A molecular view on the escape of lipoplexed DNA from the endosome
Figures
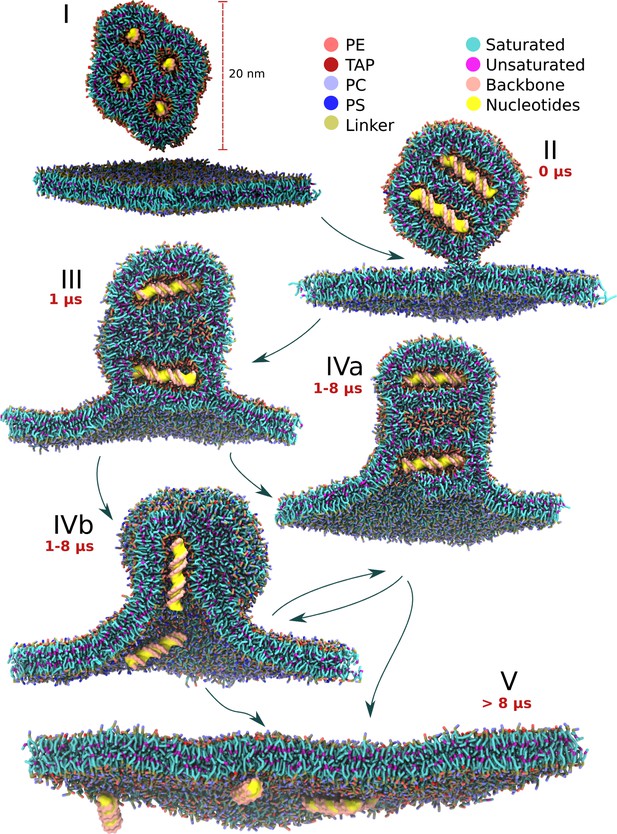
Schematic view of the lipoplex-membrane transfection pathways.
Unadhered lipoplex containing four dsDNAs sits above the endosomal bilayer (I). After initial stalk formation (II), a wide hemifusion diaphragm is formed (III). A pore is formed in one of the channels containing the DNA (IVa, IVb). The angle of the DNA with respect to the average bilayer normal can be either perpendicular or parallel, resulting in subsequently zipper (IVa) or ejection (IVb) like release of the DNA (V). The pathways indicated are based on 25 independent fusion experiments, each 10 µs long, leading to 34 dsDNA transfection events. The time scales indicated are typical for successful transfection events.
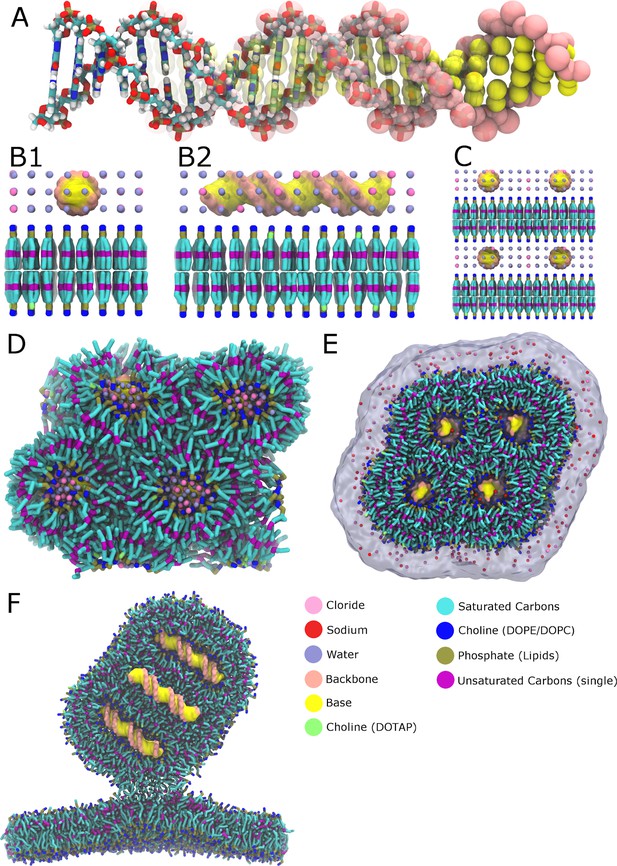
Building the small lipoplex.
First the AA geometry and force field of the dsDNA was translated into their CG counterpart (A). Then the dsDNA was stacked on top of a bilayer composed of DOTAP and DOPE in a 1:4 ratio respectively, and the system was solvated (B1/2). This DNA-bilayer complex was stacked in the z and x dimensions (C), and was simulated until a stable HII lipid phase formed (D). The naked complex was placed in a larger simulation box and lipids were added around the lipoplex, to allow for formation of the outer leaflet (E). The lipoplex, initially placed above the bilayer, diffused to the membrane and the fusion stalk was initiated (F). This figure was reproduced from Bonomi and Camilloni (2019).
© 2019 Springer Science+Business Media LLC. All rights reserved. Panel F is reproduced from Bonomi and Camilloni (2019) with permission. It is not covered by the CC-BY 4.0 licence and further reproduction of this panel would need permission from the copyright holder.
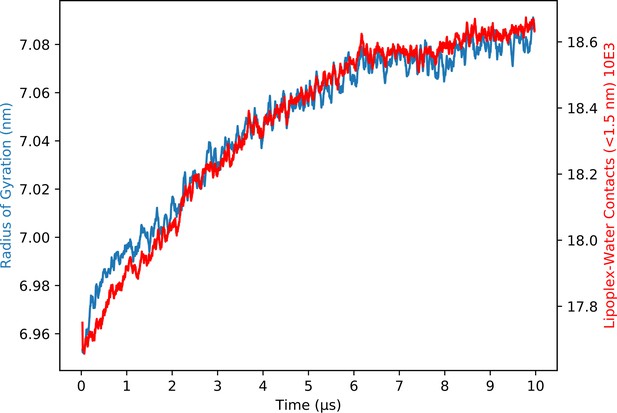
Radius of gyration for the solvated lipoplex.
The radius of gyration for the solvated small lipoplex over time. Over the last 2 µs of the simulation the radius of gyration increases less than a quarter of an Ångstrom. We therefore concluded that the lipoplex complexes are close enough to equilibrium to use for transfection experiments. The water contacts (<1.5 nm) with the lipoplex were calculated using the inhouse VMD counting script. The solvation of the lipoplex shows the same trend as the radius of gyration also confirming that the lipoplex is stable and can be used for transfection. Since the hydration and gyration show the same (minor) growth, we think that the observed growth in size of the lipoplex is due to a minor underhydration upon construction of the periodic lipoplex.
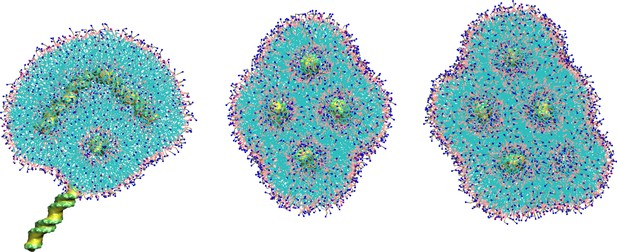
Coating the lipoplex.
Upon solvating the periodic lipoplex an extra layer of coating lipids was added. We used the approximate surface area of the lipoplex and divided it by the APL of the DOTAP:DOPE mixture (1:4; 0.63 nm2). From left to right we have 1/2, 1/1, 2/1 times the calculated amount. It is clear that underestimating the amount of coating lipids leads to loss of core lipids resulting in the loss of the internal structure and internalized dsDNA (left image). Oversaturating the core with coating lipids results in internalization of coating lipids, resulting in a new channel within the crystal lattice (right image). The expected amount results in no loss of internal structure of dsDNA, and no extra channels are formed (central image). Each condition was simulated for 10 µs after adding the coating.
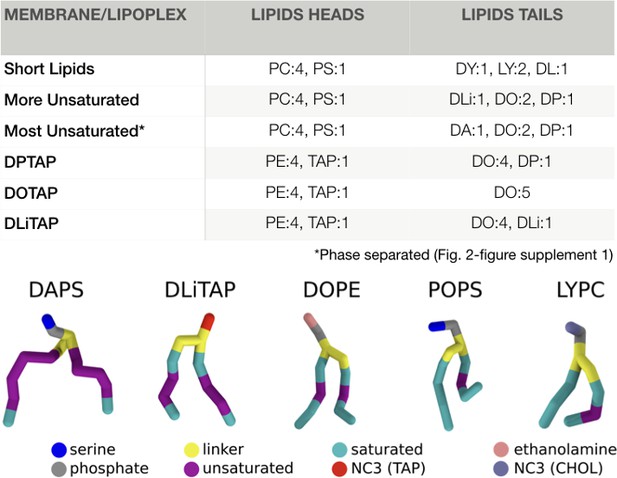
Membrane and lipoplex compositions.
The membrane compositions for the fusion experiments, and the lipoplex formulations are indicated in molar ratios. All PC/PS heads were combined with all tails, resulting in six different lipids per membrane composition (first 3). For the lipoplex formulations all PE was linked to DO and all TAP to either DP/DO/DLi (last 3). The nomenclature follows the default Martini lipids abbreviations as described by T.A. Wassenaar et al. (2015). An example of each tail and headgroup is displayed under the table (the tails are in the order of occurrence in their name). The complete nomenclature can also be found online at: ‘http://cgmartini.nl/index.php/force-field-parameters/lipids2/350-lipid-details’.

Lipid mixing in all endosome bilayer compositions.
Snapshots of endosome bilayers after 1 µs of equilibration after being generated by insane. From left to right we show the most, more unsaturated and short lipids bilayer respectively (see Figure 2). In purple the unsaturated carbons are displayed. Teal represents the saturated carbons. Only the most unsaturated bilayer shows phase separation (left, DA).
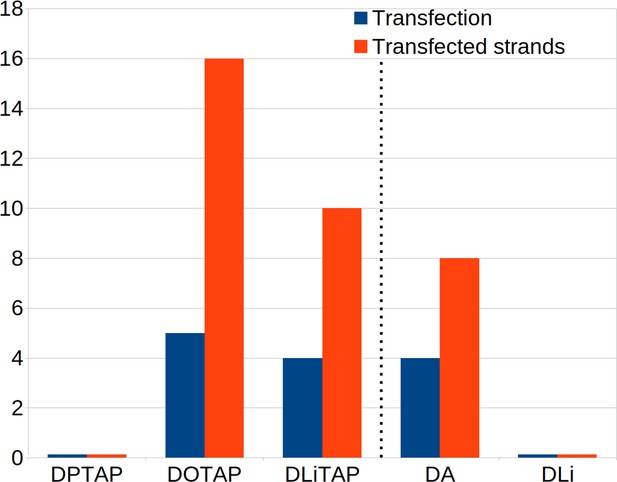
Effect of lipid composition on transfection efficiency.
The number of successful transfections and amount of transfected DNA considering different lipid compositions. Each composition had 5 repeats with four dsDNA fragments each. Therefore the maximum number of transfections is five and the maximum amount of transfected fragments is 20 per condition. Left of the dotted line are the fusion results for varying lipoplex formulations on a short lipid membrane. For these formulations the lipoplex formulation was 4:1 DOPE/DxTAP. On the right of the dotted line the results for varying endosomal membrane compositions with constant lipoplex formulation (4:1 DOPE/DOTAP) are displayed. The alternative endosomal membrane models are enriched in poly-unsaturated DA or di-unsaturated DLi lipids (Figure 2).
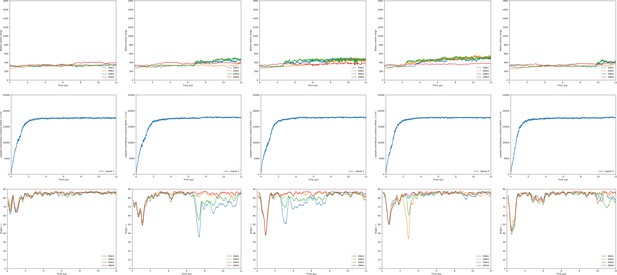
Hydration/transfection, lipid mixing and orientation of the dsDNA DOPE:DPTAP + short lipid endosome bilayer.
Shown are the observables for the lipoplex-membrane fusion experiments. Each experiment was performed five times (one repeat per column). For each simulation the first 10 µs were taken into account. The three rows in the figures show subsequently the hydration of the dsDNA with its fusion moments indicated as dotted lines; the lipoplex-membrane lipid mixing; and the angle of each dsDNA with respect to the bilayer. The first three groups of graphs are for varying lipoplex composition and the last two are for varying membrane composition.
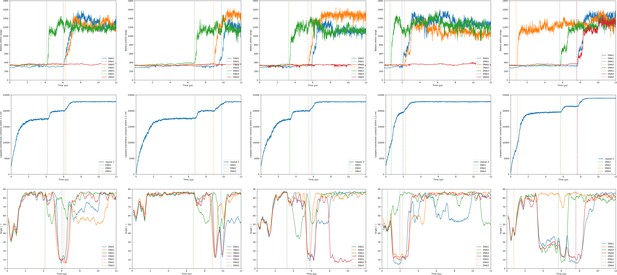
Hydration/transfection, lipid mixing and orientation of the DOPE:DOTAP + short lipid endosome bilayer.
Shown are the observables for the lipoplex-membrane fusion experiments. Each experiment was performed five times (one repeat per column). For each simulation the first 10 µs were taken into account. The three rows in the figures show subsequently the hydration of the dsDNA with its fusion moments indicated as dotted lines; the lipoplex-membrane lipid mixing; and the angle of each dsDNA with respect to the bilayer. The first three groups of graphs are for varying lipoplex composition and the last two are for varying membrane composition.
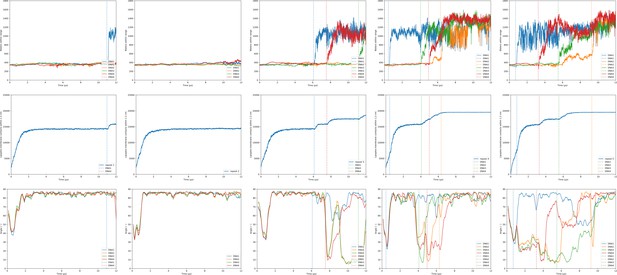
Hydration/transfection, lipid mixing and orientation of the DOPE:DLiTAP + short lipid endosome bilayer.
Shown are the observables for the lipoplex-membrane fusion experiments. Each experiment was performed five times (one repeat per column). For each simulation the first 10 µs were taken into account. The three rows in the figures show subsequently the hydration of the dsDNA with its fusion moments indicated as dotted lines; the lipoplex-membrane lipid mixing; and the angle of each dsDNA with respect to the bilayer. The first three groups of graphs are for varying lipoplex composition and the last two are for varying membrane composition.
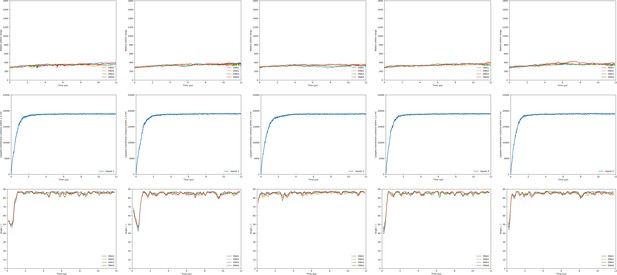
Hydration/transfection, lipid mixing and orientation of the DOPE:DOTAP + DLi containing endosome bilayer.
Shown are the observables for the lipoplex-membrane fusion experiments. Each experiment was performed five times (one repeat per column). For each simulation the first 10 µs were taken into account. The three rows in the figures show subsequently the hydration of the dsDNA with its fusion moments indicated as dotted lines; the lipoplex-membrane lipid mixing; and the angle of each dsDNA with respect to the bilayer. The first three groups of graphs are for varying lipoplex composition and the last two are for varying membrane composition.
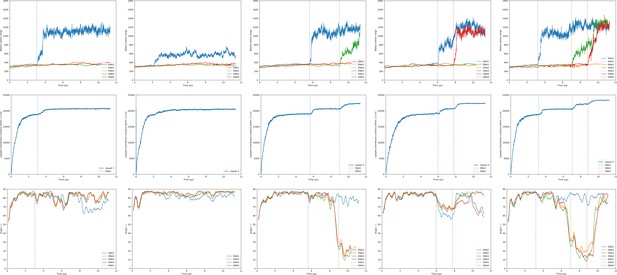
Hydration/transfection, lipid mixing and orientation of the DOPE:DOTAP + DA containing endosome bilayer.
Shown are the observables for the lipoplex-membrane fusion experiments. Each experiment was performed five times (one repeat per column). For each simulation the first 10 µs were taken into account. The three rows in the figures show subsequently the hydration of the dsDNA with its fusion moments indicated as dotted lines; the lipoplex-membrane lipid mixing; and the angle of each dsDNA with respect to the bilayer. The first three groups of graphs are for varying lipoplex composition and the last two are for varying membrane composition.
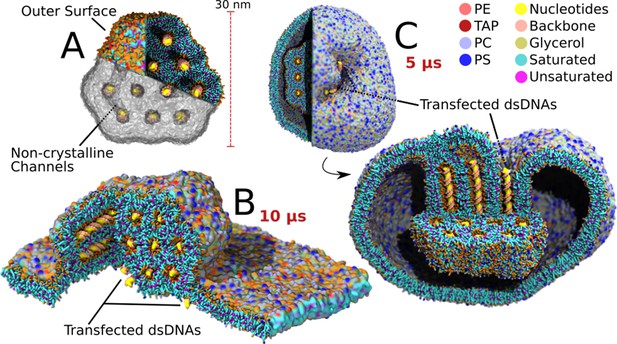
Transfection of large lipoplexes.
The large lipoplex (A) showed the same stable HII structure as the small lipoplex with dsDNA inside the aqueous channels. Small additional connecting channels are also present, as indicated. Transfection was performed on top of a large endosomal model bilayer patch (B) and from within a model endosomal vesicle (C). In contrast to the small lipoplex, fusion did not spontaneously occur and had to be initiated using a biasing potential on one of the channels. After release of the biasing potential the dsDNA inside the pulled channel, and all dsDNA connected through connecting channels, transfected (3 DNA fragments transfected in both lamellar and vesicular system upon pulling of the same initial channel).
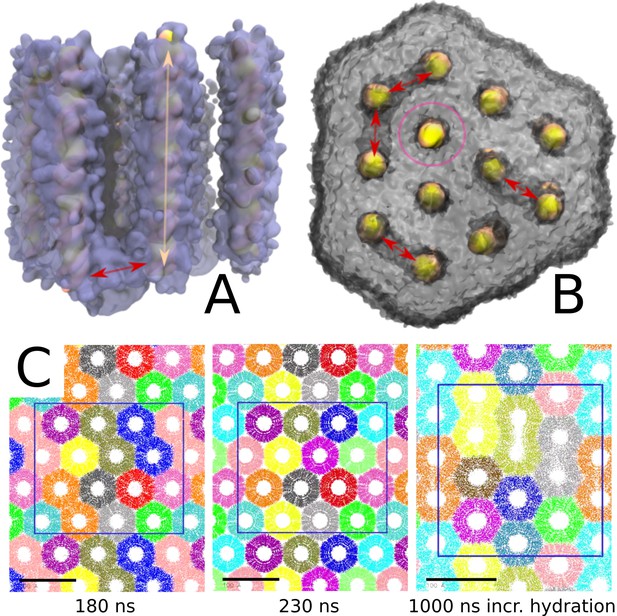
Large lipoplex channel structures with dsDNA inside.
The solvated large lipoplex contained channels: regular dsDNA/aqueous channels are indicated in peach (A); an open channel is depicted with a magenta circle (B); and aqueous connecting channels are indicated with red arrows (A/B). The water is drawn in ice-blue (A) and the lipid density in grey (B). The connecting channels were mainly perpendicular to the HII channels and positioned at the end point(s) of the dsDNA (A). All HII channels but two were closed on both ends by lipid density. One channel remained open on both sides over a period of 10 µs, with the dsDNA in the open channel positioned in the lipoplex in the same manner as the dsDNA in the closed channels. Apparently opening a channel is not enough for dsDNA release from the complex. The bottom panel shows, from left to right, the channel connectivity for the large periodic phase under standard, elongated, and extra hydrated conditions (C). The spacing of the HII phase is already finished at 180 ns (the used equilibration time), however, some connective channels still exist, indicated by neighboring channels having the same color. These connections could be closed by equilibrating slightly longer (230 ns in total). To inspect if these channels could be stabilized, we increased the hydration (lipids:watersAA 1:12). This indeed resulted in connective channels being present until the end of the simulation.
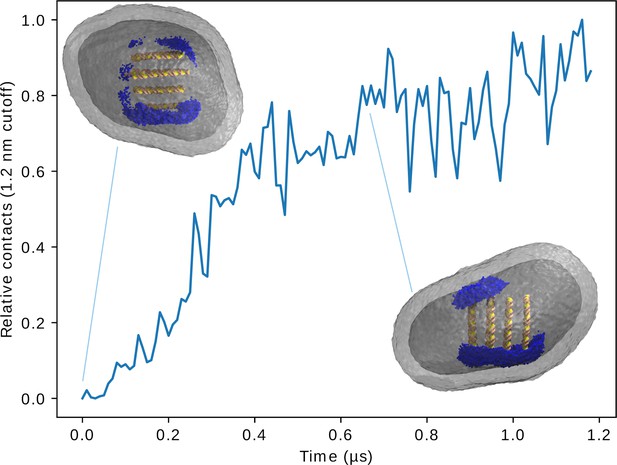
Reorientation of the large lipoplex in the endosome.
The graph shows the lipoplex-endosome lipid contacts (<1.2 nm) over time. Within 0.5 µs the lipoplex reorients itself such that its HII-channels are roughly aligned with the membrane normal in the contact region. Components of the lipoplex closer than 4 nm to the endosome are colored blue (a longer cut-off was used to help with perception of orientation), the endosomal lipids are grey and the dsDNA inside the lipoplex is yellow/pink.
Videos
Small lipoplex-bilayer transfection.
Fusion of a small lipoplex containing 4 fragments of dsDNA. The headgroups of the lipoplex lipids (DOTAP:1, DOPE:4) are depicted in blue and the bilayer lipids (short lipids Figure 2) are depicted in red. Yellow and pink are used for subsequently the bases and backbone of the dsDNA. No bias was applied during this simulation.
Large lipoplex-vesicle transfection.
Fusion of the large lipoplex from inside the endosomal vesicle. The headgroups of the endosomal and lipoplex lipids are subsequently blue and orange. The endosomal lipids have dark grey tails and the lipoplex lipids have light grey tails. Yellow and pink are used for subsequently the bases and backbone of the dsDNA. No bias was applied during this simulation, though opening of the first channel occurred by a biasing potential as described in the methods section.
Reorientation of lipoplex in vesicle.
Components of the lipoplex closer than 4 nm to the endosome are colored blue, the endosomal lipids are grey and the dsDNA inside the lipoplex is yellow/pink. Within 1 µs the lipoplex reorients itself such that its HII-channels are roughly aligned with the membrane normal in the contact region.
Additional files
-
Supplementary file 1
DOTAP_validation.
- https://cdn.elifesciences.org/articles/52012/elife-52012-supp1-v2.docx
-
Supplementary file 2
lipoplex_lipid-properties.
- https://cdn.elifesciences.org/articles/52012/elife-52012-supp2-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/52012/elife-52012-transrepform-v2.pdf





