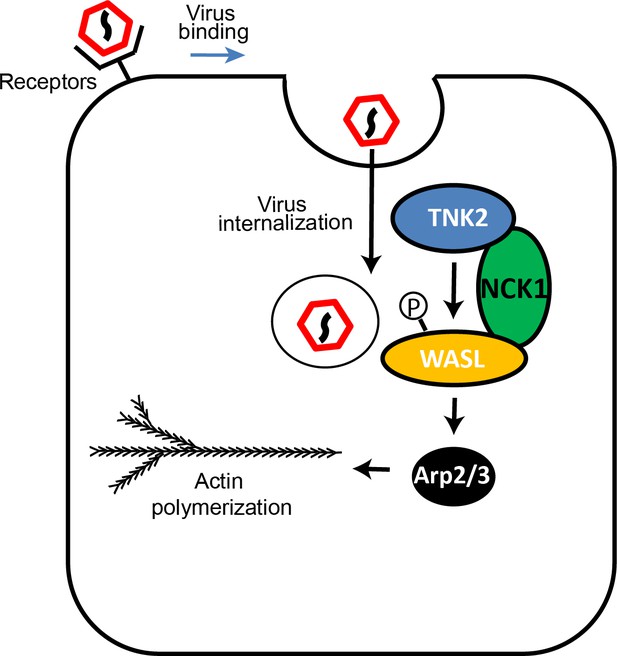
Entry by multiple picornaviruses is dependent on a pathway that includes TNK2, WASL, and NCK1
Figures
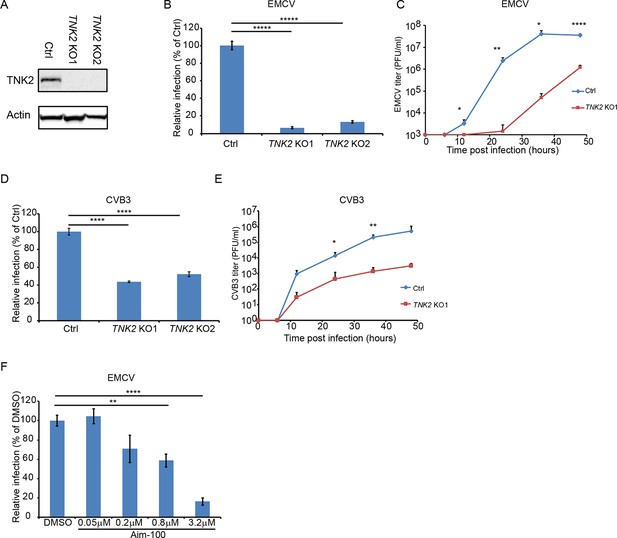
TNK2 is critical for multiple picornavirus infections.
(A) TNK2 protein expression in TNK2 KO1, TNK2 KO2, and Ctrl (control) cells generated by CRISPR-Cas9 genome editing with either specific targeting or non-specific targeting sgRNA in A549 cells. Cells lysates were analyzed by Western blot. (B) FACS quantification of EMCV positive cells for TNK2 KO1, TNK2 KO2, and Ctrl cells 10 hr post infection at an MOI of 1. (C) Multi-step growth curve for EMCV multiplication on TNK2 KO1 and Ctrl cells at an MOI of 0.01. Virus titers in the culture supernatant were quantified by plaque assay at 0, 6, 12, 24, 36, and 48 hr post infection. (D) FACS quantification of CVB3 virus positive cells for TNK2 KO1, TNK2 KO2, and Ctrl cells 8 hr post infection at an MOI of 1. (E) Multi-step growth curve for CVB3 multiplication on TNK2 KO1 and Ctrl cells at an MOI of 0.01. Virus titers in the culture supernatant were quantified by plaque assay at 0, 6, 12, 24, 36, and 48 hr post infection. (F) Aim-100 inhibition of EMCV infection on naïve A549 cells. A549 cells were pre-treated with Aim-100 at indicated concentrations and infected with EMCV at an MOI of 1. Virus positive cells were quantified by FACS. (B, D, F) Error bars represent standard deviation of three replicates. The data shown are representatives of three independent experiments. *: p<0.05, ***: p<0.001, ****: p<0.0001, *****: p<0.00001.
-
Figure 1—source data 1
Source data for Figure 1B, D and F: FACS quantification of virus infected cells.
- https://cdn.elifesciences.org/articles/50276/elife-50276-fig1-data1-v2.xlsx
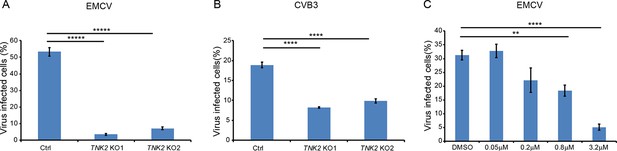
TNK2 is critical for multiple picornavirus infections.
(A) FACS quantification of EMCV virus positive cells (without normalization) for TNK2 KO1, TNK2 KO2, and Ctrl cells 10 hr post infection at an MOI of 1. (B) FACS quantification of CVB3 virus positive cells (without normalization) for TNK2 KO1, TNK2 KO2, and Ctrl cells 8 hr post infection at an MOI of 1. (C) Aim-100 inhibition of EMCV infection on naïve A549 cells. A549 cells were pre-treated with Aim-100 at indicated concentrations and infected with EMCV at an MOI of 1. Virus positive cells were quantified by FACS and plotted (without normalization). (A–C) Error bars represent standard deviation of three replicates. The data shown are representatives of three independent experiments. **: p<0.01, ***: p<0.001, ****: p<0.0001, *****: p<0.00001.
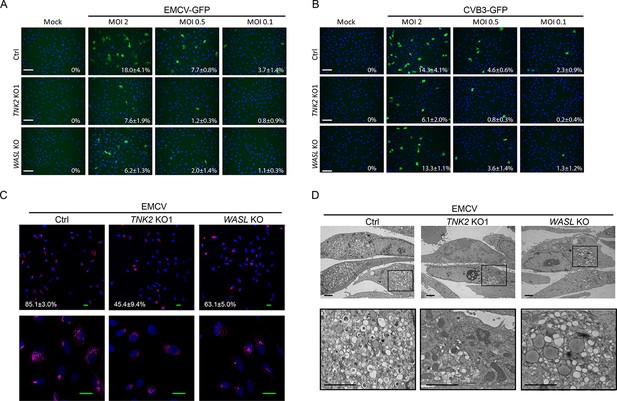
TNK2 and WASL are critical for multiple picornavirus infection on A549 cells.
(A) EMCV-GFP infection on Ctrl, TNK2 KO1 and WASL KO A549 cells at MOI 2, 0.5 and 0.1. Percentage of positive cells were quantified (values denote mean ±s.d., n = 3 images quantified). Scale bars represent 20 μm. (B) CVB3-GFP infection on Ctrl, TNK2 KO1 and WASL KO A549 cells at MOI 2, 0.5 and 0.1. Percentage of positive cells were quantified (values denote mean ±s.d., n = 3 images quantified). Scale bars represent 20 μm. (C) dsRNA immunostaining of EMCV infection on Ctrl, TNK2 KO1, and WASL KO A549 cells. Percentage of positive cells were quantified (values denote mean ±s.d., n = 3 images quantified). Scale bars represent 10 μm. (D) Electron microscopy detection of EMCV replication complex on Ctrl, TNK2 KO1, and WASL KO A549 cells. Scale bars represent 2 μm.
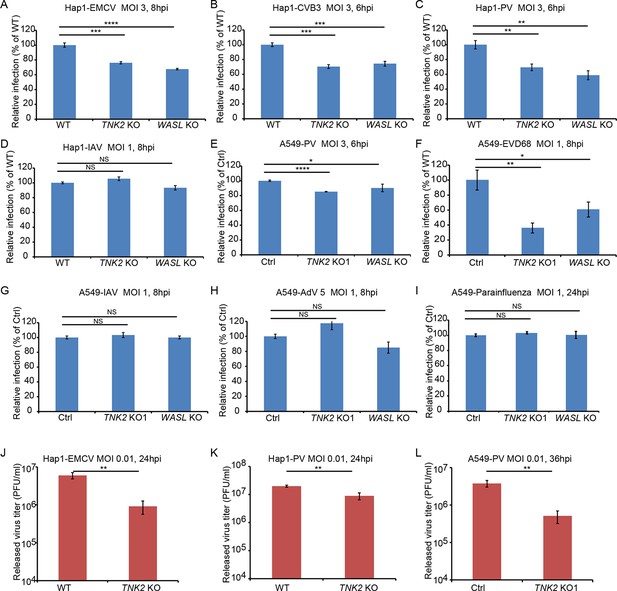
TNK2 and WASL are critical for multiple picornavirus infection on both A549 and Hap1 cells.
(A–D) FACS quantification of EMCV, CVB3, Poliovirus (PV), and influenza A virus (IAV) virus infection on WT (wild type), TNK2 KO and WASL KO Hap1 cells. (E–I) FACS quantification of poliovirus (PV), enterovirus D68 (EVD68), influenza A virus (IAV) virus, Adenovirus type 5 (AdV5) and parainfluenza virus infection on Ctrl, TNK2 KO and WASL KO A549 cells. (J) EMCV growth titration on WT and TNK2 KO Hap1 cells at 24 hr post infection. (K) Poliovirus growth titration on WT and TNK2 KO Hap1 cells at 24 hr post infection. (L) Poliovirus growth titration on Ctrl and TNK2 KO1 A549 cells at 36 hr post infection. (A–L) Error bars represent standard deviation of three replicates. The data shown are representative of at least two independent experiments. *: p<0.05, **: p<0.01, ***: p<0.001, ****: p<0.0001, NS: not significant (p>0.05).
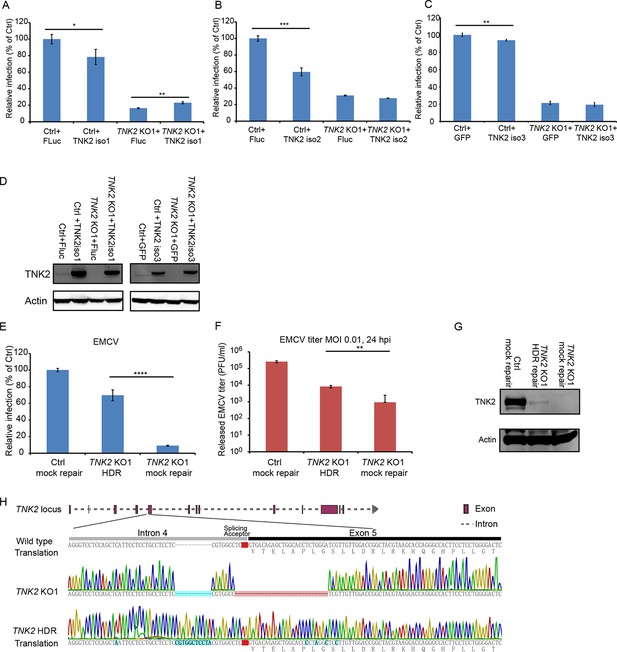
EMCV virus infection of TNK2 rescue on knockout and control cells.
(A–C) Quantification of EMCV infection on TNK2 KO1 and Ctrl cells transduced with TNK2 isoform 1, 2 and 3 and Fluc. Cells were infected with EMCV at an MOI of 1 and quantified at 10 hr post infection. (D) Expression of TNK2 in Ctrl and TNK2 KO1 cells transduced with lentivirus expressing TNK2 isoform 1, isoform 3, and Fluc or GFP by Western blot. (E) FACS quantification of EMCV positive cells for Ctrl mock repair, TNK2 KO1 HDR (homologous template-directed recombination), and TNK2 KO1 mock repair cells 10 hr post infection at an MOI of 1. Mock repair: cells subjected to the same HDR genome editing but with non-specific targeting sgRNA. (F) EMCV growth titration on Ctrl mock repair, TNK2 KO1 HDR, and TNK2 KO1 mock repair cells at 24 hr post infection. (G) TNK2 protein expression in Ctrl mock repair, TNK2 KO1 HDR cells and TNK2 KO1 mock repair cells. Cells lysates were analyzed by Western blot. (H) Sequence alignment of TNK2 KO1 cells and TNK2 KO1 HDR repaired cells. Splice acceptor is marked with red; synonymous mutations introduced by design and random insertion mutations are marked with blue. (A–C, E, F) Error bars represent standard deviation of three replicates. The data shown are representative of at least two independent experiments. Fluc: firefly luciferase. *: p<0.05, **: p<0.01, ***: p<0.001, NS: not significant (p>0.05).
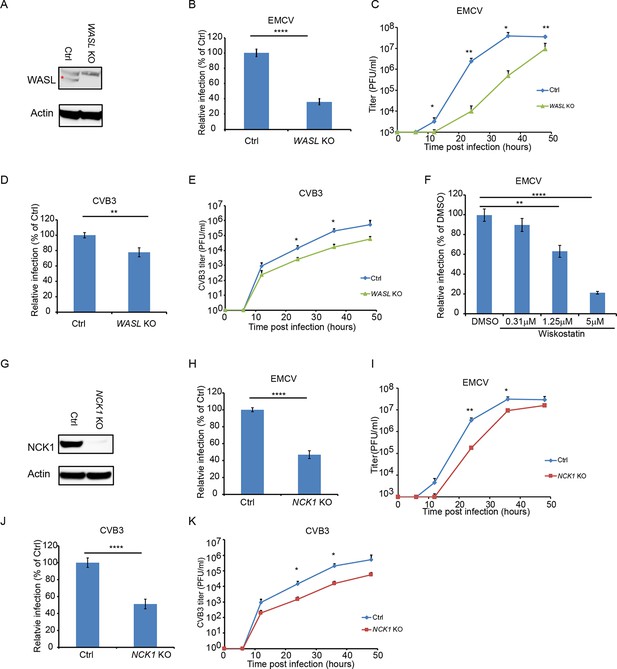
WASL and NCK1 are critical for multiple picornavirus infections.
(A) WASL protein expression in WASL KO and Ctrl cells generated by CRISPR-Cas9 genome editing with either specific targeting or non-specific targeting sgRNA in A549 cells. Cells lysates were analyzed by Western blot. (B) FACS quantification of EMCV positive cells for WASL KO and Ctrl cells 10 hr post infection at an MOI of 1. (C) Multi-step growth curve for EMCV multiplication on WASL KO and Ctrl cells infected at an MOI of 0.01. (D) FACS quantification of CVB3 positive cells for WASL KO and Ctrl cells 8 hr post infection at an MOI of 1. (E) Multi-step growth curve for CVB3 multiplication on WASL KO and Ctrl cells infected at an MOI of 0.01. (F) Wiskostatin inhibition of EMCV infection on naïve A549 cells. A549 cells were pre-treated with Wiskotstatin at indicated concentrations and infected with EMCV at an MOI of 1. Virus positive cells were quantified by FACS. (G) NCK1 protein expression in NCK1 KO and Ctrl cells generated by CRISPR-Cas9 genome editing with either specific targeting or non-specific targeting sgRNA in A549 cells. Cells lysates were analyzed by Western blot. (H) FACS quantification of EMCV positive cells for NCK1 KO and Ctrl cells 10 hr post infection at an MOI of 1. (I) Multi-step growth curve for EMCV multiplication on NCK1 KO and Ctrl cells infected at an MOI of 0.01. (J) FACS quantification of CVB3 positive cells for NCK1 KO and Ctrl cells 8 hr post infection at an MOI of 1. (K) Multi-step growth curve for CVB3 multiplication on NCK1 KO and Ctrl cells infected at an MOI of 0.01. (A) The red asterisk indicates WASL protein band. (B, D, F, H, J) Error bars represent standard deviation of three replicates. The data shown are representatives of at least two independent experiments. **: p<0.01, ***: p<0.001, ****: p<0.0001, *****: p<0.00001, NS: not significant (p>0.05).
-
Figure 2—source data 1
Source data for Figure 2B, D, F, H and J: FACS quantification of virus infected cells.
- https://cdn.elifesciences.org/articles/50276/elife-50276-fig2-data1-v2.xlsx
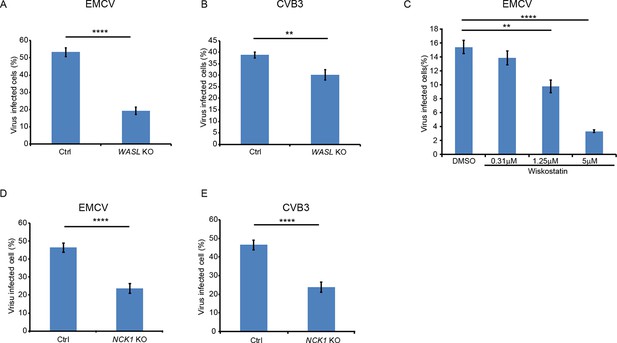
WASL and NCK1 are critical for multiple picornavirus infections.
(A) FACS quantification of EMCV positive cells (without normalization) for WASL KO and Ctrl cells 10 hr post infection at an MOI of 1. (B) FACS quantification of CVB3 positive cells (without normalization) for WASL KO and Ctrl cells 10 hr post infection at an MOI of 1. (C) Wiskostatin inhibition of EMCV infection on naïve A549 cells. A549 cells were pre-treated with Wiskotstatin at indicated concentrations and infected with EMCV at an MOI of 1. Virus positive cells were quantified by FACS and plotted (without normalization). (D) FACS quantification of EMCV positive cells (without normalization) for NCK1 KO and Ctrl cells 8 hr post infection at an MOI of 1. (E) FACS quantification of CVB3 positive cells (without normalization) for NCK1 KO and Ctrl cells 8 hr post infection at an MOI of 1. Error bars represent standard deviation of three replicates. The data shown are representatives of at least two independent experiments. **: p<0.01, ***: p<0.001, ****: p<0.0001, *****: p<0.00001, NS: not significant (p>0.05).
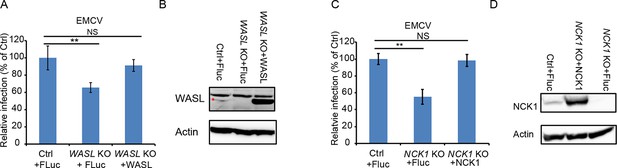
EMCV infection of WASL and NCK1 rescued knock out cells.
(A) FACS quantification of EMCV positive cells for lentivirus-mediated WASL rescue in WASL KO and Ctrl cells at an MOI of 1. Fluc: firefly luciferase. (B) Western blot detection of lentivirus-mediated WASL expression in WASL KO and Ctrl cells. (C) FACS quantification of EMCV positive cells for lentivirus-mediated NCK1 rescue in NCK1 KO and Ctrl cells at an MOI of 1. (D) Western blot detection of lentivirus-mediated NCK1 expression in NCK1 KO and Ctrl cells. Error bars represent standard deviation of three replicates. The data shown are representatives of at least two independent experiments. **: p<0.01, NS: not significant (p>0.05).
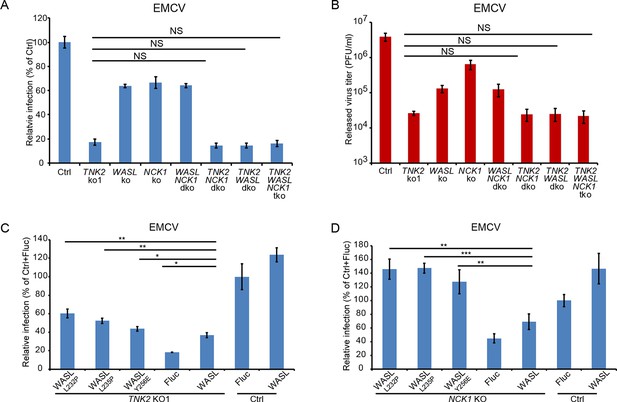
TNK2, WASL, and NCK1 are in a pathway supporting virus infection.
(A) FACS quantification of EMCV positive cells for TNK2, WASL, NCK1 single, double, triple gene knockout and Ctrl cells 10 hr post infection at an MOI of 1. (B) Virus titer for EMCV multiplication on TNK2, WASL, and NCK1 single, double, triple gene knockout and Ctrl cells at 24 hr post infection at an MOI of 0.01. (C) FACS quantification of EMCV positive cells for TNK2 KO1 cells that were transduced with constitutively active WASL constructs 10 hr post infection at an MOI of 1. (D) FACS quantification of EMCV positive cells for NCK1 KO cells that were transduced with constitutively active WASL constructs 10 hr post infection at an MOI of 1. (A, B) dko: double knockout, tko: triple knockout. (A–D) Error bars for virus infection represent standard deviation of three replicates. The data shown are representatives of two independent experiments.
-
Figure 3—source data 1
Source data for Figure 3A, C and D: FACS quantification of virus infected cells.
- https://cdn.elifesciences.org/articles/50276/elife-50276-fig3-data1-v2.xlsx
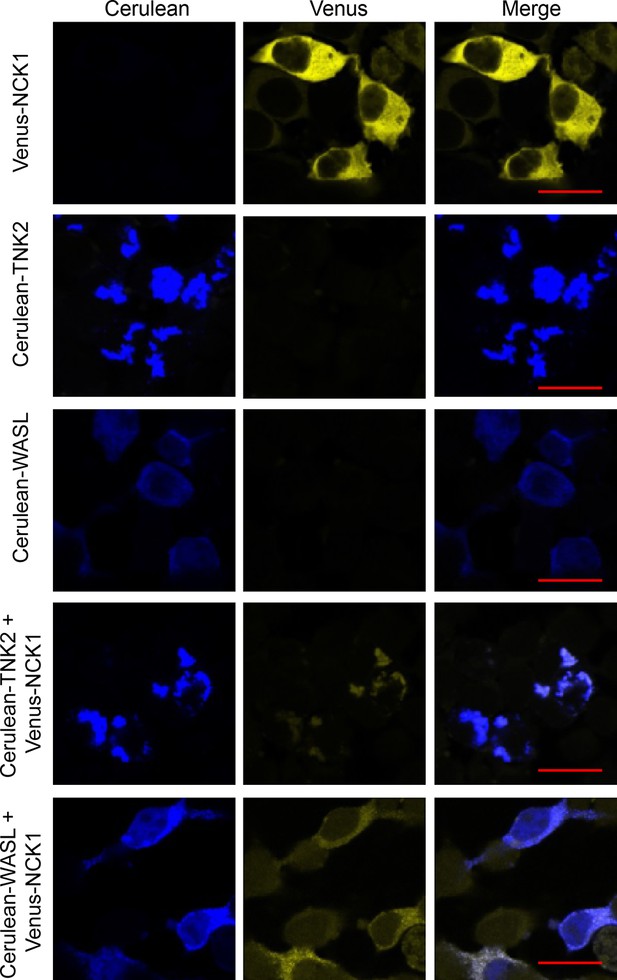
Co-localization of fluorescently tagged NCK1, TNK2, and WASL expressed in 293 T cells by confocal imaging.
293 T cells were transfected with Venus-NCK1, Cerulean-TNK2, Cerulean-WASL individually, Venus-NCK1 with Cerulean-TNK2 or Venus-NCK1 with Cerulean-WASL. Cells were imaged by confocal 24 hr after transfection. Scale bars represent 20 μm.
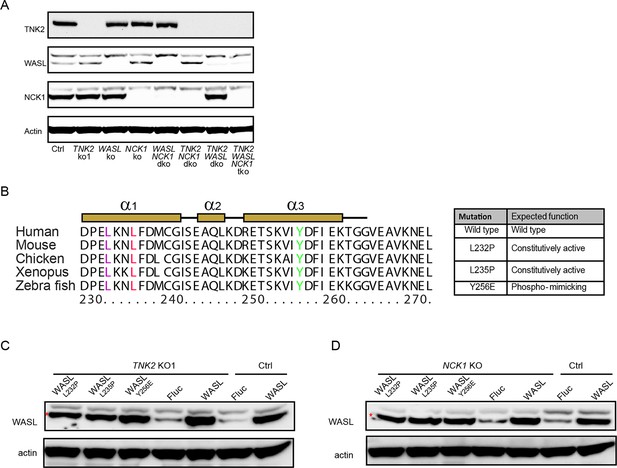
Gene expression in knock out cells and constitutively active WASL expression.
(A) Western blot detection of TNK2, WASL, and NCK1 expression on single, double, triple gene knockout and Ctrl cells. (B) Sequence alignment of WASL protein sequences from different vertebrate species. Table indicates specific mutation that has constitutive activity. (C) Expression of WASL in Ctrl and TNK2 KO1 cells transduced with lentivirus expressing constitutively active WASL, wild type WASL, or Fluc. Cell lysates were analyzed by Western blot. (D) Expression of WASL in Ctrl and NCK1 KO cells transduced with lentivirus expressing constitutively active WASL, wild type WASL, or Fluc. Cell lysates were analyzed by Western blot. (C, D) The red asterisks indicate WASL protein band.
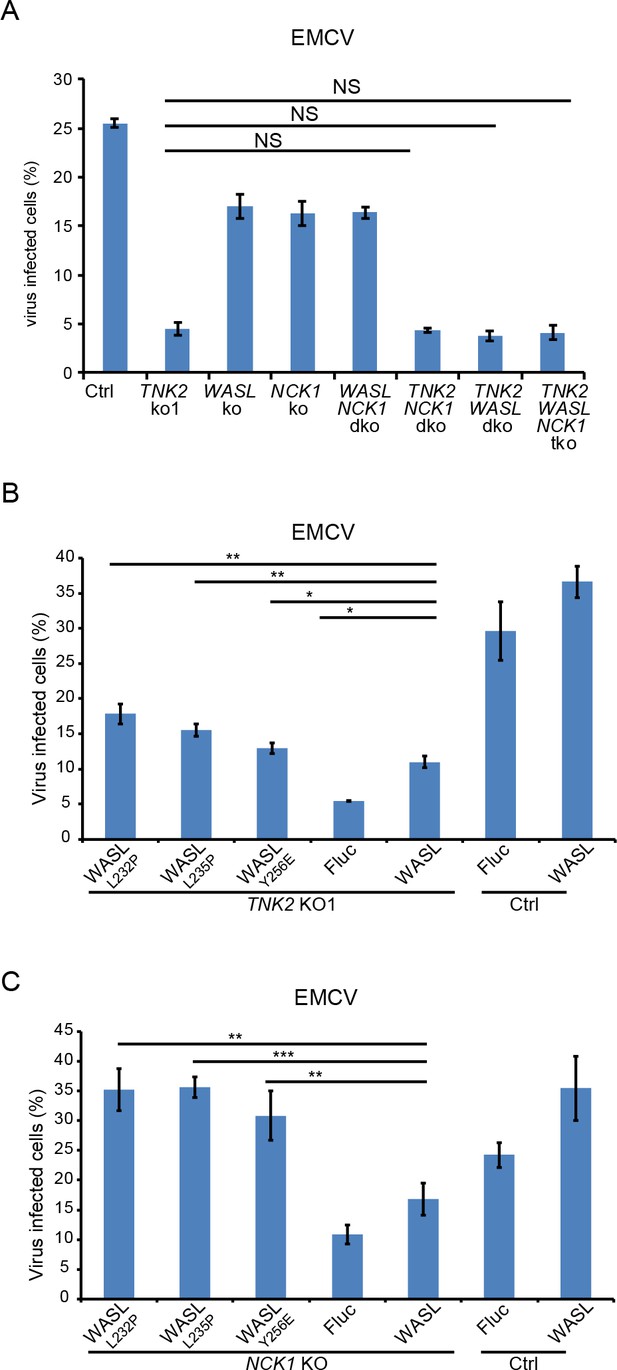
TNK2, WASL, and NCK1 are in a pathway supporting virus infection.
(A) FACS quantification of EMCV positive cells (without normalization) for TNK2, WASL, NCK1 single, double, triple gene knockout and Ctrl cells 10 hr post infection at an MOI of 1. (B) FACS quantification of EMCV positive cells (without normalization) for TNK2 KO1 cells that were transduced with constitutively active WASL constructs 10 hr post infection at an MOI of 1. (C) FACS quantification of EMCV positive cells (without normalization) for NCK1 KO cells that were transduced with constitutively active WASL constructs 10 hr post infection at an MOI of 1. (A, B, C) dko: double knockout, tko: triple knockout. (A–C) Error bars for virus infection represent standard deviation of three replicates. The data shown are representatives of two independent experiments.
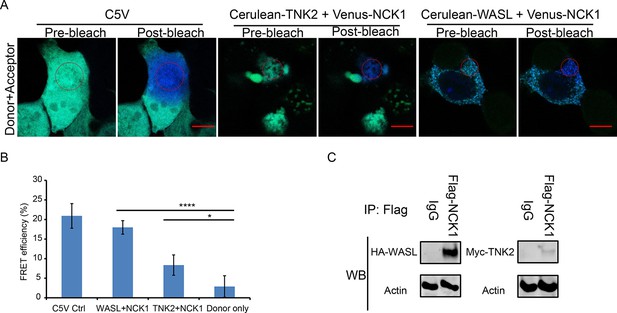
TNK2 and WASL directly interact with NCK1.
(A) FRET imaging of C5V positive control, NCK1 and TNK2, and NCK1 and WASL expression in 293 T cells before and after acceptor photo bleach. The red circles indicate photo bleached areas. Scale bars represent 10 μm. (B) Quantification of FRET efficiency of C5V positive control, NCK1 and TNK2, and NCK1 and WASL expression in 293 T cells. Error bars for FRET efficiency represent standard deviation of average from four individually bleached images. *: p<0.05, ****: p<0.0001, NS: not significant (p>0.05). (C) Immunoprecipitation of FLAG-tagged NCK1 with HA-tagged WASL and FLAG-tagged NCK1 with Myc-tagged TNK2.
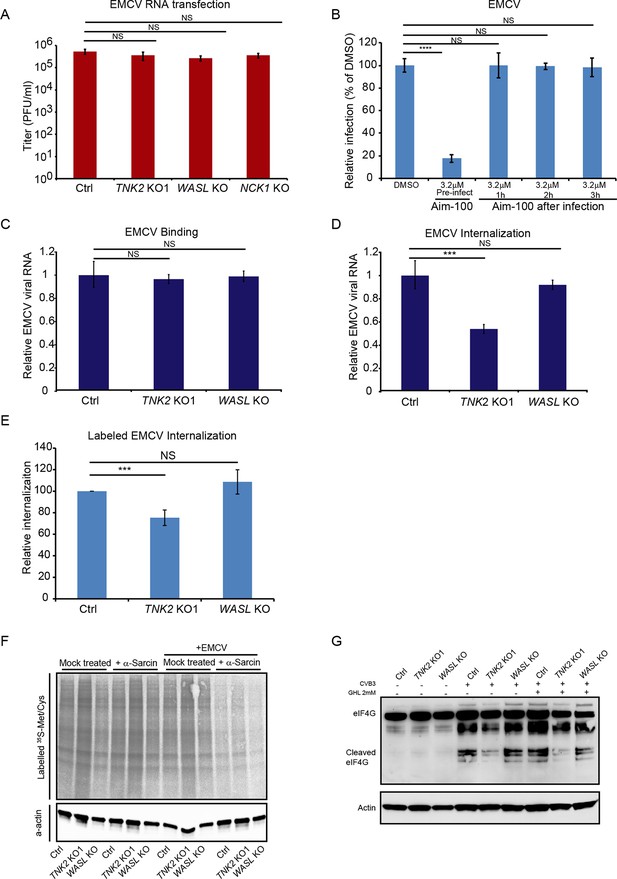
TNK2, WASL, and NCK1 function at an early stage of virus lifecycle.
(A) EMCV released from viral RNA transfected Ctrl, TNK2 KO1, WASL KO, and NCK1 KO cells 10 hr post transfection was quantified by plaque assay. (B) Time-dependent addition of Aim-100 on EMCV infection on naïve A549 cells. A549 cells were treated with 3.2 μM Aim-100 at different time points before and after EMCV infection at an MOI of 1. EMCV positive cells were then quantified by FACS. (C) Quantification of EMCV virus binding on TNK2 KO1 and WASL KO cells by qRT-PCR expressed as relative change to Ctrl cell binding. (D) Quantification of EMCV virus internalization in TNK2 KO1 and WASL KO cells by qRT-PCR expressed as relative change to Ctrl cell internalization. (E) FACS quantification of labeled EMCV internalization in TNK2 KO1, WASL KO and Ctrl cells. (F) a-sarcin pore forming assay performed on TNK2 KO1, WASL KO and Ctrl cells. Translation was measured by phosphorimaging of 35S-methionine/cysteine incorporation. (G) eIF4G cleavage by CVB3 infection for 2 hr with or without 2 mM guanidine hydrochloride. (A–E) Error bars represent standard deviation of three replicates. The data shown are representative of two independent experiments. ***: p<0.001, ****: p<0.0001, NS: not significant (p>0.05).
-
Figure 4—source data 1
Source data for Figure 4B: FACS quantification of virus infected cells.
- https://cdn.elifesciences.org/articles/50276/elife-50276-fig4-data1-v2.xlsx
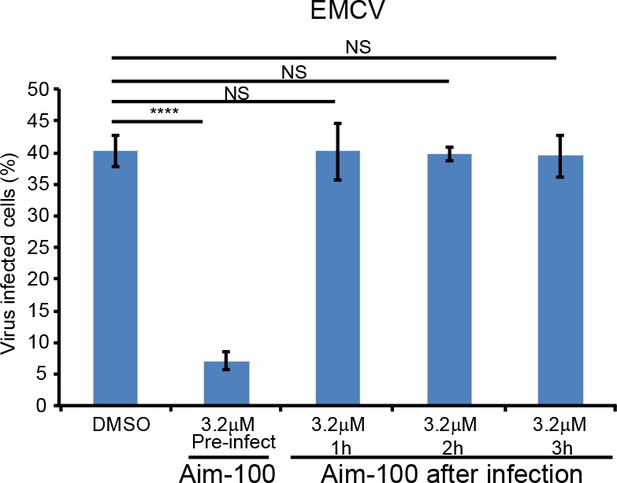
TNK2, WASL, and NCK1 function at an early stage of virus lifecycle.
Time dependent addition of Aim-100 on EMCV infection on naïve A549 cells. A549 cells were treated with 3.2 μM Aim-100 at different time points before and after EMCV infection at an MOI of 1. EMCV positive cells were then quantified by FACS and plotted without normalization. ****: p<0.0001, NS: not significant (p>0.05).

EMCV internalization in Ctrl, TNK2 KO1 and WASL KO cells.
(A) Labeled EMCV viruses bind and internalize in TNK2 KO1, WASL KO, and Ctrl cells. Scale bars represents 20 μm. (B) FACS plot of labeled EMCV internalization in TNK2 KO1, WASL KO, and Ctrl cells. (C–E) Trypsin treatment removes viral particles bound on the cell surface equally. qRT-PCR quantification of virus bound with Ctrl, TNK2 KO1 and WASL KO cell either treated or mock treated with trysin for 6 min after virus binding on ice for 1 hr, then washed five times with cold PBS. **: p<0.01, ***: p<0.001.
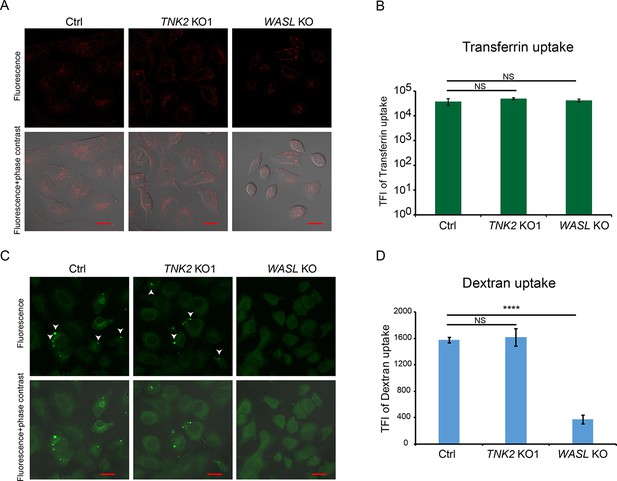
Transferrin, and dextran uptake in Ctrl, TNK2 KO1, and WASL KO cells.
(A) Fluorescent images of transferrin uptake in Ctrl, TNK2 KO1, and WASL KO cells. Scale bars represent 20 μm. (B) Quantification of transferrin uptake by flow cytometry. Error bars represent standard deviation of three replicates. The data shown are representative of two independent experiments. TFI: total fluorescence intensity. NS: not significant (p>0.05). (C) Fluorescent images of dextran uptake in Ctrl, TNK2 KO1, and WASL KO cells. White arrows indicate macropinosomes. Scale bars represent 20 μm. (D) Quantification of Dextran uptake by flow cytometry. Error bars represent standard deviation of three replicates. The data shown are representative of two independent experiments. TFI: total fluorescence intensity. NS: not significant (p>0.05), ****: p<0.0001.
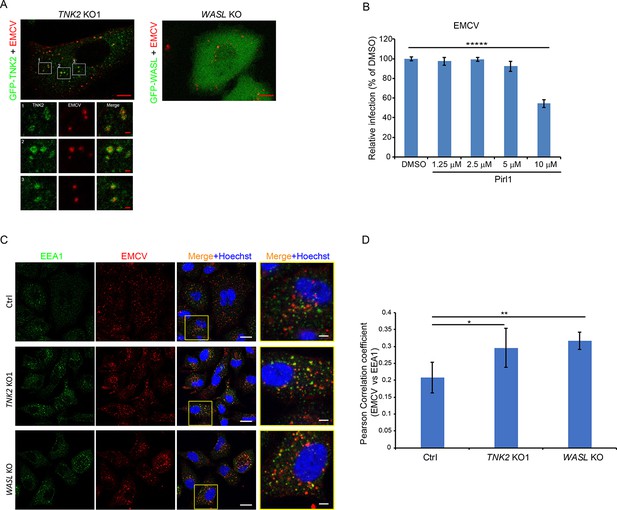
TNK2 mediates virus infection through endosomal trafficking pathways.
(A) Confocal imaging of GFP-tagged TNK2 localization with fluorescently labeled EMCV virus in TNK2 KO1 cells and GFP-tagged WASL localization with fluorescently labeled EMCV virus in WASL KO cells. Scale bars represent 10 μm. Individual channels of different insets were shown. Scale bars represent 2 μm. (B) FACS quantification of EMCV infection on pirl1 treated A549 cells at 10 hr post infection at an MOI of 1. Error bars represent standard deviation of three replicates. (C) EEA1 staining of fluorescently labeled EMCV infected Ctrl, TNK2 KO1, and WASL KO cells. Scale bars represent 20 μm. Insets represent magnification of the boxed region. Scale bars represent 5 μm. (D) Quantification of Pearson correlation coefficient of EMCV and EEA1 colocalization in Ctrl, TNK2 KO1, and WASL KO cells infected with fluorescently labeled EMCV. 33 cells for Ctrl, 37 cells for TNK2 KO1 and 19 cells for WASL KO were quantified. Error bars represent standard deviation of images quantified. (B, D) The data shown are representative of two independent experiments. *: p<0.05, **: p<0.01.
-
Figure 5—source data 1
Source data for Figure 5B: FACS quantification of virus infected cells.
- https://cdn.elifesciences.org/articles/50276/elife-50276-fig5-data1-v2.xlsx
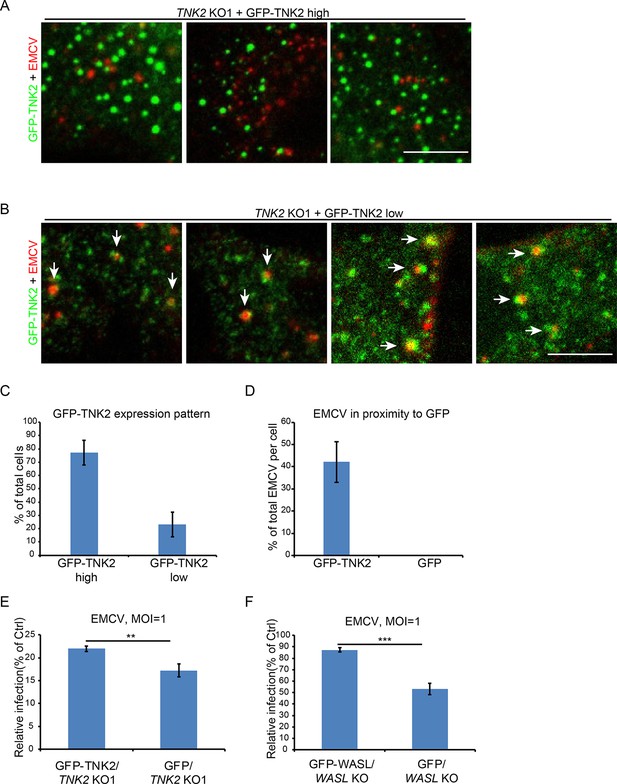
Localization of GFP-TNK2 with fluorescently labeled EMCV.
(A) Fluorescent images of high GFP-TNK2 expression patterns in TNK2 KO1 cells and its localization with fluorescently labeled EMCV. Scale bar represents 10 μm. (B) Fluorescent images of low GFP-TNK2 expression patterns in TNK2 KO1 cells and its localization with fluorescently labeled EMCV. White arrow heads indicate EMCV particles in proximity to GFP-TNK2. Scale bar represents 10 μm. (C) Quantification of GFP-TNK2 high and low expression patterns in TNK2 KO1 cells. 10 images with an average of 12 cells per image were quantified. (D) Quantification of EMCV particles in proximity to GFP-TNK2 in TNK2 KO1 cells express low GFP-TNK2 compared to EMCV particles in proximity to GFP in TNK2 KO1 cells express GFP. 14 GFP-TNK2 expressing cells and 24 GFP expressing cells were quantified. (E) Quantification of EMCV infection on TNK2 KO1 cells transduced with lentivirus expressing GFP-TNK2 or GFP at 10 hr post infection. Error bars represent standard deviation of three replicates. The data shown are representative of two independent experiments. **:p<0.01. (F) Quantification of EMCV infection on WASL KO cells transduced with lentivirus expressing GFP-WASL or GFP at 10 hr post infection. Error bars represent standard deviation of three replicates. The data shown are representative of two independent experiments. ***: p<0.001.
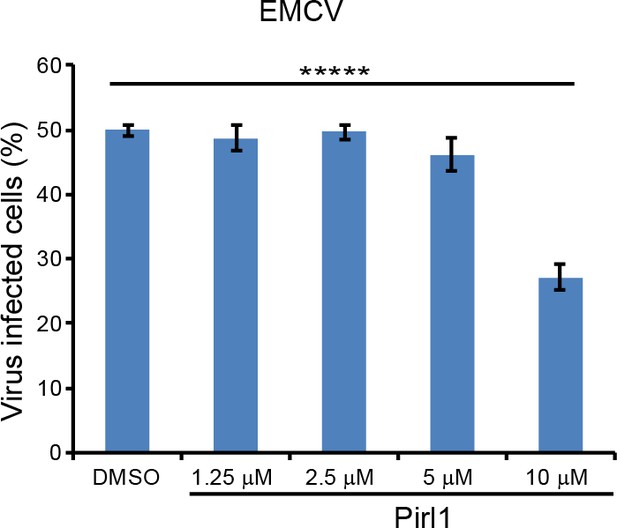
TNK2 mediates virus infection through endosomal trafficking pathways.
FACS quantification of EMCV infection on pirl1 treated A549 cells at 10 hr post infection at an MOI of 1. Error bars represent standard deviation of three replicates. ****: p<0.0001.
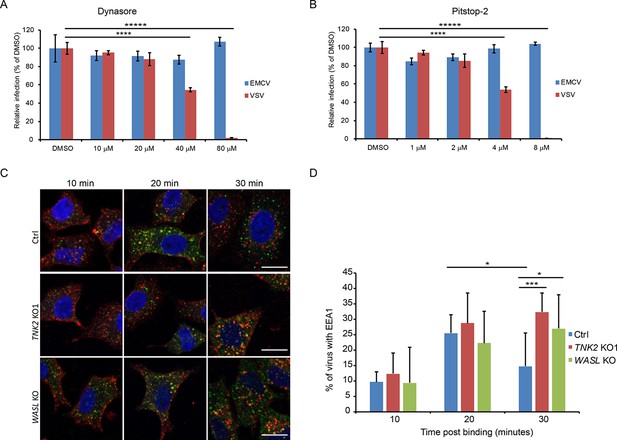
The endocytosis pathway inhibition on EMCV infection and localization of EEA1 with fluorescently labeled EMCV.
(A) Dynasore inhibition of EMCV and VSV infection on naïve A549 cells. ****: p<0.0001, *****: p<0.00001. (B) Pitstop-2 inhibition of EMCV and VSV infection on naïve A549 cells. ****: p<0.0001, *****: p<0.00001. (C) Fluorescent images of EEA1 co-localization with fluorescently labeled EMCV at 10, 20, and 30 min post internalization. Scale bars represent 20 μm. (D) Quantification of EEA1 co-localization with fluorescently labeled EMCV at 10, 20, and 30 min post internalization. Percentage of EMCV co-localized with EEA1 were quantified by image analysis. For each cell type, 10 images each at 10, 20 min post internalizaiton were analyzed, 15 images for Ctrl, TNK2 KO1 and 14 images for WASL KO cells at 30 min post internalizaiton were analyzed (7 cells per image in average). Error bars represent standard deviation of all images analyzed. *:p<0.05, ***: p<0.001.
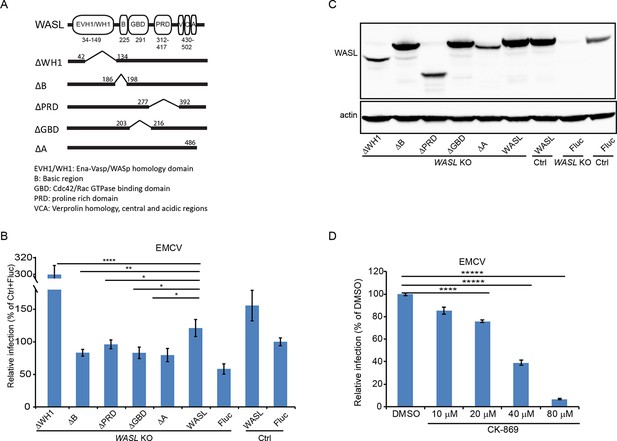
WASL activation and its actin modulation are critical for EMCV virus infection.
(A) Schematic representation of different WASL domain truncations. Each truncation is indicated by amino acid position on the constructs. (B) FACS quantification of EMCV infection in WASL KO cells transduced with different WASL domain truncations. (C) Western blot detection of WASL domain truncation expression constructs in lentivirus transduced WASL KO cells. (D) CK-869 inhibition of EMCV infection on naïve A549 cells at 10 hr post infection at an MOI of 1. (B, D) Error bars represent standard deviation of three replicates. The data shown are representative of two independent experiments. *: p<0.05, **: p<0.01, ***: p<0.001, ****: p<0.0001.
-
Figure 6—source data 1
Source data for Figure 6B and D: FACS quantification of virus infected cells.
- https://cdn.elifesciences.org/articles/50276/elife-50276-fig6-data1-v2.xlsx
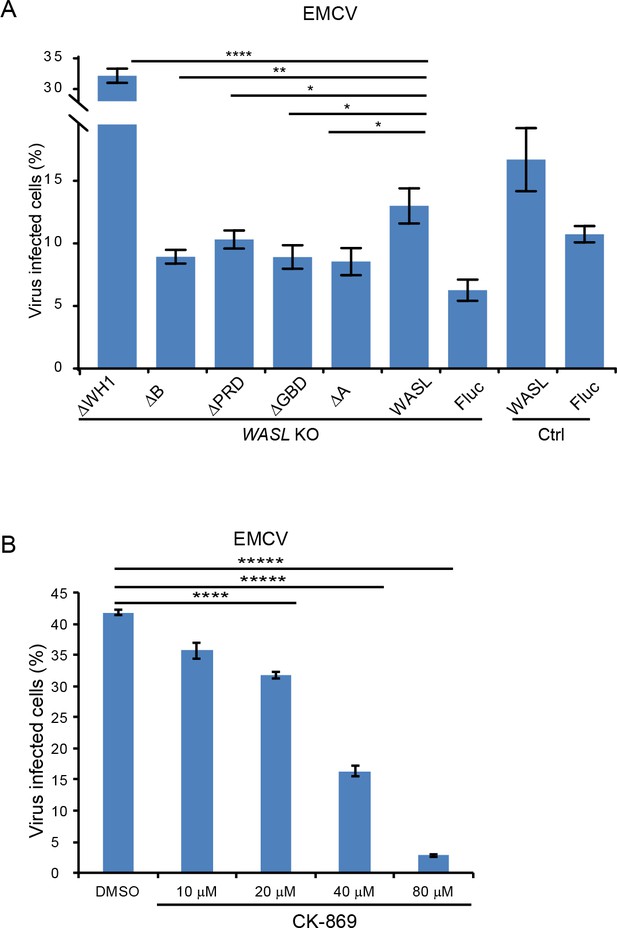
WASL activation and its actin modulation are critical for EMCV virus infection.
(A) FACS quantification of EMCV infection in WASL KO cells (without normalization) transduced with different WASL domain truncations. (B) CK-869 inhibition of EMCV infection on naïve A549 cells (without normalization) at 10 hr post infection at an MOI of 1. Error bars represent standard deviation of three replicates. The data shown are representative of two independent experiments. *: p<0.05, **: p<0.01, ***: p<0.001, ****: p<0.0001.
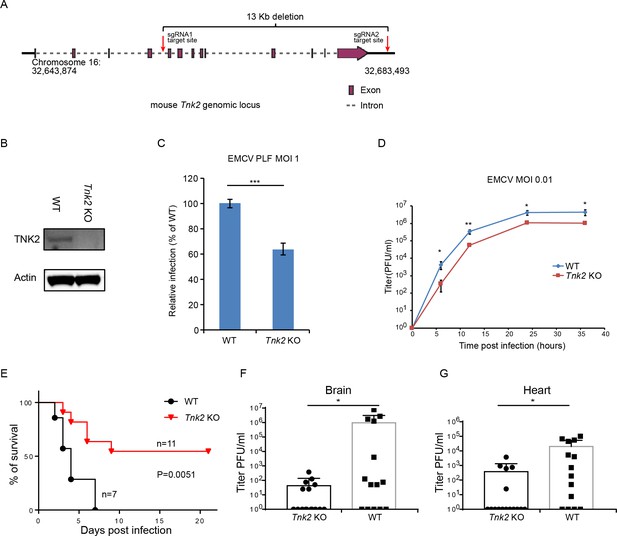
TNK2 is required for EMCV infection in vivo.
(A) Schematic representation of Tnk2 knockout design by CRIPSR-Cas9 genome editing in mouse. Exon, intron and genomic position are indicated. (B) TNK2 expression in mouse primary lung fibroblast cells derived from Tnk2 knockout and wild type animals. Cell lysates were analyzed by Western blot. (C) FACS quantification of EMCV infection in mouse primary lung fibroblast cells derived from Tnk2 knockout and wild type animals 6 hr post infection at an MOI of 1. Error bars represent standard deviation of three replicates. The data shown are representative of two independent experiments. ***: p<0.001. (D) Multi-step growth of EMCV in mouse primary lung fibroblast cells from Tnk2 knock out and wild type animals. *: p<0.05, **: p<0.01. (E) Survival curve of EMCV infection via oral gavage in Tnk2 knockout and wild type mice. p=0.0051 by log-rank test. (F, G) EMCV titer in infected mouse brain and heart. *: p<0.05 by Mann-Whitney test, n = 16 for Tnk2 KO and n = 14 for WT.
-
Figure 7—source data 1
Source data for Figure 7C, E, F, G.
Source data for Figure 7C: FACS quantification of virus infected cells; Source data for Figure 7E: death report for EMCV-infected mice; Source data for Figure 7F and G: virus titer in EMCV-infected mouse brain and heart.
- https://cdn.elifesciences.org/articles/50276/elife-50276-fig7-data1-v2.xlsx

TNK2 is required for EMCV infection in vivo.
FACS quantification of EMCV infection in mouse primary lung fibroblast cells (without normalization) derived from Tnk2 knockout and wild type animals 6 hr post infection at an MOI of 1. Error bars represent standard deviation of three replicates. The data shown are representative of two independent experiments. ***: p<0.001.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (human) | TNK2 isoform1 | GenBank | NM_005781.4 | |
| Gene (human) | TNK2 isoform2 | GenBank | BC028164.1 | |
| Gene (human) | TNK2 isoform3 | GenBank | NM_001308046.1 | |
| Gene (human) | WASL | GenBank | NM_003941.3 | |
| Gene (human) | NCK1 | GenBank | NM_006153.5 | |
| Strain, strain background (Mouse) | C57BL/6J | The Jackson Laboratory | 000664 | |
| Strain, strain background (Mouse) | C57BL/6J TNK2 KO | This paper | Generated at the Genome Engineering and iPSC Center (GEiC) at Washington University. David Wang lab. | |
| Strain, strain background (Virus) | EMCV VR-129 strain | Michael Diamond lab | Michael Diamond lab | |
| Strain, strain background (Virus) | Coxsackie B3 Virus Nancy strain | Julie Pfeiffer lab | Julie Pfeiffer lab | |
| Strain, strain background (Virus) | Poliovirus Mahony strain | Nihal Altan-Bonnet lab | Nihal Altan-Bonnet lab | |
| Strain, strain background (Virus) | Adenovirus A5 | David Curiel lab | David Curiel lab | |
| Strain, strain background (Virus) | Influenza A virus WSN strain | Adrianus Boon lab | Adrianus Boon lab | |
| Strain, strain background (Virus) | enterovirus D68 | ATCC | ATCC VR-1826 | |
| Strain, strain background (Virus) | Parainfluenza | Robert A Lamb lab | Robert A. Lamb lab | |
| Strain, strain background (Virus) | GFP-EMCV | Frank JM van Kuppeveld lab | Frank JM van Kuppeveld lab | |
| Strain, strain background (Virus) | GFP-CVB3 | Frank JM van Kuppeveld lab | Frank JM van Kuppeveld lab | |
| Cell line (human) | A549 | ATCC | ATCC CCL-185 | |
| Cellline (human) | A549 TNK2 KO1 | This paper | Generated by CRISPR at David Wang lab. | |
| Cell line (human) | A549 TNK2 KO2 | This paper | Generated by CRISPR at David Wang lab. | |
| Cell line (human) | A549 WASL KO | This paper | Generated by CRISPR at David Wang lab. | |
| Cellline (human) | A549 TNK2 WASL dKO | This paper | Generated by CRISPR at David Wang lab. | |
| Cell line (human) | A549 TNK2 NCK1 dKO | This paper | Generated by CRISPR at David Wang lab. | |
| Cell line (human) | A549 WASL NCK1 dKO | This paper | Generated by CRISPR at David Wang lab. | |
| Cell line (human) | A549 TNK2 WASL NCK1 tKO | This paper | Generated by CRISPR at David Wang lab. | |
| Cell line (human) | Hela | ATCC | ATCC CCL-2 | |
| Cellline (human) | 293T | ATCC | ATCC CRL-3216 | |
| Cell line (human) | Hap1 WT | Horizon | C631 | |
| Cellline (human) | Hap1 TNK2 KO | Horizon | HZGHC002454c026 | |
| Cell line (human) | Hap1 WASL KO | Horizon | HZGHC002632c003 | |
| Cellline (human) | RD (rhabdomyosarcoma) | ATCC | ATCC CCL-136 | |
| Cell line (mouse) | C57BL/6J primary lung fibroblast | This paper | Generated by lung digestion at David Wang lab. | |
| Cell line (mouse) | C57BL/6J Tnk2 KO primary lung fibroblast | This paper | Generated by lung digestion at David Wang lab. | |
| Cellline (hamster) | BHK-21 | ATCC | ATCC CCL-10 | |
| Antibody | Mouse monoclonal anti-TNK2 clone A11 | Santa Cruz | sc-28336 | WB (1:1000) |
| Antibody | Rabbit monoclonal anti-WASL | Abcam | ab126626 | WB (1:1000) Used in Figure 2A, Figure 3—figure supplement 2A,C,D |
| Antibody | Rabbit polyclonal anti-WASL | Sigma | HPA005750 | WB (1:1000) Used in Figure 6C |
| Antibody | Rabbit polyclonal anti-NCK1 | Millipore | 06–288 | WB (1:1000) |
| Antibody | Mouse monoclonal anti-actin | Sigma | MAB1501 | WB (1:1000) |
| Antibody | Mouse polyclonal anti-EMCV antibodies | This paper | Antibodies generated in Michael Diamond lab. ICC (1:1000) | |
| Antibody | Mouse monoclonal anti-Coxsakie B3 virus | ThermoFisher | MAB948 | ICC (1:1000) |
| Antibody | Mouse monoclonalanti-Avenovirus A5 | ThermoFisher | MA5-13643 | ICC (1:1000) |
| Antibody | Mouse Monoclonal anti-influenza virus NP | Millipore | MAB8258B | ICC (1:2000) |
| Antibody | Rabbit polyclonal anti-Enterovirus D68 | GeneTex | GTX132313 | ICC (1:1000) |
| Antibody | Mouse monoclonal anti-poliovirus antibodies | Nihal Altan-Bonnet lab | Antibodies generated in Nihal Altan-Bonnet lab. ICC (1:2000) | |
| Antibody | Mouse monoclonal anti-HA | ThermoFisher | 26183 | WB (1:1000) |
| Antibody | Mouse monoclonal anti-Flag | GenScript | A00187-100 | WB (1:1000) |
| Antibody | Mouse monoclonal anti-c-Myc | Invitrogen | 13–2500 | WB (1:1000) |
| Antibody | Mouse monoclonal anti-double stranded J2 antibodies | Scicons | 10010200 | IFA (1:500) |
| Recombinant DNA reagent | pReceiver-TNK2 | Genecopia | EX-Y4392-M02 | |
| Recombinant DNA reagent | pReceiver-WASL | Genecopia | EX-I2067-M68 | |
| Recombinant DNA reagent | Lenti CRISPR v2 | Addgene | 98290 | |
| Recombinant DNA reagent | pSPAX2 | Addgene | 12260 | |
| Recombinant DNA reagent | pMD2.G | Addgene | 12259 | |
| Recombinant DNA reagent | pCW57-GFP-P2A-MCS (Neo) | Addgene | 89181 | |
| Recombinant DNA reagent | pFCIV | PMID: 27384652 | Michael Diamond lab | |
| Recombinant DNA reagent | C5V | Addgene | 26394 | |
| Recombinant DNA reagent | pcDNA-NCK1 | Genscript | OHu24619D | |
| Recombinant DNA reagent | pcDNA-TNK2 isoform 3 | Genescript | OHu13497C | |
| Sequence-based reagent(primers and Oligonucleotides) | All cloning primers and oligonucleotides used are in supplemental table | This paper | Synthesized by IDT. David Wang lab. | |
| Peptide, recombinant protein | Alt-R S.p. Cas9 nuclease 3NLS protein | IDT | 1081058 | |
| Commercial assay or kit | BCA assay | ThermoFisher | 23235 | |
| Commercial assay or kit | Cell titer-glo assay | Promega | G7570 | |
| Commercial assay or kit | QuikChange II Site-Directed Mutagenesis Kits | Agilent | 200521 | |
| Commercial assay or kit | TaqMan Fast Virus 1-Step Master Mix | ThermoFisher | 4444434 | |
| Commercial assay or kit | Lipofectamine CRISPR MAX | ThermoFisher | CMAX00001 | |
| Commercial assay or kit | CoIP | Pierce | 88804 | |
| Commercial assay or kit | Zymo RNAeasy Miniprep | Zymo research | R2052 | |
| Commercial assay or kit | Zymo RNAeasy 96-well extraction | Zymo research | R2056 | |
| Commercial assay or kit | Nap-5 desalting column | GE Healthcare | GE17-0853-01 | |
| Chemical compound, drug | Aim-100 | Apexbio | 4946 | |
| Chemical compound, drug | Wiskotstatin | Sigma | 681525 | |
| Chemical compound, drug | a-sarcin | Santa Cruz | CAS 86243-64-3 | |
| Chemical compound, drug | METHIONINE,L-[35S]- | Perkin Elmer | NEG009T005MC | |
| Chemical compound, drug | CYSTEINE, L-[35S]- | Perkin Elmer | NEG022T005MC | |
| Chemical compound, drug | Alexa FluorA647 succinimidyl ester | ThermoFisher | A20106 | |
| Chemical compound, drug | Polybrene | Millipore | TR-1003-G | |
| Chemical compound, drug | Puromycin | Sigma | P8833 | |
| Chemical compound, drug | Doxycyclin | Sigma | D1822 | |
| Chemical compound, drug | Dynasore | Sigma | D7693 | |
| Chemical compound, drug | Pitstop-2 | Sigma | SML1169 | |
| Chemical compound, drug | CK-869 | Sigma | C9124 | |
| Chemical compound, drug | Pirl1 | Hit2leads | SC-5137877 | |
| Software, algorithm | Flowjo V10 | FlowJo, LLC | Commercial software for flow cytometry analysis | |
| Software, algorithm | Prism V7 | GraphPad Software, Inc | Commercial software for statistical analysis | |
| Software, algorithm | Volocity V6.3 | PerkinElmer | Commercial software for image analysis |
Additional files
-
Supplementary file 1
Primers and oligonucelotides used in this study.
- https://cdn.elifesciences.org/articles/50276/elife-50276-supp1-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/50276/elife-50276-transrepform-v2.docx