Peptidoglycan-dependent NF-κB activation in a small subset of brain octopaminergic neurons controls female oviposition
Figures
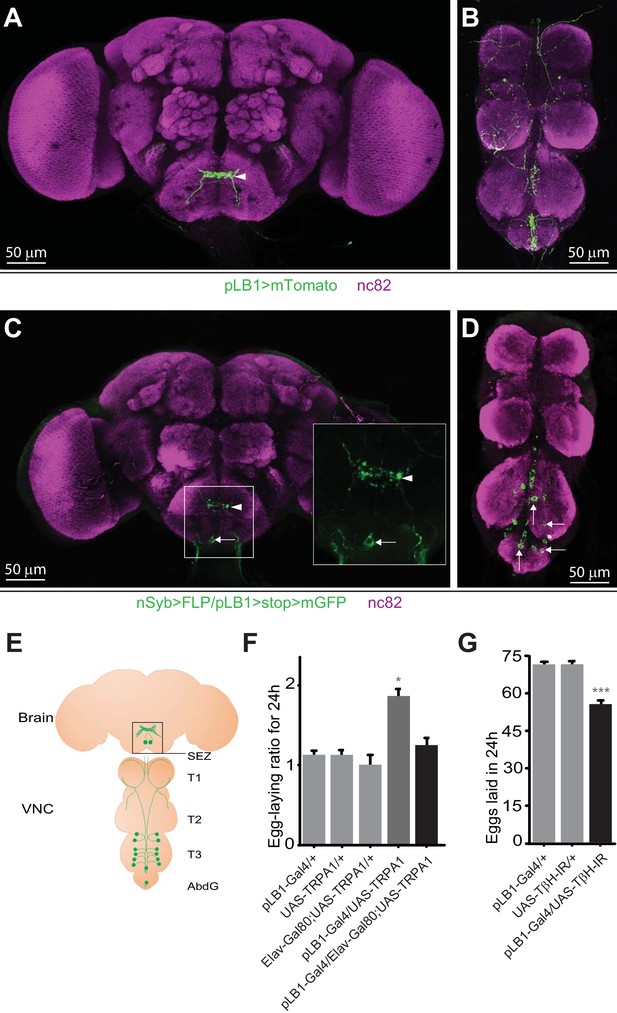
pLB1 is expressed in neurons modulating egg-laying via octopamine.
(A, B); Immunodetection of cells expressing pLB1-Gal4/UAS-Tomato-mCD8 (pLB1>mTomato) in females. For the homogeneity of the different images, the red signal corresponding to Tomato-mCD8 was converted in green. In the brain (A), pLB1 is expressed in the Sub Esophageal Zone (SEZ) (arrowhead). In the ventral nerve cord (VNC) (B), the network links the brain to T1, T2, T3 and the Abdominal Ganglia (AbdG).(C, D); Pattern of cells co-expressing the neuronal markers nSyb and the pLB1 driver (nSyb>FLP/pLB1>stop>mGFP). The GFP can only be expressed if the stop sequence inserted upstream of the gfp gene is flipped-out in pLB1+/nSyb+ cells. In the brain (C), neuronal projections are found in the SEZ (arrowhead) as well as a cell body (arrow). The inserted box is a magnification of the SEZ. In the VNC (D), 8 to 14 cell bodies are revealed in T3 and AbdG.(E); Map representing projections and cell bodies of neurons expressing pLB1 in brain and VNC. (F); Preventing the expression of the transient receptor potential cation channel, subfamily A, member 1 (TRPA1) in pLB1+ neurons impairs the egg-laying increase in non-infected mated females. The egg-laying ratio for 24 hours (24 h) corresponds to the number of eggs laid by a female at the restrictive temperature (29°C) over the average number of eggs laid by females of the same genotype at the permissive temperature (23°C). (G); RNAi-mediated inactivation of the octopamine-producing enzyme TβH in pLB1+ cells reduces egg-laying. For (A–D), immuno-staining against the neuropil marker nc82 was used to stain the organ. For (F), shown is the average egg-laying ratio per 24 hours ± SEM (29°C/23°C) from at least two independent trials with at least 16 females per genotype and condition used. An egg-laying ratio of 1 indicates an absence of difference between the test and the control. For (G), shown is the average number of eggs laid per fly per 24 hours ± SEM from at least three independent trials with at least 58 females per genotype and condition used. * indicates p<0.05, *** indicates p<0.0001; non-parametric ANOVA, Dunn’s multiple comparison test. Details including n values and genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
-
Figure 1—source data 1
Egg laying raw data for Figure 1F.
- https://doi.org/10.7554/eLife.50559.003
-
Figure 1—source data 2
Egg laying raw data for Figure 1G.
- https://doi.org/10.7554/eLife.50559.004
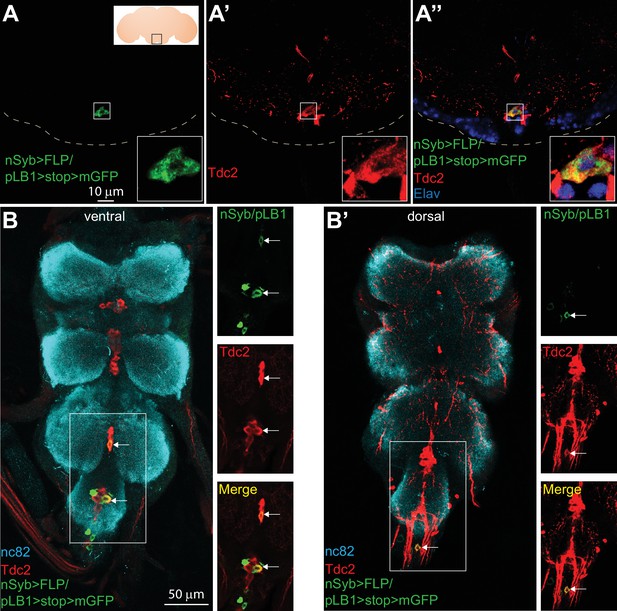
Some of the pLB1+ neurons are octopaminergic.
(A–B’); Immuno-detection in the brain Sub Esophageal Zone (SEZ; A–A’’) and ventral nerve cord (VNC; B–B’) of neurons expressing pLB1 (nSyb>FLP/pLB1>stop>mGFP) (A,B and B’) and producing the Tdc2 enzyme (A’, B and B’). In (A’’), the nuclear neuronal marker Elav was also immuno-detected. For (A), the inserted scheme represents the brain and the empty black square delineates the observed area. For (A-A’’), the inserted box is a magnification of the outlined box and the dashed line represents the ventral limit of the brain. For (B-B’), staining against nc82 was used to delineate the shape of the VNC. For (B–B’), the merged channels of the outlined box are separated on the images on the right. Arrows point to pLB1+/Tdc2+ cells in the VNC. Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
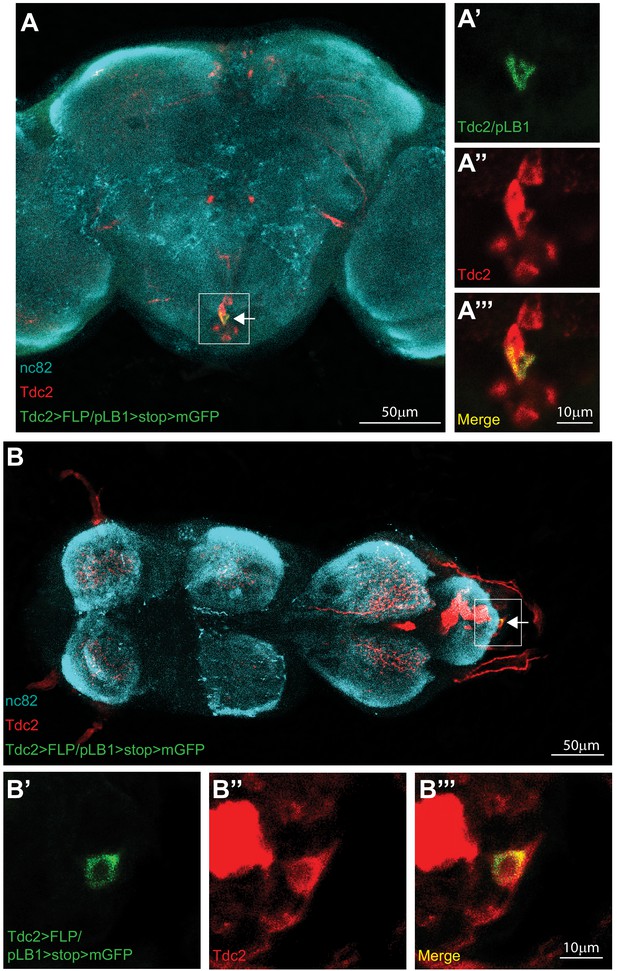
Some octopaminergic neurons are pLB1+.
Immuno-detection in the brain (A-A’’’) and ventral nerve cord (VNC; B-B’’’) of cells co-expressing Tdc2 and pLB1 (Tdc2>FLP/pLB1>stop>mGFP). The GFP can only be expressed if the stop sequence inserted upstream of the gfp gene is flipped-out in pLB1+/Tdc2+ cells. For (A and B), the area outlined in the box is magnified in (A’-A’’’) and (B’-B’’’), respectively. Staining against nc82 was used to delineate the shape of the brain (A) and VNC (B). The arrow indicates neurons co-expressing pLB1 and Tdc2. Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
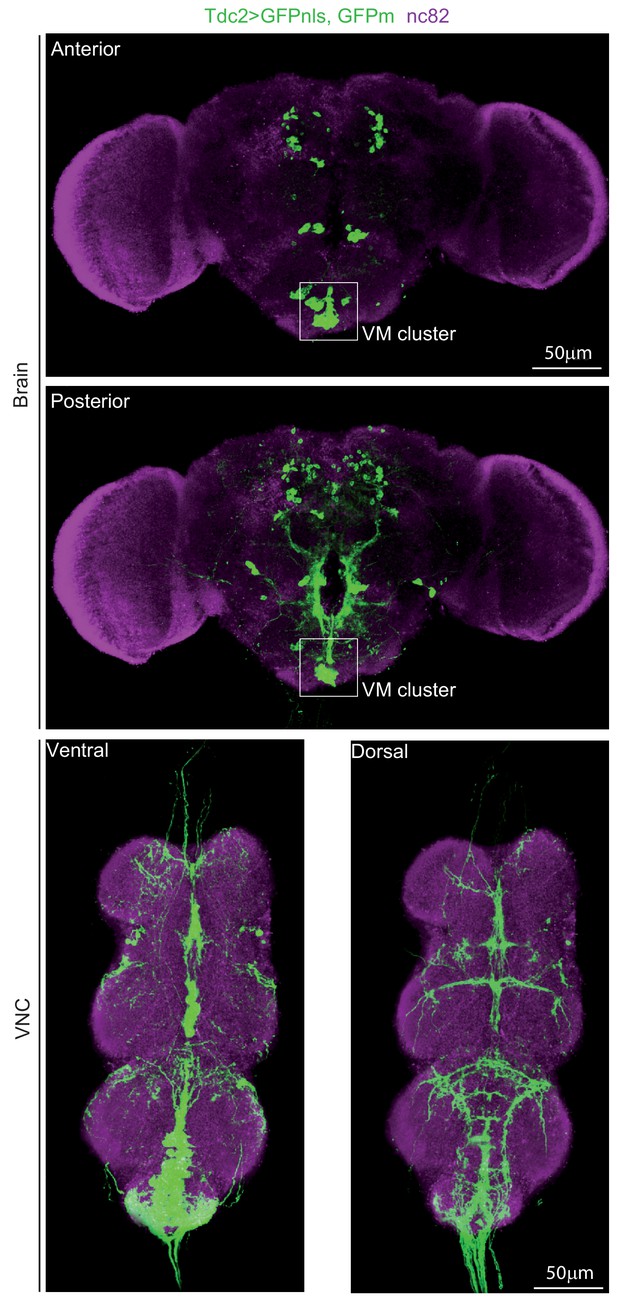
Map of Tdc2 expressing neurons in the brain and VNC.
Immunodetection in the brain and ventral nerve cord (VNC) of cells expressing Tdc2. The area in the brain outlined in the box corresponds to the VM cluster. In the projection of the anterior view of the brain, VM I and VM II sub-clusters are visible. In the projection of the posterior view of the brain, the VM III sub-cluster is visible. Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
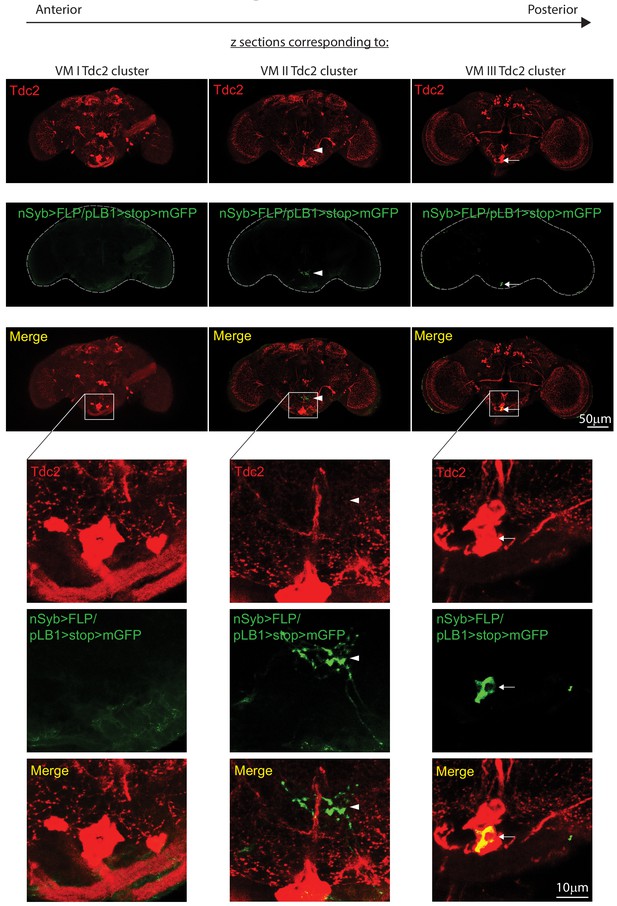
The pLB1+ octopaminergic neurons in the brain belong to the VM III sub-cluster.
Immuno-detection in the brain of neurons expressing pLB1 (nSyb>FLP/pLB1>stop>mGFP) and producing the enzyme Tdc2. Brain stacks from anterior to posterior with focal plane corresponding to VM I, VM II and VM III octopaminergic sub-clusters are shown. The sub-clusters delimited by the outlined box are magnified and the individual channels shown. Arrows indicate cell bodies and arrowheads projections of pLB1+ neurons. Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
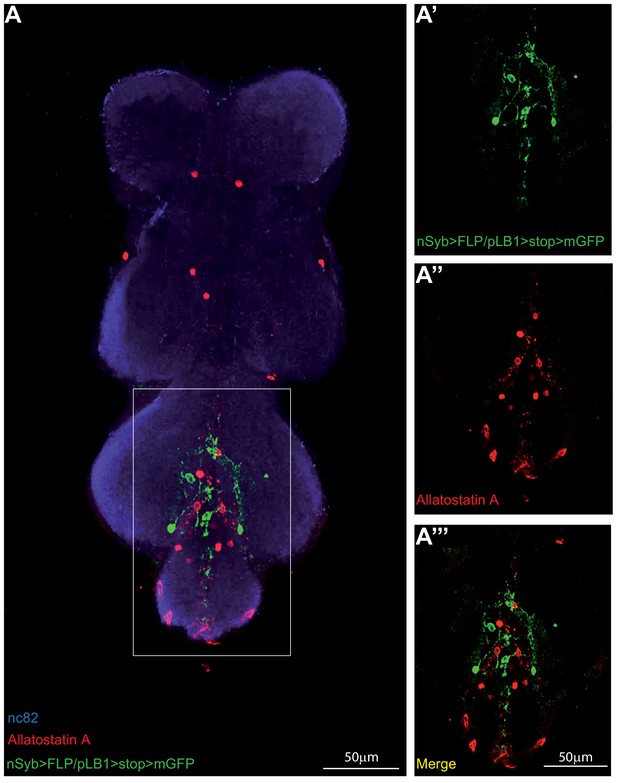
pLB1-expressing neurons in the VNC do not produce Allatostatin A.
(A-A’’’); Immuno-detection in the ventral nerve cord (VNC) of neurons expressing pLB1 (nSyb>FLP/pLB1>stop>mGFP) and producing the Allatostatin A neuropeptide. Shown is a maximum intensity projection. The area outlined in the box is magnified in (A’-A’’’). Staining against nc82 was used to delineate the shape of the VNC (A). Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
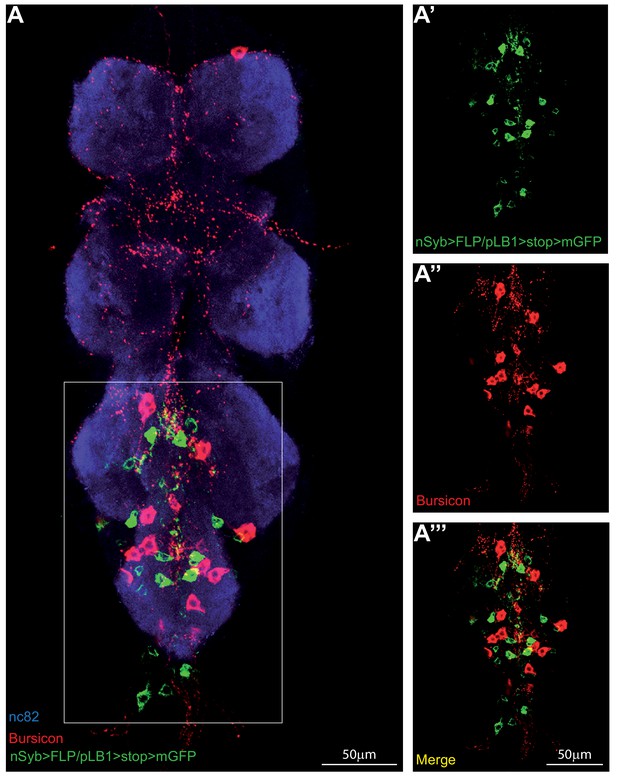
pLB1+ neurons in the VNC do not produce Bursicon.
(A-A’’’); Immuno-detection in the ventral nerve cord (VNC) of neurons expressing pLB1 (nSyb>FLP/pLB1>stop>mGFP), and producing the Bursicon neuropeptide. Shown is a maximum intensity projection. The area outlined in the box is magnified in (A’-A’’’). Staining against nc82 was used to delineate the shape of the VNC (A). Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
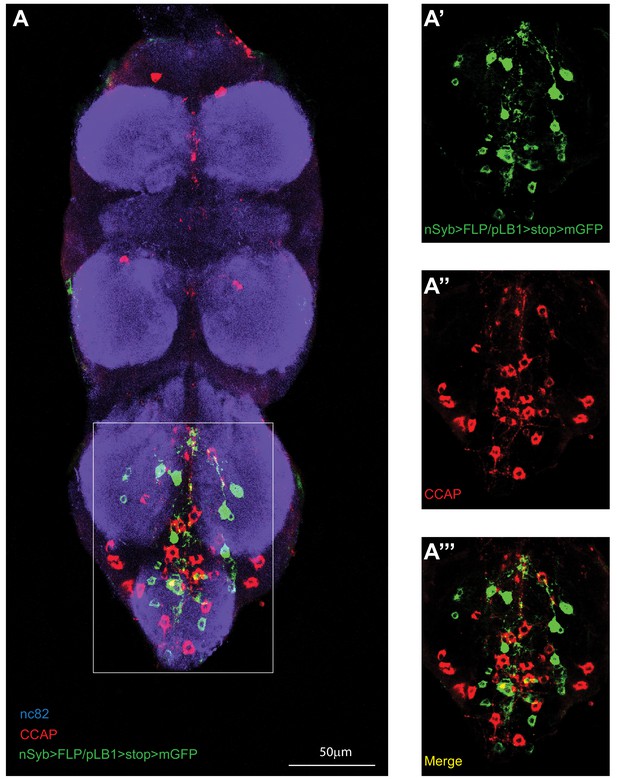
pLB1+ neurons in the VNC do not produce CCAP.
(A-’’’); Immuno-detection in the ventral nerve cord (VNC) of neurons expressing pLB1 (nSyb>FLP/pLB1>stop>mGFP)and producing the Crustacean cardioactive peptide (CCAP). Shown is a maximum intensity projection. The area outlined in the box is magnified in (A’-A’’’). Staining against nc82 was used to delineate the shape of the VNC (A). Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
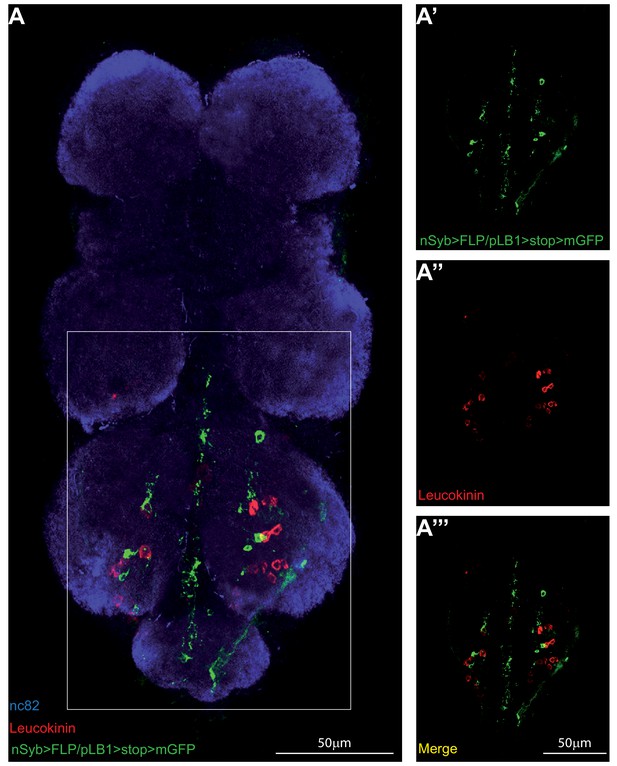
pLB1+ neurons in the VNC do not produce Leucokinin.
(A-A’’’); Immuno-detection in the ventral nerve cord (VNC) of neurons expressing pLB1 (nSyb>FLP/pLB1>stop>mGFP) and producing the Leucokinin neuropeptide. Shown is a maximum intensity projection. The area outlined in the box is magnified in (A’-A’’’). Staining against nc82 was used to delineate the shape of the VNC (A). Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
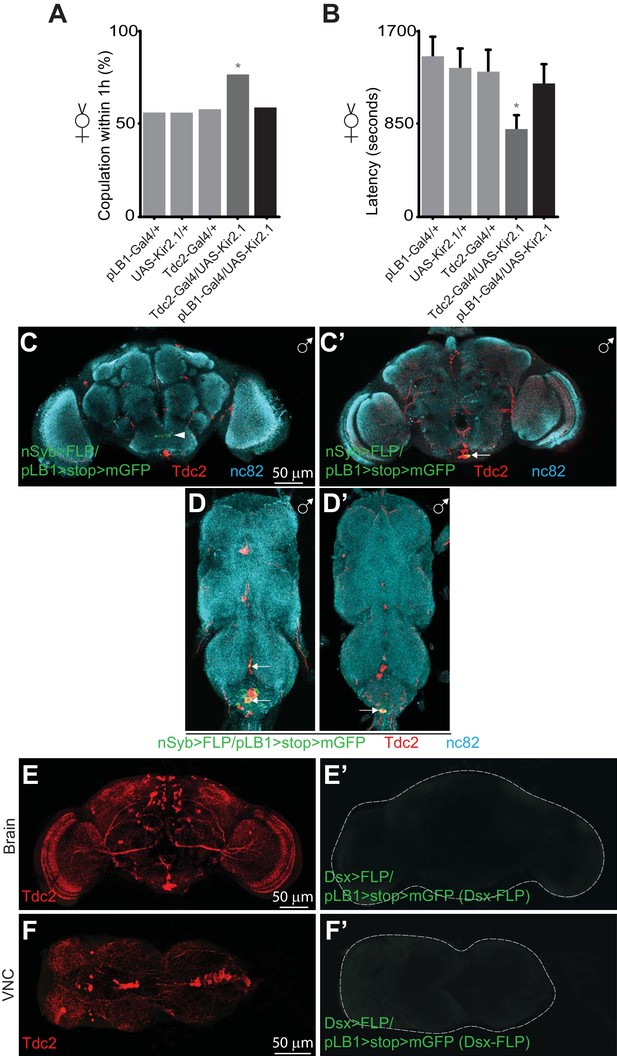
pLB1+ neurons are different from Tdc2+/Dsx+ neurons controlling receptivity.
Impairing the activity of pLB1+ neurons via UAS-Kir2.1 neither reduces virgin copulation percentage (A) nor mating latency (B). (C-D’); In adult males, immuno-detection in the CNS of neurons expressing pLB1 and producing the Tdc2 enzyme. (C and C’) are the brain anterior and posterior views, respectively. (D and D’) are the ventral nerve cord (VNC) ventral and dorsal views, respectively. In adult females, immuno-detection in the brain (E-E’) and VNC (F-F’) of Dsx+/pLB1+ cells and producing the Tdc2 enzyme; no signal for Dsx+/pLB1+ cells is detectable. For (A), shown is the copulation percentage for virgins within 1 hour (1h) from six independent trials with a total of 70–80 females per genotype and condition used. All the tested flies were pooled for the calculation and error bars are not appropriate for this kind of representation. For (B), shown is the average latency time before mating for virgins from four independent trials with a total of 24 to 40 females per genotype and condition used. For (C-D’), staining against nc82 was used to delineate the shape of the brain and VNC. Arrows indicate the position of pLB1+ cell bodies. Arrowheads indicate projections. * indicates p<0.05; Fisher exact t-test (A) and non-parametric t-test, Mann-Whitney test (B). Details including n values and genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
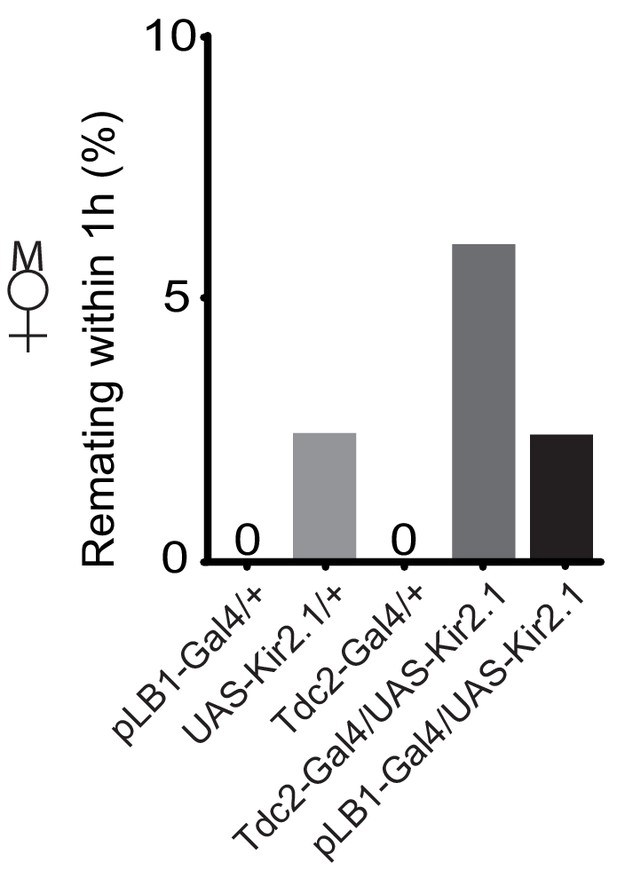
pLB1+ neurons may not control remating behavior.
Impairing the activity of pLB1 neurons via UAS-Kir2.1 does not increase the remating percentage. Shown is the average remating percentage for mated females per 1 hour (1h) from four independent trials with a total of 42 to 55 females per genotype and condition used. For statistics, Fisher exact t-test was used and despite the trends, there were no statistically significant differences. Details including n values and genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
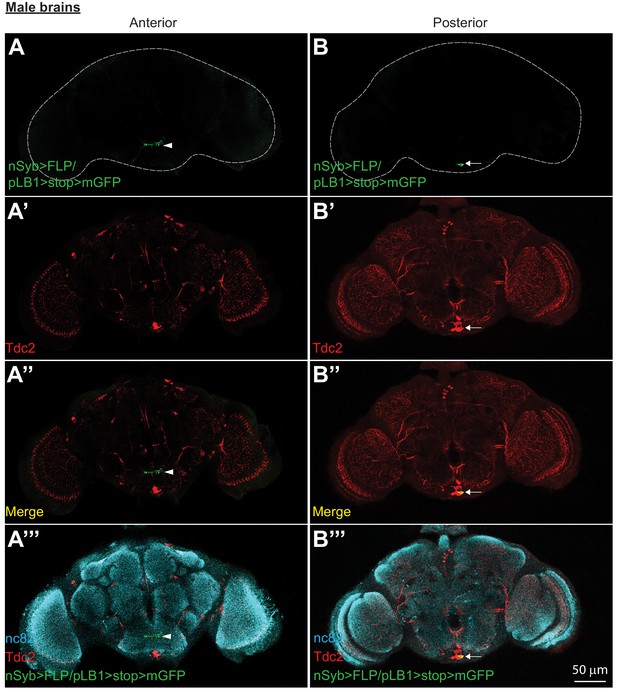
pLB1+/Tdc2+ neurons are present in male brains.
In adult males, immuno-detection in the anterior (A-A’’’) and posterior (B-B’’’) part of the brain of neurons expressing pLB1 and producing the enzyme Tdc2. For (A’’’ and B’’’), staining against nc82 was used to delineate the shape of the brain. Arrows indicate position of pLB1+ cell bodies. Arrowheads indicate projections. Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
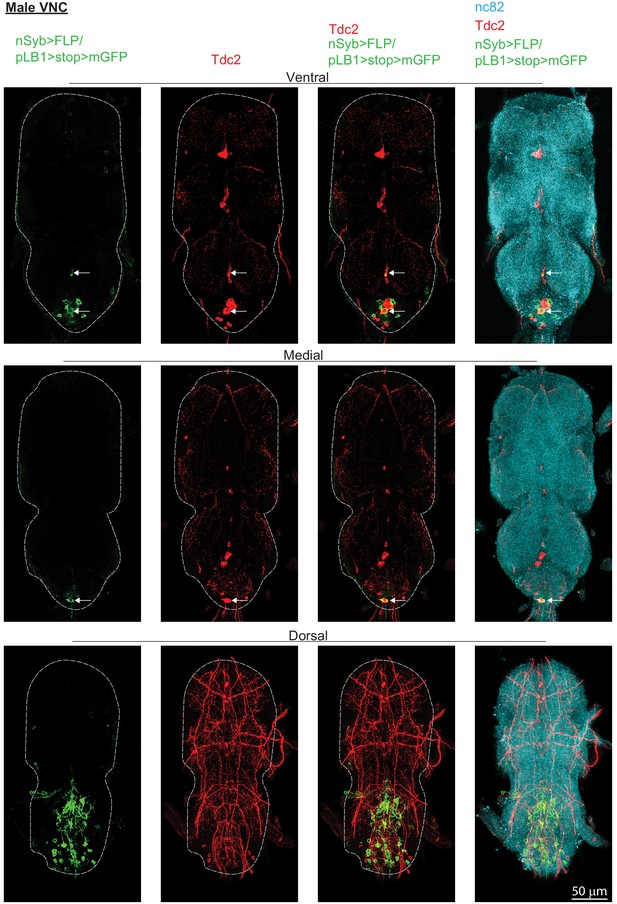
pLB1+/Tdc2+ neurons are present in male VNC.
In adult males, immuno-detection in the ventral, medial and dorsal part of the ventral nerve cord (VNC) of neurons expressing pLB1 and producing the enzyme Tdc2. Staining against nc82 was used to delineate the shape of the VNC. Arrows indicate the positions of pLB1+ cell bodies. Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
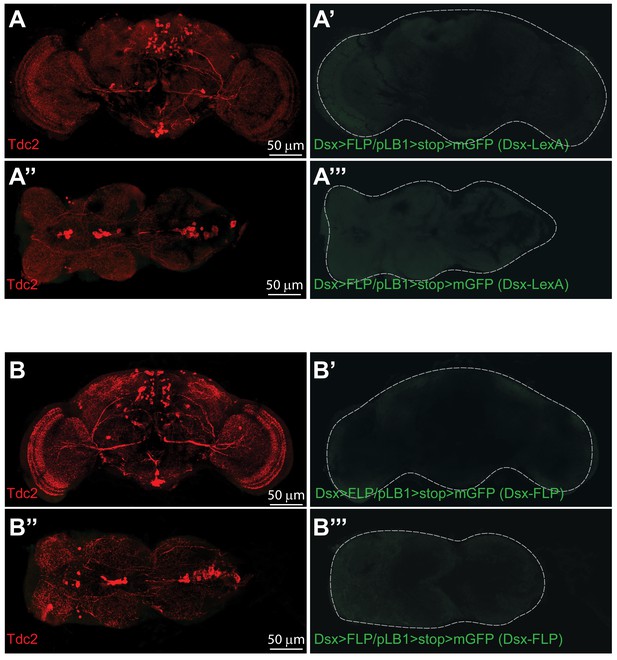
pLB1+ neurons are not Dsx+.
In adult females, immuno-detection in the brain (A-A’, B-B’) and ventral nerve cord (A’’-A’’’, B’’-B’”) of Dsx+ cells expressing pLB1 and producing the enzyme Tdc2 via intersectional strategy and the use of either Dsx-LexA (A’ and A’’’) or Dsx-FLP (B’ and B’’’); no signal for Dsx+/pLB1+ cells is detectable. Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
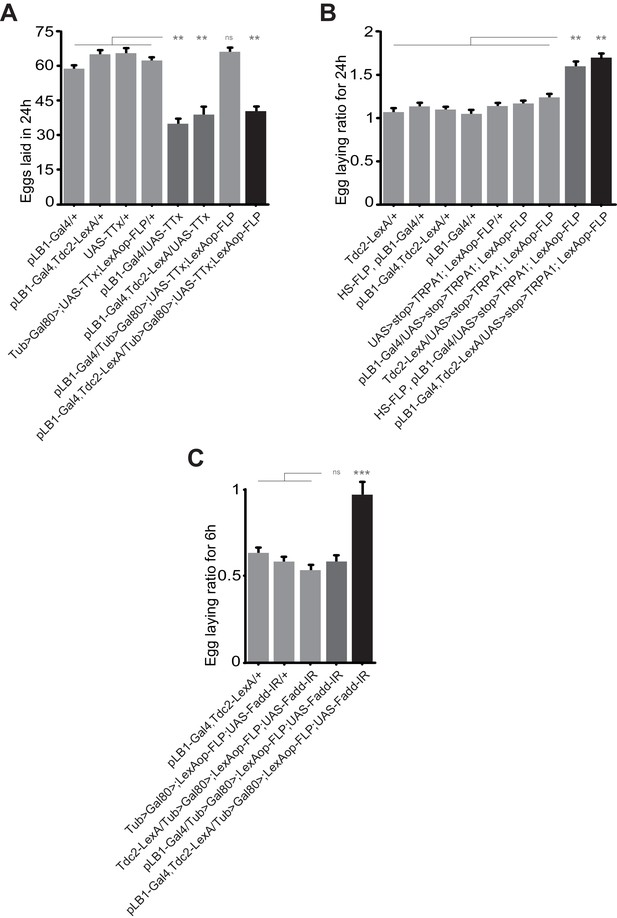
Egg-laying drop post peptidoglycan exposure is mediated by pLB1+/Tdc2+ neurons via the NF-κB pathway.
(A); Impairing the activity of octopaminergic pLB1+ neurons reduces egg-laying. The ubiquitously expressed Tub-Gal80 that inhibits the activity of Gal4 can be flipped-out in cells expressing the LexA. Thus, only in cells co-expressing the Gal4 and the LexA the UAS-TTx will be expressed. (B); Increasing the activity of pLB1+ neurons augments egg-laying. The ubiquitously expressed Tub-Gal80 can be flipped-out only in cells expressing the heat shock (HS) flippase or the LexA. Thus, only in cells co-expressing the Gal4 and the LexA the UAS-TRPA1 will be expressed. (C); Octopaminergic pLB1+ neurons control the egg-laying drop post-peptidoglycan injection via Fadd. Only cells co-expressing the Gal4 and LexA express the Fadd RNAi (UAS-Fadd-IR) transgene. For (A), shown are the average numbers of eggs laid per fly per 24 hours ± SEM from at least two independent trials with at least 20 females per genotype and condition used. For (B), shown are the average egg-laying ratios per fly per 24 hours ± SEM from three independent trials with at least 35 females per genotype and condition used. For (C), shown are the average egg-laying ratios per fly per 6 hours ± SEM from at least two independent trials with at least 20 females per genotype and condition used. * indicates p<0.05; ** indicates p<0.001; *** indicates p<0.0001; n.s. indicates p>0.05, non-parametric ANOVA, Dunn’s multiple comparison test. Details including n values and genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
-
Figure 4—source data 1
Egg laying raw data for Figure 4A.
- https://doi.org/10.7554/eLife.50559.019
-
Figure 4—source data 2
Egg laying raw data for Figure 4B.
- https://doi.org/10.7554/eLife.50559.020
-
Figure 4—source data 3
Egg laying raw data for Figure 4C.
- https://doi.org/10.7554/eLife.50559.021
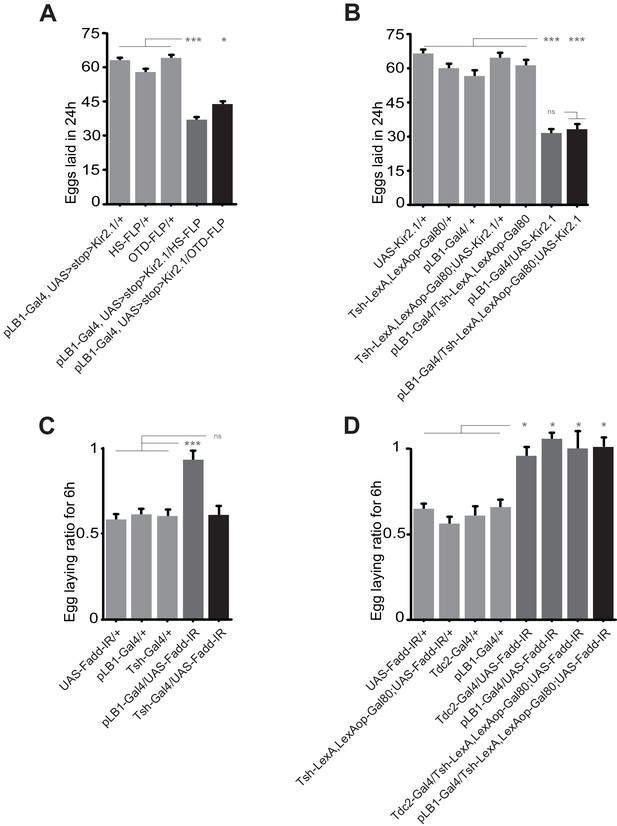
Egg-laying drop post peptidoglycan exposure is mediated by the brain, but not the VNC pLB1+ neurons.
(A); Impairing the activity of pLB1+ cells of the brain reduces egg-laying. Only in cells co-expressing the FLP and the Gal4 will the UAS>stop>Kir2.1 be effective. Heat shock (HS) is ubiquitous while OTD is brain-restricted. (B); pLB1+ cells of the VNC are dispensable for the modulation of the egg-lay. The expression of LexAop-Gal80 antagonizes the activity of Gal4, thus preventing the effects of UAS-Kir2.1. Tsh-LexA drives the expression of Gal80 in the fly thorax, including the VNC. (C); RNAi-mediated Fadd (Fadd-IR) inactivation in the Tsh-Gal4+ cells does not prevent egg-lay drop post peptidoglycan injection. (D); RNAi-mediated Fadd inactivation in pLB1+ cells of the brain, but not of the VNC prevents egg-lay drop post peptidoglycan injection. The expression of LexAop-Gal80 antagonizes the activity of Gal4, thus preventing the effects of UAS-Fadd-IR, only in cells co-expressing the Gal4 and the LexA. For (A and B), shown are the average numbers of eggs laid per fly per 24 hours ± SEM from at least two independent trials with at least 20 females per genotype and condition used. For (C and D), shown are the average egg-laying ratios per fly per 6 hours ± SEM from at least two independent trials with at least 20 females per genotype and condition used. * indicates p<0.05; ** indicates p<0.001; *** indicates p<0.0001; n.s. indicates p>0.05, non-parametric ANOVA, Dunn’s multiple comparison test. Details including n values and genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
-
Figure 5—source data 1
Egg laying raw data for Figure 5A.
- https://doi.org/10.7554/eLife.50559.024
-
Figure 5—source data 2
Egg laying raw data for Figure 5B.
- https://doi.org/10.7554/eLife.50559.025
-
Figure 5—source data 3
Egg laying raw data for Figure 5C.
- https://doi.org/10.7554/eLife.50559.026
-
Figure 5—source data 4
Egg laying raw data for Figure 5D.
- https://doi.org/10.7554/eLife.50559.027
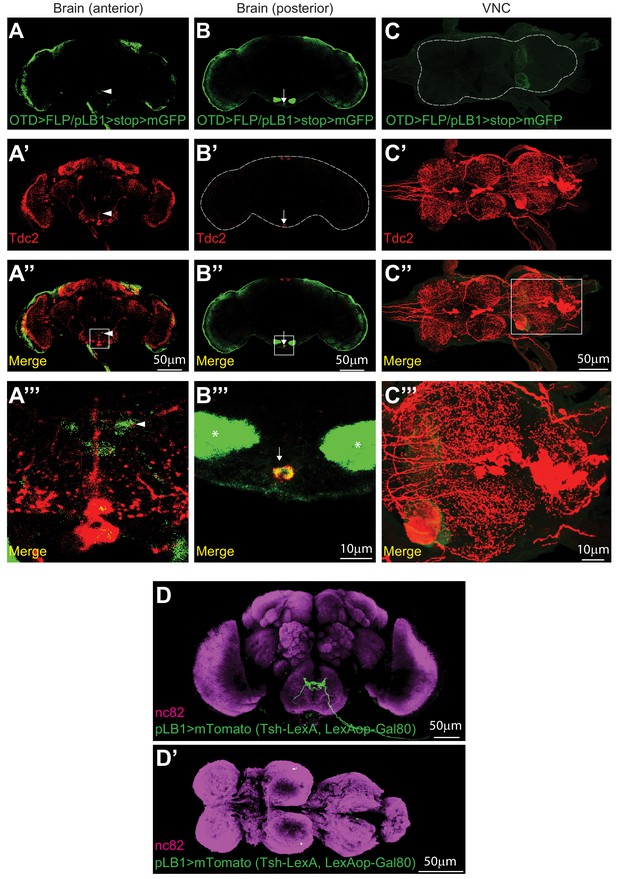
OTD-FLP is expressed in pLB1+ neurons of the head, but not of the thorax whileTsh-LexA, LexAop-Gal80 efficiently silences pLB1-Gal4 in the thorax.
(A–C’’’); Immunodetection of cells co-expressing pLB1-Gal4 and OTD-FLP (OTD>FLP/pLB1>stop >mGFP) and producing the enzyme Tdc2 in the anterior (A–A’’’) and posterior (B–B’’’) parts of the brain as well as in the VNC (C–C’’’). (D–D’); Immunodetection of cells expressing pLB1-Gal4/UAS-Tomato-mCD8 (pLB1>mTomato) in the brain, but not the VNC of flies co-expressing Tsh-LexA/LexAop-Gal80. For the homogeneity of the different images, the red signal corresponding to Tomato-mCD8 was converted in green. For (A–C’’’), pLB1+/OTD+/Tdc2+ neurons were detected in the brain. Arrows indicate cell bodies and arrowheads projection of pLB1+ neurons; no signal was detected in the VNC. In (B’’’), asterisks correspond to background. (A’’’, B’’’ and C’’’) correspond to magnification of the area delimited by the box in (A’’, B’’ and C’’). For (D), pLB1 is expressed in the brain. For (D’), pLB1 was not detected in the VNC. Details including n values and genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
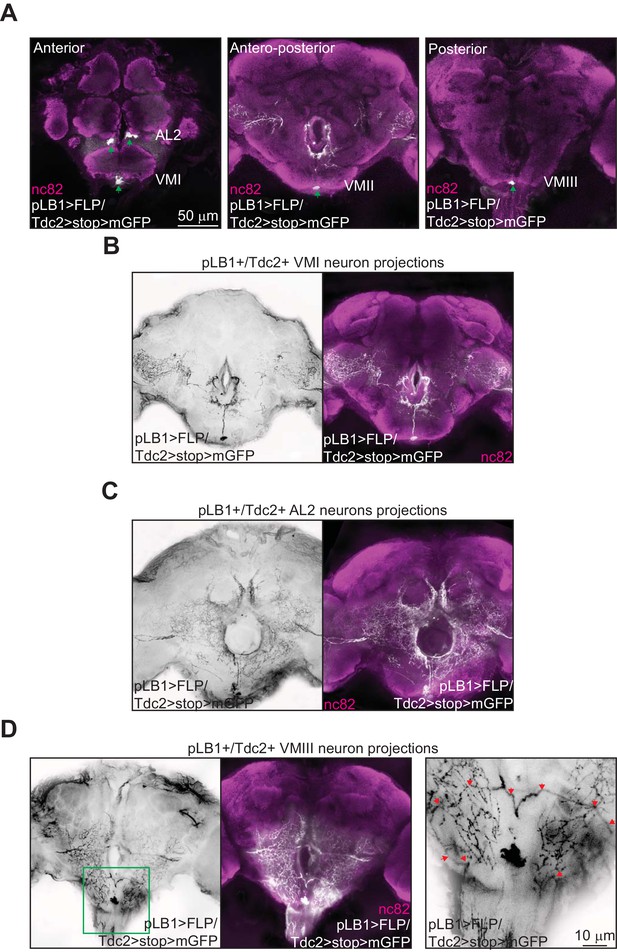
Brain pLB1+/Tdc2+ neurons project to the esophagus as well as to the VNC.
(A–D); Immuno-detection in the brain of cells co-expressing the Tdc2-Gal4 and the pLB1-LexA drivers (pLB1>FLP/Tdc2>stop>mGFP). The GFP can only be expressed under the control of Tdc2 if the stop sequence inserted upstream of the gfp gene is flipped-out in pLB1+/Tdc2+ cells. (A); Five cellular bodies (green arrows) are detected in anterior, antero-posterior and posterior parts of the brain. (B–D); Specific stacks of the brain showing the projection patterns (in black) of the neurons present in VM I (B), AL2 (C) and VM III (D). In (D), the area delimited by the green box in the left panel is magnified on the right panel to show the descending projections (red arrows). Staining against nc82 was used to delineate the shape of the brain. Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
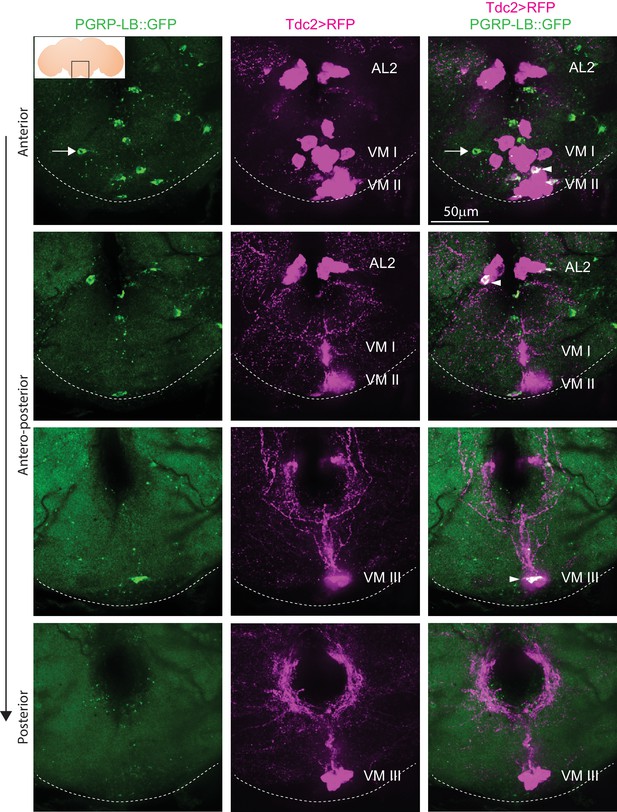
Endogenous PGRP-LB::GFP is expressed in Tdc2+ cells of the AL2 and VM clusters.
Detection of PGRP-LB::GFP fusion protein as well as Tdc2 cells expressing RFP (Tdc2>RFP) without immunostaining . Only the area of the brain containing the octopaminergic AL2, VM I, VM II, and VM III clusters is shown with stacks corresponding to anterior, antero-medial, postero-medial and posterior views. The inserted scheme represents the brain, the empty black square delineates the area observed and the dashed line represents the ventral limit of the brain. Arrowheads point to Tdc2+/PGRP-LB::GFP+ cells and the arrow points to a Tdc2 negative cell containing PGRP-LB::GFP proteins. Details including genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
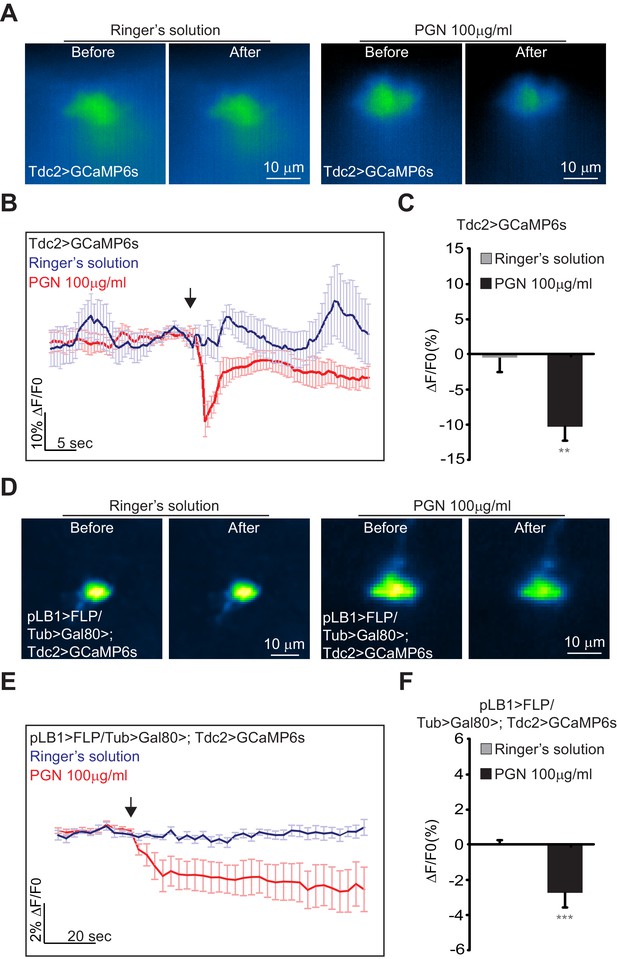
Real-time calcium imaging of Tdc2 VM II/III and pLB1+/Tdc2+ VM III neurons exposed to peptidoglycan.
(A–F); Real-time calcium imaging using the calcium indicator GCaMP6s to reflect the in vivo neuronal activity of Tdc2 neurons (Tdc2>GCaMP6s) in VM II/VM III sub-clusters (A–C) or the ex vivo neuronal activity of Tdc2+/pLB1+ VM III neuron pLB1>FLP/Tub>Gal80>,Tdc2>GCaMP6s (D–F). (A); Representative images showing the GCaMP6s intensity before and after addition of either the control Ringer’s solution (left panels) or the peptidoglycan (right panels). The images represent the average intensity of 4 frames before or after Ringer or peptidoglycan solution. (B); Averaged ± SEM time course of the GCaMP6s intensity variations (ΔF/F0 %) for Tdc2+ neurons of the VM II/VM III sub-clusters. The addition of Ringer’s solution (n=8 flies) or peptidoglycan (n=13 flies) at a specific time is indicated by the arrow. (C); Averaged fluorescence intensity of negative peaks ± SEM for control (n=8) and peptidoglycan-treated flies (n=13). (D); Representative images showing the GCaMP6s intensity before and after the addition of either the control Ringer’s solution (left panels) or the peptidoglycan (right panels). The images represent the average intensity of 4 frames before or after Ringer or peptidoglycan solution. (E); Averaged ± SEM time course of the GCaMP6s intensity variations (ΔF/F0 %) for Tdc2+/pLB1+ neuron of the VM III sub-cluster. The addition of Ringer’s solution (n=10 flies) or peptidoglycan (n=12 flies) at a specific time is indicated by the arrow. (F); Averaged fluorescence intensity of negative peaks ± SEM for control (n=10) and peptidoglycan-treated flies (n=12) In (C), ** indicates p=0.001; in (F), *** indicates p=0.0001 non-parametric t-test, Mann-Whitney test. Details including n values and genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
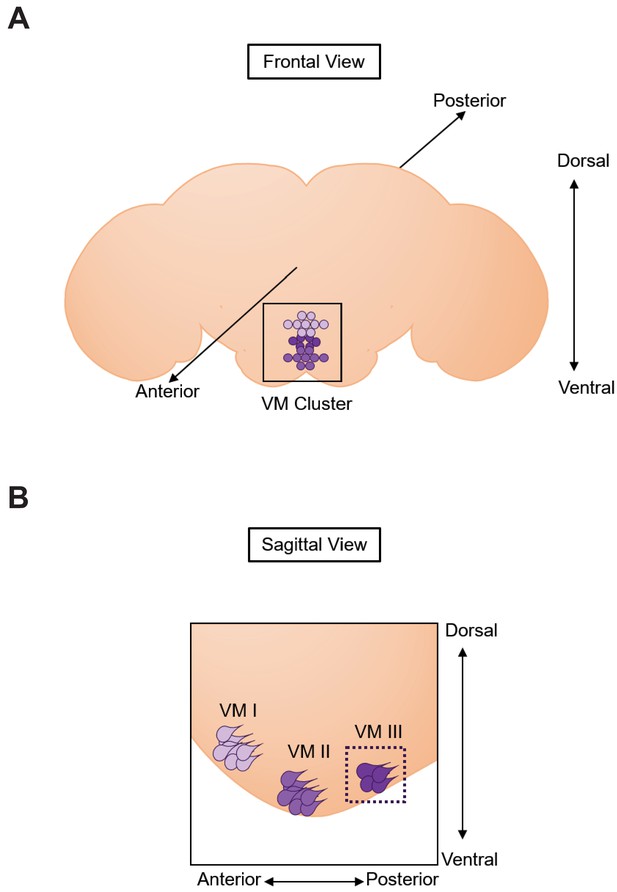
The in vivo and ex vivo real-time Calcium imaging approaches focused on neurons present in the VM II/III octopaminergic sub-cluster.
Scheme representing a frontal view (A) of a brain and a sagittal view (B) of the octopaminergic VM cluster. The VM cluster is located ventrally along the midline in the frontal view (A). Three main sub-clusters (neurons represented with colored circles) can be delineated along the antero-posterior axis as represented in the sagittal view (B). We focused on the VM II/III sub-clusters for the in vivo approach and only on the most posterior group (VM III sub-cluster), delineated by a dashed line in (B) for the ex-vivo assays.
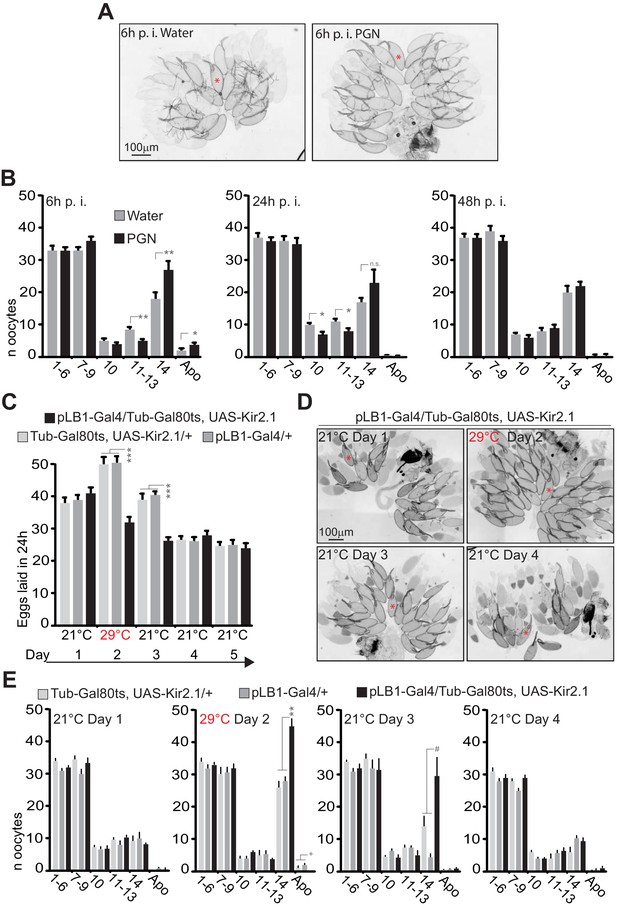
Peptidoglycan exposure as well as pLB1+ neurons conditional inactivation lead to a reversible mature oocyte accumulation.
(A–B); Injection of peptidoglycan triggers a reversible accumulation of mature oocytes (stage14). (A); 6 hours (6h) post-treatment (p.i.), stage 14 oocytes accumulate in the ovaries of peptidoglycan-injected flies. Transmission light microscopy images of ovaries dissected from control flies (water injected) or peptidoglycan-injected animals 6h post-treatment. (B); peptidoglycan injection modifies the quantity and quality of oocytes by 6 h, leading to an accumulation of mature stage (stage14) eggs. The dynamic over three different time points (6h-24h-48h) post-treatment was assayed. (C–E); pLB1+ neurons reduced activity leads to stage 14 oocyte accumulation. (C); The conditional inactivation of pLB1+ neurons reduces egg-laying. At 21°C, the ubiquitously produced Gal80ts inactivates the Gal4 and thus the Kir 2.1 protein expression. At 29°C, the Gal80ts doesn’t inactivate the Gal4, leading to the inhibited activity of pLB1+ neurons via Kir2.1. Switching back the animals to 21°C inhibits the Gal4 activity via Gal80ts. (D); Conditional inactivation of the pLB1+ neurons triggers a reversible stage 14 oocytes accumulation. Ovaries images of pLB1-Gal4/Tub-Gal80ts, UAS-Kir2.1 flies were acquired with transmission light microscopy for 4 days at two different temperatures. (E); pLB1+ neurons conditional inactivation modifies the quantity and quality of oocytes,leading to an accumulation of stage 14 oocytes. The dynamic over four different time points and two different temperatures is shown. It is important to note that the switch from 21°C to 29°C might be stressful for all the lines since stage 14 oocytes accumulated in all of them. In (A and D), a prototypical stage 14 oocyte is indicated with a red asterisk. In (B), shown are the average numbers over time of different oocyte stages ± SEM from two cumulated independent trials with at least 18 females per genotype and condition used. In (C), shown are the average numbers of eggs laid per fly per 24h ± SEM over 5 days from two cumulated independent trials with at least 59 females per genotype and condition used. In (E), shown are the average numbers of different oocyte stages ± SEM over 4 days for one representative assay out of two independent trials with at least 10 females per genotype and condition used. For (B) and (E), on the x axis, 1–6 corresponds to early stages (from stage 1 to stage 6)oocytes ; 7–9 corresponds to the sum of stages 7, 8 and 9; 10 is for stage 10; 11–13 is for the sum of stages 11, 12 and 13; 14 is for stage 14 and Apo is for apoptotic events, all identified via DAPI staining. * indicates p<0.05; ** indicates p<0.01; *** indicates p<0.0001; + and # indicate statistically significant differences between the test and the controls, but not all of them (see detailed statistics for Figure 9E). In (B), non-parametric t-test, Mann-Whitney test; in (C and E), non-parametric ANOVA, Dunn’s multiple comparison test between the genotypes or treatments. Details including n values and genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
-
Figure 9—source data 1
Egg laying raw date for Figure 9C.
- https://doi.org/10.7554/eLife.50559.039
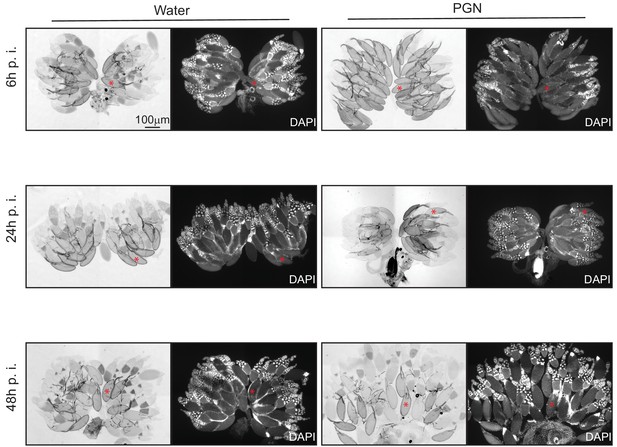
Peptidoglycan exposure leads to a reversible accumulation of stage 14 oocytes.
Peptidoglycan injection triggers stage 14 oocyte accumulation that is reversed by 24 hours (24h). Ovaries seen with transmission light microscopy and DAPI staining from control flies (water injected) or peptidoglycan-injected animals 6h, 24h and 48h post-treatment (p.i.). Prototypical stage 14 oocyte is indicated with a red asterisk. Details including n values and genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
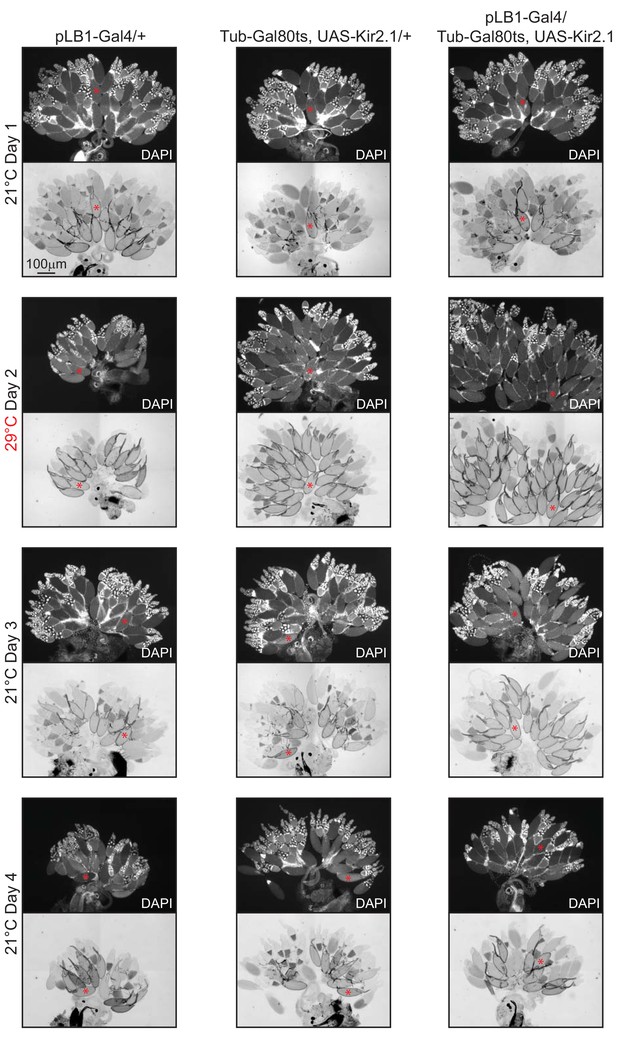
Conditional inactivation of pLB1+ neurons leads to a reversible accumulation of stage 14 oocytes.
Conditional inactivation of the pLB1+ neurons in pLB1-Gal4/Tub-Gal80ts, UAS-Kir2.1 flies triggers a reversible stage 14 oocytes accumulation. Ovaries from control flies (pLB1-Gal4/+ and Tub-G80ts, UAS-Kir2.1/+) and test animals (pLB1-Gal4/Tub-Gal80ts, UAS-Kir2.1) are seen with transmission light microscopy and DAPI staining over the course of 4 days and two different temperatures. Prototypical stage 14 oocyte is indicated with a red asterisk. Details including n values and genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
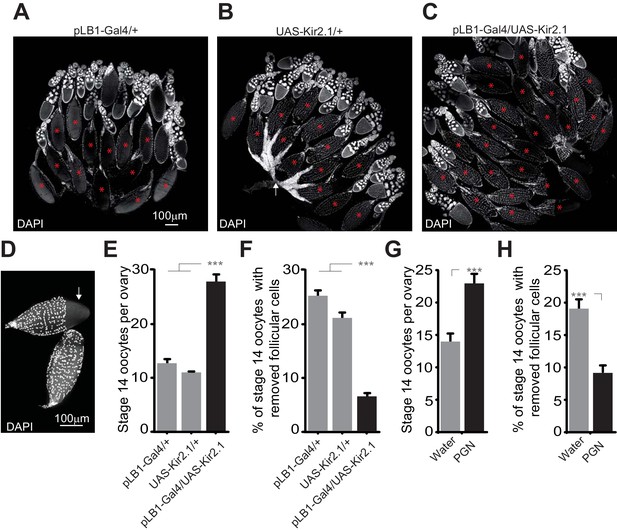
Impairing the activity of pLB1+ cells or injecting peptidoglycan leads to an accumulation of mature oocytes and a defect in follicular cells rupture.
(A–C); Reducing the activity of pLB1+ cells leads to an accumulation of mature oocytes (stage14). DAPI staining of ovaries from control flies (A and B) or animals with reduced activity of the pLB1+ cells (C). Mature oocytes are indicated with a red asterisk and an oocyte with follicular cells ruptured is indicated with a white arrow. (D); Follicular cells surrounding the mature oocytes are removed before the entry in lateral oviducts. DAPI staining of stage 14 oocytes with follicular cells partly removed (white arrow) or fully covering the oocyte (bottom). (E and G); Stage 14 oocytes accumulate in the ovaries when pLB1+ cells activity is impaired (E) or when peptidoglycan is injected (G). (F and H); The ratio of mature oocytes with removed follicular cells is decreased when pLB1+ cells activity is impaired (F) or when peptidoglycan is injected (H). For (E and G), shown is the average number of stage 14 oocytes per ovary ± SEM in 5 day-old females (E) or 6 hours post-treatment (G) from three independent trials with at least 50 ovaries per genotype and condition used. For (F and H), shown are the ratios of stage 14 oocytes with removed follicular cells ± SEM in 5 day-old females (F) or 6 hours post-treatment (H) from three independent trials with at least 50 ovaries per genotype and condition used. For (E and F), *** indicates p<0.0001; non-parametric ANOVA, Dunn’s multiple comparison test. For (G and H), *** indicates p<0.0001; non-parametric t-test, Mann-Whitney test. Details including n values and genotypes can be found in the detailed lines, conditions and, statistics for the figure section.
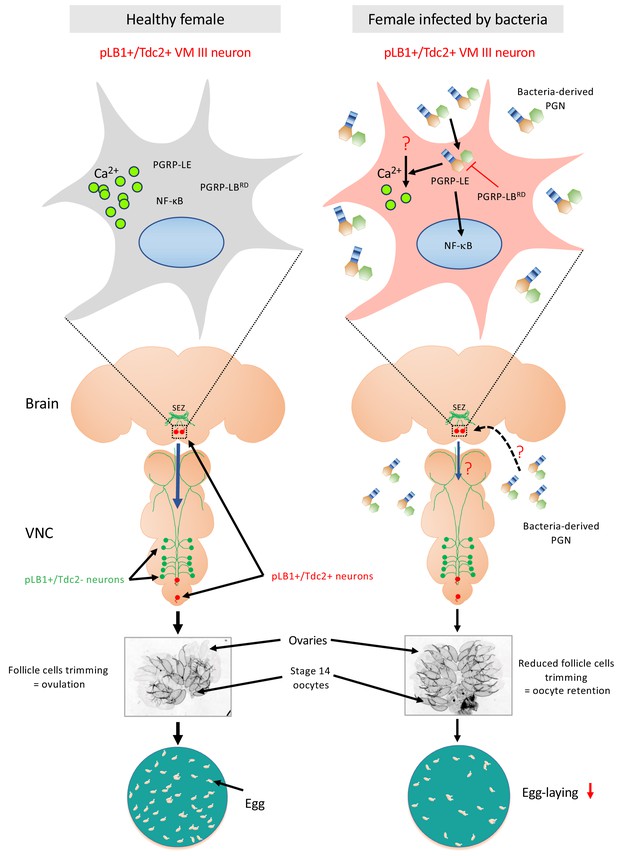
Diagram summarizing the effect of peptidoglycan sensing by pLB1+/Tdc2+ brain neurons on female oviposition.
(Left) Around 20 neurons distributed in the brain and the ventral nerve cord (VNC) express immune genes such as PGRP-LB (called pLB1+ neurons, labeled in green and red). Among them, very few are also expressing the enzyme Tdc2 indicating that they are octopaminergic neurons (labeled in red), with the most robust pLB1 expression for those localized in the brain ventral midline, the VM cluster (delineated by the box with dashed line and schematically magnified in the top box). In homeostasic conditions, mated females produce mature oocytes surrounded by follicular cells. Rupture of these follicular cells (trimming) leads to egg ovulation. (Right) Upon bacterial infection either systemic or enteric, cell wall peptidoglycan (PGN) is released by proliferating bacteria and transported into the hemolymph. Intracytosolic peptidoglycan sensing via the PGRP-LE Rc, in the very few brain pLB1+/Tdc2+ neurons (but not in the VNC pLB1+/Tdc2+ neurons), leads to cell-autonomous NF-κB pathway activation. This causes a reduction of follicular cells trimming in mature (stage 14) oocytes, hence a reduction of egg-laying in infected females. Since some pLB1+/Tdc2+ neurons of the VM cluster (VM III sub-cluster specifically) are sending descending projections toward the VNC and that the direct addition of peptidoglycan reduces their intracytosolic calcium level, we believe that peptidoglycan directly reduces the activity of these cells. Then, these brain pLB1+/Tdc2+ cells could mediate their effect via secondary neurons present in the VNC. It remains to be understood how PGN from the hemolymph is reaching brain neurons, how PGN stimulation reduces calcium levels, how this is linked to NF-κB pathway signaling and whether secondary neurons modulate ovulation. Moreover, we believe that pLB1+/Tdc2- neurons control other behaviors that are probably modulated by infection.
Videos
Effect of Ringer's solution stimulation on Tdc2>GCaMP6s VM II/III neurons in vivo.
GFP recording of an in vivo Tdc2>GCaMP6s fly brain in the VM II/III octopaminergic sub-clusters region. Brains of flies from which the head capsule has been removed were exposed to Ringer's solution. GFP signal was recorded every 500 ms.
Effect of peptidoglycan solution stimulation on Tdc2>GCaMP6sVM II/III neurons in vivo.
GFP recording of an in vivo Tdc2>GCaMP6s fly brain in the VM II/III octopaminergic sub-clusters region. Brains of flies from which the head capsule has been removed were exposed to peptidoglycan solution. GFP signal was recorded every 500 ms.
Effect of Ringer's solution stimulation on pLB1>FLP/Tub>Gal80>,Tdc2>GCaMP6s VM III neurons ex vivo.
GFP recording of an ex vivo pLB1>FLP/Tub>Gal80>,Tdc2>GCaMP6s fly brain in the VM III octopaminergic sub-cluster region. Dissected brains were mounted in Ringer's solution and stimulated with the same control solution. GFP signal was recorded every 2 s.
Effect of peptidoglycan solution stimulation on pLB1>FLP/Tub>Gal80>,Tdc2>GCaMP6s VM III neurons ex vivo.
GFP recording of an ex vivo pLB1>FLP/Tub>Gal80>,Tdc2>GCaMP6s fly brain in the VM III octopaminergic cluster region. Dissected brains were mounted in Ringer's solution and stimulated with peptidoglycan solution. GFP signal was recorded every 2 s.
Tables
Number and position of GFP-positive cells in the CNS of nSyb>FLP/pLB1>stop>mGFP flies.
The cells positive for GFP are counted.
| Organ | pLB1+ neurons | N events/total flies |
|---|---|---|
| Brain | 0 1 in the SEZ 2-5 in the SEZ 6-10 in the SEZ | 3/20 12/20 3/20 2/20 |
| VNC | 8-14 in T3 and AbdG | 13/13 |
Number and position of pLB1+/Tdc2+ neurons.
(A); pLB1-Gal4, UAS>stop>GFPmCD8; LexAop-FLP/nSyb-LexA brains and ventral nerve cords (VNCs) stained with an anti-Tdc2 Ab. The cells positive for GFP and stained with the anti-Tdc2 Ab (pLB1+/Tdc2+) are counted (left). The cells positive for GFP and negative for the Tdc2 Ab (pLB1+/Tdc2-) are counted. (B); pLB1-Gal4, UAS>stop>GFPmCD8; LexAop-FLP/Tdc2-LexA brains and VNCs stained with an anti-Tdc2 Ab. The GFP+ cells (pLB1+/Tdc2+) also positive for Tdc2 Ab are counted (left). NR = non relevant. This intersectional strategy only reveals pLB1+/Tdc2+ cells.
| (A) Organ | pLB1+/Tdc2+ neurons | pLB1+/Tdc2- neurons | N events/total flies |
|---|---|---|---|
| (Strategy 1, see legend for details) | |||
| Brain | 0 1 in the VMV III cluster 1 in the VMV III cluster 2-5 in the VMV III cluster | 0 0 2-4 in the SEZ 0 | 1/18 6/18 3/18 8/18 |
| VNC | 0 1 in the AbdG and 2 in T3 | 8-14 in T3 and AbdG 8-14 in T3 and AbdG | 11/14 3/14 |
| (B) Organ | pLB1+/Tdc2+ neurons | pLB1+/Tdc2- neurons | N events/total flies |
| (Strategy 2, see legend for details) | |||
| Brain | 0 1 in the VMV III cluster 2 in the VMV III cluster | NR NR NR | 4/19 13/19 2/19 |
| VNC | 0 1 in the AbdG 2 in the AbdG | NR NR NR | 9/18 7/18 2/18 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (D. melanogaster) | pLB1-Gal4 | (Kurz et al., 2017) | ||
| Genetic reagent (D. melanogaster) | UAS-Kir2.1 | Bloomington Drosophila Stock Center (Hardie et al., 2001) | BDSC Cat# 6595, RRID:BDSC_6595 | |
| Genetic reagent (D. melanogaster) | UAS-TTx | Bloomington Drosophila Stock Center (Sweeney et al., 1995) | BDSC Cat# 28838, RRID:BDSC_28838 | |
| Genetic reagent (D. melanogaster) | UAS-TRPA1 | Bloomington Drosophila Stock Center | BDSC Cat# 26264, RRID:BDSC_26264 | |
| Genetic reagent (D. melanogaster) | UAS-Fadd-IR | (Khush et al., 2002) | ||
| Genetic reagent (D. melanogaster) | UAS> stop> GFPmCD8 | Bloomington Drosophila Stock Center | BDSC Cat# 30125, RRID:BDSC_30125 | |
| Genetic reagent (D. melanogaster) | nSyb-LexA | Bloomington Drosophila Stock Center | BDSC Cat# 51951, RRID:BDSC_51951 | |
| Genetic reagent (D. melanogaster) | Tdc2-LexA | Bloomington Drosophila Stock Center | BDSC Cat# 52242, RRID:BDSC_52242 | |
| Genetic reagent (D. melanogaster) | Tub>Gal80> | Bloomington Drosophila Stock Center | BDSC Cat# 38879, RRID:BDSC_38879 | |
| Genetic reagent (D. melanogaster) | LexAop-FLP | Bloomington Drosophila Stock Center | BDSC Cat# 55819, RRID:BDSC_55819 | |
| Genetic reagent (D. melanogaster) | 8XLexAop2-FLP | Bloomington Drosophila Stock Center | BDSC Cat# 55820, RRID:BDSC_55820 | |
| Genetic reagent (D. melanogaster) | UAS>stop > Kir2.1 | |||
| Genetic reagent (D. melanogaster) | UAS>stop> TRPA1 | Bloomington Drosophila Stock Center | BDSC Cat# 66871, RRID:BDSC_66871 | |
| Antibody | Rabbit polyclonal anti-Tdc2 | Abcam | Cat# ab128225, RRID:AB_11142389 | 1:1000 |
| Chemical compound | PGN fromE. coli | Invivogen | 14C14-MM | |
| Software, algorithm | Fiji | NIH | https://fiji.sc/ | |
| Software, algorithm | GraphPad Prism 6 | GraphPad | RRID:SCR_002798 |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.50559.073





