Chronic muscle weakness and mitochondrial dysfunction in the absence of sustained atrophy in a preclinical sepsis model
Figures
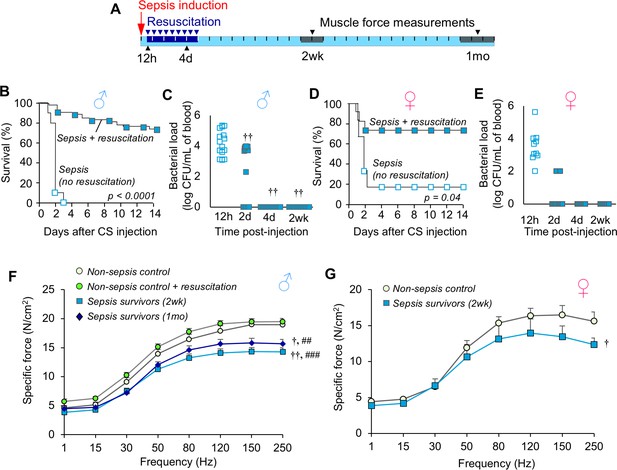
Mice exhibit chronic muscle weakness after recovering from sepsis induced by a severe model with ICU-like resuscitation.
(A) Schematic diagram of the protocol which allows for long-term assessments in sepsis survivors. Sepsis is induced by cecal slurry (CS) injection (i.p.) and therapeutic resuscitation is delayed until 12h. Resuscitation includes antibiotics and fluid administration which is continued twice daily for 5 days. Each segment on the line corresponds to one day. (B, D) After sepsis induction, animals received either no intervention (n = 10 ♂, n = 6 ♀) or therapeutic resuscitation (n = 54 ♂, n = 11 ♀); survival was monitored for 14 days. Kaplan-Meier Log-rank test was performed. (C, E) Circulating bacterial load was assessed immediately before initiation of resuscitation (12h), and at 2d, 4d, and 2wk (n = 15 ♂, n = 7 ♀). Repeated measures ANOVA was performed (†† p<0.01 compared to 12h). (F, G) Specific force of the extensor digitorum longus was measured to assess muscle strength of non-sepsis control (NSC; n = 9 ♂, n = 5 ♀), NSC + resuscitation (n = 7 ♂), and sepsis surviving mice at 2wk (n = 7 ♂, n = 5 ♀) and 1mo (n = 7 ♂) after sepsis. Data are expressed as means ± SEM, † p<0.05, †† p<0.01 compared to NSC, ## p<0.01, ### p<0.001 compared to NSC + resuscitation.
-
Figure 1—source data 1
Mice exhibit chronic muscle weakness after recovering from sepsis induced by a severe model with ICU-like resuscitation.
- https://doi.org/10.7554/eLife.49920.004
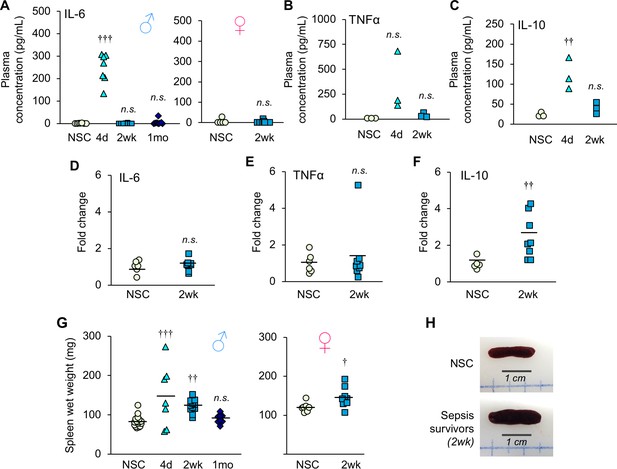
Inflammation is resolved by two weeks after sepsis.
(A) Plasma IL-6 concentration was measured in non-sepsis control (NSC) and sepsis survivors (n = 7–8 ♂, n = 6–9 ♀, per group). (B) Plasma TNFα and (C) IL-10 concentrations were quantified in non-sepsis control (NSC), and sepsis surviving mice (n = 3 per group). Relative gene expression of (D) IL-6, (E) TNFα, and (F) IL-10 in gastrocnemius of male NSC (n = 6) and sepsis survivors (n = 8). (G) Spleen wet weight was recorded at time of sacrifice at 4d (n = 7 ♂), 2wk (n = 12 ♂, n = 9 ♀), and 1mo (n = 10 ♂) alongside non-sepsis controls (n = 16 ♂, n = 7 ♀). One-way ANOVAs were performed. n.s. not significant, † p<0.05, †† p≤0.01, ††† p<0.001 compared to NSC. (H) Representative macroscopic images of the spleen.
-
Figure 2—source data 1
Inflammation is resolved by two weeks after sepsis.
- https://doi.org/10.7554/eLife.49920.006
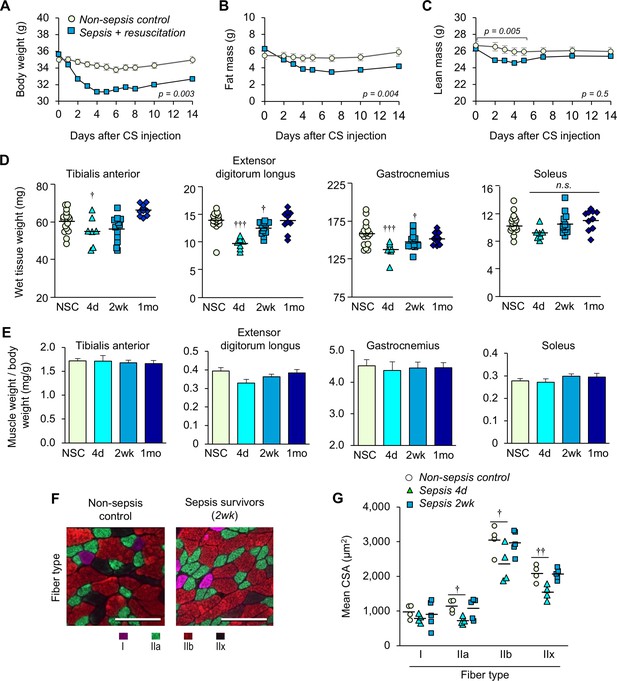
Sepsis-induced muscle wasting is evident during the acute phase but is later recovered.
(A) Body weight, (B) fat mass, and (C) lean mass were assessed in non-sepsis controls (NSC, n = 7) alongside animals with sepsis + resuscitation (n = 11). Repeated Measures ANOVAs were conducted (A–C). (D) Wet tissue weight of hindlimb skeletal muscles was measured from NSC (n = 17) and sepsis mice at 4d, (n = 7), 2wk (n = 12), and 1mo (n = 10). One-way ANOVAs were performed. n.s. not significant, † p<0.05, ††† p<0.001 compared to NSC. (E) Skeletal muscle weight/body weight ratio. Data are presented as means ± SEM. One-way ANOVAs were performed, no statistical difference was found across the groups. Similar trends were observed in female sepsis survivors (Figure 3—figure supplement 1). (F) Representative images of fiber-type staining of the gastrocnemius show myosin heavy chain type I fibers (pink), IIa (green), IIb (red), and IIx (unstained/black); scale bars represent 100 µm. (G) Cross-sectional area (CSA) analysis by fiber type (n = 4–5 per group). Staining and CSA analysis on soleus muscles are provided in Figure 3—figure supplement 2. One-way ANOVAs were performed, † p<0.05, †† p<0.01, compared to NSC.
-
Figure 3—source data 1
Sepsis-induced muscle wasting is evident during the acute phase but is later recovered.
This data file includes data presented in main Figure 3 as well as Figure 3—figure supplements 1 and 2.
- https://doi.org/10.7554/eLife.49920.010
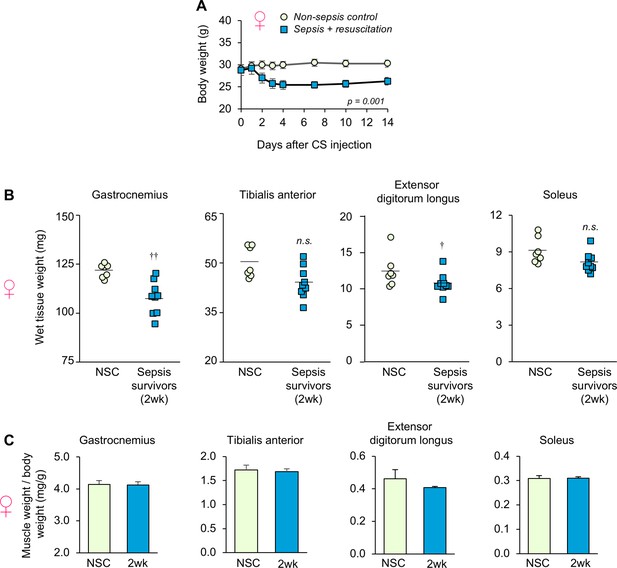
Sepsis-induced body weight and skeletal muscle weight in female animals.
Sepsis was induced in female mice (16 mo-old) as described in Figure 1. (A) Body weight was monitored regularly for non-sepsis controls (NSC, n = 7) and sepsis survivors (2wk, n = 9). (B) Hindlimb skeletal muscle wet weight at time of sacrifice. (C) Skeletal muscle weight/body weight ratio. Data are presented as means ± SEM. Repeated Measures ANOVA (A) and one-way ANOVAs (B, C) were performed. n.s. not significant, † p<0.05, †† p<0.01 compared to NSC.
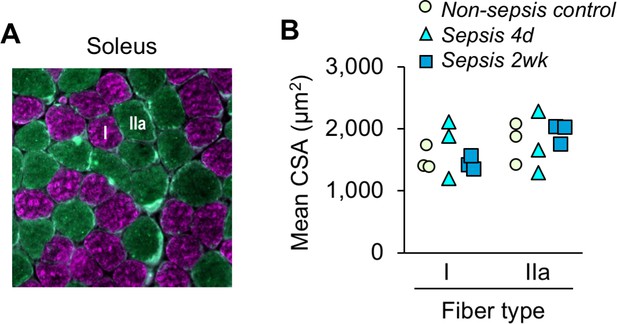
Sepsis-induced atrophy is not evident in soleus muscle.
(A) Representative image of fiber-type staining showing myosin heavy chain type I fibers (pink) and type IIa (green). (B) Fiber-type-specific cross-sectional area analysis (described in Figure 2) was performed on soleus muscles from non-sepsis control, sepsis at 4d and 2wk animals (n = 3 per group). Data are expressed as individual data points with means displayed as bars. One-way ANOVAs were performed, no statistical differences were found across the groups.
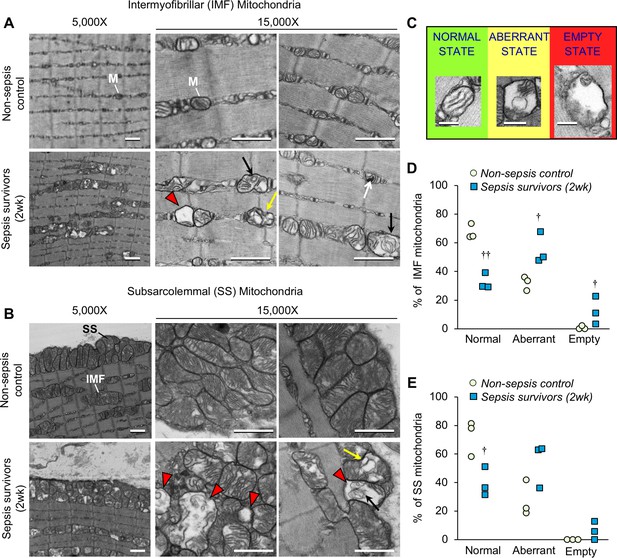
Sepsis triggers profound long-lasting ultrastructural defects in skeletal muscle mitochondrial populations.
The extensor digitorum longus was harvested from murine sepsis survivors (2wk) and non-sepsis controls (n = 3 per group) and processed for transmission electron microscopy observation of mitochondria (M). Representative micrographs of (A) intermyofibrillar (IMF) and (B) subsarcolemmal (SS) mitochondria at 5,000X and 15,000X magnifications are shown (scale bars represent 1,000 nm). Abnormal mitochondrial structures are indicated with the following symbols: destruction of cristae with expanded matrix space (red arrowheads), concentric ‘onion shaped’ cristae (black arrows), compartmentalization into vacuolar structures (yellow arrows), and densely compacted cristae (white arrows). (C) Representative images for classification of normal, aberrant, or empty mitochondria for morphometric analyses of (D) intermyofibrillar and (E) subsarcolemmal mitochondrial populations which are shown as percent for non-sepsis control and sepsis survivors (2wk). Repeated-measures ANOVAs were performed, each considering a two-way interaction between group and category (normal/aberrant/empty). n.s. not significant, † p<0.05, †† p<0.01 compared to NSC.
-
Figure 4—source data 1
Sepsis triggers profound long-lasting ultrastructural defects in skeletal muscle mitochondrial populations.
- https://doi.org/10.7554/eLife.49920.013
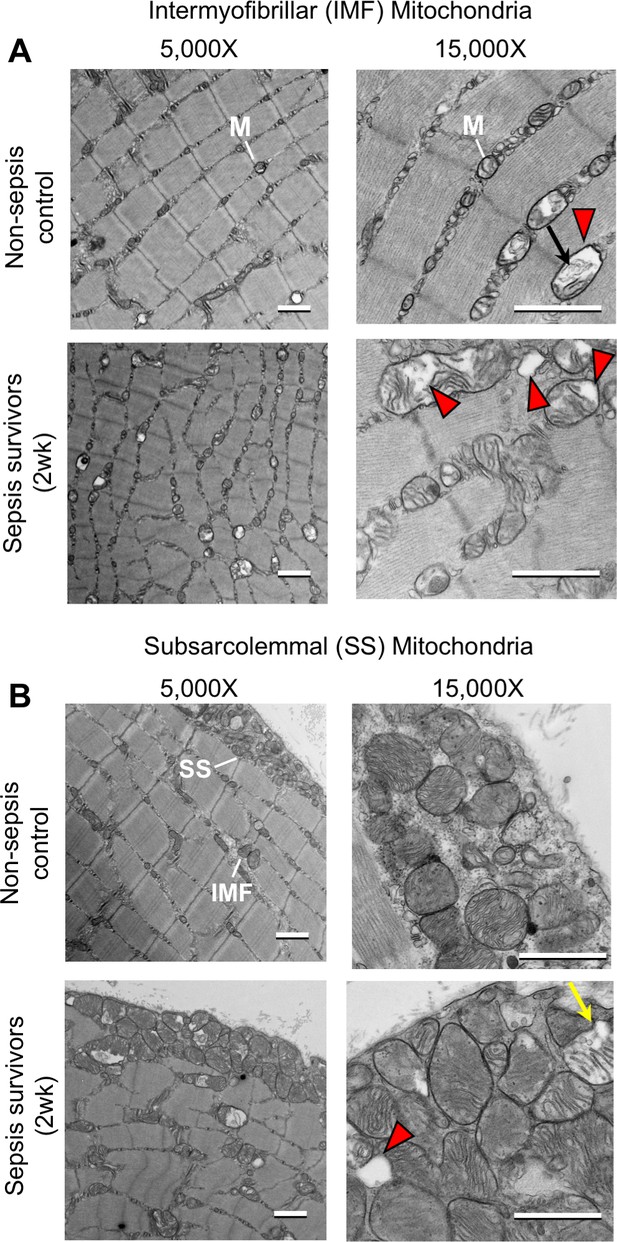
Mitochondrial ultrastructure in tibialis anterior skeletal muscle of murine sepsis survivors.
Transmission electron microscopy was performed on tibialis anterior specimens from murine sepsis survivors (2wk) and non-sepsis controls (n = 3 per group) as an additional observation from the experiment shown in Figure 4. Representative micrographs of (A) intermyofibrillar (IMF) and (B) subsarcolemmal (SS) mitochondria at 5,000X and 15,000X magnifications are shown (scale bars represent 1,000 nm). Abnormal mitochondrial structures are indicated with the following symbols: destruction of cristae with expanded matrix space (red arrowheads), concentric ‘onion shaped’ cristae (black arrows), and compartmentalization into vacuolar structures (yellow arrows).
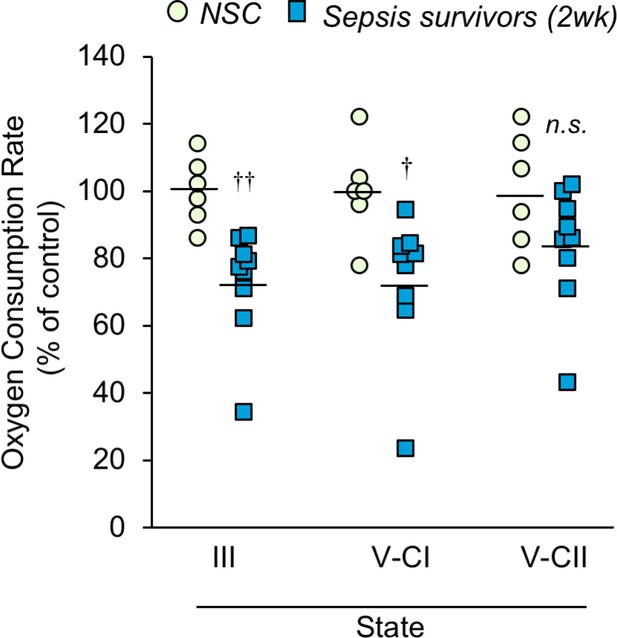
Mitochondrial respiration capacity is reduced in skeletal muscle of sepsis survivors.
Mitochondria were isolated from the tibialis anterior of non-sepsis control (NSC, n = 6) and sepsis survivors (2wk, n = 9) and oxygen consumption rate was measured in triplicate for each sample. State III refers to the ADP phosphorylation rate. States V-CI and V-CII refer to complex I and complex II driven maximum electron transport, respectively. Data are represented as individual data points with mean values shown. Two-sample t-tests were performed. n.s. not significant, † p<0.05, †† p<0.01 compared to NSC.
-
Figure 5—source data 1
Mitochondrial respiration capacity is reduced in skeletal muscle of sepsis survivors.
This data file includes data presented in main Figure 5 as well as Figure 5—figure supplement 1.
- https://doi.org/10.7554/eLife.49920.016
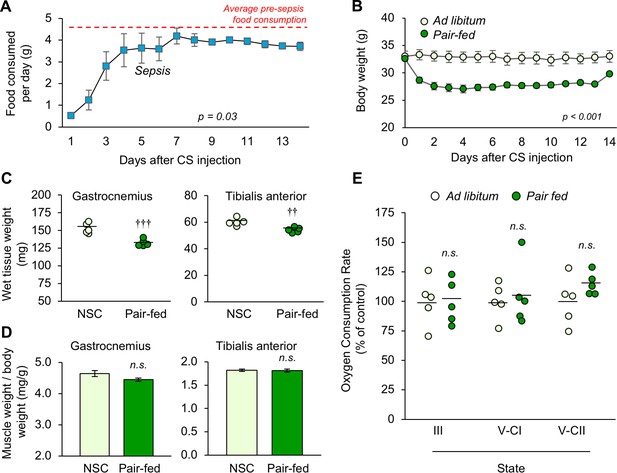
Mitochondrial respiration is unchanged in pair-fed food-restricted mice.
Daily food consumption was measured pre-sepsis for 5 days, and for 2 weeks following sepsis induced by cecal slurry injection (n = 5 sepsis survivors, (A). A set of non-sepsis animals was pair-fed based on these data alongside ad libitum (freely-fed) mice (n = 5 per group). Body weight was monitored regularly (B). At euthanasia, hindlimb skeletal muscles were weighed (C), and weights are shown respective to body weight (D). Mitochondria were isolated from the tibialis anterior of pair-fed and ad libitum mice, and oxygen consumption rate was measured in triplicate for each sample (E). Refer to Figure 5 for details. Repeated measures ANOVA (A–B), one-way ANOVA (C–D), and two-sample t-tests (E) were performed. n.s. not significant, †† p<0.01, ††† p<0.001 compared to average baseline food consumption (A) or to ad libitum controls (B–E).
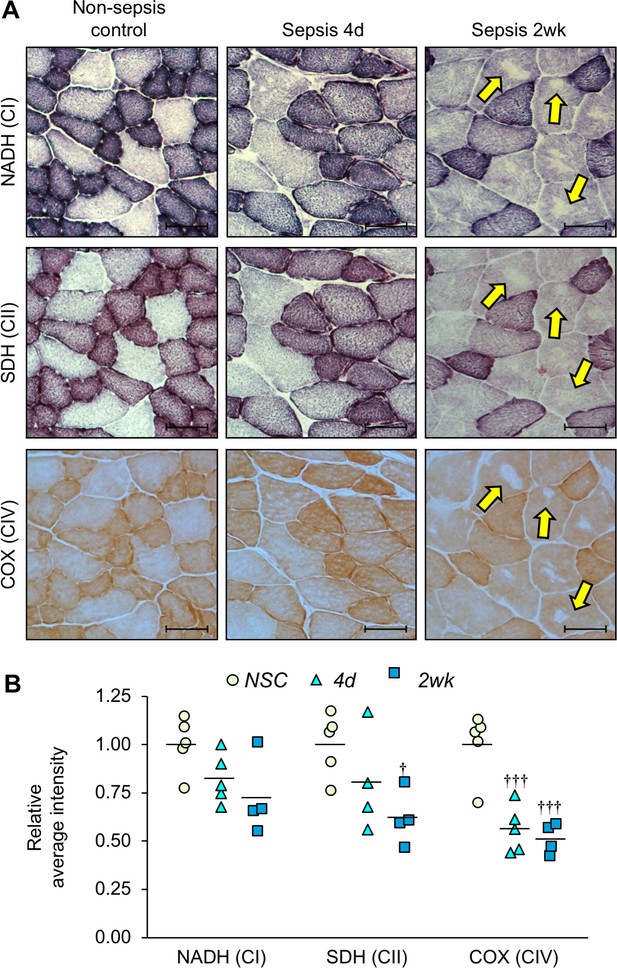
Mitochondrial enzyme activities are progressively reduced over time in sepsis survivors.
Tibialis anterior specimens were utilized for histochemical staining of oxidative phosphorylation enzyme activities in non-sepsis controls (n = 5), and after sepsis at 4d (n = 4–5) and 2wk (n = 4). (A) Representative images of staining for nicotinamide adenine dinucleotide dehydrogenase (NADH; complex I, top), succinate dehydrogenase (SDH; complex II, middle), and cytochrome C oxidase (COX; complex IV, bottom) enzyme activities conducted on serial sections. Scale bars represent 50 µm; arrows indicate areas devoid of mitochondrial enzyme activity and correspond to the same fiber in serial sections. (B) Average intensities were quantified using Aperio ImageScope software and normalized to the intensity of the controls. Data were analyzed by one-way ANOVA. † p<0.05, ††† p<0.001 compared to NSC.
-
Figure 6—source data 1
Mitochondrial enzyme activities are progressively reduced over time in sepsis survivors.
- https://doi.org/10.7554/eLife.49920.018
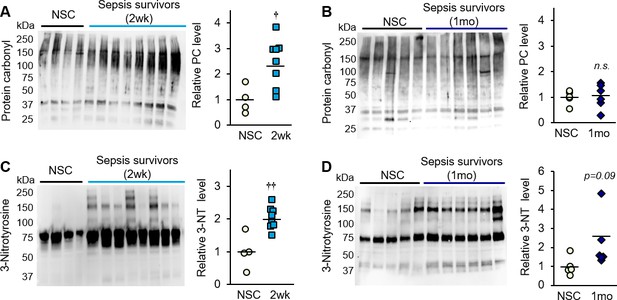
Oxidative damage is evident in skeletal muscle of sepsis survivors.
Protein isolated from whole tibialis anterior muscles of non-sepsis control (NSC) and sepsis survivors was used to assess markers of oxidative damage by Western blot. Protein carbonylation (PC) and nitrotyrosination (3-NT) were detected in NSC (n = 4–5), and sepsis survivors at (A, C) 2 weeks (n = 8) and (B, D) 1 month (n = 6). Uncropped blots are shown. Total intensity of each lane was quantified by densitometric analysis and normalized to total protein content. Data are expressed as individual data points with mean values shown. Two-sample t-tests were performed. n.s. not significant, † p<0.05, †† p<0.01 compared to NSC.
-
Figure 7—source data 1
Oxidative damage is evident in skeletal muscle of sepsis survivors.
- https://doi.org/10.7554/eLife.49920.020
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic Reagent (M. musculus; male and female) | C57BL/6 mice | National Institute on Aging | RRID:SCR_007317 | 16 months old |
| Biological sample (M. musculus) | Cecal slurry; CS | Starr and Saito, 2014; Steele et al., 2017 | Donor mice: 16 week old male C57BL/6 mice obtained from The Jackson Laboratory | |
| Antibody | Mouse monoclonal myosin heavy chain type I (MIgG2b) | Developmental Studies Hybridoma Bank | Cat# BA-D5; RRID: AB_2235587 | 1:100 |
| Antibody | Mouse monoclonal myosin heavy chain type IIa (MIgG1) | Developmental Studies Hybridoma Bank | Cat# SC-71; RRID: AB_2147165 | 1:100 |
| Antibody | Mouse monoclonal myosin heavy chain type IIb (MIgM) | Developmental Studies Hybridoma Bank | Cat# BF-F3; RRID: AB_2266724 | 1:100 |
| Antibody | Goat anti-mouse IgG2b, Alexa Fluor 647 conjugated | Life Technologies- Thermo Fisher | Cat# A-21242; RRID: AB_2535811 | 1:250 |
| Antibody | Goat anti-mouse IgG1, Alexa Fluor 488 conjugated | Life Technologies- Thermo Fisher | Cat# A-21121; RRID: AB_2535764 | 1:500 |
| Antibody | Goat anti-mouse IgM, Alexa flour 555 conjugated | Life Technologies- Thermo Fisher | Cat# A-21426; RRID: AB_2535847 | 1:250 |
| Antibody | Mouse monoclonal 3-Nitrotyrosine (39B6) | Abcam | Cat# ab61392; RRID: AB_942087 | 1:3000 |
| Antibody | Goat anti-mouse IgG-HRP | Santa Cruz | Cat# SC-2005 | 1:10,000 |
| Commercial assay or kit | High sensitivity IL-6 mouse ELISA kit | eBIOSCIENCES- Thermo Fisher | Cat# BMS603HS; RRID: AB_2575654 | Sepsis 4d plasma samples diluted 5X for assay |
| Commercial assay or kit | V-PLEX multiplex assay | V-Plex Proinflammatory Panel 1 Mouse Kit, customized | Cat# K15048D | |
| Commercial assay or kit | PureLink RNA Mini Ki | Invitrogen-Thermo Fisher | Cat# 12183025 | |
| Commercial assay or kit | SuperScript III First-Strand Synthesis SuperMix | Life Technologies- Thermo Fisher | Cat# 18080400 | |
| Commercial assay or kit | TaqMan mouse IL-6 gene expression assay | Thermo Fisher | Assay ID: Mm00446190_m1 | |
| Commercial assay or kit | TaqMan mouse TNFα gene expression assay | Thermo Fisher | Assay ID: Mm00443258_m1 | |
| Commercial assay or kit | TaqMan mouse IL-10 gene expression assay | Thermo Fisher | Assay ID: Mm01288386_m1 | |
| Commercial assay or kit | TaqMan mouse HPRT gene expression assay | Thermo Fisher | Assay ID: Mm03024075_m1 | |
| Commercial assay or kit | BCA protein assay kit | Thermo Fisher | Cat# 23225 | |
| Commercial assay or kit | RC DC protein assay kit II | Bio-Rad | Cat# 5000122 | |
| Commercial assay or kit | OxyBlot Protein Oxidation Detection Kit | EMD Millipore | Cat# S7150 | |
| Chemical compound, drug | Imipenem; IPM | Primaxin IV; imipenem 500 mg stabilized in cilastatin | NDC 0006-3516-59 | 1.5 mg/mouse |
| Chemical compound, drug | Physiological Saline (0.9%) | Abbott Laboratories | Ref. # 04930-04-10 | |
| Software | ZEN (blue edition) imaging software | Zeiss | RRID:SCR_013672 | |
| Software | ImageJ software (version 1.46 r) | National Institutes of Health | RRID:SCR_003070 | |
| Software | Image Lab software (2017) | Bio-Rad | Cat# #1709690 | |
| Software | SAS 9.4 | SAS Institute Inc | ||
| Software | Aperio ImageScope software (12.4) | Leica | RRID:SCR_014311 | Positive-pixel algorithm |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.49920.021





