An alternatively spliced, non-signaling insulin receptor modulates insulin sensitivity via insulin peptide sequestration in C. elegans
Figures
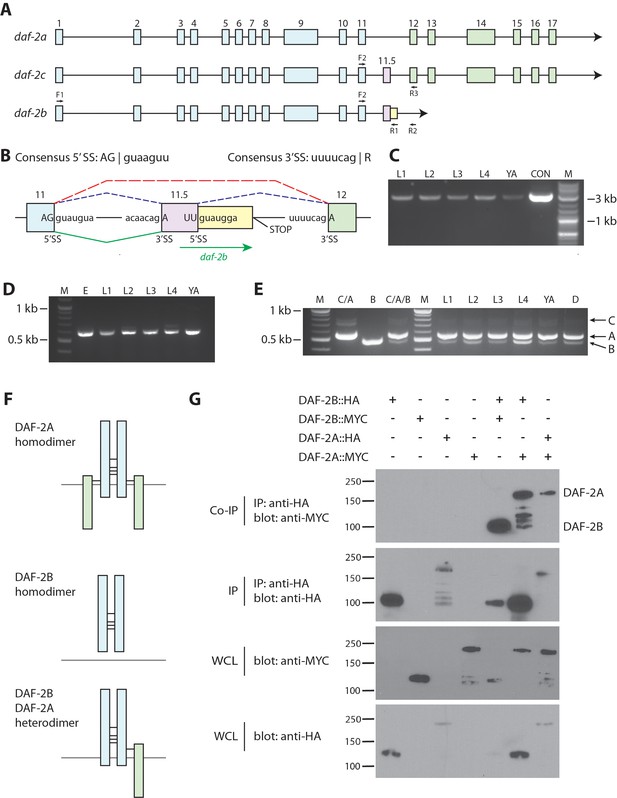
daf-2b encodes a truncated insulin receptor that is capable of dimerization.
(A) Genomic organization of the daf-2 locus. Exons shaded in blue encode the α subunit (extracellular domain) and those in green encode the β subunit (transmembrane and tyrosine kinase domains). The alternate cassette exon utilized in daf-2c (exon 11.5) is shown in pink. The daf-2b transcript is predicted to arise if splicing at the exon 11.5 5’SS is skipped leading to the addition of 46 bp of intronic sequence (shown in yellow) before an in-frame stop codon is reached. F1, F2, R1, R2 and R3 indicate the location of primers used in cDNA amplification. (B) Details of the daf-2 genomic locus from exon 11 to exon 12. The sequence of 5’ and 3’ splice sites (SS) are indicated. Dotted lines indicate splicing events for daf-2a (red) and daf-2c (blue). Green solid lines indicate splicing events for generation of the daf-2b transcript. (C) PCR amplification of full-length daf-2b cDNA using primers F1 and R1 (panel A). M - molecular weight markers, L – larval stage, YA – young adults, CON – daf-2b cDNA from plasmid template. (D) PCR amplification of a daf-2b cDNA fragment encompassing exon 11 and the predicted 3’ UTR using primers F2 and R2 (panel A). M - molecular weight markers, E – embryos, L – larval stage, YA – young adults. (E) Multiplex PCR of daf-2a, daf-2b and daf-2c from pooled cDNA (lanes marked C/A, B and C/A/B) and larval stages including dauer (D) using primers F2, R1 and R3 (panel A). (F) Schematic illustrating the possible formation of DAF-2A and DAF-2B homodimers and DAF-2A/DAF-2B heterodimers via formation of disulfide bonds at conserved cysteine residues. (G) Coimmunoprecipitation of epitope tagged DAF-2A and DAF-2B indicates the capacity to dimerize. Immunoprecipitates were subjected to SDS PAGE and blotted with anti-MYC (top panel) and anti-HA (second panel). Whole cell lysates (WCL) were blotted with anti-MYC and anti-HA (bottom two panels). Coimmunoprecipitation data are representative of 3 independent experiments.
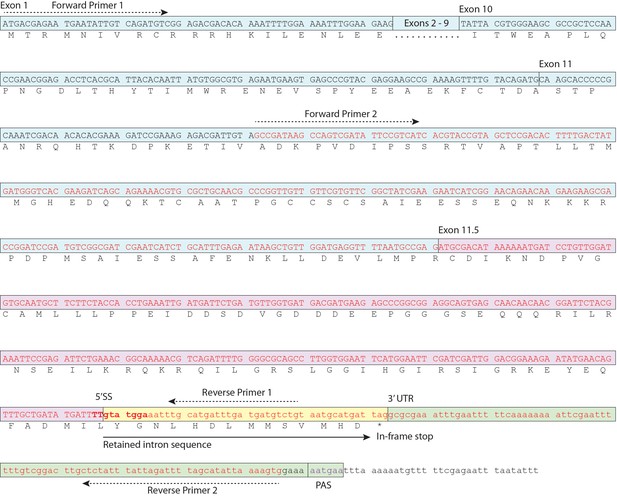
Schematic of daf-2b cDNA.
Nucleotide sequence corresponding to EC004351 EST cDNA is indicated in red text and the exon 11.5 5’ splice site is highlighted in bold. Primer sequences for amplification of the cDNA fragment shown in Figure 1C and D are indicated. PAS indicates the variant polyA signal.
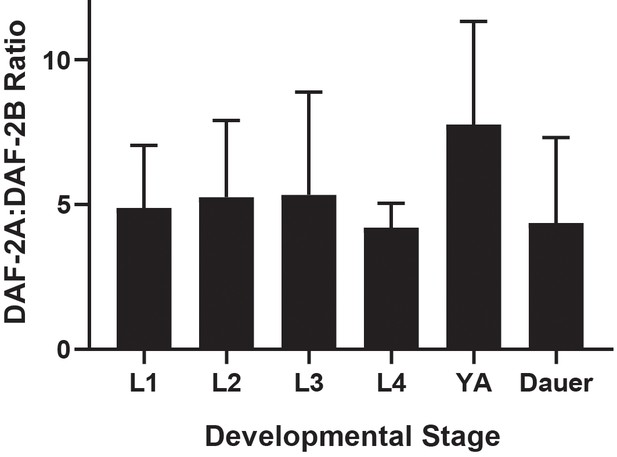
Ratio of daf-2a to daf-2b.
Multiplex PCR was performed within the linear range of amplification and band intensities were measured using Image J. Each larval stage was examined in triplicate except dauer (duplicate) and data are presented as mean+ sd.
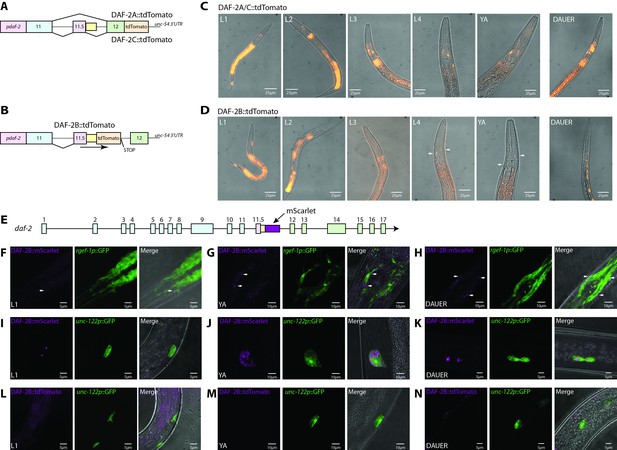
daf-2b splicing and expression are subject to temporal and spatial regulation.
(A) Schematic illustrating the organization of the daf-2a/c splicing reporter. (B) Schematic illustrating the organization of the daf-2b splicing reporter. (C) Representative fluorescence images showing the expression of DAF-2A/C::tdTomato through reproductive development and in dauers in animals bearing extrachromosomal arrays. All images were taken with 1 s exposure. (D) Representative fluorescence images showing the expression of DAF-2B::tdTomato through reproductive development and in dauers in animals bearing an integrated array. All images were taken with 1.5 s exposure. (E) Schematic illustrating the location of mScarlet insertion into the daf-2 genomic locus. Neuronal expression of DAF-2B::mScarlet in L1 (F), young adults (G) and dauers (H). Accumulation of DAF-2B::mScarlet in coelomocytes in L1 (I), young adults (J) and dauer (K) animals. DAF-2B::tdTomato from the transgenic splicing reporter is not expressed in coelomocytes in L1 (L), young adults (M) or dauers (N). Arrows indicate neuronal expression.
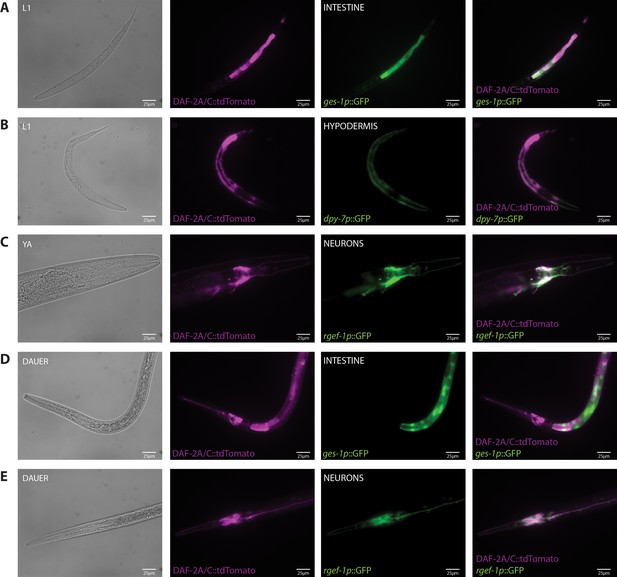
Tissue localization of the DAF-2A/C::tdTomato splicing reporter.
(A) Expression of DAF-2A/C::tdTomato with ges-1p::GFP in the intestine in an L1 animal. (B) Expression of DAF-2A/C::tdTomato with dpy-7p::GFP in the hypodermis in an L1 animal. (C) Expression of DAF-2A/C::tdTomato with rgef-1p::GFP in neurons in a young adult animal. (D) Expression of DAF-2A/C::tdTomato with ges-1p::GFP in the intestine in a dauer. (E) Expression of DAF-2A/C::tdTomato with rgef-1p::GFP in neurons in a dauer.
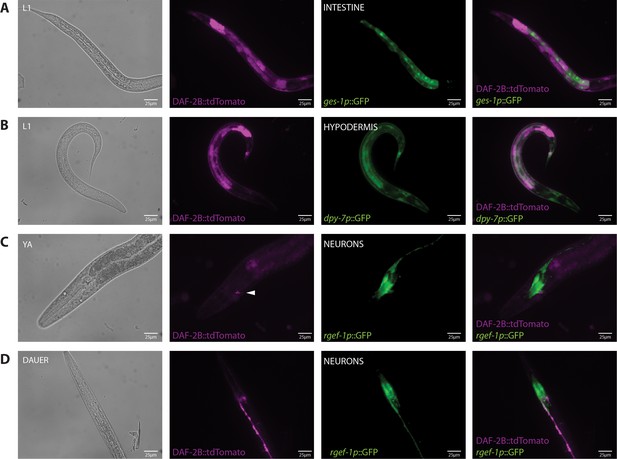
Tissue localization of the DAF-2B::tdTomato splicing reporter.
(A) Expression of DAF-2B::tdTomato with ges-1p::GFP in the intestine in an L1 animal. (B) Expression of DAF-2B::tdTomato with dpy-7p::GFP in the hypodermis in an L1 animal. (C) Expression of DAF-2B::tdTomato with rgef-1p::GFP in neurons in a young adult animal. (D) Expression of DAF-2B::tdTomato with rgef-1p::GFP in neurons in a dauer.
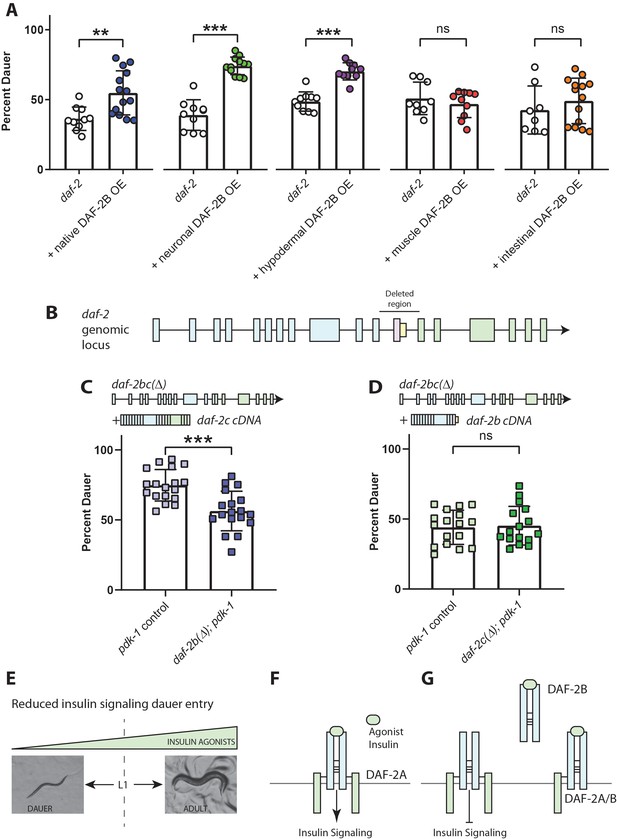
DAF-2B influences dauer entry in temperature-sensitive insulin-signaling mutants.
(A) DAF-2B overexpression (OE) from the native daf-2 promoter, in the nervous system or in the hypodermis enhances dauer formation in daf-2 mutants at 23.2°C. There was no effect of DAF-2B expressed in the intestine or in the body wall muscle. Data are pooled from 2 (hypodermal promoter) or 3 (all other promoters) transgenic lines with at least four biological replicates per transgene. Students t-test ns – not significant, **p<0.01, ***p<0.001. Two additional trials with native, neuronal, muscle and intestinal DAF-2B OE showed similar effects. Raw data can be found in Figure 3—source data 1. (B) Schematic illustrating the region of the daf-2 genomic locus deleted to generate daf-2bc(Δ). (C) Genetic deletion of daf-2b suppresses dauer formation in pdk-1 mutants at 26.8°C. Data are pooled from three independent trials with six biological replicates from one control and one daf-2b(Δ) strain per trial. Students t-test ***p<0.0001. Raw data can be found in Figure 3—source data 2. (D) Genetic deletion of daf-2c has no effect on dauer formation in pdk-1 mutants at 26.8°C. Data are pooled from three independent trials with three biological replicates from each of two control and daf-2c(Δ) strains per trial. Students t-test ns – not significant. Raw data can be found in Figure 3—source data 3. (E) In temperature-sensitive hypomorphic insulin signaling mutants, the decision to enter dauer or develop into reproductive adults is dependent on the level of insulin agonism. (F) Under semi-permissive conditions, agonism at the insulin receptor promotes insulin signaling and reproductive growth. (G) DAF-2B, acting as a homodimer or a heterodimer, is predicted to sequester agonist insulin peptides, thereby inhibiting insulin signaling and promoting dauer formation. Error bars represent mean ± sd and data are shown as independent replicates from multiple transgenic lines.
-
Figure 3—source data 1
DAF-2B overexpression enhances dauer formation indaf-2(e1368).
- https://cdn.elifesciences.org/articles/49917/elife-49917-fig3-data1-v1.xlsx
-
Figure 3—source data 2
daf-2bdeletion suppresses dauer formation inpdk-1(sa709).
- https://cdn.elifesciences.org/articles/49917/elife-49917-fig3-data2-v1.xlsx
-
Figure 3—source data 3
daf-2cdeletion has no effect on dauer formation inpdk-1(sa709).
- https://cdn.elifesciences.org/articles/49917/elife-49917-fig3-data3-v1.xlsx
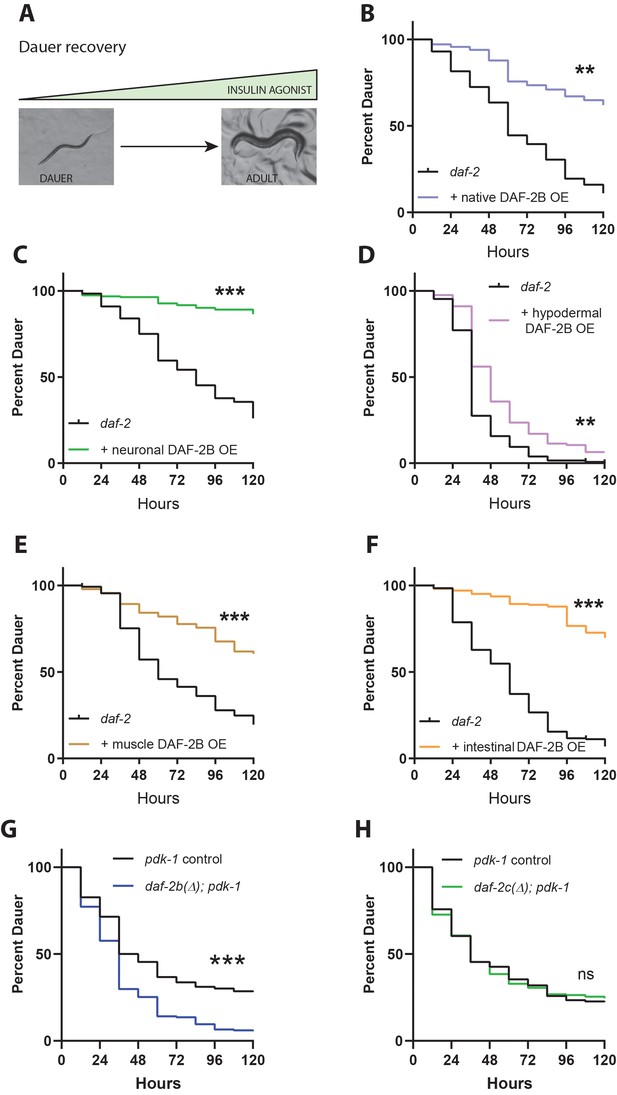
DAF-2B influences dauer recovery in temperature-sensitive insulin-signaling mutants.
(A) Exit from the dauer larva into a reproductive adult requires an increase in the activity of insulin agonists. Overexpression of DAF-2B from the native daf-2 promoter (B), in the nervous system (C), in the hypodermis (D), in muscle (E) or in the intestine (F) inhibits dauer recovery in daf-2 mutants under permissive conditions. Data are pooled from three transgenic lines (except two for hypodermal and muscle promoters) with at least three technical replicates per transgene. Two additional trials with native, neuronal, muscle and intestinal DAF-2B OE showed similar effects. Raw data can be found in Figure 4—source data 1. (G) Genetic deletion of daf-2b promotes dauer exit in pdk-1 mutants. Data are pooled from six technical replicates from one control and one daf-2b(Δ) strain. Two biological repeats showed similar effects. Raw data can be found in Figure 4—source data 2. (H) Genetic deletion of daf-2c has no effect on dauer exit in pdk-1 mutants. Data are pooled from three technical replicates from each of 2 control and daf-2c(Δ) strains per trial. Two biological repeats showed similar effects. Log Rank test ns – not significant, **p<0.01, ***p<0.001. Raw data can be found in Figure 4—source data 2.
-
Figure 4—source data 1
DAF-2B overexpression inhibits dauer recovery indaf-2(e1368).
- https://cdn.elifesciences.org/articles/49917/elife-49917-fig4-data1-v1.xlsx
-
Figure 4—source data 2
daf-2bdeletion enhances dauer recovery inpdk-1(sa709).
- https://cdn.elifesciences.org/articles/49917/elife-49917-fig4-data2-v1.xlsx
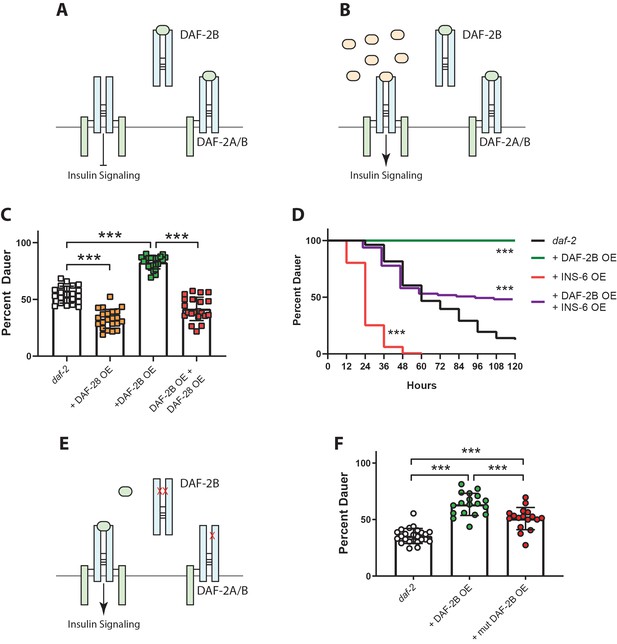
DAF-2B acts via sequestration of insulin peptides.
(A) Model for the action of DAF-2B. Increased expression of DAF-2B leads to sequestration of insulin peptides away from full length receptors resulting in reduced insulin signaling. In this respect, DAF-2B may act as a homodimer or as a heterodimer with a full-length isoform. (B) A prediction of the model is that increased availability of insulin peptide agonists should restore insulin signaling. (C) Overexpression of the DAF-28 agonist peptide from the native daf-28 promoter suppresses dauer formation in daf-2 mutants, while overexpression of DAF-2B from an integrated neuronal rab-3 promoter enhances dauer formation at 23°C. Combined overexpression of DAF-28 and DAF-2B reduces dauer formation compared with DAF-2B overexpression alone. Data are pooled from two independent trials with 12 biological replicates for daf-2(e1368) and the integrated neuronal DAF-2B OE line and two biological replicates from each of 5 or 6 DAF-28 OE extrachromosomal array lines. Post-hoc pairwise comparisons following one-way ANOVA ***p<0.0001. Raw data can be found in Figure 5—source data 1. (D) Overexpression of the INS-6 insulin peptide from the rgef-1 promoter accelerates dauer recovery in daf-2 mutants at 22.5°C. Combined overexpression of INS-6 and DAF-2B in daf-2 mutants reverses the inhibitory effect of integrated neuronal DAF-2B expression on daf-2 dauer recovery. Data are pooled from six technical replicates for daf-2(e1368) and the integrated neuronal DAF-2B OE line. For extrachromosomal array INS-6 OE lines, one replicate from each of 6 lines were pooled and at least three lines pooled for DAF-2B OE + INS-6 OE. Log Rank test – daf-2 vs daf-2 + DAF-2B OE p<0.0001; daf-2 vs daf-2 + INS-6 OE p<0.0001; daf-2 + DAF-2B OE vs daf-2 + DAF-2B OE + INS-6 OE p<0.0001. Two additional trials showed similar effects. Raw data can be found in Figure 5—source data 2. (E) Mutations in the ligand binding domain of DAF-2B that reduce affinity for insulin peptides should limit the ability of DAF-2B to inhibit insulin signaling. (F) Overexpression of wild type DAF-2B in the nervous system enhances dauer entry in daf-2 mutants. The inhibitory effect of DAF-2B is attenuated by a point mutation that affects the L1 insulin binding domain. Data are pooled from two independent trials with six transgenic lines for DAF-2B::FLAG and DAF-2B(C196Y)::FLAG and 1–2 biological replicates per line. Post-hoc pairwise comparisons following one-way ANOVA ***p<0.0001. Error bars represent mean ± sd and data are shown as independent replicates from multiple transgenic lines. Raw data can be found in Figure 5—source data 3.
-
Figure 5—source data 1
Overexpression of the DAF-28 agonist insulin peptide suppresses the effects of DAF-2B OE indaf-2(e1368).
- https://cdn.elifesciences.org/articles/49917/elife-49917-fig5-data1-v1.xlsx
-
Figure 5—source data 2
Overexpression of the INS-6 agonist insulin peptide suppresses the effects of DAF-2B OE indaf-2(e1368).
- https://cdn.elifesciences.org/articles/49917/elife-49917-fig5-data2-v1.xlsx
-
Figure 5—source data 3
Overexpression of mutant DAF-2B indaf-2(e1368).
- https://cdn.elifesciences.org/articles/49917/elife-49917-fig5-data3-v1.xlsx
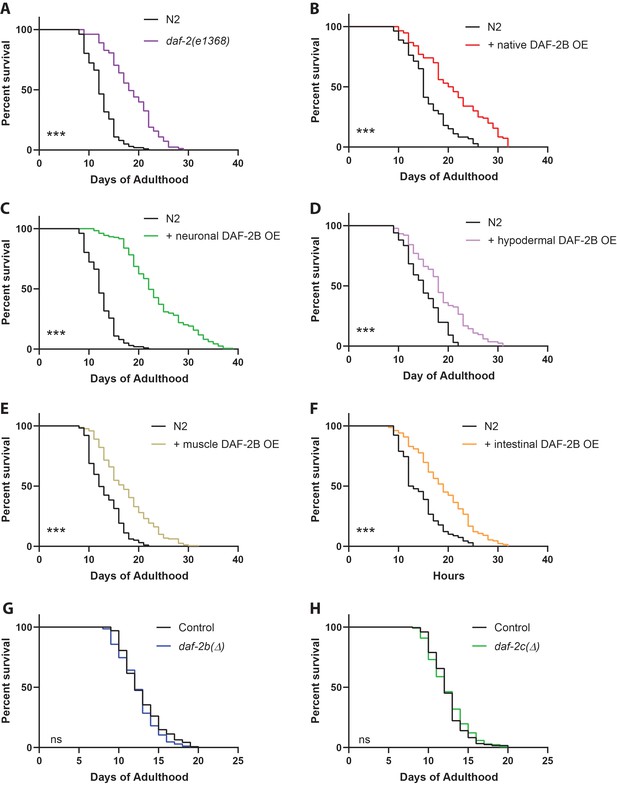
DAF-2B overexpression extends lifespan.
(A) The daf-2(e1368) hypomorphic mutation confers lifespan extension. Overexpression of DAF-2B from the native daf-2 promoter (B), in the nervous system (C), in the hypodermis (D), in the muscle (E) or in the intestine (F) extends lifespan in wild type N2 animals. Genetic deletion of daf-2b (G) and daf-2c (H) has no effect on lifespan in wild type N2 animals. Log Rank test ns – not significant, ***p<0.001. Lifespan data are representative of n = 2 biological replicates for DAF-2B overexpressers and n = 3 for deletion strains. Summary data for all replicates are presented in Supplementary file 1.
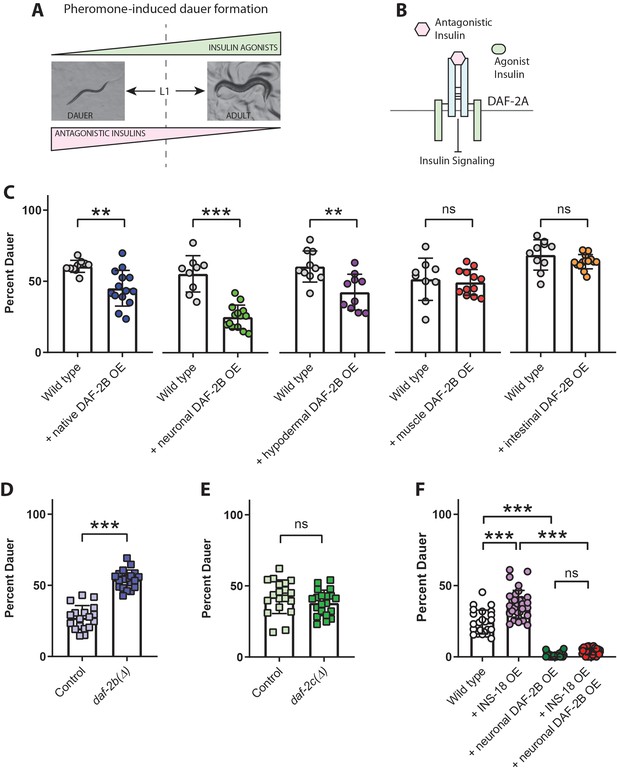
DAF-2B attenuates the activity of antagonistic insulin peptides.
(A) In the presence of the dauer-inducing pheromone extracts, the decision in the first larval stage (L1) to proceed with reproductive growth or enter the dauer stage is dependent on the balance between antagonistic and agonistic insulin peptides. (B) Increased activity of antagonistic insulin peptides inhibits insulin signaling to promote dauer entry. (C) DAF-2B overexpression (OE) from the native daf-2 promoter, in the nervous system, or in the hypodermis suppresses pheromone-induced dauer formation in wild type animals. There was no effect of DAF-2B expressed in the intestine or in the body wall muscle. Data are pooled from 2 (hypodermal promoter) or 3 (all other promoters) transgenic lines with at least four biological replicates per transgene. Students t-test ns – not significant, **p<0.01, ***p<0.001. Similar effects were observed in an additional trial for muscle, intestine and neuronal promoters and 1/2 additional trials for the native promoter. Raw data can be found in Figure 7—source data 1. (D) Genetic deletion of daf-2b enhances dauer formation in the presence of pheromone extracts. Data are pooled from two independent trials with three biological replicates from each of two control and two daf-2b(Δ) strains per trial. Students t-test ***p<0.001. Raw data can be found in Figure 7—source data 2. (E) Genetic deletion of daf-2c has no effect on dauer formation in the presence of pheromone extracts. Student t-test ns – not significant. Raw data can be found in Figure 7—source data 3. (F) Overexpression of the INS-18 antagonist insulin peptide from the native ins-18 promoter enhances pheromone-induced dauer formation in wild type animals, while overexpression of DAF-2B from an integrated neuronal rab-3 promoter suppresses dauer formation. Combined overexpression of INS-18 and DAF-2B also suppresses dauer formation compared with INS-18 overexpression alone. Data are pooled from two independent trials with 12 biological replicates for wild type, 11 or 12 biological replicates for the integrated neuronal DAF-2B OE line and 2 or three biological replicates from each of 6 INS-18 OE extrachromosomal array lines. Post-hoc pairwise comparisons following one-way ANOVA ***p<0.0001. Raw data can be found in Figure 7—source data 4. Error bars represent mean ± sd, and data are shown as independent replicates from multiple transgenic lines.
-
Figure 7—source data 1
DAF-2B overexpression suppresses pheromone-induced dauer formation in wild type.
- https://cdn.elifesciences.org/articles/49917/elife-49917-fig7-data1-v1.xlsx
-
Figure 7—source data 2
daf-2bdeletion enhances pheromone-induced dauer formation in wild type.
- https://cdn.elifesciences.org/articles/49917/elife-49917-fig7-data2-v1.xlsx
-
Figure 7—source data 3
daf-2cdeletion has no effect on pheromone-induced dauer formation in wild type.
- https://cdn.elifesciences.org/articles/49917/elife-49917-fig7-data3-v1.xlsx
-
Figure 7—source data 4
DAF-2B overexpression suppresses the effect of INS-18 OE in pheromone-induced dauer formation in wild type.
- https://cdn.elifesciences.org/articles/49917/elife-49917-fig7-data4-v1.xlsx
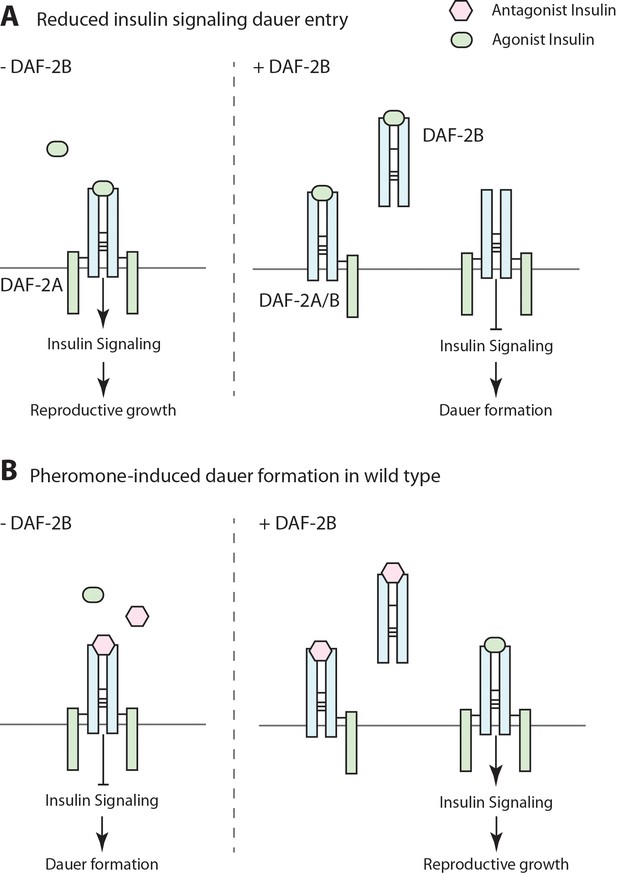
Model for the effect of DAF-2B on dauer formation.
(A) In temperature-sensitive hypomorphic mutants that affect the insulin signaling pathway, such as daf-2(e1368) and pdk-1(sa709), dauer entry is driven by a reduction in agonist insulin peptides. Under these conditions, loss of daf-2b promotes reproductive growth by enhancing insulin sensitivity, while extra-copies of DAF-2B enhance dauer entry by sequestering agonist insulins. (B) Pheromone-induced dauer formation in wild type worms is driven by an increase in activity of antagonist insulin peptides. In this paradigm, loss of daf-2b enhances dauer formation. Conversely, overexpression of DAF-2B promotes reproductive growth via sequestration of insulin antagonists.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Caenorhabditis elegans) | daf-2 | www.wormbase.org | WBGene00000898 | |
| Strain, strain background (Escherichia coli)) | OP50 | Caenorhabditis Genetics Center (CGC) | OP50 | |
| Strain, strain background (C. elegans) | C. elegans strains used and generated in this study | Caenorhabditis Genetics Center (CGC) and this paper | Supplementary file 4 | |
| Cell line (human) | HEK293T/17 | ATCC | CRL-11268 | This cell line has been re-verified and has tested negative for mycoplasma |
| Antibody | Rabbit polyclonal anti-HA | Life Technologies | 715500 | IP: 3 ul for 20 mg protein |
| Antibody | Mouse monoclonal anti-myc | Cell Signaling Technology | 2276S | Dilution 1:1000 |
| Antibody | Mouse monoclonal anti-HA | Cell Signaling Technology | 2367S | Dilution 1:1000 |
| Antibody | Goat HRP-conjugated anti-mouse IgG | Fisher Scientific | Cat #9491974 | Dilution 1:4000 |
| Peptide, recombinant protein | Protein G agarose | Sigma | 11719416001 | |
| Recombinant DNA reagent | Plasmids generated in this study | This paper | Supplementary file 3 | |
| Recombinant DNA reagent | Primers used in this study | This paper | Supplementary file 2 | |
| Software, algorithm | Snapgene | GSL Biotech | RRID:SCR_015052 | https://www.snapgene.com/ |
| Software, algorithm | Prism 8 | Graphpad Software | RRID:SCR_002798 | https://www.graphpad.com/scientificsoftware/prism/ |
| Software, algorithm | Illustrator CS5 | Adobe | RRID:SCR_010279 | https://www.adobe.com/products/illustrator.html |
| Software, algorithm | Photoshop CS5 | Adobe | RRID:SCR_014199 | https://www.adobe.com/products/photoshop.html |
| Software, algorithm | Zen Digital Imaging for Light Microscopy | Zeiss | RRID:SCR_013672 | http://www.zeiss.com/microscopy/en_us/products/microscope-software/zen.html#introduction |
| Software, algorithm | ImageJ | ImageJ | RRID:SCR_003070 | https://imagej.net/ |
Additional files
-
Supplementary file 1
Summary data for all replicates of lifespan experiments.
- https://cdn.elifesciences.org/articles/49917/elife-49917-supp1-v1.docx
-
Supplementary file 2
List of primers used in this study.
- https://cdn.elifesciences.org/articles/49917/elife-49917-supp2-v1.docx
-
Supplementary file 3
List of plasmids used in this study.
- https://cdn.elifesciences.org/articles/49917/elife-49917-supp3-v1.docx
-
Supplementary file 4
List of C. elegans strains used in this study.
- https://cdn.elifesciences.org/articles/49917/elife-49917-supp4-v1.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/49917/elife-49917-transrepform-v1.docx





