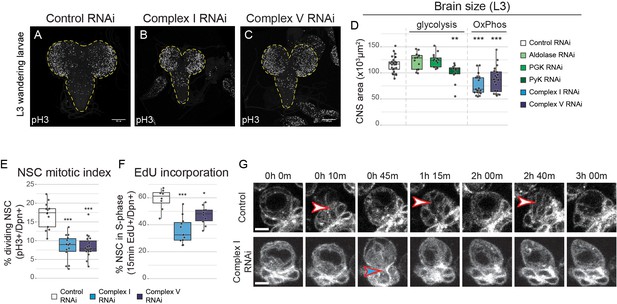
OxPhos inhibition decreases NSC proliferation.
(a–c) pH3 staining in the CNS of L3 larvae. Maximum intensity projections through the entire CNS; dashed lines outline the CNS. (d) Brain size from L3 larvae. (e,f) Mitotic index (e) and 15 min EdU incorporation (f) in NSCs expressing the indicated RNAi (Wor-GAL4;Tub-GAL80ts). (g) Stills from time-lapse imaging of NSCs (Figure 2—video 1) in the early third instar larval VNC with NSC-specific expression of GFP or Complex I RNAi. Arrowheads indicate mitoses of selected NSCs. Datapoints indicate individual brains from four (e), one (f) and two to four (d) biological replicates. Scale bars are 5 µm (g) or 100 µm (a–c).
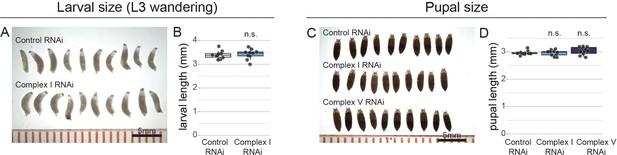
OxPhos inhibition does not affect body size.
Wandering larvae (a,b) or pupae (c,d) with NSC-specific expression of the indicated RNAi. Length is shown in millimetres (mm). Datapoints indicate individual organisms from one biological replicate. Scale bars are 5 mm.
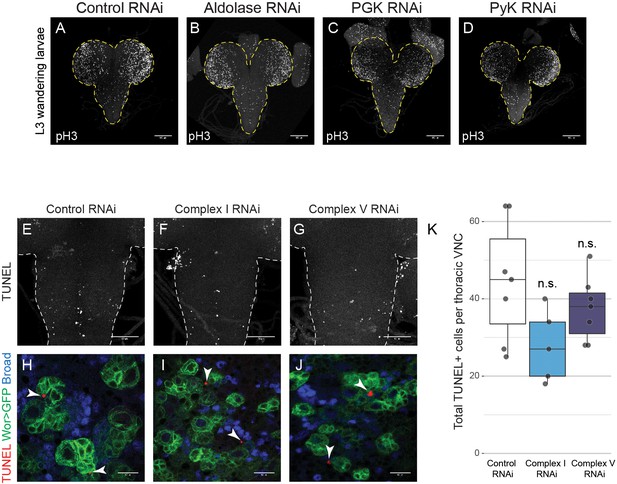
OxPhos inhibition does not increase apoptosis.
(a–d) pH3 staining in the CNS of L3 larvae with NSC-specific expression of the indicated RNAi. Maximum intensity projections through the entire CNS; dashed lines outline the CNS. (e–j) TUNEL, GFP (NSCs, Wor-GAL4 >mCD8 GFP) and Broad staining in the VNC of L3 larvae. Maximum intensity projections through the VNC (e–g); dashed lines outline the CNS; arrowheads indicate TUNEL-positive cells. (k) Total number of TUNEL-positive cells in the thoracic VNC. Datapoints indicate individual brains from one biological replicate. Scale bars are 100 µm (a–d), 50 µm (e–g) or 10 µm (h–j).
Mitochondrial dysfunction increases cell cycle length.
Time-lapse imaging of NSCs in the early third instar larval VNC (48 hr ALH at 29°C) expressing either two copies of UAS-mCD8-GFP (Control, left), or one copy of UAS-mCD8-GFP and UAS-ND75-TRIP (Complex I RNAi, right) under the control of Wor-GAL4. Images were taken every 5 min over a period of 3 hr. Red arrows indicate mitoses of selected NSCs. Scale bars are 20 µm.
Mitochondrial dysfunction increases cell cycle length.
Time-lapse imaging of NSCs in the late third instar larval VNC (wandering, 72 hr ALH at 29°C) expressing either one copy of UAS-mCD8-GFP (Control, left), or one copy of UAS-mCD8-GFP and UAS-ND75-TRIP (Complex I RNAi, right) under the control of Wor-GAL4. Images were taken every 3 min and 50 s over a period of 3 hr. Red arrows indicate mitoses of selected NSCs. Scale bars are 20 µm.
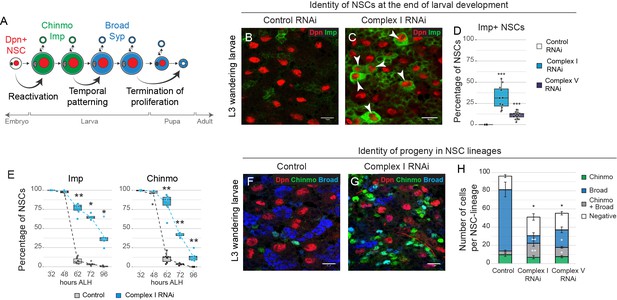
OxPhos is required for temporal patterning of NSC and their progeny.
(a) Scheme of the major temporal transitions in larval NSCs. (b–d) Dpn and Imp expression in the VNC of L3 larvae. Arrowheads indicate Imp-positive NSCs. (e) Percentage of Dpn-positive NSCs in the thoracic VNC that express the indicated temporal marker at different time points after larval hatching (ALH) at 25°C. (f,g) Dpn (NSCs), Chinmo and Broad in the VNC of L3 larvae. (h) Absolute number of cells per NSC lineage in the VNC that express Chinmo or Broad; graph indicates mean + /- s.e.m. of 6 clones. Datapoints indicate individual brains from four (d), two (e) and one (h) biological replicates. Scale bars are 10 µm.
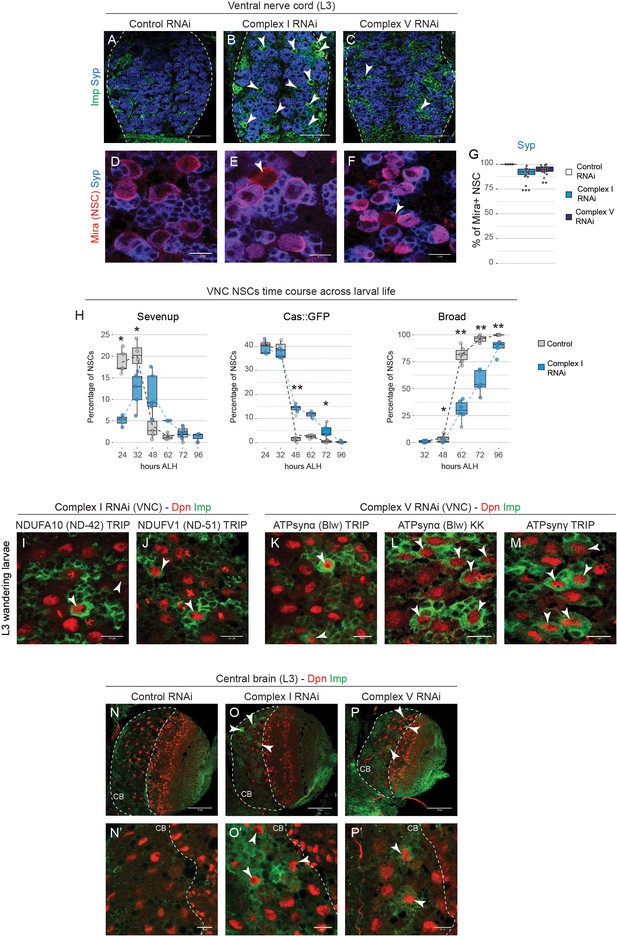
Mitochondrial dysfunction in NSCs delays temporal patterning.
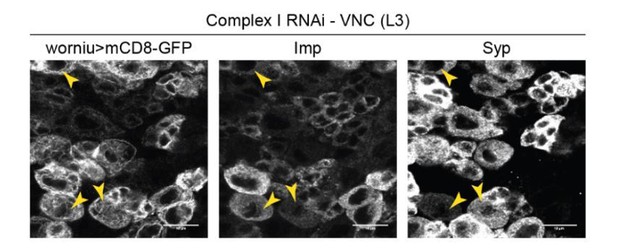
(a–c) Overview of Imp and Syp in the thoracic VNC of L3 larvae. Single confocal sections at comparable depths through the ventral side of the VNC; dashed lines outline the VNC; arrowheads indicate Imp-positive, Syp-negative NSCs. (d–g) Mira and Syp expression in the VNC of L3 larvae. Arrowheads indicate Syp-negative NSCs. (h) Percentage of Dpn- or Mira-positive NSCs in the thoracic VNC that express the indicated temporal marker at different time points after larval hatching (ALH) at 25°C. (i–m) Dpn and Imp in the VNC of L3 larvae. Arrowheads indicate Imp-positive NSCs. (n–p) Dpn and Imp in the CB of L3 larvae; (n’–p’) higher magnifications of the same genotypes as in (n–p). Single confocal sections at comparable depths through the ventral side of the CB; dashed lines outline the CB; arrowheads indicate Imp-positive NSCs. Datapoints indicate individual brains from two (g) or one (h) biological replicates. Scale bars are 50 µm (a–c,n–p) or 10 µm (d–f,i–m,n’–p’).
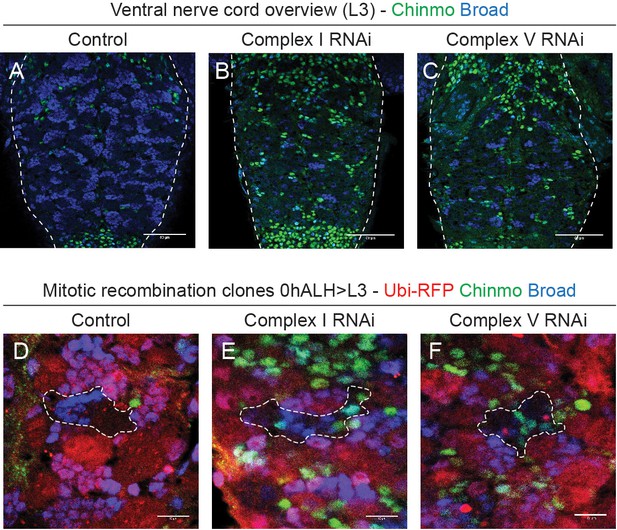
Delayed temporal patterning of NSCs affects their progeny.
(a–c) Overview of Chinmo and Broad in the thoracic VNC of L3 larvae. Single confocal sections at comparable depths through the ventral side of the VNC; dashed lines outline the VNC. (d–f) RFP (clones, negatively marked), Chinmo and Broad in the VNC of L3 larvae. Representative single confocal sections through a clone; dashed lines indicate the outline of a clone. Scale bars are 50 µm (a–c) or 10 µm (d–f).
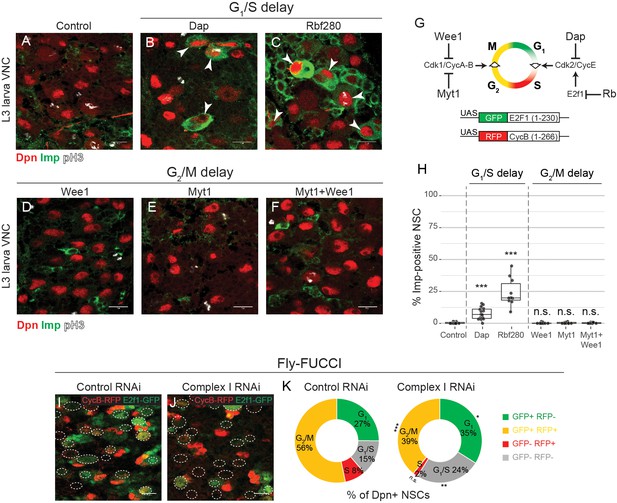
G1/S progression drives temporal patterning.
(a–f) Dpn (NSCs), pH3 (mitosis) and Imp in the VNC of L3 larvae after NSC-specific expression of the indicated transgene. Arrowheads indicate Imp-positive NSCs. (g) Scheme depicting activity of the regulators of the G1/S and G2/M transitions that are used for misexpression in this study, and the Fly-FUCCI transgenes. (h) Percentage of Dpn-positive NSCs that express Imp in L3 larvae. (i–k) L3 larvae with NSC-specific expression of the Fly-FUCCI system, together with control RNAi or complex I RNAi. Outlines indicate Dpn-positive nuclei (i,j). The percentage of Dpn-positive NSCs in the VNC that are positive for either GFP (G1), RFP (S), a combination of GFP and RFP (G2/M) or none (G1/S transition); graphs indicate mean of 8 and 9 brains from one biological replicate (k). Datapoints indicate individual brains from two or three biological replicates (h). Scale bars are 10 µm.
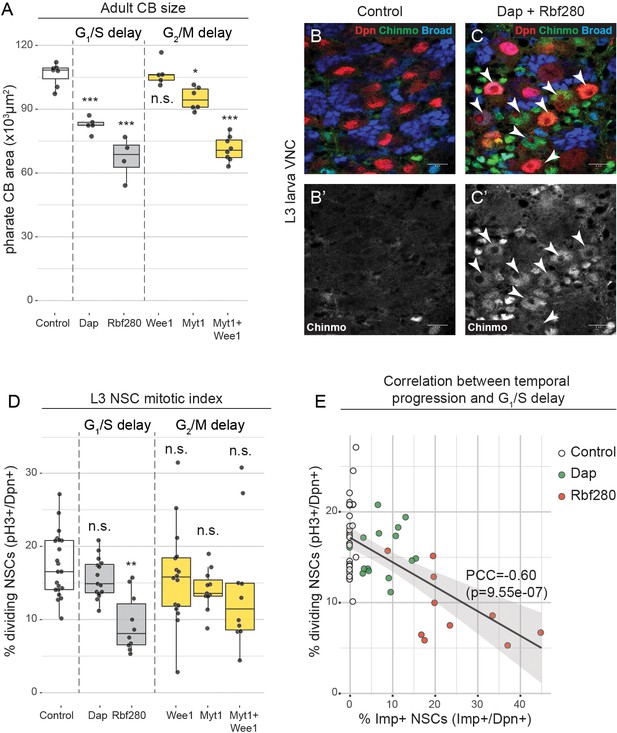
G1/S and G2/M delay results in smaller brains.
(a) size (area of CNS maximum intensity projections) of pharate adult CB. (b,c) Immunostaining for Dpn (NSCs), Chinmo and Broad in the VNC of wildtype L3 larvae or larvae with NSC-specific expression of Dap and Rbf280. Arrowheads indicate Chinmo-positive NSCs. (d) Mitotic index (percentage of pH3+ cells among all Dpn+ cells) of larval VNC NSCs. (e) Correlation between mitotic index and temporal delay with linear regression and 95% confidence interval. Pearson’s correlation coefficient = −0.6058; p-value=9,55e-07. Datapoints indicate individual brains from one to two biological replicates (a,d,e). Scale bars are 10 µm.
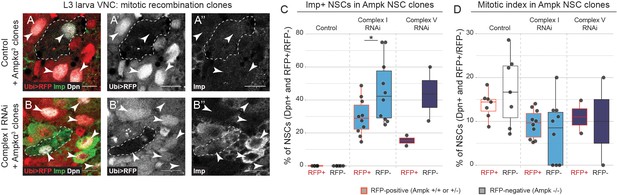
AMPK deletion does not rescue the temporal patterning defect caused by OxPhos inhibition.
(a,b) Dpn (NSCs), RFP (negatively marked AMPK-/- clones) and Imp in the VNC of L3 larvae after heatshock at 0hALH. Arrowheads indicate Dpn-positive NSCs. Dashed outlines mark RFP-negative clones. RNAi is not affected by mitotic recombination and is present both in RFP-positive and RFP-negative cells. (c,d) Percentage of all Dpn-positive and either RFP-positive (AMPK+/- or AMPK+/+; red outline) or RFP-negative (AMPK-/-; black outline) NSCs in the VNC that are Imp-positive (c) or pH3-positive (d). Datapoints indicate individual brains from one biological replicate. n = 123.1 ± 3.2; 11.7 ± 1.1; 122.1 ± 4.9; 8.2 ± 1.0; 127.5 ± 4.3; 8 ± 1.7 (average ± s.e.m) NSCs per brain of the respective genotypes. Scale bars are 10 µm.
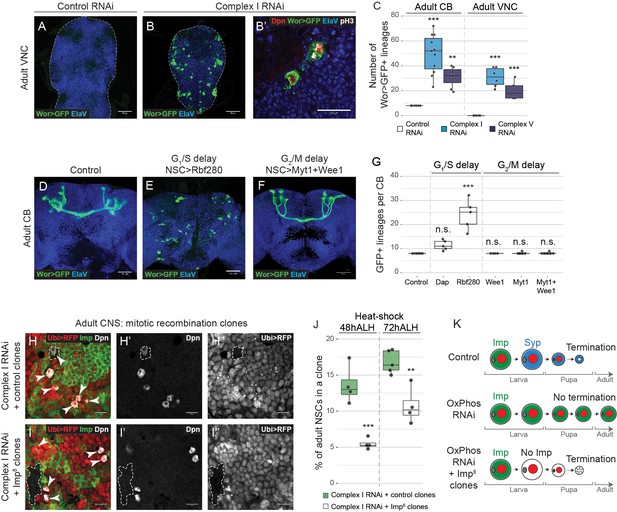
NSCs require OxPhos for termination of proliferation.
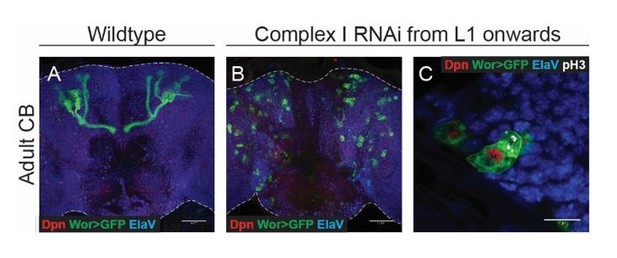
(a–g) ElaV (neurons), GFP (NSCs, Wor-GAL4 >mCD8 GFP), Dpn (NSCs) and pH3 (mitosis) in the pharate adult CB or VNC. Maximum intensity projections through the CB or VNC; dashed lines mark the outline of the CNS. (c,g) Total number of GFP-expressing NSCs in the pharate adult CB or VNC. (h,i) Dpn (NSCs), RFP (negatively marked clones) and Imp in the pharate adult CNS. Arrowheads indicate Dpn-positive NSCs. Dashed outlines mark RFP-negative clones. (j) Percentage of all Dpn-positive NSCs in the pharate adult CNS (CB and VNC) that are part of an RFP-negative clone. (k) OxPhos inhibition prevents terminal differentiation; this is rescued by timely removal of Imp. Datapoints indicate individual brains (c,g) or clones (j) from one biological replicate. Scale bars are 50 µm (a,b), 20 µm (b’) or 10 µm (h, i).
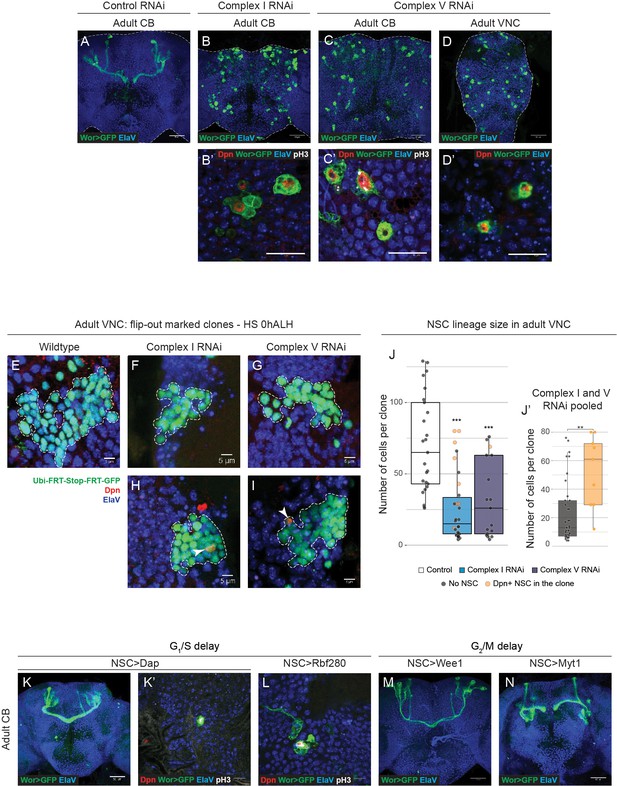
Adult neurogenesis upon OxPhos knockdown and G1/S delay.
(a–d) ElaV (neurons), GFP (NSCs, Wor-GAL4 >mCD8 GFP), Dpn (NSCs) and pH3 (mitosis) in the pharate adult CB (a–c) or VNC (d). Maximum intensity projections through the CB or VNC; dashed lines outline the CNS. (e–i) Dpn (NSCs), GFP (clones) and ElaV (neurons) in the pharate adult VNC. Arrowheads indicate Dpn-positive NSCs within the clone; dashed lines mark the outline of clones. Maximum intensity projections of z-stacks through the clone. (j) Number of cells per GFP-positive clone in the pharate adult VNC. Only those clones of ≥2 cells were quantified. Datapoints indicate single clones from three to five CNSs from one biological replicate; yellow datapoints indicate presence of a Dpn+ NSC in the clone. (j’) Clones from both complex I or V RNAi-expressing NSCs which still contain a Dpn+ NSC at pharate adult stage (yellow) are on average larger than those that do not (grey). (k–n) ElaV (neurons) and GFP (NSCs, Wor-GAL4 >mCD8 GFP), Dpn (NSCs) and pH3 (mitosis) in the pharate adult CB. Maximum intensity projections through the CB. Scale bars are 50 µm (a–d,k–n), 20 µm (b’,c’,d’), 10 µm (k’,l) or 5 µm (e–i).
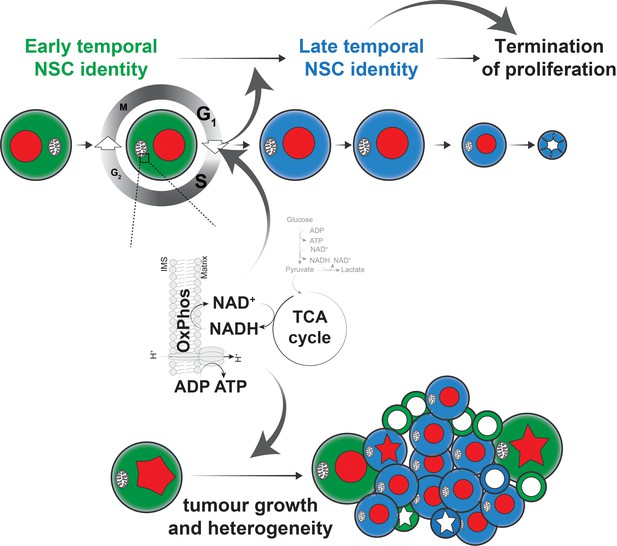
Model of the role of OxPhos in Drosophila NSCs and tumour cells.
We propose a model, whereby highly proliferative Drosophila NSCs also rely on OxPhos for most aspects of their behaviour. In particular, the G1/S transition depends on OxPhos activity and perturbation of this transition, either directly, or indirectly through OxPhos inhibition, results in delayed temporal patterning. This in turn prevents NSCs from terminating proliferation at the appropriate time, causing neurogenesis to persist into the adult. A similar dependence on OxPhos can be seen in brain tumours, where both proliferation and differentiation require mitochondrial activity, presumably through a similar mechanism to that found in normal NSCs.
Tables
| Reagent type (species) | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (D. melanogaster) | mCherry-TRIP | BDSC | RRID: BDSC_35785 | Control RNAi |
| Genetic reagent (D. melanogaster) | w1118;+;+ | BDSC | RRID: BDSC_3605 | |
| Genetic reagent (D. melanogaster) | ND75-TRIP | BDSC | RRID: BDSC_33911 | Complex I RNAi |
| Genetic reagent (D. melanogaster) | Blw-TRIP | BDSC | RRID: BDSC_28059 | Complex V RNAi |
| Genetic reagent (D. melanogaster) | Pros-TRIP | BDSC | RRID: BDSC_42538 | |
| Genetic reagent (D. melanogaster) | Brat-TRIP | BDSC | RRID: BDSC_28590 | |
| Genetic reagent (D. melanogaster) | ND42-TRIP | BDSC | RRID: BDSC_32998 | |
| Genetic reagent (D. melanogaster) | ND51-TRIP | BDSC | RRID: BDSC_36701 | |
| Genetic reagent (D. melanogaster) | Blw-RNAi-KK | VDRC | 34663 | |
| Genetic reagent (D. melanogaster) | ATPsynβ-TRIP | BDSC | RRID: BDSC_28056 | |
| Genetic reagent (D. melanogaster) | ATPsynγ-TRIP | BDSC | RRID: BDSC_28723 | |
| Genetic reagent (D. melanogaster) | ATPsynO-TRIP | BDSC | RRID: BDSC_43265 | |
| Genetic reagent (D. melanogaster) | PFK-TRIP | BDSC | RRID: BDSC_34336 | |
| Genetic reagent (D. melanogaster) | Aldolase-TRIP | BDSC | RRID: BDSC_26301 | |
| Genetic reagent (D. melanogaster) | PyK-TRIP | BDSC | RRID: BDSC_35218 | |
| Genetic reagent (D. melanogaster) | PGK-RNAi-KK | VDRC | 110081 | |
| Genetic reagent (D. melanogaster) | UASp-EGFP-Myt1 | BDSC | RRID: BDSC_65393 | |
| Genetic reagent (D. melanogaster) | UASt-dWee1 | (Price et al., 2002) PMID: 12072468 | ||
| Genetic reagent (D. melanogaster) | UASt-Dap | (Lane et al., 1996) PMID: 8980229 | ||
| Genetic reagent (D. melanogaster) | UAS-Rbf-280 | (Duman-Scheel et al., 2002) PMID: 12015606 | ||
| Genetic reagent (D. melanogaster) | UASt-aPKC.CAAXWT | (Lee et al., 2006; Sotillos et al., 2004) PMID: 16357871, 15302858 | ||
| Genetic reagent (D. melanogaster) | UAS-mito-HA-GFP,e1 | BDSC | RRID: BDSC_8443 | |
| Genetic reagent (D. melanogaster) | UAS-AT1.03-NL on III | (Tsuyama et al., 2013) PMID: 23875533 | ||
| Genetic reagent (D. melanogaster) | UAS-AT1.03-RK on III | (Tsuyama et al., 2013) PMID: 23875533 | ||
| Genetic reagent (D. melanogaster) | UAS-GFP-E2F1.1–230, UAS-mRFP1-NLS-CycB.1–266 | (Zielke et al., 2014) PMID: 24726363 | Fly FUCCI | |
| Genetic reagent (D. melanogaster) | Worniu-GAL4 on II | (Albertson et al., 2004) PMID: 15536119 | ||
| Genetic reagent (D. melanogaster) | Cas::GFP FlyFos line | VDRC | 318476 | |
| Genetic reagent (D. melanogaster) | Ubi-FRT-Stop-FRT-GFP | BDSC | RRID: BDSC_32251 | |
| Genetic reagent (D. melanogaster) | Imp8 | (Munro et al., 2006) PMID: 16476777 | Imp mutant | |
| Genetic reagent (D. melanogaster) | Ampkα3 | (Haack et al., 2013) PMID: 24337115 | AMPK mutant | |
| Antibody | rat anti-PH3 (monoclonal) | Abcam | ab10543 RRID: AB_2295065 | IF, 1/500 |
| Antibody | rabbit anti-PH3 (polyclonal) | Merck Millipore | 06–570 RRID: AB_310177 | IF, 1/500 |
| Antibody | guinea pig anti-Dpn (polyclonal) | James Skeath | IF, 1/10,000 | |
| Antibody | rabbit anti-Imp (polyclonal) | (Geng and Macdonald, 2006) PMID: 17030623 | IF, 1/600 | |
| Antibody | guinea pig anti-Syp (polyclonal) | (McDermott et al., 2012) PMID: 23213441 | IF, 1/1000 | |
| Antibody | chicken anti-GFP (polyclonal) | Abcam | ab13970 RRID: AB_300798 | IF, 1/2000 |
| Antibody | rat anti-Mira (polyclonal) | Chris Doe | IF, 1/500 | |
| Antibody | rat anti-Chinmo (polyclonal) | (Wu et al., 2012) PMID: 22814608 | IF, 1/500 | |
| Antibody | mouse anti-Broad (monoclonal) | DSHB | 25E9.07 | IF, 1/100 |
| Antibody | rabbit anti-RFP (polyclonal) | Abcam | ab62341 RRID: AB_945213 | IF, 1/500 |
| Antibody | rat anti-ElaV (monoclonal) | DSHB | 7E8A10 | IF, 1/100 |
| Antibody | mouse anti-Sevenup (polyclonal) | (Kanai et al., 2005) PMID: 15691762 | IF, 1/200 | |
| Antibody | mouse anti-ATPsynα (monoclonal) | Abcam | ab14748 RRID: AB_301447 | IF, 1/100 |
| Antibody | GFP-booster Atto647N | Chromotek | gba647n RRID: AB_2629215 | IF, 1/500 for STED |
| Commercial assay or kit | ApopTag Red In Situ Apoptosis Detection kit | Merkc Millipore | S7165 | |
| Commercial assay or kit | Click-iT EdU Alexa Fluor 647 Imaging Kit | Invitrogen | C10340 | |
| Chemical compound, drug | 2-deoxyglucose | Sigma | D8375 | 200 mM final concentration |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.47887.019