Prolonged ovarian storage of mature Drosophila oocytes dramatically increases meiotic spindle instability
Figures
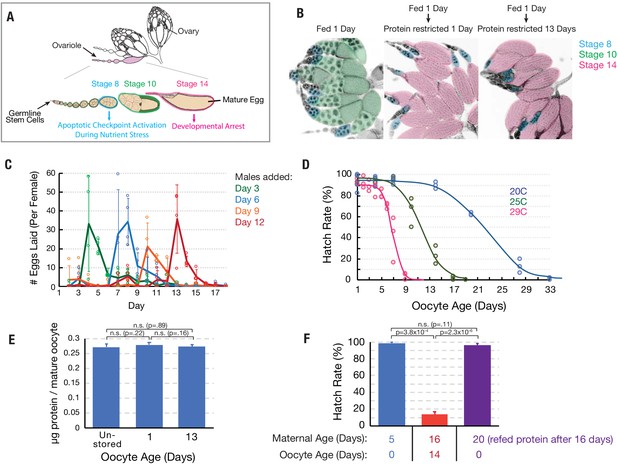
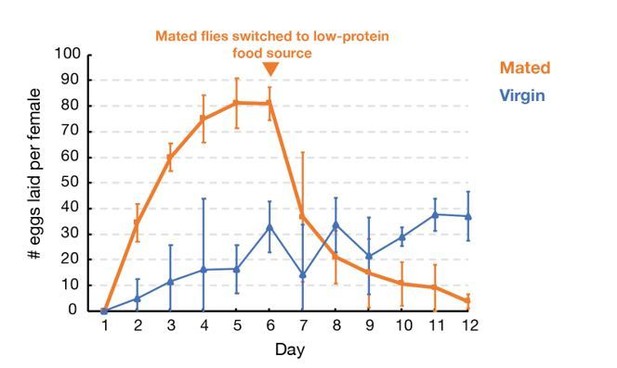
Oocytes age reproducibly in a temperature-dependent manner.
(A) A schematic of the ovarioles that make up a Drosophila ovary (above) and the structure of a single ovariole (below) showing the germline stem cells (left) and a string of increasingly mature follicles. Stages 8, 10, and 14 follicles are labeled. Prophase I arrested oocytes undergo meiotic resumption at stage 10, progressing to a secondary arrest point at metaphase I which is maintained until ovulation. (B) DAPI stained ovaries from females that were fed 1 day (left), fed 1 day then protein restricted for 1 day (middle), or fed 1 day then protein-restricted 13 days (right). Oocytes are colored as in A, revealing the stable storage of two mature stage 14 oocytes per ovariole. Each mature follicle is about 450 μM in length. (C) Eggs laid per day by females containing stored stage 14 follicles, that were provided with males after 3 (green), 6 (blue), 9 (orange) or 12 (red) days. Mating stimulates deposition of the stored oocytes as fertilized embryos. (D) Aging curves (days) for follicles stored in vivo at 29°C (magenta), 25°C (green) or 20°C (blue). For each point, stored oocytes were recovered as in (C) and the hatch rate determined (N > 100). (E) Protein content of mature oocytes that were unstored, stored in vivo for 1 day or for 13 days (‘oocyte age’). Protein restriction of mothers does not affect the protein content of stored mature oocytes. (F) Hatch rate of embryos developing from fresh mature (stage 14) oocytes from 5-day-old females, mature oocytes stored 14 days (during days 2–16) from 16 day-old females, or fresh mature oocytes from 20-day-old females. Oocyte age during storage, but not maternal age, is associated with reduced hatch rate. Error bars in (C), (E), and (F) denote SD.
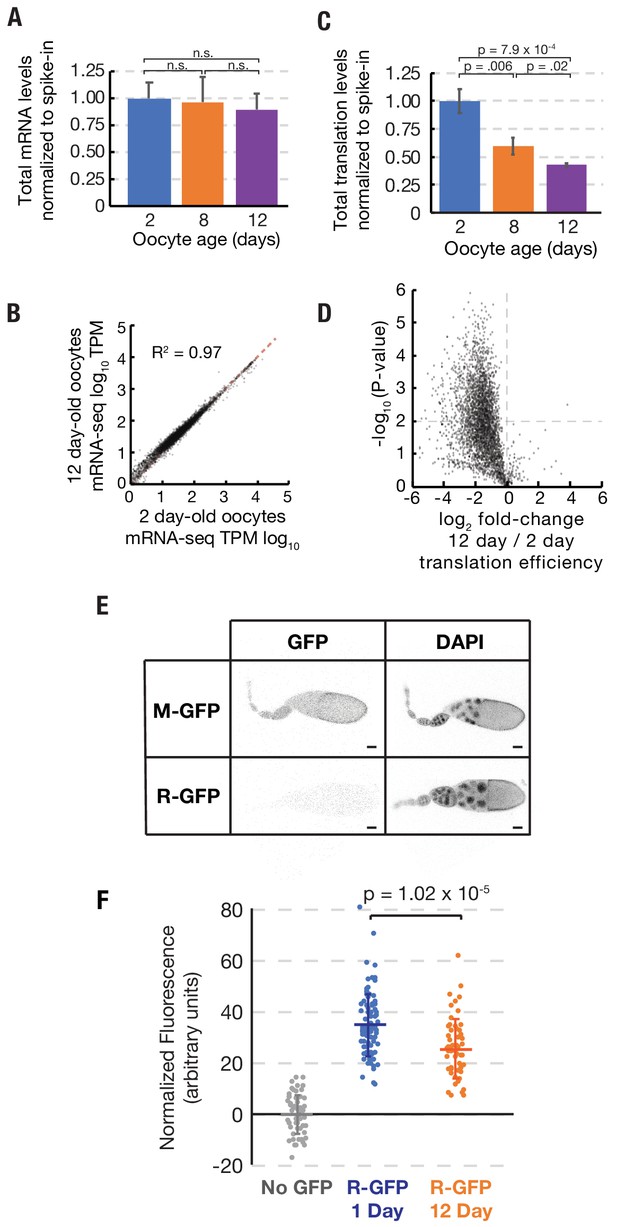
Stability of mRNA levels and translation in stored stage 14 follicles.
(A) Total mRNA per mature follicle normalized to spike-in control after oocyte aging for 2 (blue), 8 (orange) or 12 (purple) days. Differences are not significant (Student’s t-test, p=0.78, p=0.48, and p=0.49 for 2 vs. 8, 8 vs. 12, and 2 vs. 12, respectively). (B) Log-Log plot showing high correlation (R2 = 0.97) of mRNA-seq values (TPM, transcripts per million) from stage 14 follicles stored at 25°C for 2 days (96% viability) vs 12 days (44% viability). Equal expression (dashed red line). (C) Total translation levels per mature follicle normalized to spike-in control were compared between oocytes aged for 2 (blue), 8 (orange) or 12 (purple) days. Differences are significant as shown (Student’s t-test). (D) Volcano plot showing global reduction in translation efficiency in 12 day versus 2 day oocytes. (E) R-GFP serves as a reporter of nascent protein levels. Steady state levels of N-end rule proteasomal substrate R-GFP are greatly decreased as compared to stable control M-GFP, consistent with rapid degradation of R-GFP but not M-GFP. Scale bar = 50 μm. (F) Plot showing that R-GFP levels decrease ~30% during oocyte aging, consistent with reduced translation. Error bars in (A) and (C) denote SD.
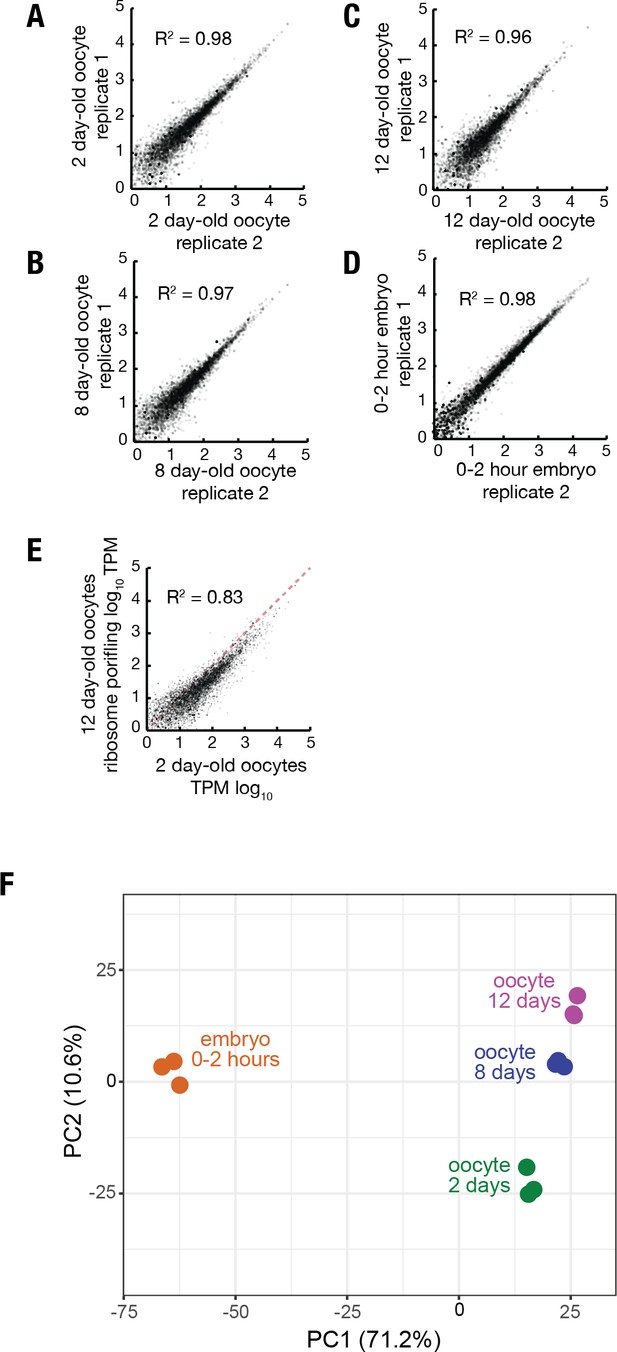
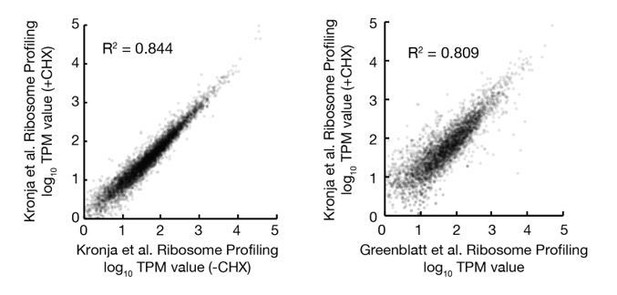
Reproducibility of ribosome profiling data.
(A–D) Log-log plots showing high reproducibility of individual gene ribosome profiling TPM values from replicate ribosome profiling experiments of 2 day (A) 8 day (B) and 12 day (C) oocytes and 0–2 hr embryos from non-aged oocytes (D). (E) Log-log plot showing that translation of individual genes is less correlated when comparing 12 day to 2 day oocytes versus replicate experiments. (F) Principle component analysis (ClustVis) showing tight clustering of replicate ribosome profiling experiments.
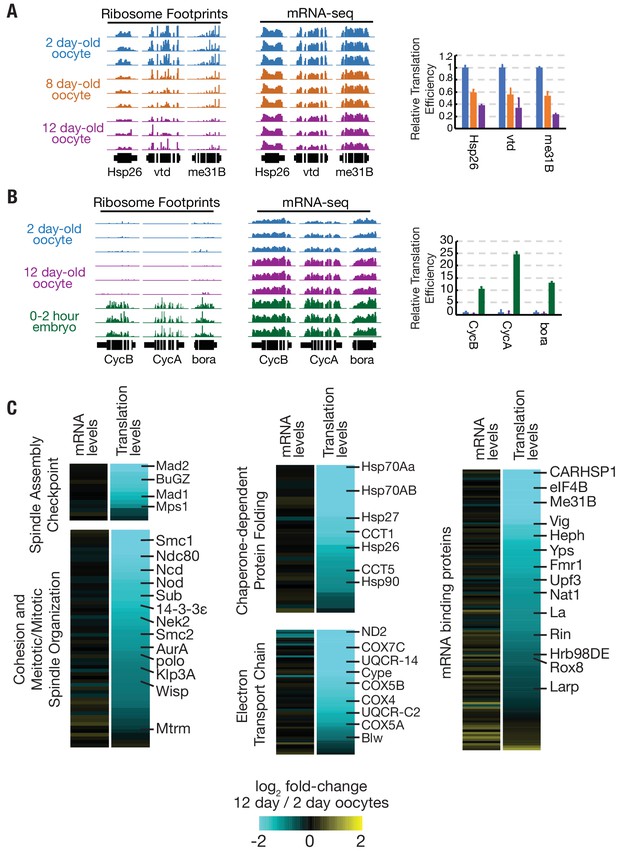
Translation is reduced globally during oocyte aging.
(A) Relative read depths from replicate ribosome footprinting and mRNA-sequencing experiments of Hsp26, vtd, and me31B from 2-, 8-, and 12-day-old oocytes. Data were normalized to spike-in controls. Relative translational efficiency (right panel) falls 2.5–5 fold between day 2 and day 12. (B) Read depths and translational efficiency values are shown as in (A) for genes preferentially translated in embryos CycB, CycA, and bora from 2-day oocytes, 12-day oocytes, and 0–2-hr embryos from non-aged oocytes. (C) Heat maps showing reduced translation, but similar mRNA levels, of genes of various GO categories in 12-day oocytes as compared to 2-day oocytes. Gene classes of interest are indicated on the left, along with specific genes on the right; see Flybase for information on each gene (http://flybase.org). Error bars in (A) and (B) denote SD.
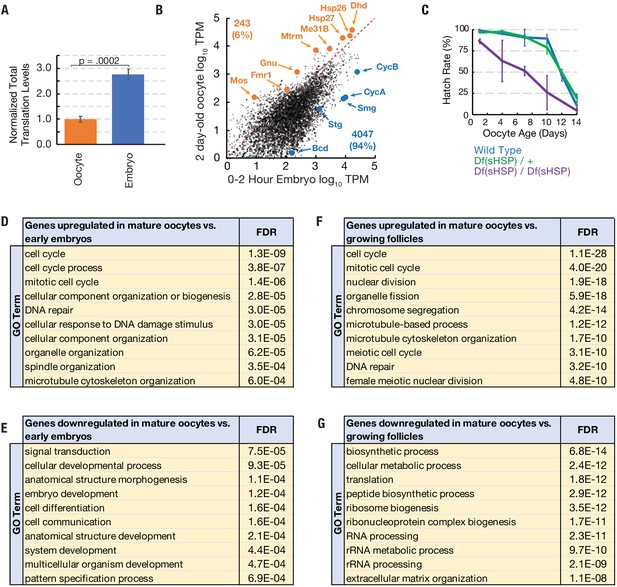
Translation of a small group of genes is boosted during oocyte arrest.
(A) Normalized total translation levels from ribosome profiling in 0–2 hr embryos compared to oocytes stored for two days. (B) Plot showing that most genes are translated at higher levels in the 0–2 hr embryo than the 2 day stored oocyte. Examples of the 243 more highly translated ‘pilot light genes’ are labeled in orange. (C) The hatch rate of stored oocytes from wild type (blue), Df(sHSP)/+ (green), or Df(sHSP)/Df(sHSP) (purple) females after indicated storage period (‘oocyte age’). Deletion of Hsp26 and Hsp27 accelerated the rate of decline during storage (N = 3 at each point). (D–G) GO analysis (PANTHER) of genes with significantly (p<0.01, Student’s t-test) increased (D,F) or decreased (E,G) translation in 2- day-old mature oocytes compared to 0–2-hr embryos (D,E) or growing follicles (F,G). Error bars in (A) and (C) denote SD. FDR = false discovery rate.
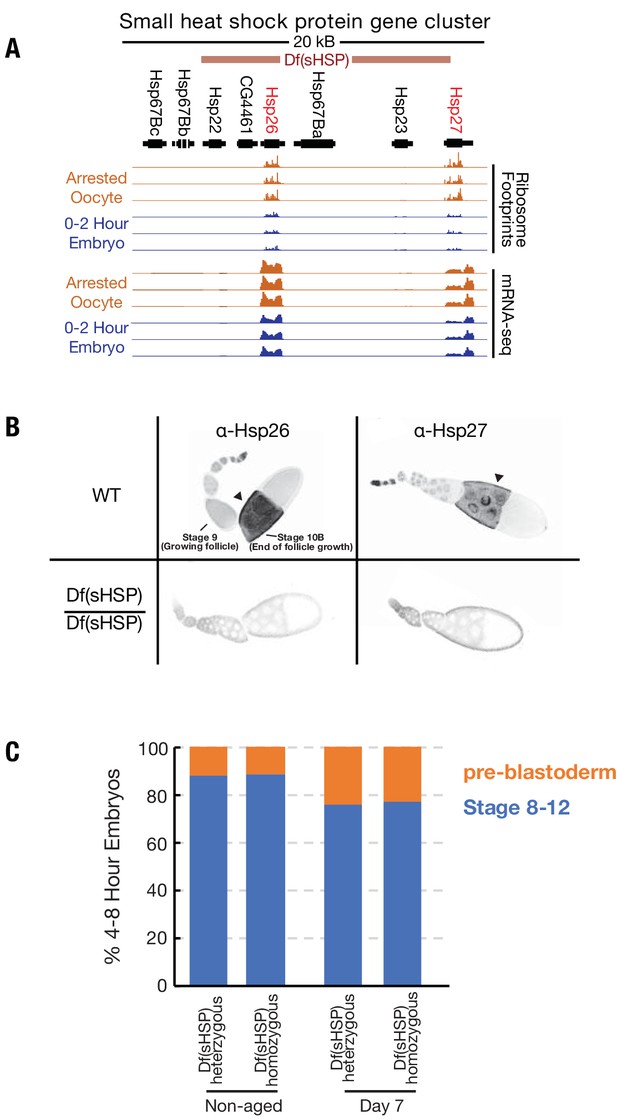
Deletion of the small heat shock protein gene locus.
(A) Normalized ribosome footprinting (upper tracks) and mRNA-seq (lower tracks) read depths in the 67B small heat shock protein gene cluster, are compared for replicate experiments from oocytes stored for 2 days (orange) and from 0–2-hr embryos (blue). Above the tracks is a map of the gene cluster, as well as the position of an FRT recombination-induced deletion of all but the left-most two genes that were generated. Two genes, Hsp26 and Hsp27, are transcribed and translated in both stored oocytes and 0–2-hr embryos. (B) Wild type (WT) and Df(sHSP)/Df(sHSP) ovarioles were stained with antibodies specific for Hsp26 and Hsp27. Both genes are expressed during oogenesis and are abundant in later stages (stage 10 shown). However, expression was absent above background in stage 10 follicles from the Df(sHSP)/Df(sHSP) ovarioles. (C) Plot showing similar stage distributions of 4–8-hr embryos derived from non-aged or aged Df(sHSP)/+ or Df(sHSP)/Df(sHSP) oocytes, suggesting that the premature loss of viability of oocytes lacking small heat shock proteins is not due to an acceleration of meiotic spindle defects observed in aged wild type oocytes.
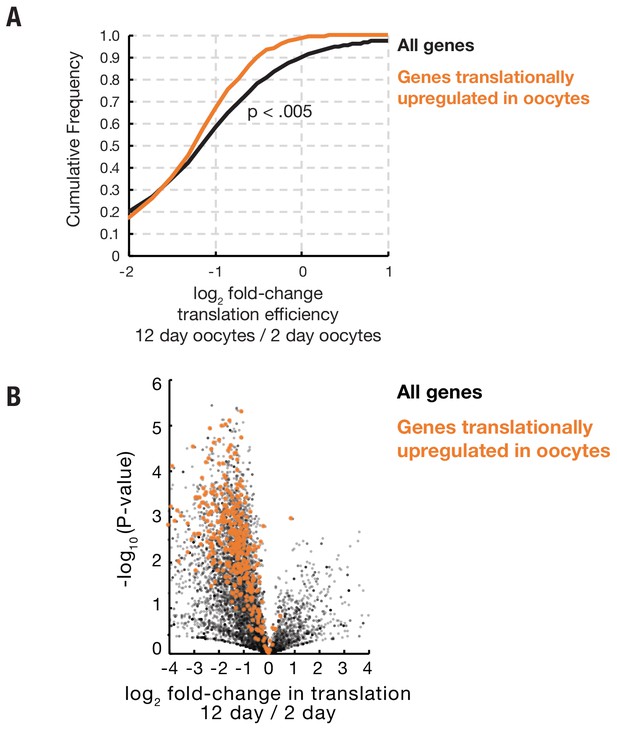
Genes preferentially translated during oocyte arrest are not protected from widespread age-associated reduced translation efficiency.
(A) Cumulative distribution plot showing that genes that are translationally upregulated in arrested oocytes (orange) as compared to 0–2-hr embryos show a slightly greater reduction in translation efficiency during aging as a group as compared to the distribution of all genes translated in oocytes. (B) Volcano plot showing that genes preferentially translated in oocytes (orange) as compared to early embryos are globally reduced in translation during oocyte aging.
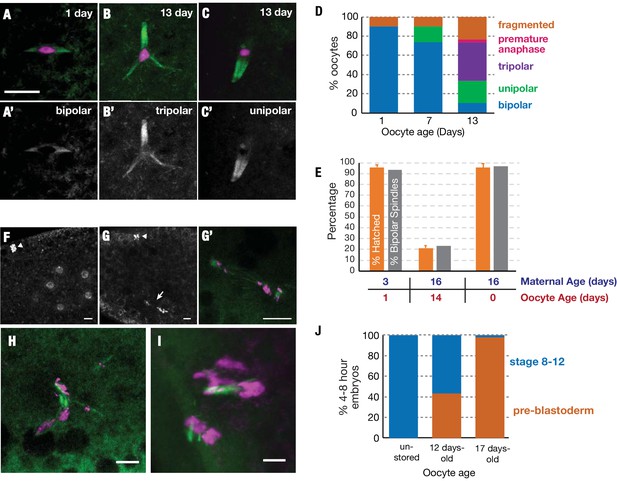
Stored oocytes lose developmental competence primarily due to problems completing meiosis.
Meiotic spindles of oocytes stored for 1, 7 or 13 days at 25°C were visualized using α-tubulin-GFP (green) and DAPI (magenta) (A–C) or using α-tubulin-GFP alone (A'–C'). Normal bipolar spindles predominate at 1 day (A) but tripolar (B) and unipolar spindles (C) increase, and predominate by 13 days (D) (30 oocytes analyzed per timepoint). (E) Spindle structure correlates closely with oocyte function. (orange bars) The hatch rate of oocytes from wild type animals with the same oocyte and maternal ages measured in parallel. (gray bars) The percentage of oocytes that contain bipolar spindles (measured using α-tubulin-GFP) at the indicated age of storage, produced by females of the indicated ages. (Hatch rates were measured in triplicate and the meiotic spindles of 30 oocytes were analyzed per timepoint). (F–H) Stored oocytes that fail to develop show problems of meiotic completion and preblastoderm arrest. (F) DAPI stained 0–1-hr embryo from an oocyte stored <1 day shows normal cleavage stage nuclei and condensed polar body (arrowhead) visible at the 8 cell stage. (G,G’) 0–1-hr embryo from 12-day-old oocyte shows arrest at the first mitotic division; arrested mitotic spindle (arrow) and polar body (arrowhead). (G') higher magnification of the spindle in (G) with tubulin-GFP (green) and DAPI (magenta). (H,I) 0–1 hr embryos from a 12-day stored oocyte showing chaotic, arrested meiotic divisions with abnormal, tripolar/fragmented spindles. (J) Stage distribution of embryos from non-stored or 12-day-old or 17 day-old stored oocytes (N>30 at each point). Embryos fell into two categories; embryos from non-aged oocytes developed to stages 8–12 (blue), whereas embryos from 12-day and 17-day oocytes either progressed to stages 8–12 or arrested (orange) during the initial meiotic/mitotic divisions (pre-blastoderm). Scale bars = 10 μM.
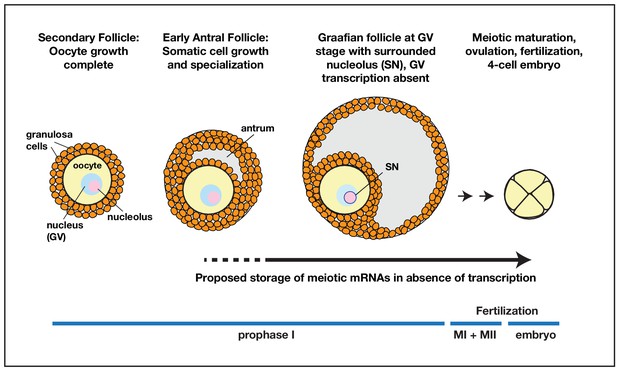
Model for prolonged mRNA storage during human oocyte development.
Accumulation of meiotic mRNAs occurs prior to the cessation of oocyte growth in pre-antral secondary follicles. Meiotic maturation of prophase I arrested human oocytes requires the translation of mRNAs that have been stored for a prolonged period of development.
Additional files
-
Supplementary file 1
Translational and mRNA changes during oocyte aging.
- https://cdn.elifesciences.org/articles/49455/elife-49455-supp1-v2.xlsx
-
Supplementary file 2
‘Pilot light’ genes translationally upregulated during oocyte arrest.
- https://cdn.elifesciences.org/articles/49455/elife-49455-supp2-v2.xlsx
-
Supplementary file 3
Genes upregulated during oocyte maturation.
- https://cdn.elifesciences.org/articles/49455/elife-49455-supp3-v2.xlsx
-
Supplementary file 4
Counts of total reads mapping to Dmel and Dpse for ribosome profiling and mRNA-sequencing experiments.
- https://cdn.elifesciences.org/articles/49455/elife-49455-supp4-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/49455/elife-49455-transrepform-v2.docx