Cycles of autoubiquitination and deubiquitination regulate the ERAD ubiquitin ligase Hrd1
Figures
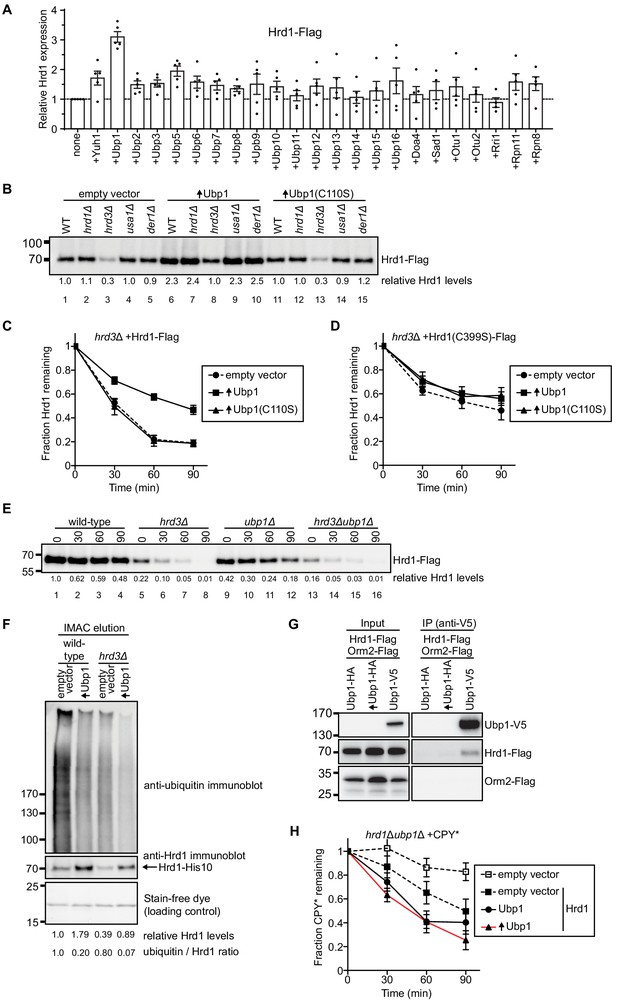
Ubp1 stabilizes Hrd1 through deubiquitination.
(A) Hrd1-Flag was expressed from a centromeric plasmid under its endogenous promoter in hrd1Δ cells together with individual deubiquitinating enzymes (DUBs). The DUBs were N-terminally tagged with hemagglutinin (HA) and overexpressed from a centromeric plasmid under the GPD-promoter. Hrd1 levels were analyzed by SDS-PAGE and immunoblotting for the Flag epitope. The data are the mean + /- SEM from at least three experiments. (B) Comparison of Hrd1-Flag levels in different strains. Where indicated, wild-type Ubp1 or an enzymatically inactive mutant (Ubp1(C110S)) were overexpressed (indicated by upwards-pointing arrows). The numbers below the lanes give quantification of Hrd1-FLAG levels relative to wild-type cells containing an empty vector. This panel is representative of at least three experiments. (C) The degradation of Hrd1-Flag was followed in a hrd3Δ strain overexpressing the deubiquitinating enzyme Ubp1 or a catalytically-inactive mutant (Ubp1(C110S)). Cycloheximide was added to the cell culture at time point zero, and samples were taken at different time points and analyzed by SDS-PAGE and immunoblotting with anti-Flag antibodies. The data are the mean + /- SEM from at least three experiments. (D) As in (C) but following the degradation of a catalytically-inactive Hrd1 mutant (Hrd1(C399S)-Flag). (E) The degradation of Hrd1-Flag was monitored following addition of cycloheximide in the indicated strains. This panel is representative of at least three experiments. (F) Hrd1-His10 was expressed from the endogenous Hrd1 promoter on a centromeric plasmid in either wild-type or hrd3Δ cells together with Ubp1 as in (C). The membrane proteins were purified with IMAC beads under denaturing conditions, separated by SDS-PAGE, and immunoblotted with anti-Hrd1 and anti-ubiquitin antibodies. The stain-free dye panel shows a non-specific protein band that co-eluted from the IMAC beads and serves to demonstrate equal loading of the samples. This panel is representative of at least three experiments. (G) Hrd1-Flag, Orm2-Flag, and Ubp1-V5 were expressed from their endogenous promoters on centromeric plasmids in ubp1Δ cells. The lysates were immunoprecipitated with anti-V5 affinity resin, separated by SDS-PAGE and immunoblotted with anti-Flag and anti-V5 antibodies. This panel is representative of at least three experiments. (H) The degradation of CPY*-HA was monitored following addition of cycloheximide in the hrd1Δubp1Δ strain with Hrd1-Flag and Ubp1-V5 expressed from either its endogenous or GPD promoter on a centromeric plasmid. The data are the mean + /- SEM from at least three experiments. See also Figure 1—figure supplements 1, 2 and 3.
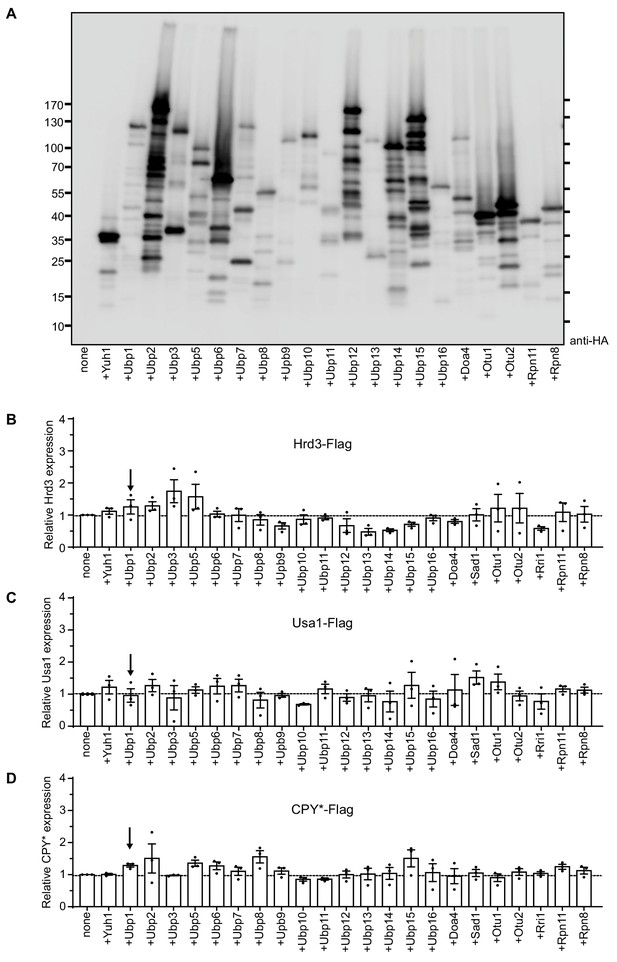
Levels of ERAD components in S. cerevisiae cells overexpressing various deubiquitinating enzymes.
(A) The steady-state levels of the HA-tagged deubiquitinating enzymes were analyzed by SDS-PAGE and immunoblotting for the HA epitope. Note that many of the deubiquitinating enzymes migrate slower than predicted from their molecular weights. (B) Hrd3-Flag was expressed from a centromeric plasmid under its endogenous promoter in hrd3Δ cells together with individual deubiquitinating enzymes (DUBs). The DUBs were N-terminally tagged with hemagglutinin (HA) and overexpressed from a centromeric plasmid under the GPD-promoter. Hrd3 levels were analyzed by SDS-PAGE and immunoblotting for the Flag epitope. The data are the mean + /- SEM from at least three experiments. (C) As in (B), but following Usa1-Flag in usa1Δ cells. (D) As in (B), but following the ERAD substrate CPY*-Flag in wild-type cells.
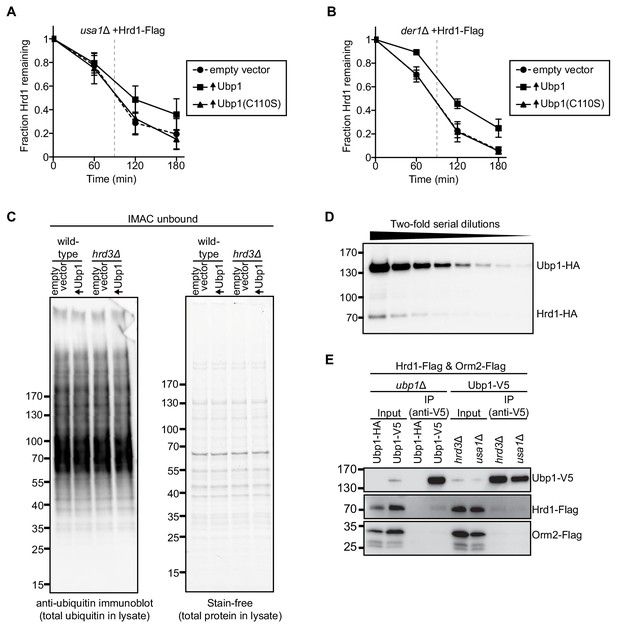
Hrd1 is stabilized by Ubp1 overexpression.
(A) The degradation of Hrd1-Flag was followed by cycloheximide-chase experiments in usa1Δ cells overexpressing the deubiquitinating enzyme Ubp1 or a catalytically-inactive Ubp1 mutant (Ubp1(C110S)). The data are the mean + /- SEM from at least three experiments. The vertical gray dashed line indicates the 90 min end-point used for CPY* chases. (B) As in (A), but with der1Δ cells. (C) The samples used for Figure 1F were analyzed for their total ubiquitin levels and total protein. The membrane fractions were solubilized in detergent and material not bound to IMAC beads was analyzed by SDS-PAGE and blotting with anti-ubiquitin antibodies (left) or by total protein visualization with stain-free dye technology (right). (D) Two-fold serial dilutions of whole cell lysate from wild-type cells expressing Hrd1-HA and Ubp1-HA from their endogenous promoters on centromeric plasmids. This panel is representative of at least three experiments. (E) Hrd1-Flag, Orm2-Flag, and Ubp1-V5 (or Ubp1-HA) were expressed from their endogenous promoters on centromeric plasmids in ubp1Δ, hrd3Δ, or usa1Δ cells. The lysates were immunoprecipitated with anti-V5 affinity resin, separated by SDS-PAGE and immunoblotted with anti-Flag and anti-V5 antibodies. This panel is representative of at least three experiments.
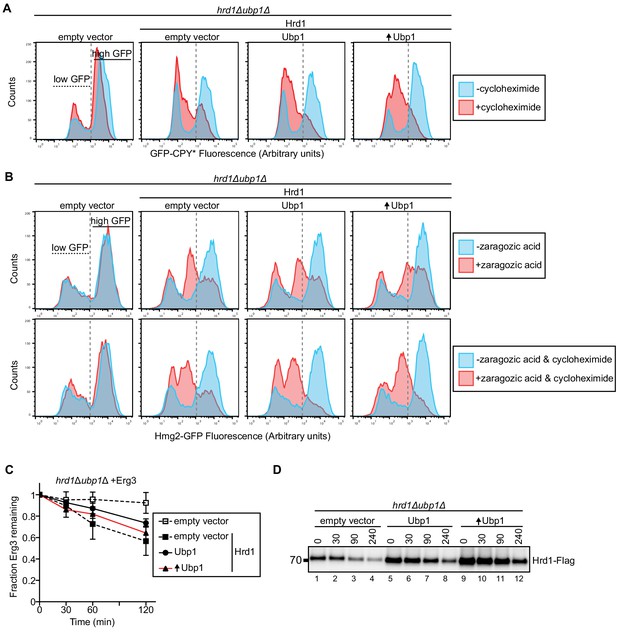
Ubp1 manipulates Hrd1 activity.
(A) GFP-CPY* was expressed from a centromeric plasmid under the GPD promoter in a hrd1Δubp1Δ strain. GFP-CPY* degradation was followed in cycloheximide-chase experiments. The indicated additional proteins were expressed from centromeric plasmids under their endogenous promoters or GPD promoter, indicated by upward arrow. Samples were analyzed by flow cytometry after 4 hr of treatment. The number of cells expressing low levels of GFP (horizontal dashed line) does not change during cycloheximide treatment, whereas those highly expressing GFP-CPY* changes (horizontal solid line). This panel is representative of at least three experiments. (B) As in (A) but with a centromeric plasmid containing Hmg2-GFP driven by the GPD promoter. Degradation was stimulated using zaragozic acid or combination of zaragozic acid and cycloheximide. This panel is representative of at least three experiments. (C) The degradation of Erg3-Flag was followed by cycloheximide chase in a hrd1Δubp1Δ strain containing the indicated proteins under endogenous promoter or GPD promoter, indicated by upward arrow. The data are the mean + /- SEM from at least three experiments. (D) As in (C) but following degradation of Hrd1-Flag in hrd1Δubp1Δ cells expressing CPY*-HA with the indicated proteins. This panel is representative of at least three experiments.
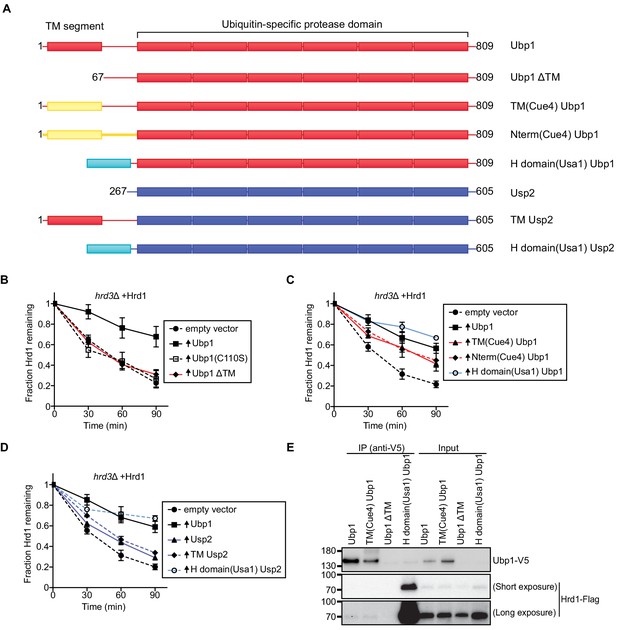
Ubp1 function requires its anchoring to the ER membrane.
(A) Schematic of chimeric deubiquitinating enzyme constructs used in (B)-(D). TM, transmembrane. Red segments are from Ubp1, yellow ones from Cue4, cyan ones from Usa1, and blue ones from Usp2. The H domain comprises amino acids 437–490 of Usa1. (B) The degradation of Hrd1-Flag was followed in a hrd3Δ strain overexpressing the indicated Ubp1 constructs. Cycloheximide was added to the cell culture at time point zero, and samples were taken at the indicated time points and analyzed by SDS-PAGE and immunoblotting with anti-Flag antibodies. The data are the mean + /- SEM from at least three experiments. (C) As in (B), but with other Ubp1 chimeras and wild-type Ubp1 as control. (D) As in (B) with Usp2 chimeras and wild-type Ubp1 as control. (E) Hrd1-Flag and Ubp1-V5 variants were expressed from their endogenous promoters on centromeric plasmids in hrd1Δubp1Δ cells. The lysates were immunoprecipitated with anti-V5 affinity resin, separated by SDS-PAGE and immunoblotted with anti-Flag and anti-V5 antibodies. This panel is representative of at least three experiments. See also Figure 2—figure supplement 1.
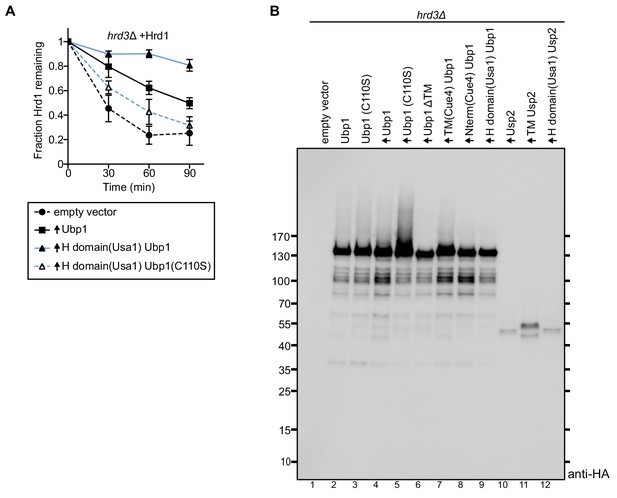
Steady-state levels of overexpressed proteins in S. cerevisiae cells.
(A) The degradation of Hrd1-Flag was followed in the indicated strains by cycloheximide-chase experiments. The data are the mean + /- SEM from at least three experiments. (B) The steady-state levels of C-terminally HA-tagged DUB constructs were analyzed by SDS-PAGE and immunoblotting for the HA epitope. The overexpressed constructs shown in lanes 4–12 were loaded at 10% compared to lanes 1–3.
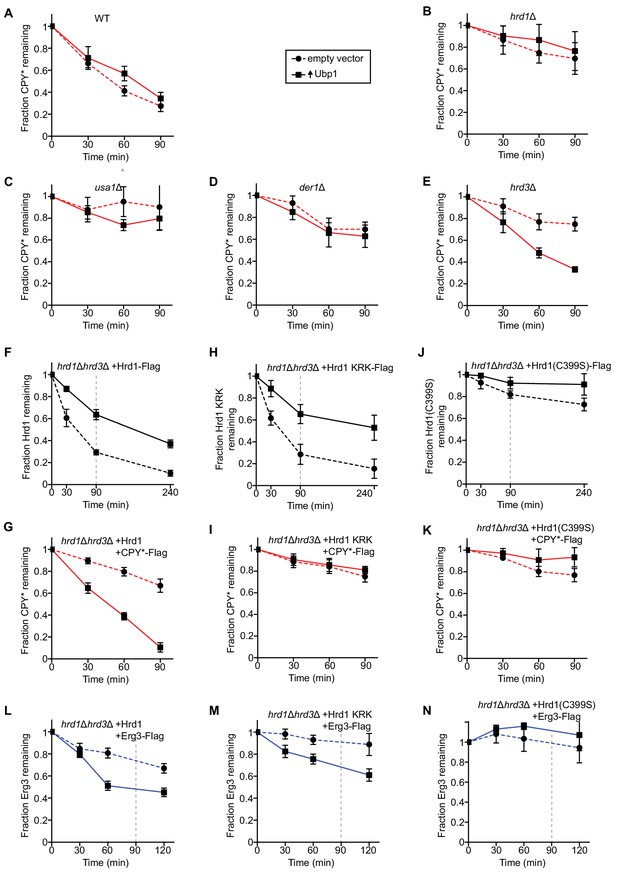
Ubp1 overexpression bypasses the requirement for Hrd3 in ERAD.
(A) The degradation of the ERAD-L substrate CPY*-Flag was tested in wild-type cells with or without overexpression of the deubiquitinating enzyme Ubp1. Cycloheximide was added to the cell culture at time point zero, and samples were taken at the indicated time points and analyzed by SDS-PAGE and immunoblotting with anti-Flag antibodies. The data are the mean + /- SEM from at least three experiments. (B) As in (A), but with hrd1Δ cells. (C) As in (A), but with usa1Δ cells. (D) As in (A), but with der1Δ cells. (E) As in (A), but with hrd3Δ cells. (F) The degradation of Hrd1-Flag was followed by cycloheximide-chase experiments in hrd1Δhrd3Δ cells with or without overexpression of the deubiquitinating enzyme Ubp1. The vertical gray dashed line indicates the 90 min end-point in CPY* chase experiments. (G) The degradation of CPY*-Flag was monitored by cycloheximide-chase experiments in hrd1Δhrd3Δ cells expressing Hrd1 from its endogenous promoter on a centromeric plasmid together with overexpressed Ubp1. (H) As in (F) but following the degradation of a Hrd1 variant that cannot be autoubiquitinated in its RING finger domain (Hrd1 KRK-Flag). (I) As in (G), but with Hrd1 KRK. (J) As in (F), but following the degradation of catalytically-inactive Hrd1 (Hrd1(C399S)-Flag). (K) As in (G), but with Hrd1(C399S). (L) The degradation of the ERAD-M substrate Erg3-Flag was monitored by cycloheximide-chase experiments in hrd1Δhrd3Δ cells expressing Hrd1 from its endogenous promoter on a centromeric plasmid together with overexpressed Ubp1. The vertical gray dashed line indicates the 90 min end-point used for CPY* chases. (M) As in (L), but with Hrd1 KRK. (N) As in (L), but with Hrd1(C399S).
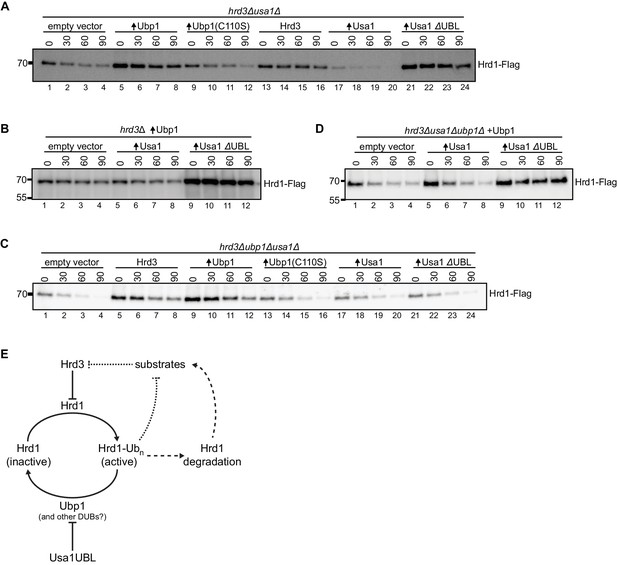
The UBL domain of Usa1 inhibits Ubp1 activity.
(A) Degradation of Hrd1-Flag in hrd3Δusa1Δ cells overexpressing the indicated proteins analyzed by cycloheximide-chase experiments. Ubp1(C110S) is an enzymatically-inactive version of Ubp1, and Usa1ΔUBL is Usa1 lacking its UBL domain. This panel is representative of at least three experiments. (B) Degradation of Hrd1-Flag in hrd3Δ cells overexpressing both Ubp1 and wild-type Usa1 or Usa1ΔUBL. This panel is representative of at least three experiments. (C) Degradation of Hrd1-Flag in hrd3Δubp1Δusa1Δ cells overexpressing the indicated proteins. This panel is representative of at least three experiments. (D) Degradation of Hrd1-Flag in hrd3Δubp1Δusa1Δ cells expressing Ubp1 and overexpressing the indicated proteins. This panel is representative of at least three experiments. (E) Model for the regulation of Hrd1 by autoubiquitination and deubiquitination. Hrd3 serves as an inhibitor of Hrd1 autoubiquitination. Binding of an ERAD substrate to Hrd3 would relieve this inhibition, resulting in Hrd1 activation and degradation of the substrate (dotted line). Hrd1 ubiquitination also leads to Hrd1 degradation, which counteracts substrate degradation (broken lines). Ubp1 reverses autoubiquitination of Hrd1 and returns Hrd1 to the inactive state. Ubp1 activity is attenuated by the UBL domain of the Hrd1-associated component Usa1. See also Figure 4—figure supplement 1.
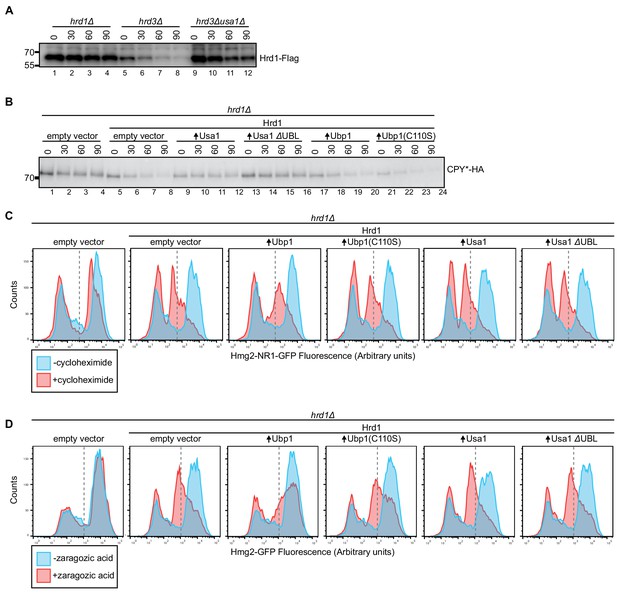
The Usa1 UBL domain influences Hrd1 stability.
(A) The degradation of Hrd1-Flag was followed in the indicated strains by cycloheximide-chase experiments. The data are the mean + /- SEM from at least three experiments. (B) The degradation of CPY*-HA was followed in a hrd1Δ strain containing the indicated proteins by cycloheximide-chase experiments. This panel is representative of at least three experiments. (C) Hmg2-non-responder-1-GFP was expressed from a centromeric plasmid under the GPD promoter in a hrd1Δ strain, and its degradation was followed after the addition of cycloheximide. The indicated additional proteins were expressed from centromeric plasmids under their endogenous promoters. Samples were analyzed by flow cytometry after 4 hr of treatment. The number of cells expressing low levels of GFP (dashed line) and does not change during cycloheximide treatment, whereas those highly expressing Hmg2-non-responder-1-GFP changes (solid line). This panel is representative of at least three experiments. (D) As in C but with Hmg2-GFP, using zaragozic acid to stimulate degradation. This panel is representative of at least three experiments.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Saccharomyces cerevisiae) | HRD1 | This study | YOL013C | Amplified from BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | HRD3 | This study | YLR207W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | USA1 | This study | YML029W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | DER1 | This study | YBR201W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | YUH1 | This study | YJR099W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP1 | This study | YDL122W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP2 | This study | YOR124C | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP3 | This study | YER151C | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP5 | This study | YER144C | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP6 | This study | YFR010W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP7 | This study | YIL156W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP8 | This study | YMR223W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP9 | This study | YER098W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP10 | This study | YNL186W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP11 | This study | YKR098C | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP12 | This study | YJL197W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP13 | This study | YBL067C | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP14 | This study | YBR058C | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP15 | This study | YMR304W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | UBP16 | This study | YPL072W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | DOA4 | This study | YDR069C | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | SAD1 | This study | YFR005C | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | OTU1 | This study | YFL044C | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | OTU2 | This study | YHL013C | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | RRI1 | This study | YDL216C | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | RPN11 | This study | YFR004W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | RPN8 | This study | YOR261C | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | ORM2 | This study | YLR350W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | ERG3 | This study | YLR056W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | HMG2 | This study | YLR450W | BY4741 (genomic DNA) |
| Gene (S. cerevisiae) | PRC1 (CPY) | This study | YMR297W | BY4741 (genomic DNA) |
| Gene (Homo sapiens) | Ubiquitin specific protease 2; Usp2 | This study | AAP36388.1 | Synthetic gene |
| Strain, strain background (S. cerevisiae) | BY4741 | GE Dharmacon | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 | |
| Strain, strain background (S. cerevisiae) | hrd1Δ | GE Dharmacon | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 hrd1::kanR | |
| Strain, strain background (S. cerevisiae) | hrd3Δ | GE Dharmacon | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 hrd3::kanR | |
| Strain, strain background (S. cerevisiae) | usa1Δ | GE Dharmacon | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 usa1::kanR | |
| Strain, strain background (S. cerevisiae) | der1Δ | GE Dharmacon | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 der1::kanR | |
| Strain, strain background (S. cerevisiae) | ubp1Δ | This study | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 ubp1::kanR | |
| Strain, strain background (S. cerevisiae) | yRB057A | This study | MATa his3Δ1 leu2Δ0 MET15 lys2Δ0 ura3Δ0 hrd1::kanR hrd3::kanR | |
| Strain, strain background (S. cerevisiae) | yBGP15A | This study | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 hrd1::kanR ubp1::kanR | |
| Strain, strain background (S. cerevisiae) | yRB0065A | This study | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 hrd3::kanR usa1::kanR | |
| Strain, strain background (S. cerevisiae) | yRB0126 | This study | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 hrd3::kanR ubp1::hphNT1 | |
| Strain, strain background (S. cerevisiae) | yRB0075C | This study | MAT? his3Δ1 leu2Δ0 MET15 ura3Δ0 hrd3::kanR usa1::kanR ubp1::hphNT1 | |
| Genetic reagent (S. cerevisiae) | CPY* | (Finger et al., 1993) | G255R point mutation in PRC1 (CPY) leading to CPY* | |
| Genetic reagent (S. cerevisiae) | GPD promoter | This study | YGR192C | 667 basepairs upstream of TDH3 (GPD) gene; BY4741 (genomic DNA) |
| Antibody | THE Anti-DYKDDDK Antibody (mAb, mouse) | Genscript | A00187 | 1:2000 (5% milk in TBST) |
| Antibody | Anti-HA High Affinity Antibody (mAb, clone 3F10, rat) | Roche | 11867423001 | 1:2000 (5% milk in TBST) |
| Antibody | THE V5 Tag Antibody(mAb, mouse) | Genscript | A01724 | 1:2500 (5% milk in TBST) |
| Antibody | Anti-PGK1 (mAb, clone22C5D8, mouse) | Invitrogen | 459250 | 1:1000 (5% milk in TBST) |
| Antibody | Anti-Ubiquitin(mAb, clone P4D1, mouse) | Santa Cruz Biotechnologies | sc-8017 | 1:500 (5% milk in TBST) |
| Antibody | Amersham ECL Rabbit IgG, HRP-linked wholeAb (donkey) | GE Healthcare | NA934 | 1:4000 (5% milk in TBST) |
| Antibody | Amersham ECL Mouse IgG, HRP-linked whole Ab (sheep) | GE Healthcare | NA931 | 1:4000 (5% milk in TBST) |
| Antibody | Amersham ECL Rat IgG, HRP-linked whole antibody (goat) | GE Healthcare | NA935 | 1:4000 (5% milk in TBST) |
| Commercial assay or kit | Western Lightning Plus-ECL, Enhanced Chemiluminescence Substrate | Perkin-Elmer | NEL103E001 | |
| Commercial assay or kit | Amersham ECL Select Western Blotting Detection Reagent | GE Healthcare | RPN2235 | |
| Chemical compound, drug | SYTOX Blue Nucleic Acid Stain | Invitrogen | S11348 | |
| Chemical compound, drug | Cycloheximide | Calbiochem | 239763 | |
| Chemical compound, drug | Decyl Maltose Neopentyl Glycol (DMNG) | Anatrace | NG322 | |
| Chemical compound, drug | Zaragozic Acid A | Cayman Chemical | 17452 | |
| Software, algorithm | Graphpad Prism 8 | Graphpad Software LLC. | version 8.3.0 | |
| Software, algorithm | FlowJo | Becton, Dickinson and Company | Version 10.6.0 | |
| Other | Dynabeads His-Tag Isolation and Pulldown | Invitrogen | 10104D | |
| Other | Anti-V5 Agarose Affinity Gel | Millipore | A7345 |
Additional files
-
Supplementary file 1
Yeast strains used in this study.
- https://cdn.elifesciences.org/articles/50903/elife-50903-supp1-v2.docx
-
Supplementary file 2
Plasmids used in this study.
- https://cdn.elifesciences.org/articles/50903/elife-50903-supp2-v2.docx
-
Supplementary file 3
1-way ANOVA and Tukey Honestly Significant Difference Test.
- https://cdn.elifesciences.org/articles/50903/elife-50903-supp3-v2.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/50903/elife-50903-transrepform-v2.pdf





