The MADS-box transcription factor PHERES1 controls imprinting in the endosperm by binding to domesticated transposons
Figures
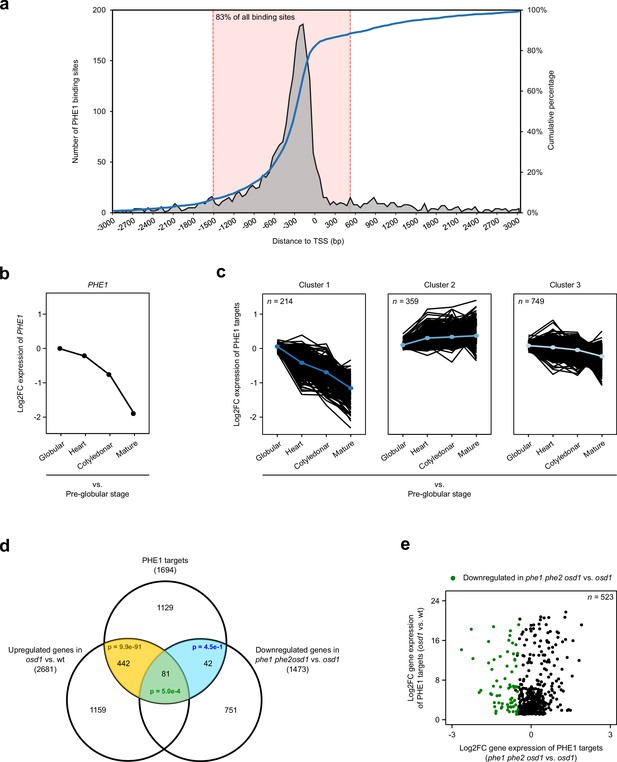
Identification and expression profile of PHE1 target genes.
(a) Spatial distribution of PHE1 binding sites around transcription start sites (TSS). The dotted pink lines indicate the spatial interval used to define PHE1 target genes. (b–c) Expression of PHE1 (b), and its target genes (c), across different stages of seed development. Gene expression is represented as a Log2-fold change between expression in the endosperm at the stages indicated on the x axis vs. expression in the pre-globular stage. A k-means clustering analysis was performed to group PHE1 targets that show similar expression trends across seed development. Gene expression data were retrieved from Belmonte et al. (2013). (d) Overlap between PHE1 targets, genes that were found to be significantly upregulated in osd1 when compared to wild-type (wt) seeds, and genes that were found to be significantly downregulated in phe1 phe2 osd1 when compared to osd1 seeds. P-values were determined using hypergeometric tests. (e) Expression of PHE1 targets that are significantly upregulated in osd1 seeds when compared to wt seeds. Genes marked in green are also significantly downregulated in phe1 phe2 osd1 seeds when compared to osd1 seeds.
-
Figure 1—source data 1
PHE1 target genes and their respective endosperm expression cluster.
- https://cdn.elifesciences.org/articles/50541/elife-50541-fig1-data1-v2.xlsx
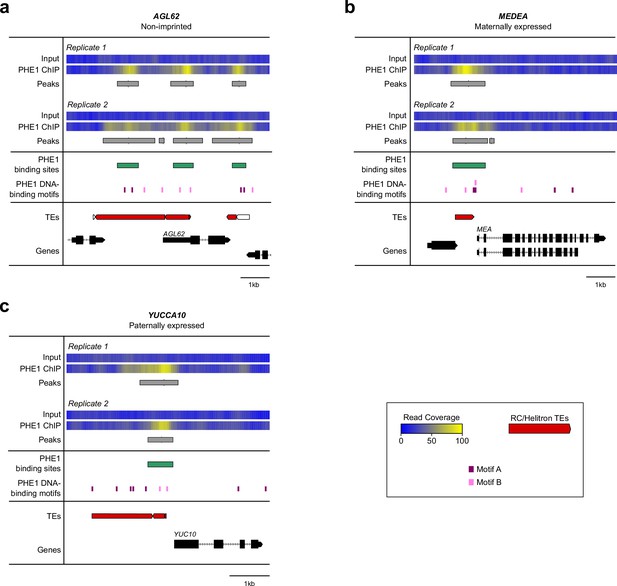
Examples of ChIP-seq read, peak, and motif distributions in PHE1 targets.
For each target gene, the read coverage for Input (as control) and PHE1–GFP ChIP samples of each replicate is shown. Called peaks in each replicate are indicated by the grey boxes. PHE1 binding sites, identified from the overlapped peak regions for replicates 1 and 2, are represented by the green boxes. PHE1 DNA-binding motifs are those defined in Figure 3a, and are represented here by the purple (Motif A) and pink boxes (Motif B). Transposable elements in the vicinity of the target gene are indicated. Red boxes correspond to TEs of the RC/Helitron superfamily, whereas white boxes represent TEs of other superfamilies. For this analysis, three different PHE1 targets with distinct imprinting statuses were selected: (a) AGL62 – non-imprinted; (b) MEDEA – a maternally expressed gene; (c) YUCCA10 – a paternally expressed gene.
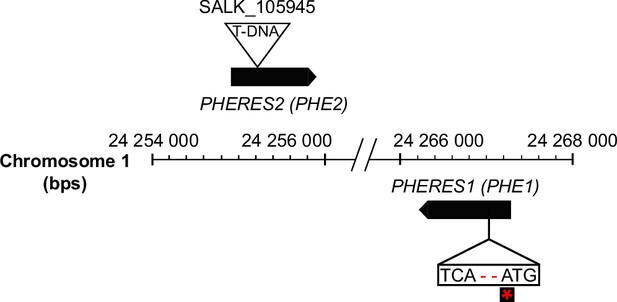
Schematic representation of the phe1 phe2 mutant.
CRISPR/Cas9 was used to generate a premature stop codon in PHE1, represented by the red asterisk. PHE1 mutagenesis was performed in the phe2 background because of the proximity and probable redundancy of the two PHE genes (see Materials and methods).
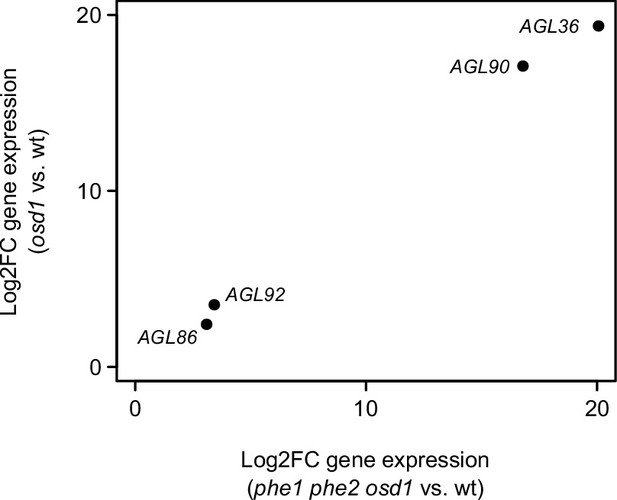
Expression of PHE1 paralogs in osd1 vs. wt and phe1 phe2 osd1 vs. wt.
Paralogous genes were selected according to the phylogenetic analysis performed by Parenicová et al. (2003). All genes are significantly upregulated in osd1 vs. wt and in phe1 phe2 osd1 vs. wt.
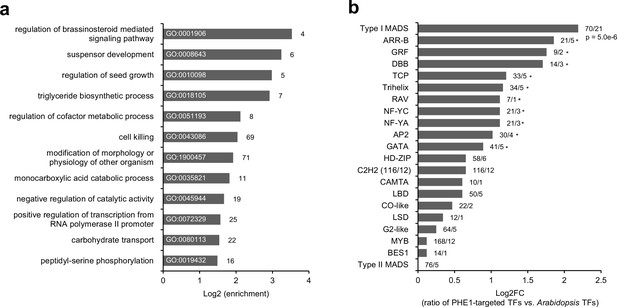
Transcription factor genes are enriched among PHE1 targets.
(a) Enriched biological processes associated with PHE1 target genes. Numbers on bars indicate number of PHE1 target genes within each GO term. (b) Enrichment of transcription factor (TF) families among PHE1 target genes (see Materials and methods). Numbers indicate the total number of Arabidopsis genes belonging to a certain TF family, and the total number of genes in that family targeted by PHE1. *, p-values <0.05. P-values were determined using the hypergeometric test.
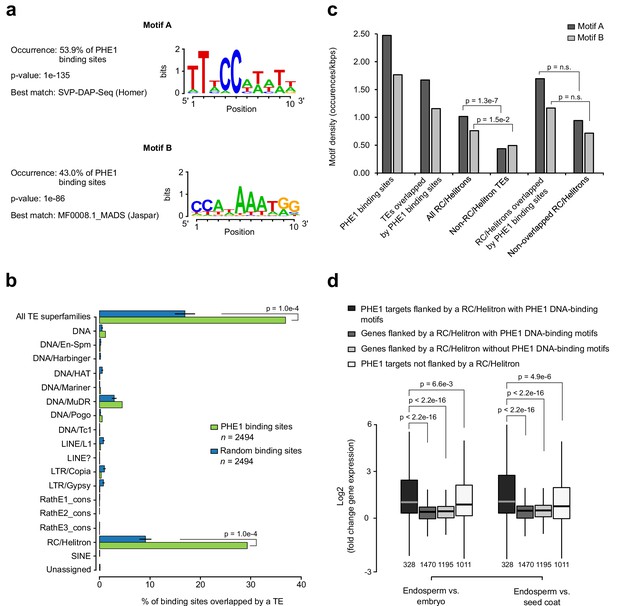
RC/Helitrons carry PHE1 DNA-binding motifs.
(a) CArG-box like DNA-binding motifs identified from PHE1 ChIP-seq data. (b) Fraction of PHE1 binding sites (green) that overlap transposable elements (TEs). Overlap is expressed as the percentage of total binding sites for which spatial intersection with features on the y-axis is observed. A set of random binding sites is used as control (blue). This control set was obtained by randomly shuffling the identified PHE1 binding sites within random A. thaliana gene promoters (see Materials and methods). P-values were determined using Monte Carlo permutation tests (see Materials and methods). Bars represent ± s.d. (n = 2494, for PHE1 binding sites and random binding sites). (c) Density of PHE1 DNA-binding motifs in different genomic regions of interest. P-values were determined using χ2 tests. (d) Fold-change in the expression of genes flanked by RC/Helitron TEs. Fold-change was determined by comparing endosperm and embryo, or by comparing endosperm and seed coat. Genes were divided into four categories depending on their PHE1 target status, and the presence of RC/Helitrons with and without PHE1 DNA-binding motifs. Gene expression data were retrieved from Belmonte et al. (2013). Pre-globular seed stage was used in this analysis. P-values were determined using two-tailed Mann-Whitney tests (n = number represented below boxplots).
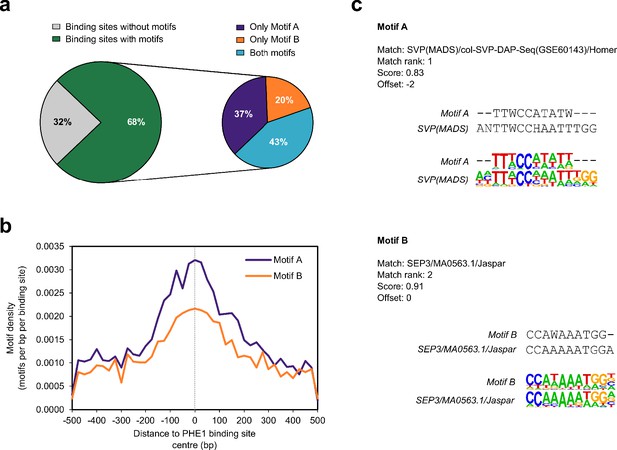
PHE1 DNA-binding motifs show similarity to type II MADS-box CArG-boxes.
(a) Occurrence of PHE1-binding motifs in PHE1 binding sites. (b) Density of PHE1-binding motifs across PHE1 binding sites. (c) Alignment between PHE1-binding motifs and known motif matches. Both motifs match with previously characterized type II MADS-box CArG-boxes.
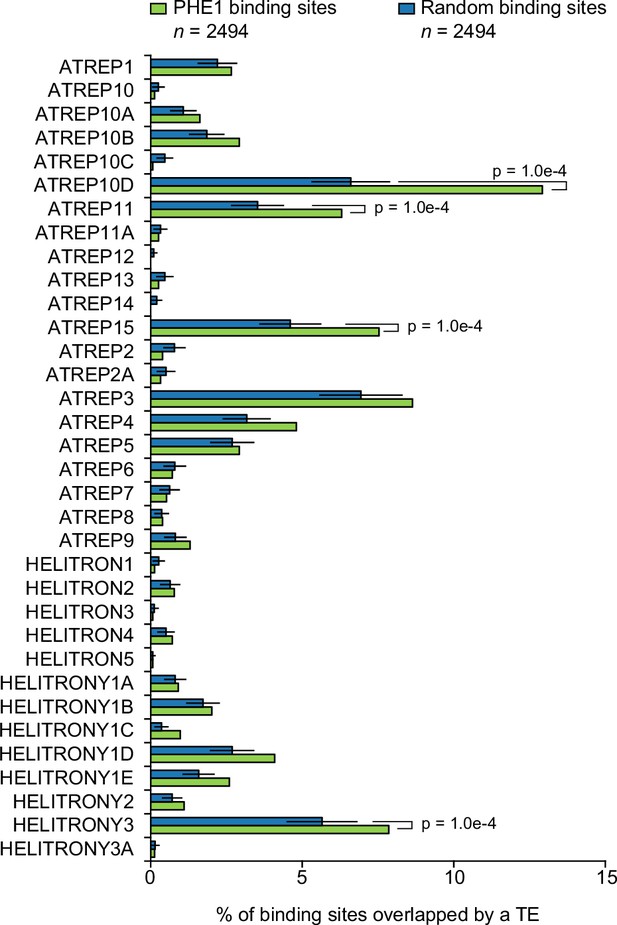
RC/Helitron family overlap with PHE1 binding sites.
Fraction of PHE1 binding sites (green) overlapping with different RC/Helitron TE families. Overlap is expressed as the percentage of binding sites where spatial intersection with the families specified on the y-axis is observed. A set of random binding sites is used as control (blue). This control set was obtained by randomly shuffling the identified PHE1 binding sites within A. thaliana gene promoters (see Materials and methods). P-values were determined using Monte Carlo permutation tests (see Materials and methods). Bars represent ± s.d., (n = 2494, for PHE1 binding sites and random binding sites).
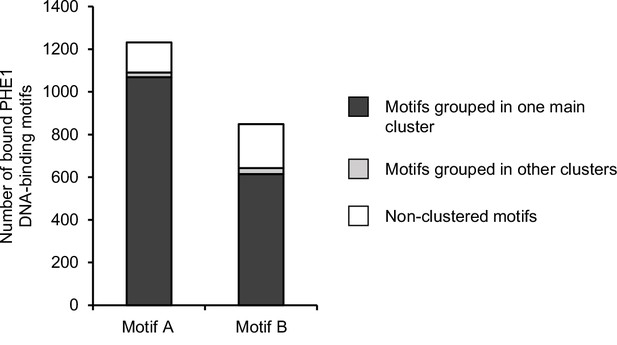
Clustering of bound PHE1 DNA-binding motifs.
PHE1-DNA binding motifs and their flanking sequences were paired on the basis of sequence homology. The pairwise homologous sequences were then merged into higher-order clusters, which were based on shared elements in the homologous pairs (see Materials and methods). Motifs are those identified in Figure 3a.
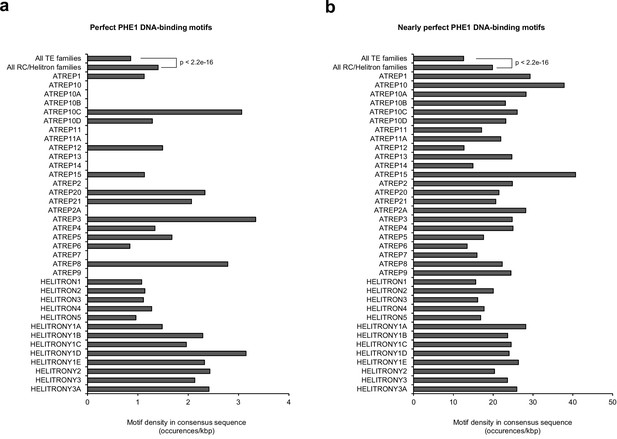
PHE1 DNA-binding motif densities in RC/Helitron consensus sequences.
(a) Density of perfect PHE1 DNA-binding motifs in the consensus sequences of different RC/Helitron families. Perfect motifs are those described in Figure 3a. As a reference, the density of perfect PHE1 DNA-binding motifs in the consensus sequences of all TE families is shown. (b) Density of nearly perfect PHE1 DNA-binding motifs in the consensus sequences of different RC/Helitron families. Nearly perfect motifs are those sequences where only one nucleotide substitution is required to generate a perfect PHE1 DNA-binding motif. As a reference, the density of nearly perfect PHE1 DNA-binding motifs in the consensus sequences of all TE families is shown. P-values were determined using χ2 tests.
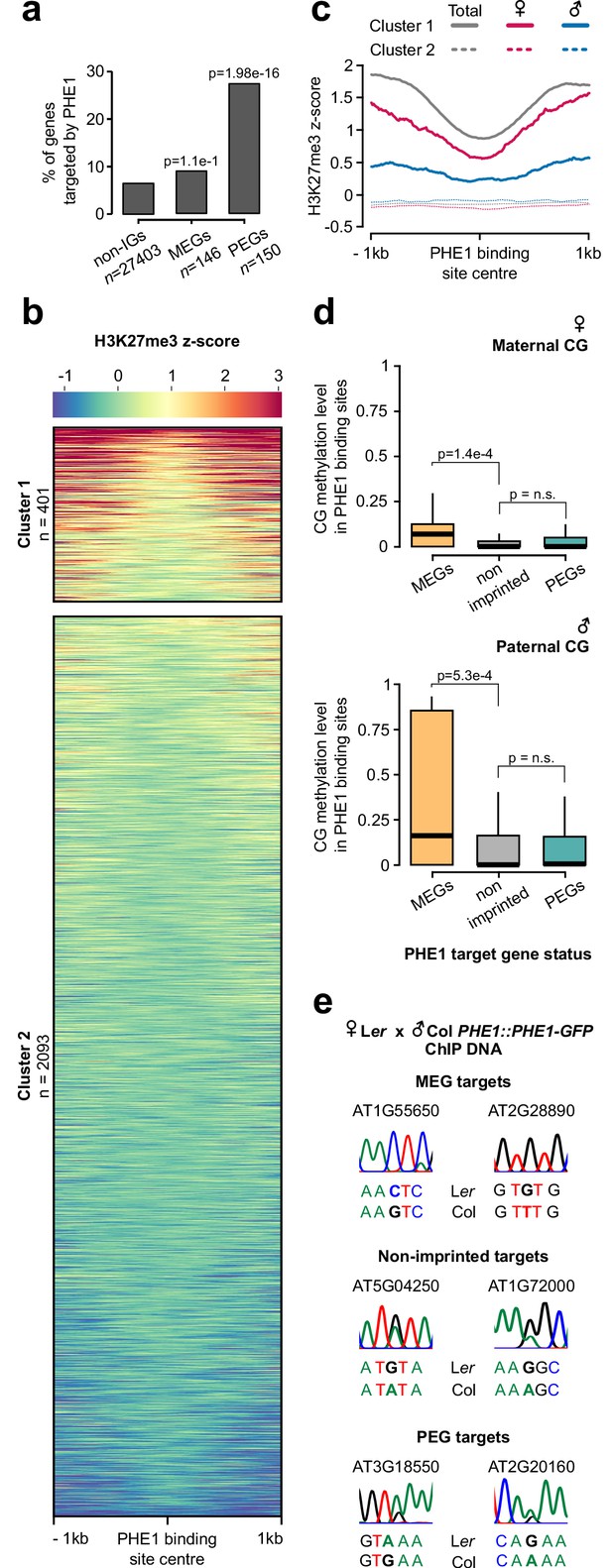
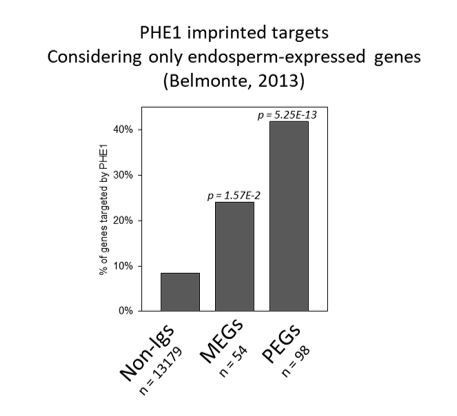
Parental asymmetry of epigenetic marks in imprinted gene promoters conditions PHE1 binding.
(a) Fraction of non-imprinted and published imprinted genes targeted by PHE1. P-values were determined using the hypergeometric test. The list of published imprinted genes used for this analysis is detailed in Figure 4—source data 1. (b) Heatmap of endosperm H3K27me3 distribution along PHE1-binding sites. Each horizontal line represents one binding site. Clusters were defined on the basis of the pattern of H3K27me3 distribution (see Materials and methods) (c) Metagene plot of average maternal (♀, pink), paternal (♂, blue) and total (grey) endosperm H3K27me3 marks along PHE1 binding sites. (d) CG methylation levels in maternal (♀, upper panel) and paternal (♂, lower panel) alleles of PHE1 binding sites associated with MEGs (yellow), PEGs (green) and non-imprinted (grey) PHE1 targets. P-values were determined using two-tailed Mann-Whitney tests. (e) Sanger sequencing of imprinted and non-imprinted gene promoters bound by PHE1. SNPs for maternal (Ler) and paternal (Col) alleles are shown (n = 1 biological replicate). Maternal:total read ratios for imprinted genes are as follows: AT1G55650 – 1.0; AT2G28890 – 0.97; AT3G18550 – 0.44; and AT2G20160 – 0.32.
-
Figure 4—source data 1
List of imprinted genes used in this study.
- https://cdn.elifesciences.org/articles/50541/elife-50541-fig4-data1-v2.xlsx
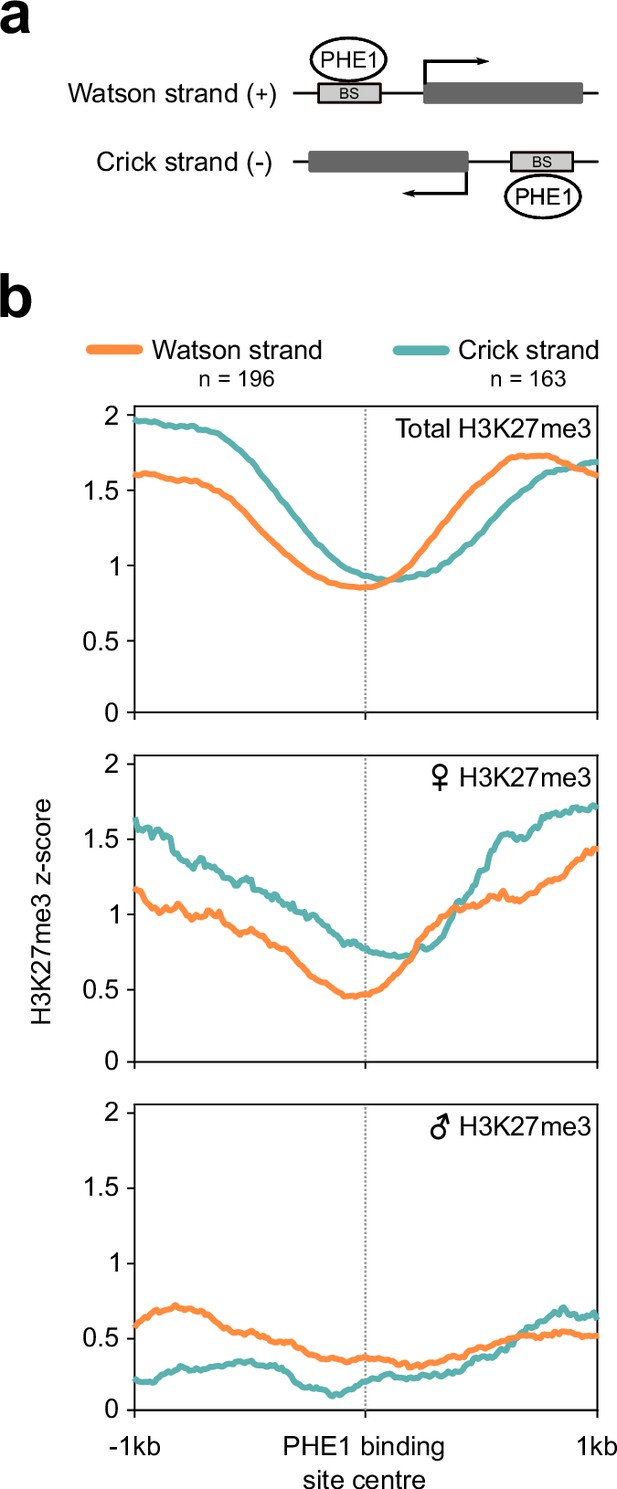
Directionality of H3K27me3 distribution in Cluster 1 binding sites.
(a) Binding sites and their respective target genes can be present in the Watson (+) or Crick (–) strand of DNA. (b) Metagene plots of H3K27me3 distribution in PHE1binding sites associated with target genes located in the Watson (orange line) or Crick strands (blue lines). For this analysis, only H3K27me3 distributions in PHE1-binding sites associated with Cluster 1 were considered (Figure 4b). Total, maternal, and paternal H3K27me3 is represented in the top, middle, and bottom panels, respectively.
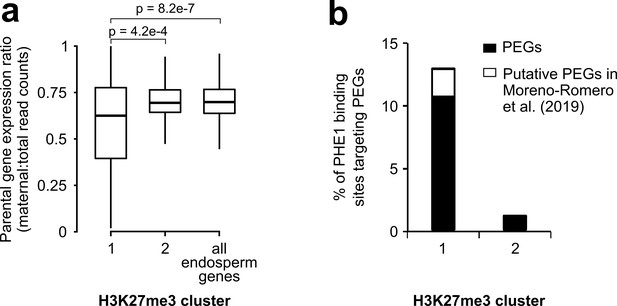
Characterization of H3K27me3 clusters.
(a) Parental gene expression ratio of PHE1 targets associated with Cluster 1 and Cluster 2 binding sites, and all endosperm expressed genes. P-values were determined using two-tailed Mann-Whitney tests. (b) Fraction of Cluster 1 and Cluster 2 binding sites that target PEGs (Figure 4—source data 1) and putative PEGs (Moreno-Romero et al., 2019). (a, b) H3K27me3 clusters are those defined in Figure 4b.
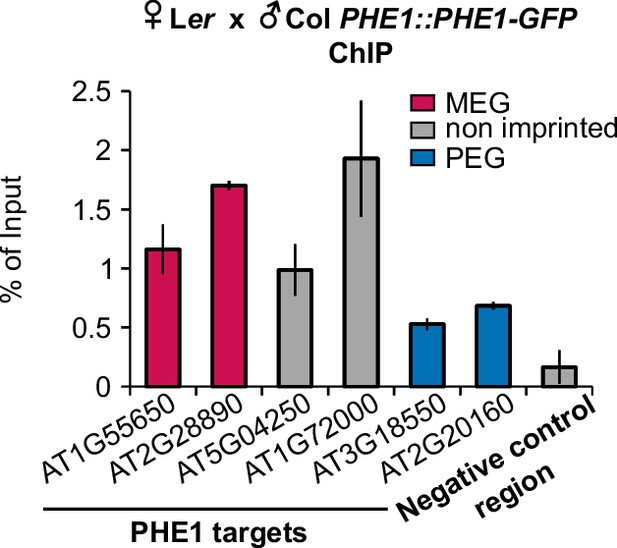
Parental-specific PHE1 ChIP.
qPCR of purified ChIP-DNA. Enrichment is shown as % of input DNA, in regions associated with MEGs (pink), PEGs (blue), and non-imprinted (grey) PHE1 target genes. Bars represent ± s.d. Data from one representative biological replicate are shown (n = 2 biological replicates).
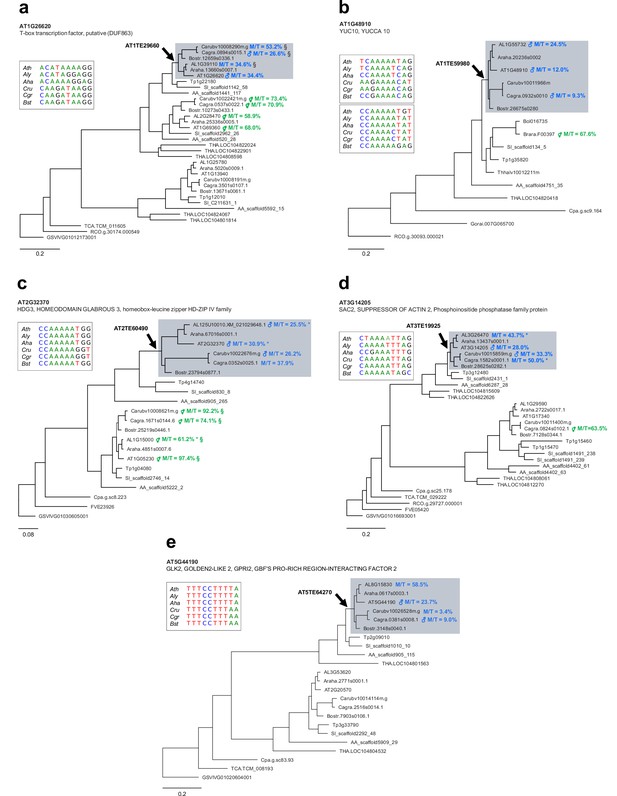
Ancestral RC/Helitron insertions are associated with gain of imprinting in the Brassicaceae.
(a–e) Phylogenetic analyses of PHE1-targeted PEGs and their homologs. Each panel represents a distinct target gene and its corresponding homologs in different species. The genes shown on a grey background have homologous RC/Helitron sequences in their promoter region. The arrow indicates the putative insertion of an ancestral RC/Helitron. The identity of the RC/Helitron identified in A. thaliana is indicated. These A. thaliana RC/Helitrons contain a PHE1 DNA-binding motif and are associated with a PHE1 binding site. The inset boxes represent the alignment between the A. thaliana PHE1 DNA-binding motif and similar DNA motifs contained in RC/Helitrons that are present in the promoter regions of orthologous genes. When available, the imprinting status of a given gene is indicated by the presence of ♂ (PEG) or ⚥ (non-imprinted), and reflects the original imprinting analyses done in the source publications (see Materials and methods). The maternal:total read ratio (M/T) for each gene is also indicated. §: potential contamination from maternal tissue. *: accession-biased expression. The scale bars represent the frequency of substitutions per site for the ML tree. The tree is unrooted. Gene identifier nomenclatures: AT, Arabidopsis thaliana; AL, Arabidopsis lyrata; Araha, Arabidopsis halleri; Bostr, Boechera stricta; Carubv, Capsella rubella; Cagra, Capsella grandiflora; Tp, Schrenkiella parvula; SI, Sisymbrium irio; Bol, Brassica oleracea; Brapa, Brassica rapa; Thhalv, Eutrema salsugineum; AA, Aethionema arabicum; THA, Tarenaya hassleriana; Cpa, Carica papaya; TCA, Theobroma cacao; Gorai, Gossypium raimondii; RCO, Ricinus communis; FVE, Fragaria vesca; and GSVIVG, Vitis vinifera.

RC/Helitrons carrying PHE1 DNA-binding motifs are more prevalent in PEGs.
Fraction of PHE1 binding sites targeting MEGs, PEGs, or non-imprinted genes where a spatial overlap between a RC/Helitron and a PHE1 DNA-binding motif is observed. P-values were determined using the hypergeometric test.
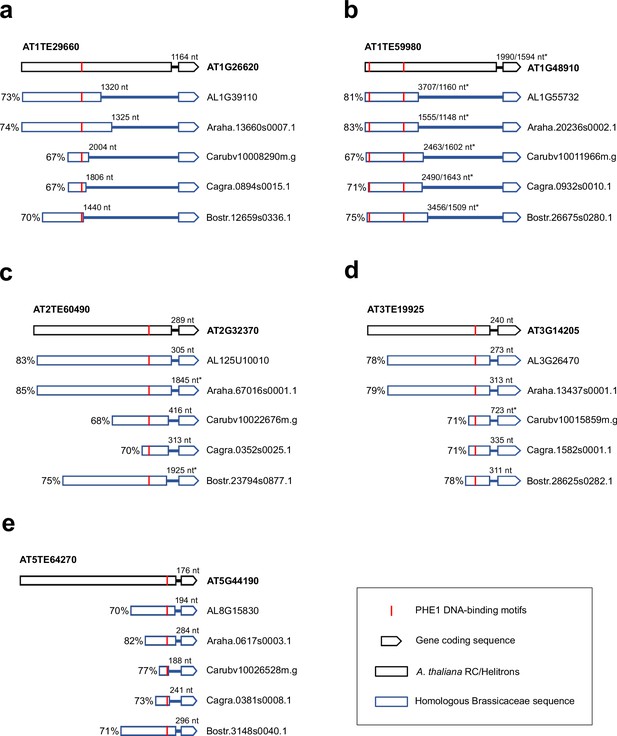
Analysis of PHE1-targeted PEG orthologs and upstream RC/Helitron sequences within Brassicaceae.
Alignments between A. thaliana RC/Helitrons associated with PEGs (Figure 5a–e) and homologous sequences in other Brassicaceae, where the putative homologous PHE1 DNA-binding motifs are present across all six species. For each aligned fragment, the percent sequence identity is indicated. The distances between the binding motifs and the TSS of PEGs are labelled. *, the distance to the TSS, in cases where the annotation for the 5′-UTR of the gene is not available.
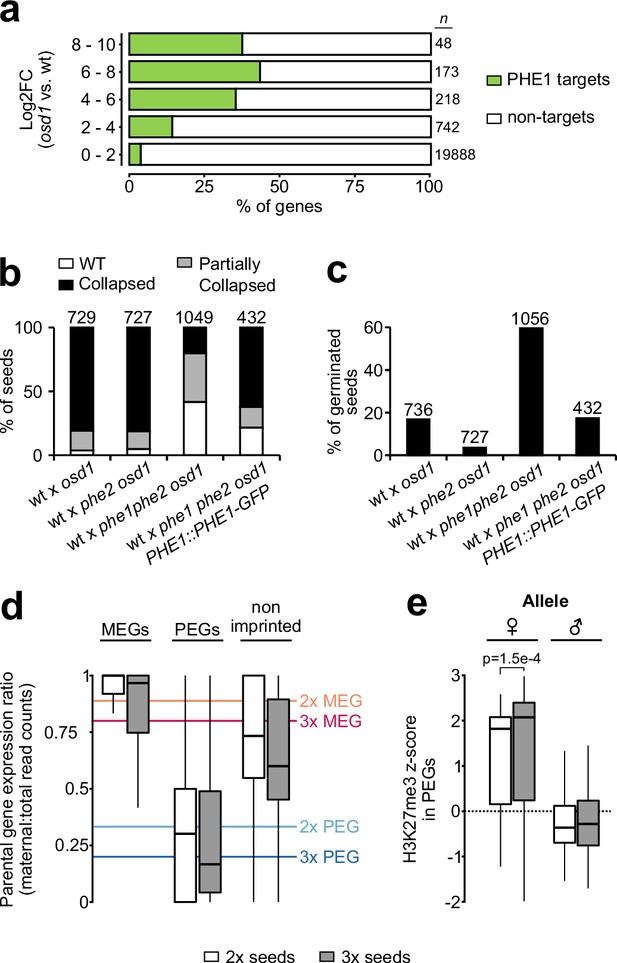
PHE1 establishes 3x seed inviability of paternal excess crosses.
(a) Target status of upregulated genes in paternal excess crosses. Highly upregulated genes in 3x seeds are more often targeted by PHE1 (p<2.2e–16, χ2 test). n = numbers on the left. (b, c) Seed inviability phenotype (b) of paternal excess crosses in wild-type (wt), phe1 phe2, and phe1 complementation lines, with their respective seed germination rates (c). The maternal parent is always indicated first. Remaining control crosses are shown in Figure 6—figure supplement 1a–b. n = numbers on top of bars (seeds). (d) Parental expression ratio of imprinted genes in the endosperm of 2x (white) and 3x seeds (grey). Solid lines indicate the ratio thresholds for the definition of MEGs and PEGs in 2x and 3x seeds. (e) Accumulation of H3K27me3 across maternal (♀) and paternal (♂) gene bodies of PEGs in the endosperm of 2x and 3x seeds (white and grey, respectively). H3K27me3 accumulation in MEGs and non-imprinted genes is shown in Figure 6—figure supplement 2. P-value was determined using a two-tailed Mann-Whitney test.
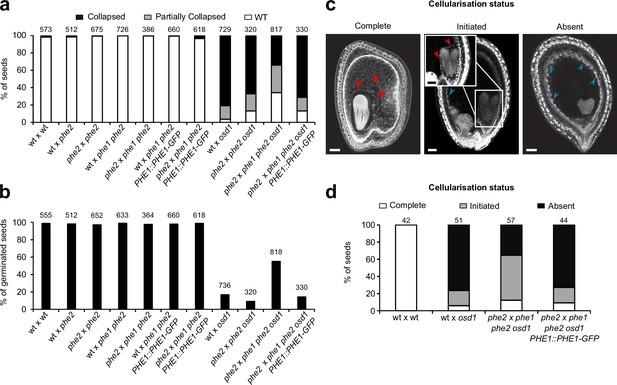
Rescue of 3x seed inviability in phe1 phe2.
(a, b) Seed inviability phenotype of wild-type (wt) and paternal excess crosses in wt, phe1 phe2, and phe1 complementation lines (a), with respective seed germination rates (b). (c, d) Status of endosperm cellularization in wt and paternal excess seeds. (a, b, d) n = numbers on top of bars (seeds).
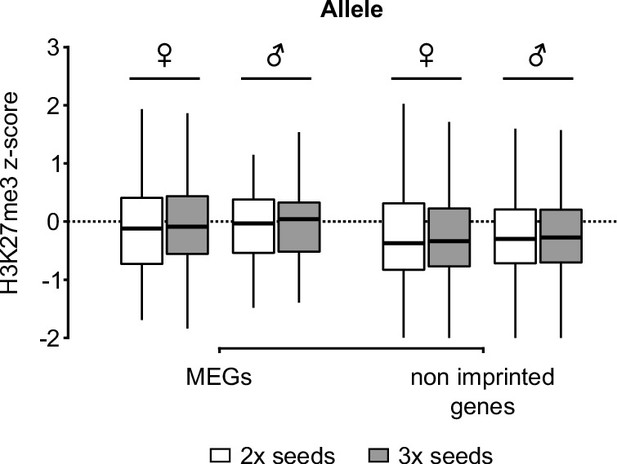
Distribution of H3K27me3 in 2x and 3x seeds.
Accumulation of H3K27me3 in maternal (♀) and paternal (♂) gene bodies of MEGs and non-imprinted genes in the endosperm of 2x and 3x seeds (white and grey, respectively).
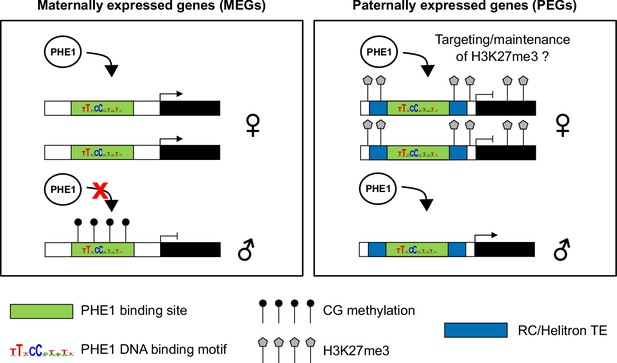
Control of imprinted gene expression by PHE1.
Schematic model of imprinted gene control by PHE1. Maternally expressed genes (left panel) show DNA hypermethylation of paternal (♂) PHE1 binding sites. This precludes PHE1 accessibility to paternal alleles, leading to predominant binding and transcription from maternal (♀) alleles. In paternally expressed genes (right panel), RC/Helitrons found in flanking regions carry PHE1 DNA-binding motifs, allowing PHE1 binding. The paternal PHE1 binding site is devoid of repressive H3K27me3, facilitating the binding of PHE1 and transcription of this allele. H3K27me3 accumulates at the flanks of maternal PHE1 binding sites, whereas the binding sites remain devoid of this repressive mark. PHE1 is able to bind maternal alleles, but fails to induce transcription. We hypothesize that the accessibility of maternal PHE1binding sites might be important for deposition of H3K27me3 during central cell development (possibly by another type I MADS-box transcription factor). It may also be required for the maintenance of H3K27me3 during endosperm proliferation.
Tables
PHE1 ChIP-seq read mapping and peak calling information.
Peak calling was done using the ChIP sample and its respective Input sample as control. The fraction of peaks present in both replicates was determined as the percentage of peaks for which spatial overlap between Replicate 1 and Replicate 2 peaks is observed (see Materials and methods).
| Sample | No. of sequenced reads | % of mapped reads | No. of called ChIP-seq peaks | % of ChIP-seq peaks present in both replicates |
|---|---|---|---|---|
| Replicate 1 PHE1::PHE1–GFP ChIP | 17,037,975 | 65.3 | 2818 | 88.5 |
| Replicate 1 PHE1::PHE1–GFP Input | 24,276,095 | 71.1 | ||
| Replicate 2 PHE1::PHE1–GFP ChIP | 21,838,147 | 70.5 | 4521 | 55.2 |
| Replicate 2 PHE1::PHE1–GFP Input | 23,372,778 | 70.7 |
Annotation of PHE1 ChIP-seq peaks within genomic features of interest.
Annotation for each individual replicate, as well as for common peaks, is presented. For target gene analysis, only common peaks located 1.5 kb upstream to 0.5 kb downstream of the TSS were considered (3rd row).
| Sample | Total no. of peaks | No. of peaks in −1.5 kb to +0.5 kb window around TSS | Average distance to nearest TSS (bp) | Associated genomic feature (% of peaks) | No. of targeted genes | ||
|---|---|---|---|---|---|---|---|
| Promoter | Gene body | Intergenic | |||||
| Replicate 1 peaks | 2818 | 2182 | 445 | 88.6 | 4.7 | 6.6 | 1985 |
| Replicate 2 peaks | 4521 | 3508 | 600 | 86.6 | 3.5 | 9.9 | 2971 |
| Common peaks (PHE1 binding sites) | 2494 | 1995 | 430 | 89.6 | 4.6 | 5.8 | 1694 |
PHE1 target genes previously implicated in endosperm development.
| Gene ID | Imprinting status | Description of function |
|---|---|---|
| AGL62 | Non-imprinted | Type I MADS-box TF involved in endosperm proliferation and seed coat development (Figueiredo et al., 2016; Figueiredo et al., 2015; Kang et al., 2008; Roszak and Köhler, 2011) |
| YUC10 | PEG | Flavin monooxygenase that catalyzes the last step of the Trp-dependent auxin biosynthetic pathway (Zhao, 2012). Involved in endosperm proliferation and cellularization (Batista et al., 2019; Figueiredo et al., 2015) |
| IKU2 | Non-imprinted | Encodes a leucine-rich repeat receptor kinase protein that, together with MINI3, is part of the IKU pathway controlling seed size. iku2 mutants show reduced endosperm growth and early endosperm cellularization (Garcia et al., 2003; Luo et al., 2005). |
| MINI3 | Non-imprinted | WRKY TF that, together with IKU2, is part of the IKU pathway controlling seed size. mini3 mutants show reduced endosperm growth and early endosperm cellularization (Luo et al., 2005). |
| ZHOUPI | Non-imprinted | Encodes a bHLH TF expressed in the embryo-surrounding region of the endosperm. It is essential for embryo cuticle formation and endosperm breakdown after its cellularization (Xing et al., 2013; Yang et al., 2008). |
| MEA | MEG | Subunit of the FIS–PRC2 complex, responsible for depositing H3K27me3 at target loci including PEGs (Moreno-Romero et al., 2016; Moreno-Romero et al., 2019). Loss of MEA and, consequently, paternally biased expression of PEGs lead to a 3x-seed-like phenotype (Grossniklaus et al., 1998; Kiyosue et al., 1999). |
| ADM | PEG | Interacts with SUVH9 and AHL10 to promote H3K9me2 deposition in TEs, influencing the expression of neighboring genes. Mutations in ADM lead to rescue of the 3x seed abortion phenotype (Jiang et al., 2017; Kradolfer et al., 2013; Wolff et al., 2015). |
| SUVH7 | PEG | Encodes a putative histone-lysine N-methyltransferase. Mutations in SUVH7 lead to rescue of the 3x seed abortion phenotype (Wolff et al., 2015). |
| PEG2 | PEG | Encodes an unknown protein, which is not translated in the endosperm. PEG2 transcripts act as a sponge for siRNA854, thus regulating UBP1 abundance (Wang et al., 2018). Mutations in PEG2 lead to rescue of the 3x seed abortion phenotype (Wang et al., 2018; Wolff et al., 2015). |
| NRPD1a | PEG | Encodes the largest subunit of RNA POLYMERASE IV, which is involved in the RNA-directed DNA methylation pathway. Mutations in NRPD1a lead to rescue of the 3x seed abortion phenotype (Erdmann et al., 2017; Martinez et al., 2018). |
H3K27me3 ChIP-seq read mapping and purity information.
| Sample | No. trimmed reads | % of mapped reads | No. of Ler reads | No. of Col reads | Purity (%) |
|---|---|---|---|---|---|
| Ler x Col 4x | 30,337,933 | 68.3 | 1,844,412 | 1,563,394 | 95.7 |
| Replicate 1 Input | |||||
| Ler x Col 4x | 22,644,505 | 73.3 | 1,439,255 | 1,211,446 | |
| Replicate 1 H3 ChIP | |||||
| Ler x Col 4x | 27,448,642 | 61.5 | 1,214,486 | 681,823 | |
| Replicate 1 H3K27me3 ChIP | |||||
| Ler x Col 4x | 40,500,367 | 66.4 | 2,720,117 | 2,483,912 | 97.7 |
| Replicate 2 Input | |||||
| Ler x Col 4x | 32,322,049 | 71.9 | 2,304,635 | 2,068,612 | |
| Replicate 2 H3 ChIP | |||||
| Ler x Col 4x | 34,978,215 | 63 | 2,681,981 | 1,636,717 | |
| Replicate 2 H3K27me3 ChIP |
-
The purity of INTACT-extracted endosperm nuclei is indicated in the last column and was calculated as described in Moreno-Romero et al. (2017).
| Capsella rubella | |||
| total number of genes | number of genes with PHE1 DNA-binding motifs | percentage (%) | |
| MEGs | 77 | 52 | 0.68 |
| PEGs | 52 | 45 | 0.87 |
| all genes | 26521 | 19333 | 0.73 |
| Arabidopsis thaliana | |||
| total number of genes | number of genes with PHE1 DNA-binding motifs | percentage (%) | |
| MEGs | 145 | 97 | 0.69 |
| PEGs | 150 | 123 | 0.82 |
| all genes | 33323 | 23784 | 0.71 |
| p-values | Capsella rubella | Arabidopsis thaliana |
| PEGs vs. all genes | 0.0269 | 0.0040 |
| MEGs vs. all genes | 0.2896 | 0.2329 |