Functional divergence of Plexin B structural motifs in distinct steps of Drosophila olfactory circuit assembly
Figures
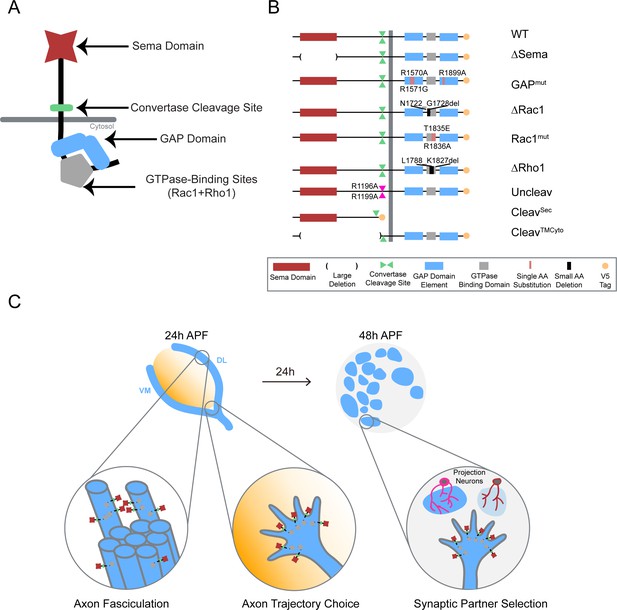
Systematic mutagenesis of PlexB structural motifs and functional interrogation in the stepwise assembly of the Drosophila olfactory map.
(A) The PlexB protein consists of several conserved structural motifs, including an extracellular Sema domain, a juxtamembrane convertase cleavage site, a cytoplasmic GTPase-binding region for Rac1 and Rho1, and a cytoplasmic bipartite GAP domain. (B) Schematic summary of PlexB variants generated in this study. Each variant encodes either a mutated form of PlexB with one structural motif disrupted or a cleaved product of PlexB. (C) In the developing antennal lobe, ORN axons first fasciculate into bundles. Each ORN axon chooses a defined trajectory along the edge of the antennal lobe, in part responding to the extracellular Sema-2a/2b gradients (orange). Subsequently, ORN axons innervate the antennal lobe to interact with dendrites of prospective projection neuron partners and thus establish specific synaptic connections. PlexB participates in all these processes (Li et al., 2018b), providing an in vivo platform for examining the functionality of PlexB variants in multiple, distinct wiring steps.
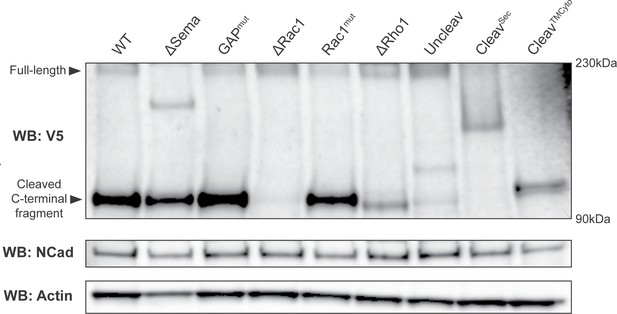
Western blot showing that PlexB variants are stably expressed in vivo.
Transgenically expressed PlexB variants, driven by pan-neural C155-GAL4, were extracted from brains and ventral nerve cords of third-instar larvae. Full-length PlexB and cleaved C-terminal fragments were detected by the C-terminal V5 tag. As in our previous observation (Li et al., 2018b), only a small fraction of PlexB proteins exist in the full-length form. Controls consisting of N-cadherin and actin were blotted after stripping the membrane.

Homology alignment of Plexins from different species and sub-families showing the conserved arginine residues (highlighted in red) of the GAP domain.
UniProt was used for alignment. Asterisk, identical residue; colon, strongly similar residue (scoring >0.5 in the Gonnet PAM 250 matrix); period, weakly similar residue (scoring =< 0.5 in the Gonnet PAM 250 matrix).
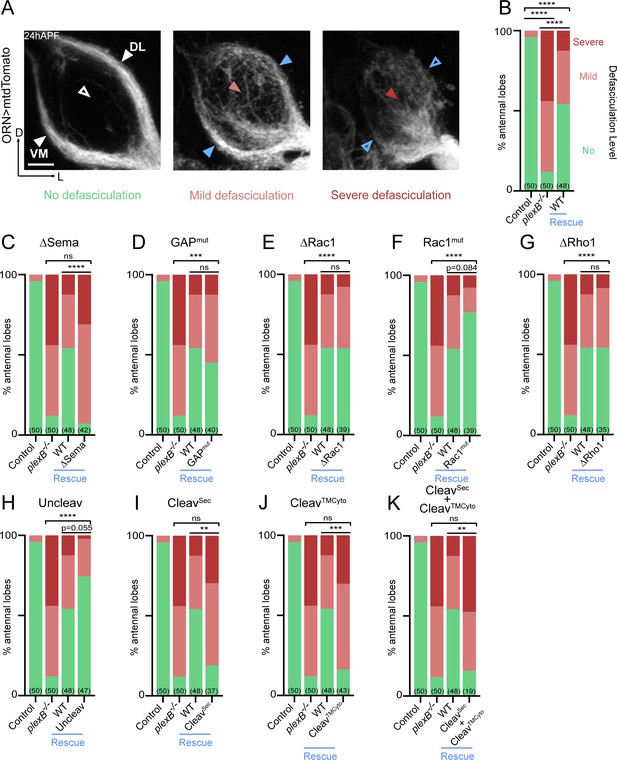
Axon fasciculation requires full-length PlexB but not its cytoplasmic motifs individually.
(A) In a wild-type fly brain at 24hAPF, ORN axons fasciculate into two bundles (left panel; white arrowheads) surrounding the antennal lobe without innervating it (left panel; empty white arrowhead). Loss of PlexB (plexB–/–) causes defasciculation of ORN axons with differing severity (middle and right panels; red arrowheads). In the severe cases, axon bundles are completely missing (right panel; empty blue arrowheads). ORN axons were labeled by pan-ORN Peb-GAL4 (Sweeney et al., 2007) driven mtdTomato expression. (B) Quantification of fasciculation defects by binning antennal lobes into three categories – no, mild, and severe defasciculation. Expressing wild-type PlexB in ORNs significantly but not completely restores ORN axon fasciculation in plexB mutant flies. ‘Rescue’ hereafter denotes ORN-specific expression of PlexB variants in plexB–/– flies. (C–K) Quantification of fasciculation defects in ORN-specific rescue experiments with respective PlexB variants. Sample sizes are noted in parentheses. Significance of the contingency tables in Figure 2B–K was determined by Fisher’s exact test. ns, not significant; **p<0.01; ***p<0.001; ****p<0.0001. Images are shown as maximum z-projections of confocal stacks. Scale bars, 10 µm. Axes, D (dorsal), L (lateral).
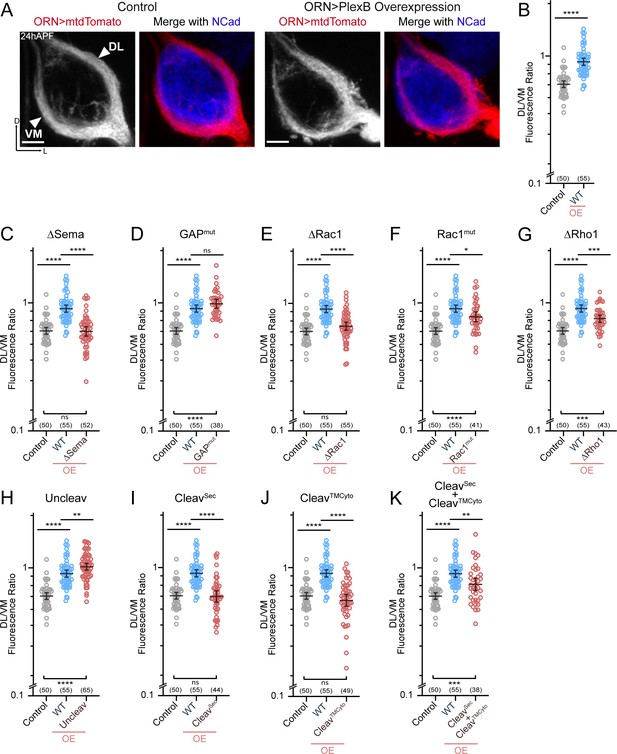
ORN trajectory choice requires both extracellular and cytoplasmic modules of PlexB.
Both full-length and reconstituted fragments of PlexB transduce signal in trajectory choice. (A) In wild-type pupal brains at 24hAPF, ORN axons form the dorsolateral (DL) and ventromedial (VM) trajectories circumnavigating the antennal lobe (left panels). Overexpression of PlexB in ORNs shifts ORN axons to the DL trajectory (right panels). ORN axons were labeled by pan-ORN Peb-GAL4 (Sweeney et al., 2007) driven mtdTomato expression. Antennal lobes were co-stained with a neuropil marker N-cadherin (NCad). (B) Fluorescence intensity ratios of ORN axon trajectories (DL/VM) in wild-type and PlexB overexpression brains at 24hAPF. Geometric means: control, 0.68; WT OE, 0.94. ‘OE’ hereafter denotes ORN-specific overexpression of PlexB variants. (C–K) Fluorescence intensity ratios of ORN axon trajectories (DL/VM) for respective PlexB variants. Geometric means: ∆Sema, 0.68; GAPmut, 1.01; ∆Rac1, 0.73; Rac1mut, 0.84; ∆Rho1, 0.81; Uncleav, 1.04; CleavSec, 0.69; CleavTMCyto, 0.65; CleavSec + CleavTMCyto, 0.82. Sample sizes are noted in parentheses. Significance among multiple groups in Figure 3B–K was determined by one-way ANOVA with Tukey’s test for multiple comparisons. ns, not significant; *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001. Images are shown as maximum z-projections of confocal stacks. Scale bars, 10 µm. Axes, D (dorsal), L (lateral).
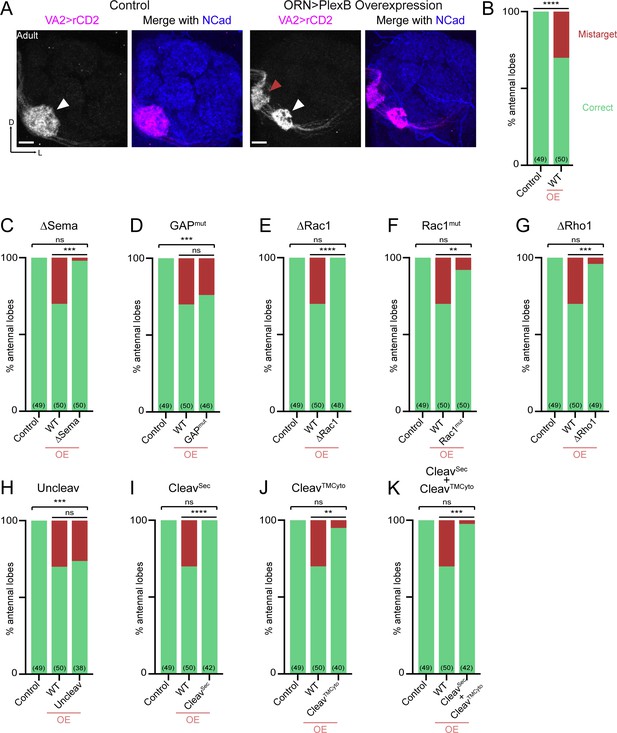
Synaptic partner selection engages both extracellular and cytoplasmic motifs of PlexB.
(A) In wild-type fly brains, Or92a+ ORN axons exclusively innervate the VA2 glomerulus at the ventromedial corner of an antennal lobe (left panels; white arrowhead). Overexpression of PlexB in ORNs causes stereotypical mistargeting to the medial DM5 glomerulus (right panels; red arrowhead). Or92a+ ORN axons were labeled by membrane-localized rCD2 driven by an Or92a promoter. Antennal lobes were co-stained with a neuropil marker N-cadherin (NCad). (B) Penetrance of glomerular mistargeting in wild-type and PlexB overexpression brains. ‘OE’ hereafter denotes ORN-specific overexpression of PlexB variants. (C–K) Penetrance of glomerular mistargeting for respective PlexB variants. Sample sizes are noted in parentheses. Significance of the contingency tables in Figure 4B–K was determined by Fisher’s exact test. ns, not significant; **p<0.01; ***p<0.001; ****p<0.0001. Images are shown as maximum z-projections of confocal stacks. Scale bars, 10 µm. Axes, D (dorsal), L (lateral).
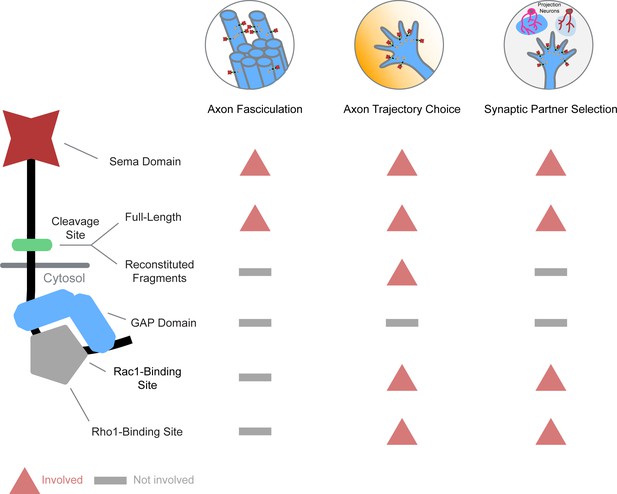
Differential engagement of PlexB structural motifs in distinct neurodevelopmental tasks.
As illustrated in columns, each distinct wiring step in the development of the fly olfactory map employs a unique combination of signaling motifs. From the perspective of individual structural motifs shown in rows, each one exhibits differing importance at different developmental stages, except the universally required Sema domain and the generally expendable GAP catalytic unit.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (Drosophila melanogaster) | C155-GAL4 | Lin and Goodman, 1994 | ||
| Genetic reagent (D. melanogaster) | Pebbled-GAL4 | Sweeney et al., 2007 | ||
| Genetic reagent (D. melanogaster) | Or92a-rCD2 | Li et al., 2018b | ||
| Genetic reagent (D. melanogaster) | UAS-mtdTomato | Potter et al., 2010 | RRID:BDSC_30124 | |
| Genetic reagent (D. melanogaster) | plexBKG00878 | Bellen et al., 2004 | RRID:BDSC_14579 | |
| Genetic reagent (D. melanogaster) | UAS-PlexB (WT) | this study | Transgenic flies of UAS-PlexB variants, described in Figure 1B | |
| Genetic reagent (D. melanogaster) | UAS-PlexB (ΔSema) | this study | ||
| Genetic reagent (D. melanogaster) | UAS-PlexB (GAPmut) | this study | ||
| Genetic reagent (D. melanogaster) | UAS-PlexB (ΔRac1) | this study | ||
| Genetic reagent (D. melanogaster) | UAS-PlexB (Rac1mut) | this study | ||
| Genetic reagent (D. melanogaster) | UAS-PlexB (ΔRho1) | this study | ||
| Genetic reagent (D. melanogaster) | UAS-PlexB (Uncleav) | this study | ||
| Genetic reagent (D. melanogaster) | UAS-PlexB (CleavSec) | this study | ||
| Genetic reagent (D. melanogaster) | UAS-PlexB (CleavTMCyto) | this study | ||
| Antibody | rat anti-Ncad | Developmental Studies Hybridoma Bank | RRID:AB_528121 | 1:40 in 5% normal donkey serum |
| Antibody | rabbit anti-DsRed | Clontech | RRID:AB_10013483 | 1:200 in 5% normal donkey serum |
| Antibody | mouse anti-rat CD2 | Bio-Rad | RRID:AB_321238 | 1:200 in 5% normal donkey serum |
| Antibody | mouse anti-V5 | Thermo Fisher | RRID:AB_2556564 | |
| Software | ZEN | Carl Zeiss | RRID:SCR_013672 | |
| Software | ImageJ | National Institutes of Health | RRID:SCR_003070 | |
| Software | Prism | GraphPad | RRID:SCR_002798 | |
| Software | Photoshop | Adobe | RRID:SCR_014199 | |
| Software | Illustrator | Adobe | RRID:SCR_010279 |
Additional files
-
Supplementary file 1
Genotypes of flies in each experiment.
- https://doi.org/10.7554/eLife.48594.009
-
Transparent reporting form
- https://doi.org/10.7554/eLife.48594.010





