Intracellular cholesterol trafficking is dependent upon NPC2 interaction with lysobisphosphatidic acid
Figures
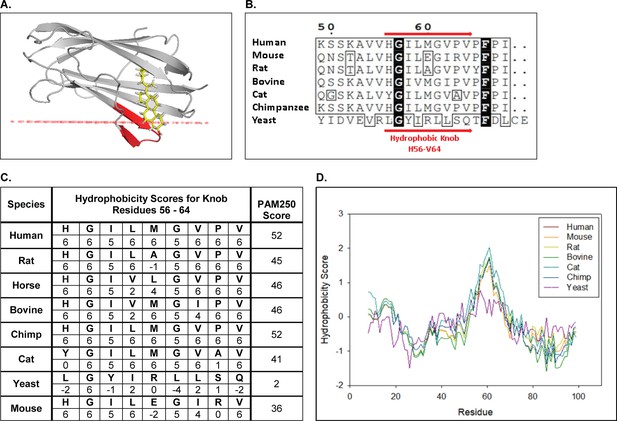
OPM predicts NPC2 stably inserts into membranes via a conserved hydrophobic knob region.
(A) Holo bovine NPC2 (PDB ID: 2HKA) is predicted by OPM to insert its prominent hydrophobic knob region (red ribbon) into the hydrophobic space of a model membrane, positioning the cholesterol in the sterol binding pocket in close proximity to the membrane surface (red line). (B) Multiple sequence alignment of human NPC2 (NCBI Accession: NP_006423.1), rat NPC2 (NP_775141.2) mouse NPC2 (NCBI Accession: NP_075898.1), bovine NPC2 (NCBI Accession: NP_776343.1), cat NPC2 (XP_003987882.1), chimpanzee NPC2 (NP_001009075.1) and the yeast NPC2 (NCBI Accession: KZV12184.1) were aligned with CLUSTAL Omega, and alignment for hydrophobic knob residues H56 to V64 are shown. Consensus sequences are in black and conserved residues are boxed. (C) Conservation scores and hydrophobicity scores for the hydrophobic knob, residues H56 to V64, were calculated based on the PAM250 scoring matrix and Kyte and Doolitle Hydrophobicity scale (Kyte and Doolittle, 1982). (D) The aligned NPC2 protein sequences were analyzed with the ProtScale Tool on ExPASy server based on the Kyte and Doolittle Amino acid Hydropathicity scale with a frame window of 15 residues (Gasteiger et al., 2005).
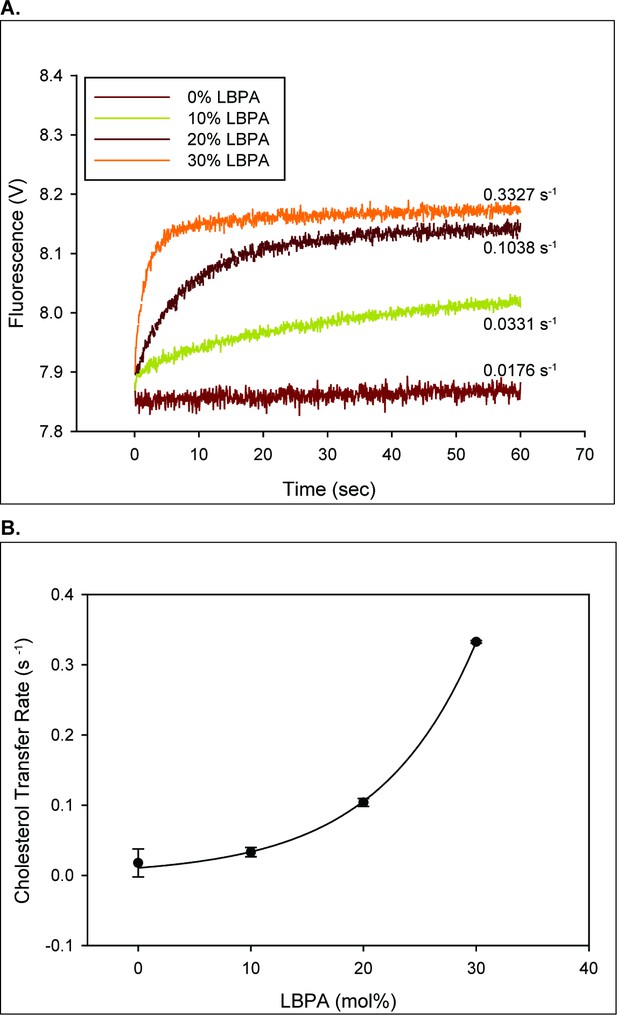
LBPA dramatically increase the rate of NPC2 mediated cholesterol transport.
(A) 2.5 µM NPC2-cholesterol complex was mixed with 250 µM of small unilamellar vesicles containing increasing mole percentages of LBPA in an SX20 stopped flow spectrofluorimeter as described under Materials and methods. The dequenching of endogenous tryptophan fluorescence was used to monitor cholesterol transfer from NPC2 to membranes. (B) The transfer rates from NPC2 to small unilamellar vesicles with various mole percentages of LBPA were fitted to a reverse single exponential function in Sigmaplot with an R-squared value of 0.9944. Data are representative of three experiments, each consisting of 2–3 individual runs ± SE.
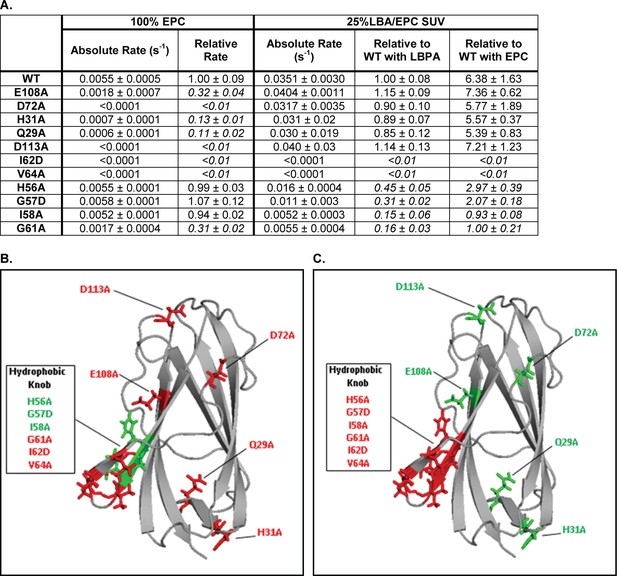
LBPA cannot reverse cholesterol transport deficiencies of NPC2 hydrophobic knob mutants.
(A) Transfer of cholesterol from 1 µM WT or mutant NPC2 to 125 µM 100% EPC or 25% LBPA/EPC vesicles was measured on an SX20 Stopped Flow Spectrofluorometer by monitoring the dequenching of NPC2 endogenous tryptophan fluorescence. All curves were well fit using a single exponential function using the Applied Photophysics Pro-Data Viewer software. Mutants with rates of cholesterol transfer less than 50% of WT NPC2 were considered to have defective transfer kinetics properties and their relative rates are italicized. Data are representative of 3 experiments, each consisting of 2–3 individual runs. Absolute and relative rates of transfer to each model membrane system, ± SE, are shown. (B) NPC2 point mutations resulting in defective cholesterol transport to 100% EPC vesicles are shown in red while mutations having little or no effect on NPC2 cholesterol transport properties relative to WT protein, are shown in green. (C) Point mutations with attenuated rates of cholesterol transport to 25% LBPA/EPC vesicles are shown in red while mutations having little or no effect, relative to WT protein, are shown in green.
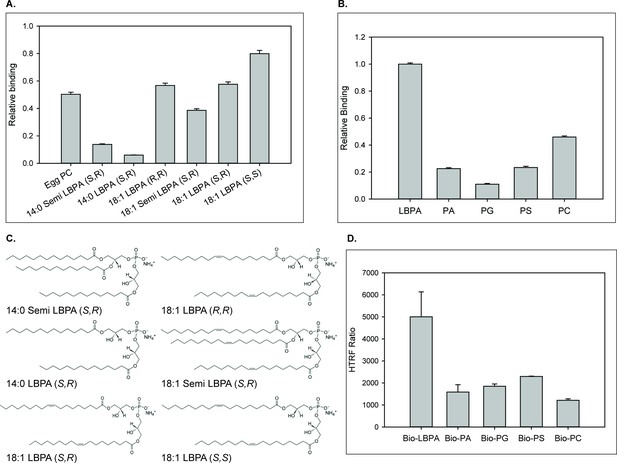
NPC2 binding to LBPA and other phospholipids.
(A) WT NPC2 protein was incubated with strips (Snoopers) containing LBPA isomers. LBPA-bound protein was detected with anti-c-myc antibody as described under Methods, and degree of binding ± SE (n = 3) is represented by the integrated density of the blots. (B) 500 pmol of various membrane phospholipids were spotted onto nitrocellulose strips and probed with WT NPC2-myc-his protein as described under Methods. Relative binding of WT NPC2 ± SE (n = 5) is shown, represented by signal intensity detected with the LI-COR system. (C) Structures of the LBPA isomers. (D) 75 nM of WT NPC2 protein was incubated with 1 μM of the indicated biotin-C12-ether phospholipid, streptavidin-d2 conjugate and europium cryptate–labeled monoclonal anti-histidine antibody in detection buffer, as described in Methods. FRET signal between europium cryptate and streptavidin was detected with a HTRF capable Envision plate reader (λex = 320 nm, λem = 615 and 665 nm; 100 μs delay time; n = 3).
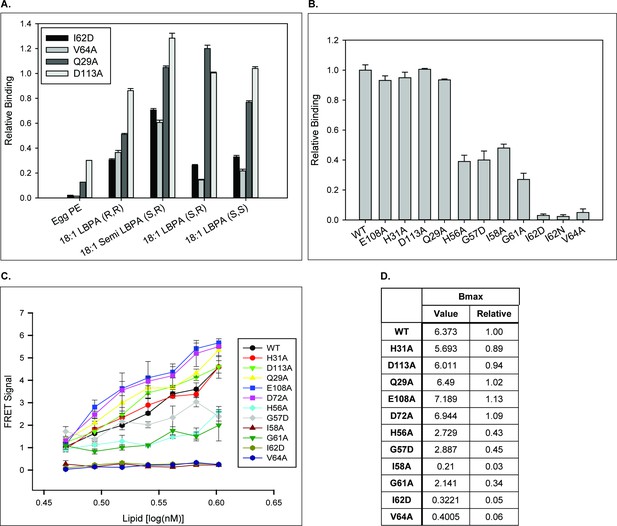
NPC2 binds to LBPA via the hydrophobic-knob domain.
Binding of NPC2 WT and mutant proteins to (A) LBPA isomers and (B), 18:1 (S,R) LBPA was detected using LBPA Snoopers and membranes spotted with 500 pmol phospholipid, respectively, as described under Methods. Relative binding is represented as (A) the integrated density of the blots, relative to WT NPC2, ± SE (n = 3) and (B) the signal intensity detected with the LI-COR system ± SE (n = 3) (C). HTRF analysis: WT or mutant NPC2 protein was incubated with biotin-C12-ether LBPA, streptavidin-d2 conjugate and europium cryptate–labeled anti-His antibody in detection buffer, as described in Methods. FRET signal between europium cryptate and streptavidin was detected with a HTRF capable Envision late reader as described in Methods. (D) FRET signal was analyzed with the One site–specific binding function (Graphpad) and the Bmax extrapolations were used to infer binding capacity between recombinant NPC2 protein and biotin-C12-ether LBPA. Results are representative of three experiments, with deviations < 20%.
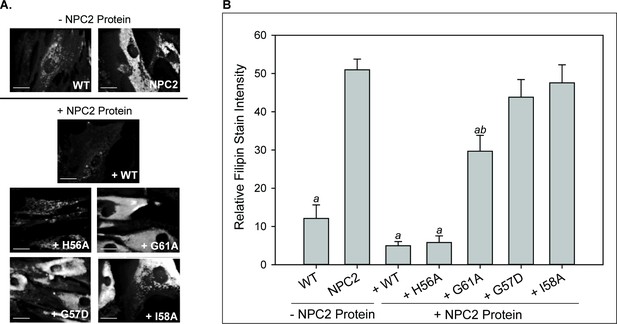
LBPA insensitive mutants are unable to rescue cholesterol accumulation in NPC2 patient fibroblasts.
NPC2-deficient fibroblasts were incubated with 0.4 nm purified WT or NPC2 mutant protein and cholesterol accumulation was quantified via filipin staining as described in the Methods. (A) Representative microscopy images of filipin stained control and treated NPC2-deficient fibroblasts. Scale bars, 70 µM. (B) Filipin stain intensity of untreated control and treated NPC2-deficient fibroblasts was quantified and are presented as percent of control. Data are representative of at least three separate experiments ± SE. a, p<0.01 vs untreated NPC2-deficient cells; b, p<0.01 vs untreated WT cells by Student’s t-test.
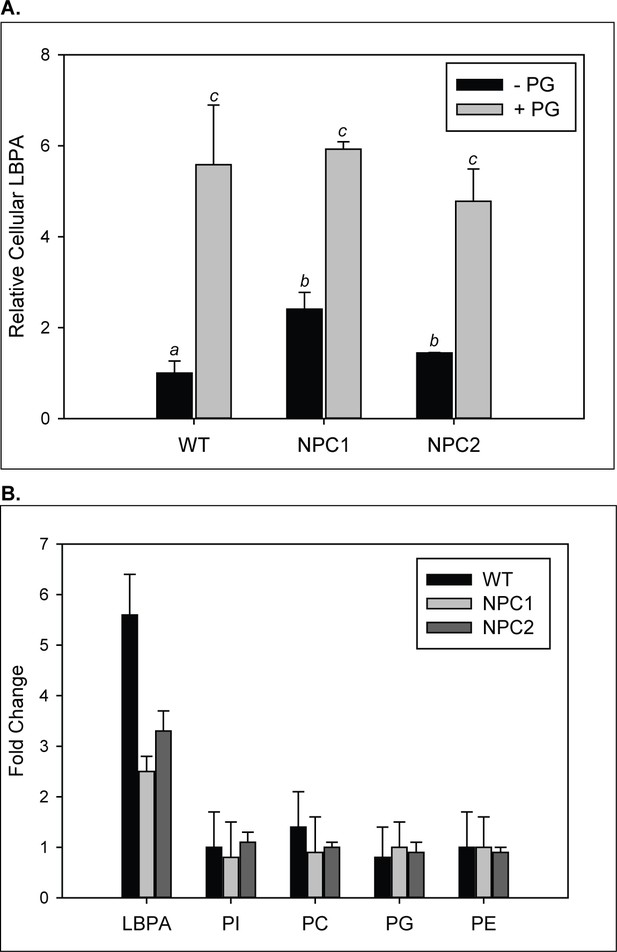
Increase in LBPA content in cells supplemented with PG.
WT, NPC1-, and NPC2 deficient fibroblasts were incubated with 100 µM PG SUVs. (A–B) Lipids were extracted and phospholipids quantified by TLC as described under Methods. Results are representative of two experiments, each conducted in duplicate, ± SE. (A) Data are normalized to LBPA levels in untreated WT cells. (B) Fold changes in PL species in PG-treated relative to untreated cells; *p<0.01 between PL species by one-way ANOVA.
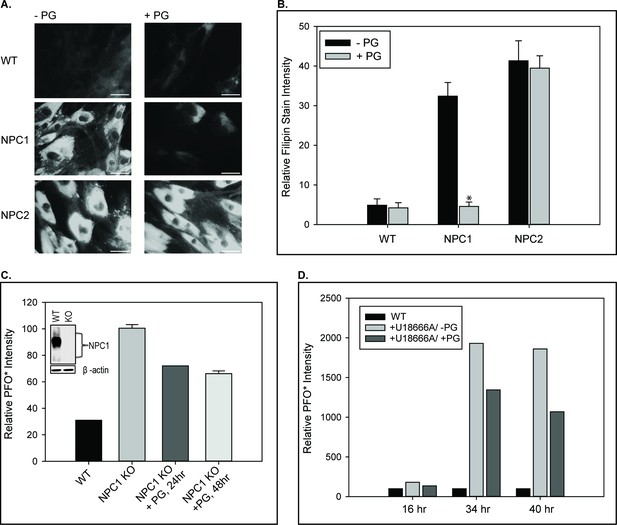
PG supplementation reverses cholesterol accumulation in NPC1– but not NPC2–deficient cells.
WT, NPC1–, and NPC2– deficient fibroblasts were incubated with 100 µM PG SUVs and cholesterol accumulation was determined by filipin staining as described under Methods. (A) Representative images of untreated and PG supplemented fibroblasts stained with filipin. Scale bars, 70 µM. (B) Percent filipin stain intensity, relative to control condition. Data are representative of three individual experiments, ± SE. *p<0.01 vs untreated cells by Student’s t-test. The effect of PG supplementation on cholesterol accumulation in NPC1-deficiency was also measured by PFO* intensity, assessed via flow cytometry, in (C) CRISPR-Cas9 mediated NPC1 knockout HeLa cells and (D) WT human fibroblasts treated with 1 µM U-18666A. Data are expressed relative to (C) untreated NPC1 knockouts and (D) untreated WT fibroblasts per incubation period.
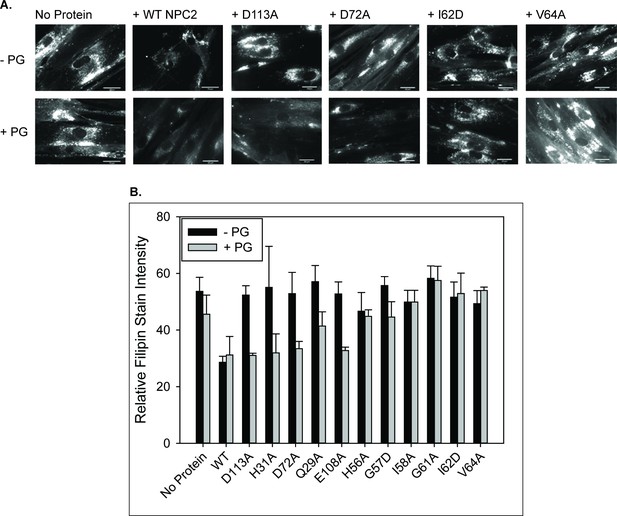
LBPA–sensitive but not insensitive NPC2 mutants reverse cholesterol accumulation in NPC2–deficient cells when co-treated with PG.
NPC2 deficient fibroblasts were incubated with purified WT or mutant NPC2 proteins alone or in the presence of PG SUVs, as described under Methods, and cholesterol accumulation was quantified by filipin staining. (A) Representative images of treated NPC2 deficient fibroblasts stained with filipin. Scale bars, 70 µM. (B) Data are from four or more individual incubations ± SE, *p<0.01 vs no PG supplementation by Student’s t-test.
Tables
Rate of NPC2-mediated membrane interaction greatly increases in the presence of LBPA.
The effect of surface residue mutations on the ability of NPC2 to induce vesicle-vesicle interactions was assessed by measuring absorbance at 350 nm (light scattering) of 200 µM LUVs in the presence of 1 µM WT or mutant NPC2 protein, as described under Materials and methods. Rates of vesicle-vesicle interactions, indicated by increases in A350nm over time, were determined by a three-parameter hyperbolic fit of the data using Sigma Plot software, and are representative of at least three individual experiments. Mutants with substantially attenuated rates of membrane aggregation are indicated in italics.
| 100% EPC | 25%LBA/EPC SUV | ||||
|---|---|---|---|---|---|
| Absolute rate (s1-) | Relative rate | Absolute rate (s−1) | Relative to WT with LBPA | Relative to WT with EPC | |
| WT | 0.0112 ± 0.0006 | 1.00 ± 0.05 | 0.1674 ± 0.0122 | 1.00 ± 0.07 | 14.95 ± 1.09 |
| H31A | 0.0018 ± 0.0005 | 0.16 ± 0.05 | 0.1262 ± 0.0079 | 0.75 ± 0.05 | 11.27 ± 0.71 |
| D113A | <0.0001 | <0.01 | 0.1573 ± 0.0112 | 0.94 ± 0.07 | 14.04 ± 1.00 |
| Q29A | 0.0029 ± 0.0008 | 0.26 ± 0.07 | 0.1337 ± 0.0039 | 0.80 ± 0.02 | 11.93 ± 0.35 |
| E108A | 0.0032 ± 0.0004 | 0.29 ± 0.03 | 0.1437 ± 0.0094 | 0.86 ± 0.06 | 12.83 ± 0.84 |
| D72A | 0.0032 ± 0.0003 | 0.29 ± 0.03 | 0.1456 ± 0.0077 | 0.87 ± 0.05 | 13.00 ± 0.69 |
| H56A | 0.0107 ± 0.0008 | 0.95 ± 0.07 | 0.0261 ± 0.0055 | 0.16 ± 0.09 | 2.33 ± 0.49 |
| G57D | 0.0103 ± 0.0016 | 0.92 ± 0.14 | 0.0202 ± 0.0066 | 0.13 ± 0.12 | 1.81 ± 0.59 |
| I58A | 0.0094 ± 0.0011 | 0.840 ± 0.10 | 0.0240 ± 0.0037 | 0.15 ± 0.06 | 2.14 ± 0.33 |
| G61A | 0.0075 ± 0.0006 | 0.67 ± 0.06 | 0.0406 ± 0.0012 | 0.25 ± 0.03 | 3.63 ± 0.11 |
| I62D | <0.0001 | <0.01 | 0.0167 ± 0.0017 | 0.10 ± 0.03 | 1.50 ± 0.15 |
| V64A | <0.0001 | <0.01 | 0.0193 ± 0.0040 | 0.12 ± 0.06 | 1.72 ± 0.36 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (C. griseus) | CHO-KI cells, NPC2-800#7 | PMID: 17018531 | Dr. Peter Lobel (Robert Wood Johnson Medical School) | |
| Cell line (Homo sapiens) | WT fibroblasts | Coriell Institute for Medical Research | Cat# GM03652; RRID:CVCL_7397 | Human skin fibroblasts from an apparently healthy 24 year old male |
| Cell line (Homo sapiens) | NPC2 fibroblasts | Coriell Institute for Medical Research | Cat# GM18455; RRID:CVCL_DA79 | Human skin fibroblasts from male identified as compound heterozygote at the NPC2 gene locus: results in E20X and C47F |
| Cell line (Homo sapiens) | NPC1 fibroblasts | Coriell Institute for Medical Research | Cat# GM03123; RRID:CVCL_7374 | Human skin fibroblasts from 9 year old female identified as compound heterozygote at the NPC1 gene locus: results in P237S and I1061T |
| Cell line (Homo sapiens) | HeLa (ATCCCCL-2) cells | ATCC | Cat # CCL-2; RRID:CVCL_0030 | Human epithelial cells from cervix of a 31 year old female with adenocarcinoma |
| Antibody | Rabbit polyclonal anti-c-myc-tag | GenScript | Cat# A00172-40; RRID:AB_914457 | IF (0.5 µg/ml) |
| Antibody | Donkey anti-rabbit IgG HRP-conjugated | GE Healthcare | Cat# NA934; RRID:AB_772206 | IF (1:20,000) |
| Antibody | Mouse monoclonal anti-myc- tag | Millipore | Cat# 05–724; RRID:AB_309938 | IF (1:2,000) |
| Antibody | Anti-mouse IgG IRDye-800CW conjugated | Li-Cor | Cat# 925–32210; RRID:AB_2687825 | IF (1:10,000) |
| Antibody | Streptavidin-d2 | Cisbio | Cat# 610SADLA | IF (50 µg/ml) |
| Antibody | Monoclonal anti-6His-Eu cryptate | Cisbio | Cat# 61HISKLA | IF (12.5 µg/ml) |
| Antibody | Rabbit monoclonal anti-NPC1 | Abcam | Cat# ab134113; RRID: AB_2734695 | WB (1:2,000) |
| Recombinant DNA reagent | Plasmid: myc 6xHis-tagged NPC2 | PMID: 12591949 | Dr. Matthew P. Scott (Stanford University) | |
| Recombinant DNA reagent | Plasmid: NPC1 CRISPR/Cas9 KO | Santa Cruz | sc-403252 | |
| Recombinant DNA reagent | Plasmid: mutant 125I-perfringolysin O (PFO*) | PMID: 23754385 | Dr. Arun Radhakrishnan (UT Southwestern) | |
| Sequence-based reagent | H31A mutant NPC2 primer: Forward, CCCACCGATCCC TGTCAGCTGGCCAAAGG; Reverse, CCTTTGGCCAGC TGAGGGATCGGTGGG | Sigma | ||
| Sequence-based reagent | D113A mutant NPC2 primer: Forward, GTGGTGGAATG GAAACTTGAAGCTGACAAAAAG; Reverse, CTTTTTGTC AGCTTCAAGTTTCCATTCCACCAC | Sigma | ||
| Sequence-based reagent | Q29A mutant NPC2 primer: Forward, CCCACCGATCCC TGTGCGCTGCACAAAGGCCAG; Reverse, CTGGCCTTT GTGCAGCGCACAGGGATCGGTGGG | Sigma | ||
| Sequence-based reagent | E108A mutant NPC2 primer: Forward, CTGGTGGTGGCA TGGAAACTTGAACTTGAAG; Reverse,CTTCAAGTTCAA GTTTCCATGCCACCACCAG | Sigma | ||
| Sequence-based reagent | D72A mutant NPC2 primer: Forward, CCCATTCCTGAG CCTGATGGTTGTAAGAGTGGAATTAAC; Reverse, GTT AATTCCACTCTTACAACCCGCAGGCTCAGGAATGGG | Sigma | ||
| Sequence-based reagent | H56A mutant NPC2 primer: Forward,GCCTTGGTCGC CGGCATCCTGGAAGGG; Reverse, CCCTTCCAGGAT GCCGGCGACCAAGGC | Sigma | ||
| Sequence-based reagent | G57D mutant NPC2 primer: Forward, CGGCCTTGGTCC ACGACATCCTGG; Reverse, CCAGGATGTCGTGGACCAAGGCCG | Sigma | ||
| Sequence-based reagent | I58A mutant NPC2 primer: Forward, GCCTTGGTCCAC GGCGCACTGGAAGGGATCC; Reverse, GGATCCCTTC CAGTGCGCCGTGGACCAAGGC | Sigma | ||
| Sequence-based reagent | G61A mutant NPC2 primer: Forward, GCATCCTGGAAG CGATCCGGGTCCC; Reverse, GGGACCCGGATC GCTTCCAGGATGC | Sigma | ||
| Sequence-based reagent | I62D mutant NPC2 primer: Forward, GCATCCTGGAAG GGGACCGGGTCCCCTTCC; Reverse, GGAAGGGGACC CGGTCCCCTTCCAGGATGC | Sigma | ||
| Sequence-based reagent | V64A mutant NPC2 primer: Forward, GGAAGGGATCCG GGCCCCCTTCCCTATTCC; Reverse, GGAATAGGGAAG GGGGCCCGGATCCCTTCC | Sigma | ||
| Chemical compound, drug | Cholesterol,>99% | Sigma Aldrich | Cat# C8667; CAS 57-88-5 | |
| Chemical compound, drug | Egg phosphatidylcholine (EPC) | Avanti Polar Lipids | Cat# 840051; CAS 97281-44-2 | |
| Chemical compound, drug | 18:1 Bismonoacylglycerol phosphate (BMP, aka LBPA) S,R isomer | Avanti Polar Lipids | Cat# 857133; CAS 799268-67-0 | |
| Chemical compound, drug | 18:1 Phosphatidic acid (PA) | Avanti Polar Lipids | Cat# 840875; CAS 108392-02-5 | |
| Chemical compound, drug | 18:1 Phosphatidylglycerol (PG) | Avanti Polar Lipids | Cat# 840475; CAS 67254-28-8 | |
| Chemical compound, drug | 18:1 Phosphatidylserine (PS) | Avanti Polar Lipids | Cat# 840035; CAS 90693-88-2 | |
| Chemical compound, drug | 18:1-12:0 Biotin PS | Avanti Polar Lipids | Cat# 860560; CAS 799812-66-1 | |
| Chemical compound, drug | 18:1-12:0 Biotin PA | Avanti Polar Lipids | Cat# 860561 | |
| Chemical compound, drug | 18:1-12:0 Biotin PG | Avanti Polar Lipids | Cat# 860581 | |
| Chemical compound, drug | 18:1-12:0 Biotin PC | Avanti Polar Lipids | Cat# 860563 | |
| Chemical compound, drug | Biotin-C12-ether LBPA | Echelon Biosciences | Cat# L-B1B12 | |
| Chemical compound, drug | Filipin III | Fisher Scientific | Cat# 62501NB; CAS 480-49-9 | Used at 0.05 mg/ml |
| Chemical compound, drug | Lipofectamine 3000 | Invitrogen | Cat# L3000- | |
| Commercial assay or kit | Stratagene QuickChange II Site Directed Mutagenesis Kit | Agilent | Cat# 200523 | |
| Commercial assay or kit | PureYield Plasmid Miniprep System | Promega | Cat# A1223 | |
| Software, algorithm | Orientation of Proteins in Membranes (OPM) | PMID: 16397007 | http://opm.phar.umich.edu/ RRID:SCR_011961 | |
| Software, algorithm | CLUSTAL Omega | PMID: 21988835 | http://www.ebi.ac.uk/Tools/msa/clustalo/ RRID:SCR_001591 | |
| Software, algorithm | PAM250 scoring matrix | PMID: 24509512 | ||
| Software, algorithm | Kyte and Doolittle Hydropathicity scale | PMID: 7108955 | ||
| Software, algorithm | ProtScale Tool | Gasteiger et al., 2005 | https://web.expasy.org/protscale/ | |
| Software, algorithm | ProData SX software, v2.5.0 | Applied Photophysics | https://www.photophysics.com | |
| Software, algorithm | NIS Elements BR software, v3.2 | Nikon Inc | https://www.nikoninstruments.com/Products/Software/NIS-Elements-Basic-Research RRID:SCR_014329 |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.50832.013





