Formation of a β-barrel membrane protein is catalyzed by the interior surface of the assembly machine protein BamA
Figures
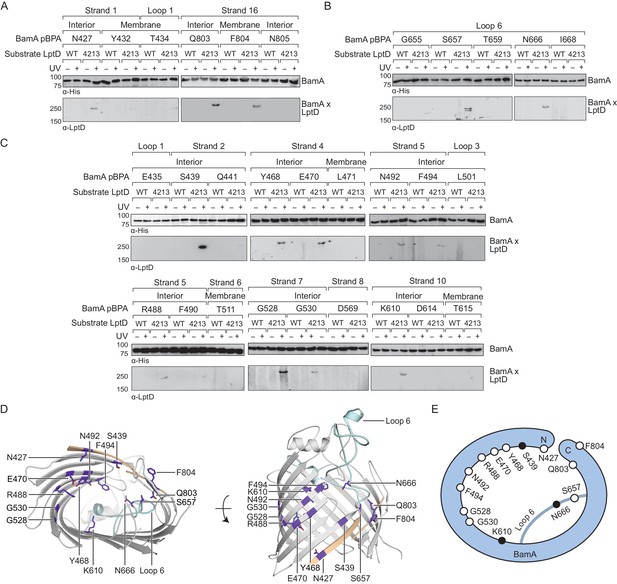
The interior wall and lateral gate of the BamA β-barrel form a substrate binding site.
(A–C) Residues at the lateral gate (A), L6 (B), and interior wall (C) of BamA interact with substrate LptD during assembly. MC4100 and lptD4213 (NR698) strains (expressing WT LptD or LptD4213, respectively) harboring the amber suppression system and expressing a His-tagged BamA (containing pBPA) were either left untreated or irradiated with UV light. Crosslinked adducts of BamA and substrate LptD/LptD4213 were identified by immunoblot analyses after Ni-NTA affinity purification. The orientation of the side chain of each residue in BamA substituted with pBPA is indicated (i.e., facing towards the membrane or interior of BamA). (D) Specific sites in the BamA β-barrel that interact with substrate LptD. Residues substituted with pBPA that crosslink to substrate are colored in purple. The first and last β-strands are colored in tan while the L6 loop is colored in cyan. Images were generated in PyMOL using the crystal structure of the BamA β-barrel from the E. coli BamABCDE complex (PDB: 5D0O). (E) Cartoon schematic of all sites in BamA that crosslink to substrate LptD. The view shown is the same as in the left panel of (D). Residues colored in black represent general substrate binding sites tested for crosslinking to full-length substrates (Figure 2). Additional views that include residues that do not form crosslinks are shown in Figure 1—figure supplement 1.
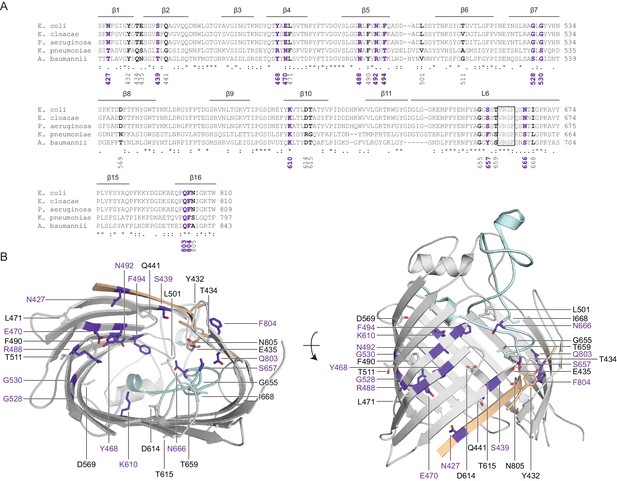
Crosslinking of the BamA interior to substrates.
(A) Sequence alignments of all identified photocrosslinking sites in the BamA β-barrel. Residues that interact with substrates are labeled in purple, while residues that do not are labeled in black. The conserved VRGF motif is boxed. Alignments were performed with Clustal Omega using BamA sequences from Escherichia coli (GI: 1036414164), Enterobacter cloacae (GI: 685450922), Pseudomonas aeruginosa (GI: 553899525), Klebsiella pneumoniae (GI: 597516803), and Acinetobacter baumannii (GI: 636479573). (B) Mapping of all photocrosslinking sites in BamA that form crosslinks to substrate LptD (purple) or do not form crosslinks (presented in same color as secondary structure).
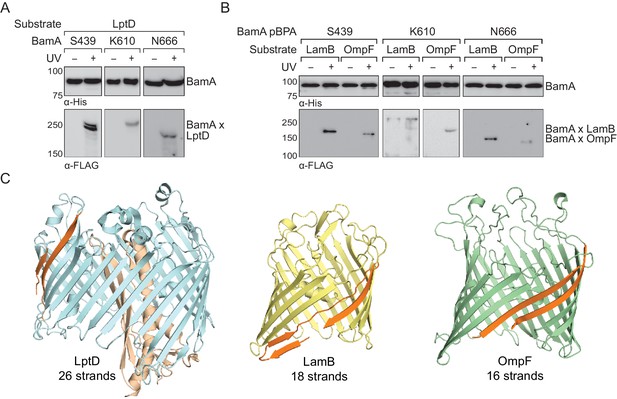
The interior of the BamA β-barrel forms a general substrate binding site.
(A–B) The interior of the BamA β-barrel interacts with a diverse number of wild-type, full-length substrates including (A) LptD and (B) LamB and OmpF. MC4100 strains harboring the amber suppression system, expressing a His-tagged BamA (containing pBPA) and a FLAG-tagged substrate were either left untreated or irradiated with UV light. Crosslinked adducts of BamA to substrate were identified by immunoblot analyses after Ni-NTA affinity purification. (C) Structural diversity of substrates of the Bam complex. The first and last β-strands of each β-barrel are colored in orange. Images were generated using PyMOL from the structures of LptD/E (PDB: 4RHB), LamB (PDB: 1MAL), and OmpF (PDB: 2OMF). The assembly efficiency of these recombinant substrates are shown in Figure 2—figure supplement 1.
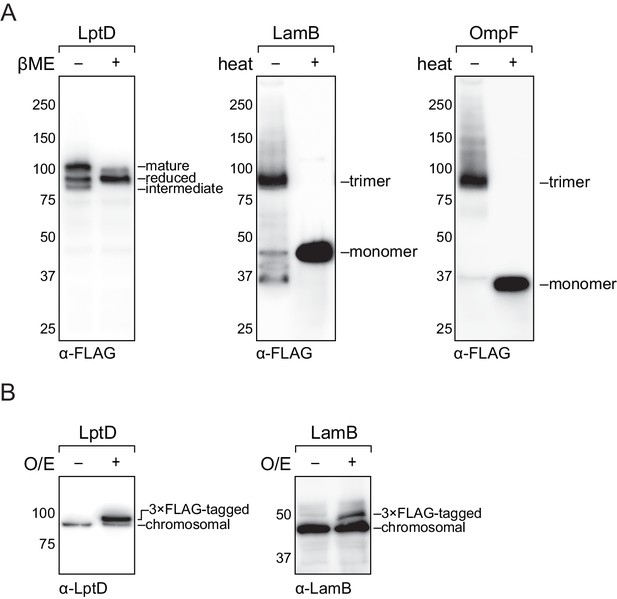
Analysis of cellular protein levels of recombinant outer membrane substrates.
(A) Recombinant LptD efficiently assembles into its mature form. Recombinant LamB and OmpF assembles into the mature trimer. MC4100 strains expressing a FLAG-tagged LptD/LamB/OmpF were harvested and analyzed by immunoblotting of cell lysates. (B) Relative expression levels of recombinant LptD/LamB as compared to their chromosomal copy.
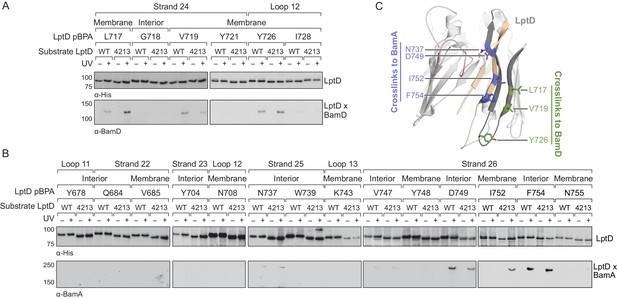
The C-terminal strands of LptD interacts with both BamA and BamD during assembly.
(A) The third-to-last β-strand and final periplasmic loop of substrate LptD interact with BamD during assembly. MC4100 strains harboring both the amber suppression system and expressing a His-tagged LptD or LptD4213 (containing pBPA) were either left untreated or irradiated with UV light. Crosslinked adducts of BamD and substrate LptD/LptD4213 were identified by immunoblot analyses after Ni-NTA affinity purification. The orientation of the side chain of each residue substituted with pBPA is indicated (i.e., facing towards the membrane or interior of the folded form of LptD). (B) As in (A), but showing crosslinking to BamA from the final two β-strands and the final extracellular loop of substrate LptD. (C) Side view of LptD (gray) mapping the residues in the C-terminal strands that interact with the Bam complex. Residues in cyan and green form crosslinks to BamA and BamD, respectively. Extracellular loop 4 of LptD, part of the region deleted in LptD4213 (pink), interacts with the ends of the LptD β-barrel (tan). Images were generated in PyMOL using the crystal structure of E. coli LptD/E (PDB: 4RHB). Additional views that include residues that do not form crosslinks are shown in Figure 3—figure supplement 1. Additional crosslinking experiments demonstrating that the BamA and BamD substrate interaction sites do not overlap are shown in Figure 3—figure supplement 2.
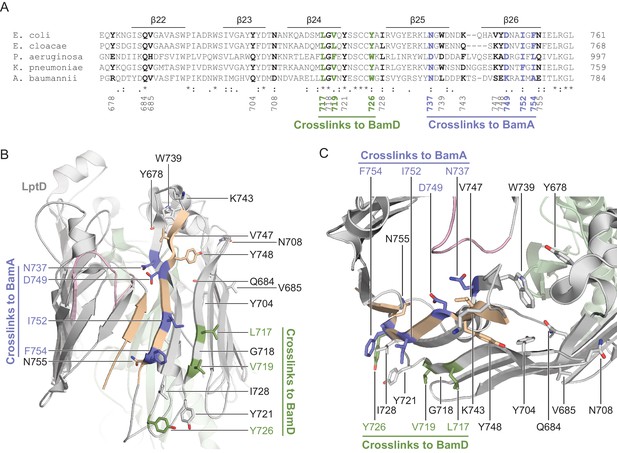
Crosslinking of the C-terminal strands of substrate LptD to BamA and BamD.
(A) Sequence alignment of the C-terminal region of LptD across representative Gram-negative bacteria. Alignments were performed by Clustal Omega using LptD sequences from Escherichia coli (GI: 2507089), Enterobacter cloacae (GI: 1017662208), Pseudomonas aeruginosa (GI: 2500883), Klebsiella pneumoniae (GI: 597516945), and Acinetobacter baumannii (GI: 751412443). Residues that interact with BamA are colored in blue, while residues that interact with BamD are colored in green. Residues that do not form crosslinks are colored in black. (B) Side view mapping the residues in the C-terminal region of substrate LptD that interact with BamA. Residues in blue and green crosslink to BamA and BamD, respectively. Residues that do not form crosslinks are presented in the same color as secondary structure. The first and last β-strands of LptD are shown in tan, the deletion in LptD4213 is shown in pink, and LptE is shown in green. (C) As in (B), but presented in a top-down view.
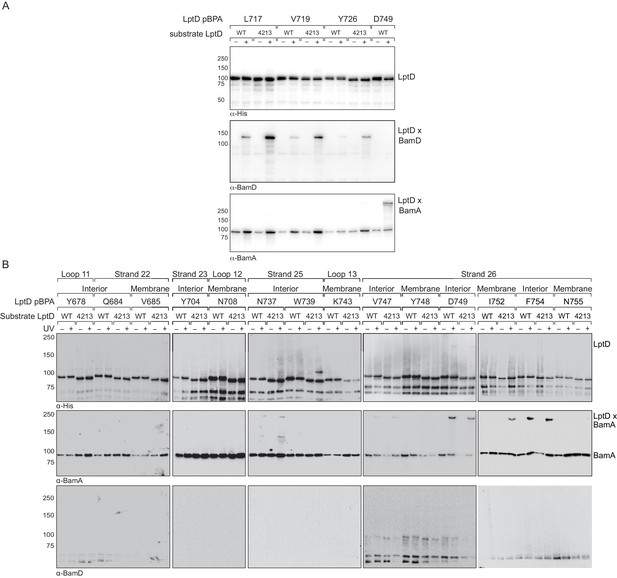
BamA and BamD bind non-overlapping regions within the C-terminal strands of substrate LptD.
(A) Residues in the third-to-last strand of LptD that interact with BamD during assembly do not interact with BamA. Crosslinking was tested as described in Figure 3. (B) Residues in the last strand of LptD that interact with BamA during assembly do not interact with BamD. The immunoblots against the His-tag and BamA are reproduced from Figure 3.
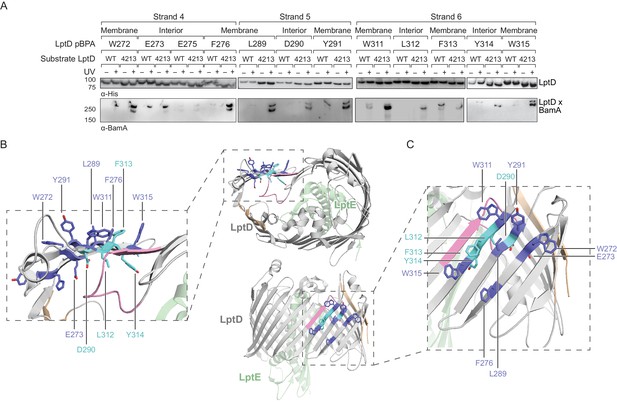
The N-terminal strands of LptD interacts with BamA during assembly.
(A) β-strands four, five, and six of substrate LptD interact with BamA. Crosslinking was tested as described in Figure 3, but with pBPA substitutions in the N-terminal portion of substrate LptD/LptD4213. ‘Membrane’ and ‘lumen’ specify where the indicated residues would face in the mature barrel. (B) Top-down view of LptD showing that residues in at least 3 β-strands in the N-terminal region of the LptD barrel interact with BamA. Residues in LptD4213 that form strong crosslinks to BamA are shown in blue, while residues that form weak crosslinks are shown in cyan. The N- and C-terminal strands of LptD are indicated in tan, and LptE is shown in green. This color scheme is maintained in the rest of the figure. (C) Side view of LptD showing crosslinking positions as depicted in (B). Additional crosslinking experiments at residues in the first three strands of substrate LptD are shown in Figure 4—figure supplement 1. Additional views that include residues that do not form crosslinks are shown in Figure 4—figure supplement 2. Note that only crosslinks within a blot can be compared, and each blot includes only proximal residues.
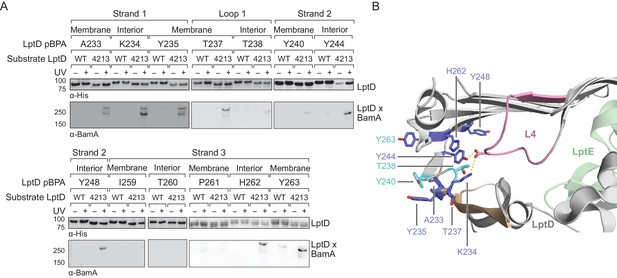
Crosslinking of the first three strands of substrate LptD to BamA.
(A) β-strands one, two, and three of substrate LptD interact with BamA. Crosslinking was tested as described in Figure 4. (B) Top-down view of the region near the N- and C-termini of LptD showing residues in LptD4213 that form crosslinks to BamA during assembly, but are important in mediating closure of LptD in the folded form. Residues K234, T238, and Y244 interact with L4 in folded LptD (within the region in pink, which is deleted in LptD4213).
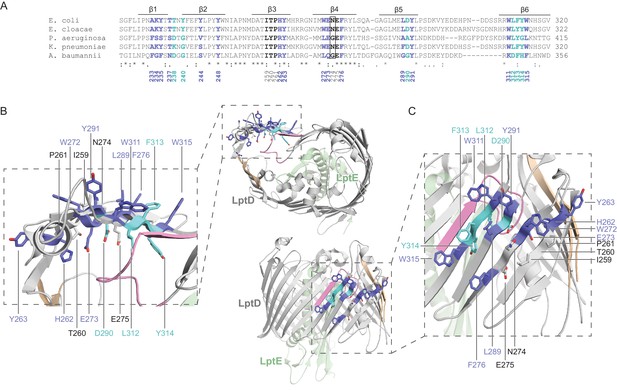
Crosslinking of the N-terminal region of substrate LptD to BamA.
(A) Sequence alignment of the N-terminal region of LptD across representative Gram-negative bacteria. Alignments were performed as in Figure 3—figure supplement 1. Residues that crosslink strongly or weakly to BamA are presented in blue or cyan, respectively, while residues that do not form crosslinks are presented in black. (B) Top-down view mapping the residues in the N-terminal strands of substrate LptD that interact with BamA. Residues in blue and cyan crosslink strongly and weakly to BamA, respectively. Residues that do not form crosslinks are presented in the same color as secondary structure. The first and last β-strands of LptD are shown in tan, the deletion in LptD4213 is shown in pink, and LptE is shown in green. (C) As in (B), but presented in a side view.
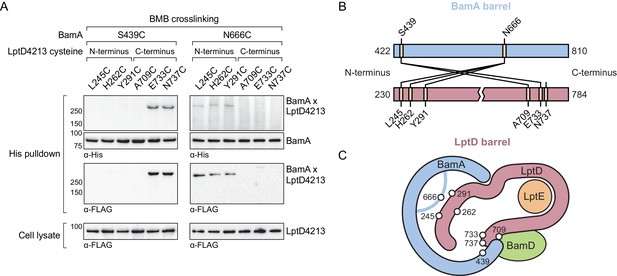
The N-terminal strands of the substrate are assembled within the BamA β-barrel.
(A) The lateral gate and interior of the BamA β-barrel interacts with the ends of the LptD substrate. The N-terminal strands of BamA (S439) interacts with the C-terminal strands of substrate LptD, while L6 (N666), within the β-barrel of BamA, interacts with the N-terminal strands of substrate LptD. MC4100 strains expressing a His-tagged BamA cysteine mutant and a FLAG-tagged LptD4213 cysteine mutant were treated with the cysteine-cysteine crosslinker 1,4-bismaleimidobutane (BMB). Crosslinked adducts of BamA and substrate LptD4213 were identified by immunoblot analyses after Ni-NTA affinity purification. Immunoblots are provided that show expression levels of cysteine-containing LptD4213 constructs from total cell lysates (bottom). (B) Map of cysteine crosslinking between the BamA and LptD4213. (C) Top-down cartoon representation of LptD engagement by BamA/D based on cysteine crosslinking data.
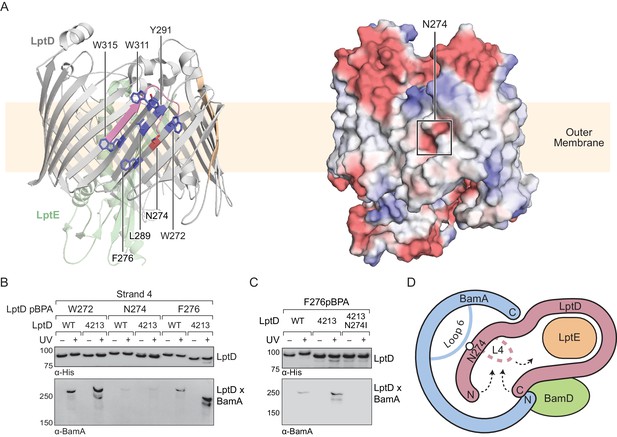
Substrate release from the interior wall of BamA allows β-barrel closure, triggering release from the Bam complex.
(A) N274 resides within a large hydrophobic patch that encompasses at least six β-strands at the N-terminal region of the LptD β-barrel. The left panel shows the structure of LptD in cartoon form. The color scheme is the same as in Figure 4, with N274 indicated in red, and six hydrophobic residues that crosslink strongly to BamA in blue. The right panel shows an electrostatic surface plot generated using APBS, presented in the same orientation as the cartoon (left). Colors in the electrostatic surface plot represent potential rather than crosslinking residues. Red represents negative potential, white represents neutral potential, and blue represents positive potential. (B) The region around N274 directly interacts with BamA. Crosslinking was tested as described in Figure 4. (C) The N274I mutation suppresses the folding defects associated with LptD4213, allowing release from BamA, as judged by a reduction in crosslinking efficiency. (D) Model for substrate β-barrel closure and release from the Bam complex. N274I suppresses the folding defect associated with LptD4213 by facilitating release of the N-terminal strands of LptD from the interior of BamA.
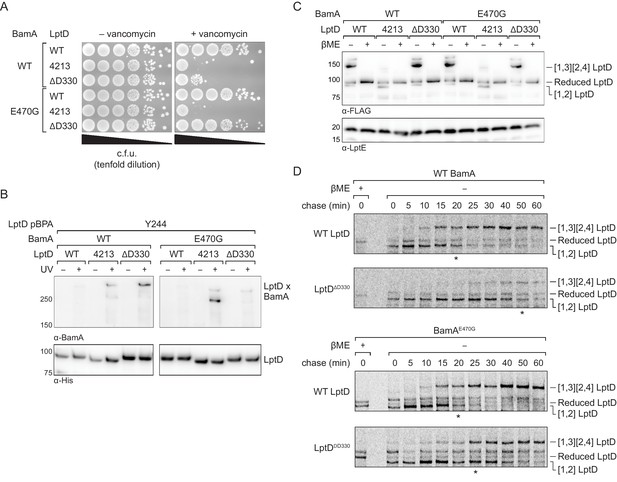
An assembly-defective LptD mutant is rescued by a compensatory mutation in the BamA interior wall.
(A) Expression of LptDΔD330 confers outer membrane permeability defects, while changes in BamA can suppress LptDΔD330 associated-defects. MC4100 or bamAE470G cells were transformed with plasmids that express WT or mutant lptD alleles. Plating assays were performed on LB supplemented with 50 μg/mL vancomycin. (B) LptDΔD330, like LptD4213, stalls on BamA during assembly. BamAE470G alleviates stalling of LptDΔD330, as judged by a reduction in crosslinking efficiency, but does not alleviate stalling of LptD4213. (C) LptDΔD330 can adopt the mature disulfide bonded state. MC4100 or bamAE470G cells expressing WT or mutant lptD alleles were harvested and analyzed via immunoblotting of cell lysates. When analyzed in the absence of reducing agent, LptD migrates at a molecular weight that reflects its state of assembly. Assembly of LptD involves the conversion of a reduced form to a form containing a disulfide bond between consecutive cysteines (designated [1,2]-LptD) that is then converted to the mature form, which contains disulfide bonds between nonconsecutive cysteines (designated [1,3][2,4]-LptD for the order in which the cysteines appear in the sequence). The [1,3][2,4] disulfide configuration reflects properly folded, functional LptD. (D) LptDΔD330 is slow to mature into the functional disulfide bond configuration, while BamAE470G alleviates LptDΔD330 assembly defects. MC4100 or bamAE470G cells expressing FLAG-tagged LptD(WT/ΔD330) were pulsed with [35S]methionine and chased with cold methionine. Samples were subsequently immunoprecipitated using α-FLAG beads and analyzed by autoradiography. The asterisk below each autoradiograph represents the time point at which approximately 50% of the substrate has converted to the mature form (containing the [1,3][2,4] disulfide bond configuration). Other compensatory mutations that restore LptDΔD330-associated defects are shown in Figure 7—figure supplement 1 and Figure 7—figure supplement 2.
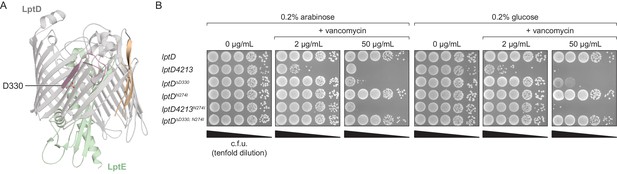
LptDΔD330 partially phenocopies LptD4213-associated assembly defects.
(A) Residue D330 in LptD resides in β-strand seven, and its side chain is oriented into the β-barrel interior. The first and last β-strands are colored in tan, the region deleted in LptD4213 is colored in pink, and LptE is colored in green. (B) LptDΔD330 confers outer membrane permeability to vancomycin even in the presence of a functional copy of LptD (+0.2% arabinose). A compensatory intragenic mutation, N274I, restores permeability defects associated with both LptD4213 and LptDΔD330.
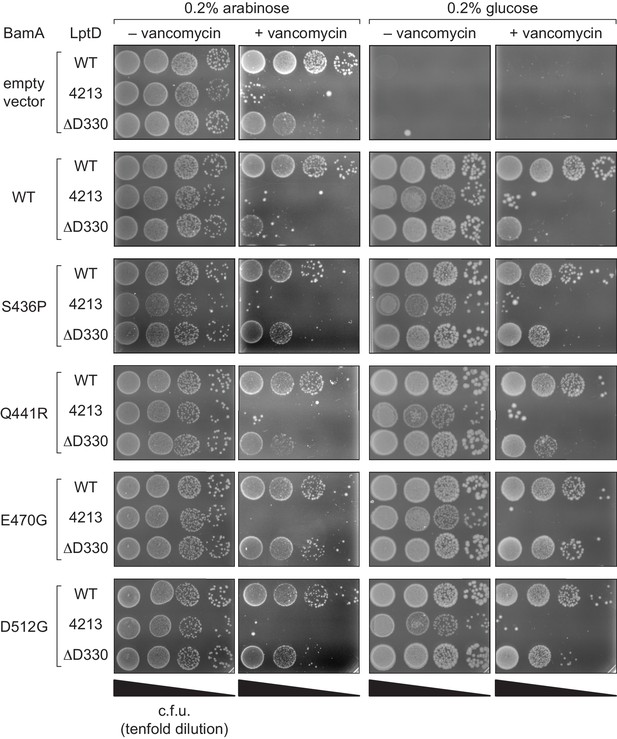
Compensatory mutations in BamA can rescue outer membrane permeability defects associated with stalled LptD substrates.
Changes in BamA can suppress LptDΔD330-associated defects. A BamA depletion strain was transformed with a plasmid encoding a BamA variant (including bamAE470G) and another plasmid encoding either WT or mutant lptD alleles. Plating assays were performed in the presence of arabinose (i.e., with expression of both the wild-type chromosomal copy of BamA and the plasmid-borne copy of BamA) or in the absence of glucose (i.e., without expression of the wild-type chromosomal copy of BamA, but with expression of the plasmid-borne copy of BamA). Strains were plated on LB supplemented with 20 μg/mL vancomycin.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain (Escherichia coli) | MC4100 | (Casadaban, 1976); PMID: 781293 | CGSC#: 6152 | See Supplementary file 1 |
| Strain (Escherichia coli) | NR698 | (Wu et al., 2005); PMID: 15851030 | See Supplementary file 1 | |
| Strain (Escherichia coli) | NR1134 | (Lee et al., 2018); PMID: 29463713 | See Supplementary file 1 | |
| Strain (Escherichia coli) | JCM166 | (Wu et al., 2005); PMID: 15851030 | See Supplementary file 1 | |
| Strain (Escherichia coli) | DEK1 | This paper | See Supplementary file 1 | |
| Strain (Escherichia coli) | DH5α λpir | (Metcalf et al., 1994; Simon et al., 1983) | See Supplementary file 1 | |
| Antibody | Mouse monoclonal Anti-Penta His HRP conjugate | Qiagen | Cat#34460 | (1:5000) |
| Antibody | Mouse monoclonal Anti-FLAG M2 HRP conjugate | Sigma | Cat#A8592; RRID:AB_439702 | (1:50,000) |
| Antibody | Rabbit polyclonal Anti-BamA primary | (Kim et al., 2007); PMID: 17702946 | (1:5000) | |
| Antibody | Rabbit polyclonal Anti-BamD primary | (Kim et al., 2007); PMID: 17702946 | (1:5000) | |
| Antibody | Rabbit polyclonal Anti-LptD primary | (Narita et al., 2013); PMID: 24003122 | (1:5000) | |
| Antibody | Rabbit polyclonal Anti-LptE primary | (Chng et al., 2010); PMID: 22936569 | (1:5000) | |
| Antibody | Rabbit IgG, HRP-linked whole Ab (from donkey) | GE Healthcare | Cat#: NA935 | (1:5000) |
| Recombinant DNA reagent | Plasmids used | This paper | See Supplementary file 2 | |
| Commercial assay | Amersham ECL Western Blotting Detection Kit | GE Healthcare | Cat#:RPN2232 | |
| Chemical compound | p-Benzoylphenylalanine | Bachem | Cat#:4017646 | |
| Chemical compound | 10X Casein blocking buffer | Sigma | Cat#:B6429 | |
| Chemical compound | Anzergent 3–14 | Anatrace | Cat#:AZ314 | |
| Chemical compound | Ni-NTA Superflow | Qiagen | Cat#:30450 | |
| Chemical compound | BMB (1,4-bismaleimidobutane) | ThermoFisher | Cat#:22331 | |
| Chemical compound | L-cysteine hydrochloride | Alfa Aesar | Cat#:L06328 | |
| Chemical compound | TCEP-HCl | VWR | Cat#:97064 | |
| Chemical compound | S35 Methionine | American Radiolabeled Chemicals | Cat#:ARS 104A | |
| Chemical compound | N-ethylmaleimide | Sigma | Cat#:E3876 | |
| Chemical compound | Anti-FLAG M2 magnetic beads | Sigma | Cat#:M8823 | |
| Software | PyMol | Schrodinger | http://pymol.org; RRID:SCR_000305 |
Additional files
-
Supplementary file 1
List of strains used.
- https://cdn.elifesciences.org/articles/49787/elife-49787-supp1-v2.docx
-
Supplementary file 2
List of plasmids used.
- https://cdn.elifesciences.org/articles/49787/elife-49787-supp2-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/49787/elife-49787-transrepform-v2.docx





