PACT-mediated PKR activation acts as a hyperosmotic stress intensity sensor weakening osmoadaptation and enhancing inflammation
Figures
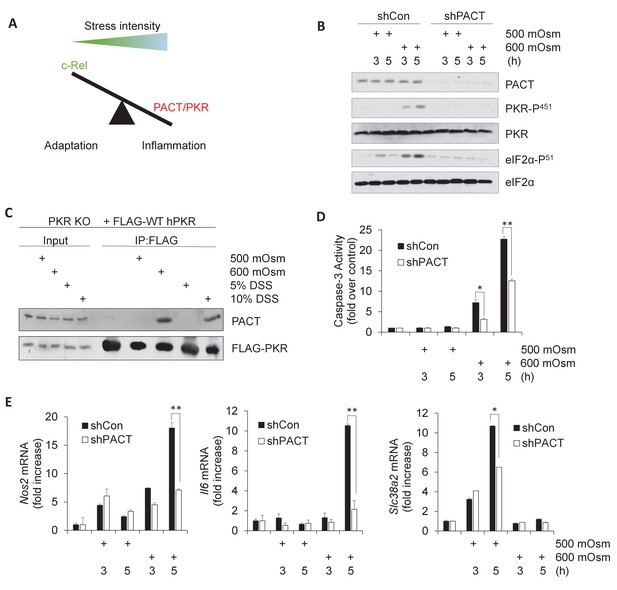
Activation of PACT-PKR axis promotes proinflammatory gene expression in response to high-intensity hyperosmotic stress.
(A) As stress intensity increases, signaling shifts from c-Rel-mediated adaptation to PACT/PKR-mediated inflammation. (B) MEFs selected for shRNA-mediated knockdown of PACT and control vector were treated with 500 or 600 mOsm sucrose for the indicated durations. Lysates were analyzed via western blot for the indicated proteins. (C) PKR KO MEFs reconstituted with FLAG-tagged human PKR were treated with sucrose or DSS for 3 hr. Co-immunoprecipitation with FLAG antibody was analyzed by western blot. Inputs are 5% of immunoprecipitated sample. (D) Control and shPACT MEFs were treated with 500 or 600 mOsm sucrose. Lysates were assayed for caspase-3 enzymatic activity. (E) Control and shPACT MEFs were treated with 500 or 600 mOsm sucrose. RNA was isolated, and transcripts were analyzed via RT-qPCR for the indicated mRNAs.
-
Figure 1—source data 1
Graph values for caspase-3 activity assays and RT-qPCR experiments in Figure 1.
- https://cdn.elifesciences.org/articles/52241/elife-52241-fig1-data1-v2.xlsx
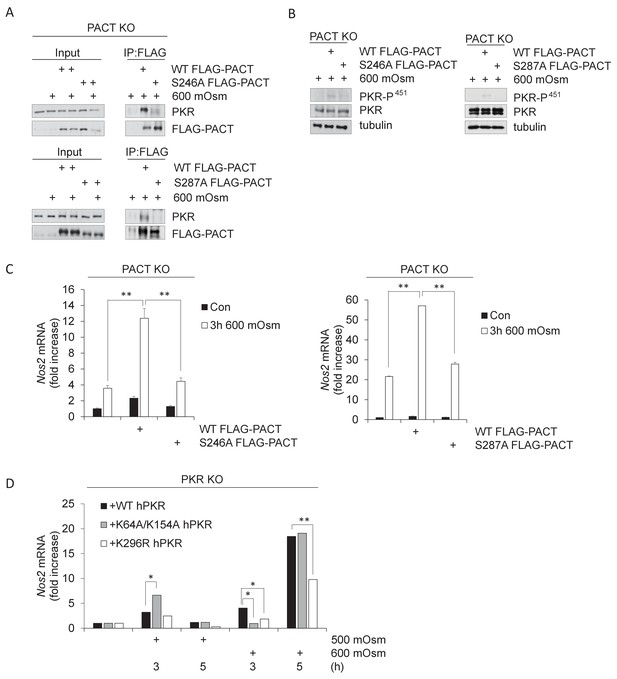
PACT Ser-246 and Ser-287 are necessary for PKR interaction and downstream inflammatory signaling.
(A) PACT KO MEFs were reconstituted with FLAG-WT PACT or FLAG-S246A mutant PACT constructs, then treated with 600 mOsm sucrose for 3 hr. Proteins from total cell extracts or co-immunoprecipitated with the FLAG antibody were analyzed via western blot. (B) PACT KO MEFs were reconstituted with FLAG-WT PACT, FLAG-S246A, or FLAG-S287A mutant PACT constructs, then treated with 600 mOsm sucrose for 3 hr. Total lysates were analyzed via western blot. (C) PACT KO MEFs were reconstituted with FLAG-WT PACT or FLAG-S246A mutant PACT constructs, then treated with 600 mOsm sucrose for 3 hr. RNA was isolated, and Nos2 transcript levels were analyzed via qPCR. (D) PKR KO MEFs reconstituted with wild type PKR, PKR mutated at the RNA-binding residues (K64R/K154R), or kinase activity-deficient PKR (K296R) were treated with 500 or 600 mOsm sucrose for the indicated durations. RNA was isolated, and Nos2 transcript levels were analyzed via qPCR.
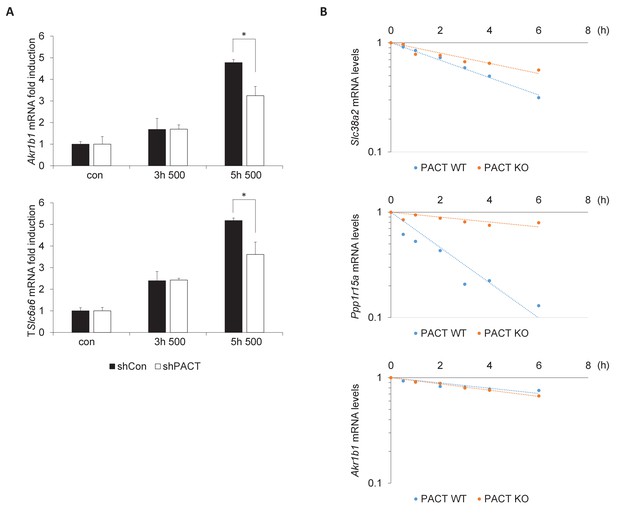
PACT influences the TonEBP-dependent adaptive transcription program.
(A) MEFs deficient in PACT were treated with 500 mOsm hyperosmotic media for the indicated durations. Total mRNA was isolated, and target mRNA levels analyzed via RT-qPCR. (B) MEFs deficient in PACT were treated with 500 mOsm hyperosmotic media for 3 hr. Actinomycin D was added, and total mRNA was harvested after the indicated duration. mRNA half-life was calculated using linear regression analysis.
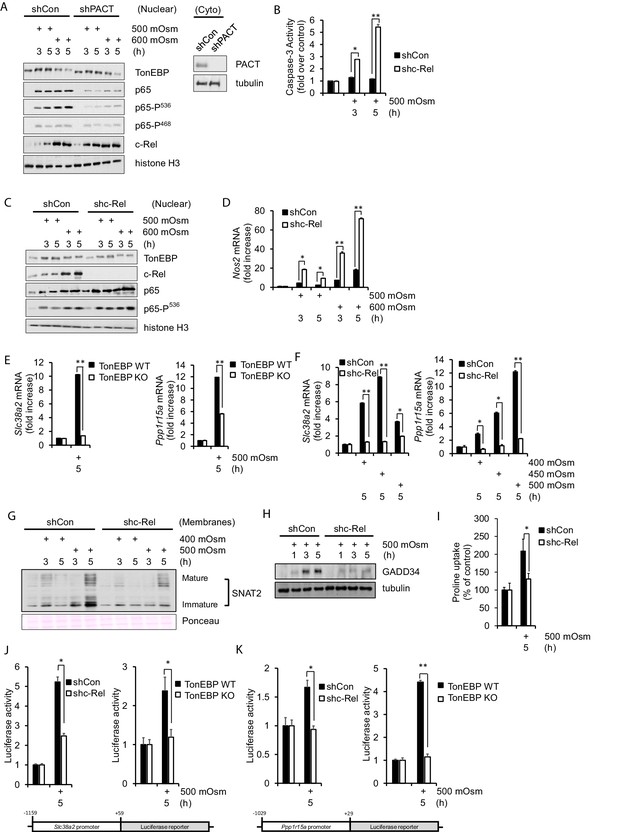
NF-κB c-Rel is a novel factor in the adaptive response to hyperosmotic stress.
(A) Control and shPACT MEFs were treated with 500 or 600 mOsm. Nuclear fractions were isolated, and protein levels were analyzed via western blot. Insert indicates efficiency of PACT depletion from cytoplasmic extracts. (B) Control and shc-Rel MEFs were treated with 500 mOsm sucrose. Lysates were assayed for caspase-3 enzymatic activity. (C) Control and shc-Rel MEFs were treated with 500 or 600 mOsm sucrose. Nuclear fractions were isolated and protein levels were analyzed via western blot. (D) Control and shc-Rel MEFs were treated with 500 or 600 mOsm sucrose. RNA was isolated, and mRNA transcripts were analyzed via RT-qPCR. (E) TonEBP WT and TonEBP KO MEFs were treated with 500 mOsm sucrose. RNA was isolated, and mRNA transcripts were analyzed via RT-qPCR. (F) Control and shc-Rel MEFs were treated with varying intensities of sucrose. RNA was isolated, and mRNA transcripts were analyzed via RT-qPCR. (G) After indicated treatments, cytoplasmic membrane fractions were isolated and analyzed via western blot. (H) After indicated treatments, total lysates were isolated and analyzed via western blot. (I) Control and shc-Rel MEFs were treated with 500 mOsm sucrose. Intracellular levels of the amino acid proline were analyzed via amino acid uptake assay. (J) Control and shc-Rel MEFs and TonEBP WT and KO MEFs were transfected with the Slc38a2-promoter luciferase reporter construct, then treated with 500 mOsm sucrose. Luciferase activity was normalized to co-transfected Renilla luciferase activity. (K) Control and shc-Rel MEFs and TonEBP WT and KO MEFs were transfected with the Ppp1r15a-promoter luciferase reporter construct, then treated with 500 mOsm sucrose. Luciferase activity was normalized to co-transfected Renilla luciferase activity.
-
Figure 2—source data 1
Graph values for caspase-3 activity assays and RT-qPCR experiments in Figure 2.
- https://cdn.elifesciences.org/articles/52241/elife-52241-fig2-data1-v2.xlsx
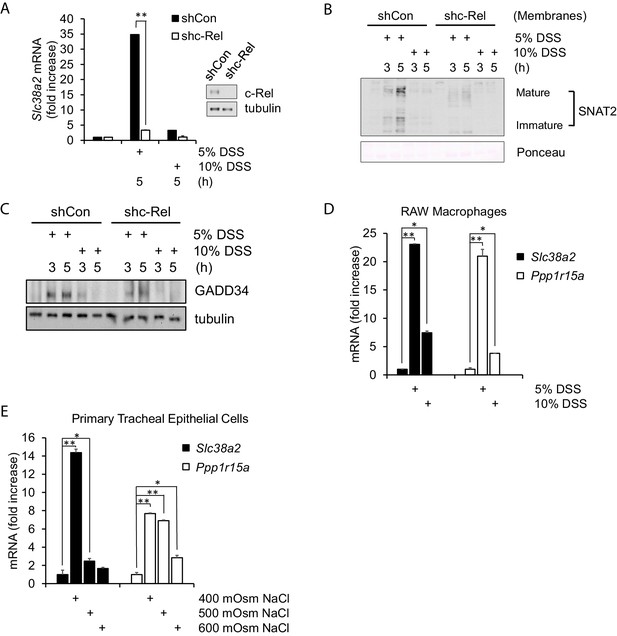
Hyperosmotic stress-induced adaptive gene expression occurs independent of cell type and osmolyte used.
(A) Control and shc-Rel MEFs were treated with 5% or 10% DSS. RNA was isolated, and mRNA transcripts analyzed via RT-qPCR. (B) Control and shc-Rel MEFs were treated with 5% or 10% DSS. Cytoplasmic membrane fractions were isolated, and protein levels analyzed via western blot. (C) Control and shc-Rel MEFs were treated with 5% or 10% DSS. Total lysates were analyzed via western blot. (D) RAW macrophages were treated with 5% or 10% DSS for the indicated durations. RNA was isolated, and transcripts were analyzed via qPCR. (E) Primary tracheal epithelial cells cultured from human patients were treated with the indicated concentration of NaCl. RNA was isolated, and transcripts were analyzed via qPCR.
-
Figure 3—source data 1
Graph values for RT-qPCR experiments in Figure 3.
- https://cdn.elifesciences.org/articles/52241/elife-52241-fig3-data1-v2.xlsx
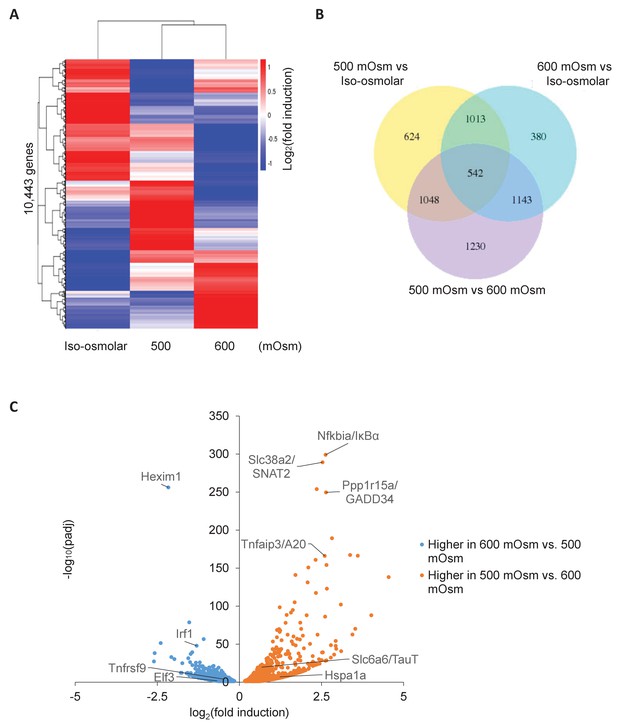
Differential gene expression in MEFs treated with low- or high-intensity hyperosmotic stress.
(A) WT MEFs were treated with 500 or 600 mOsm sucrose for 3 hr. RNA was isolated, libraries prepared, and total RNAs sequenced on an Illumina platform. Heatmap shows differences in expression for 10,443 unique genes between each treatment. (B) Venn diagram of differentially expressed genes shared between control, low intensity, and high intensity-treated cells. (C) Volcano plot analysis of upregulated and downregulated genes between low intensity and high intensity-treated cells.
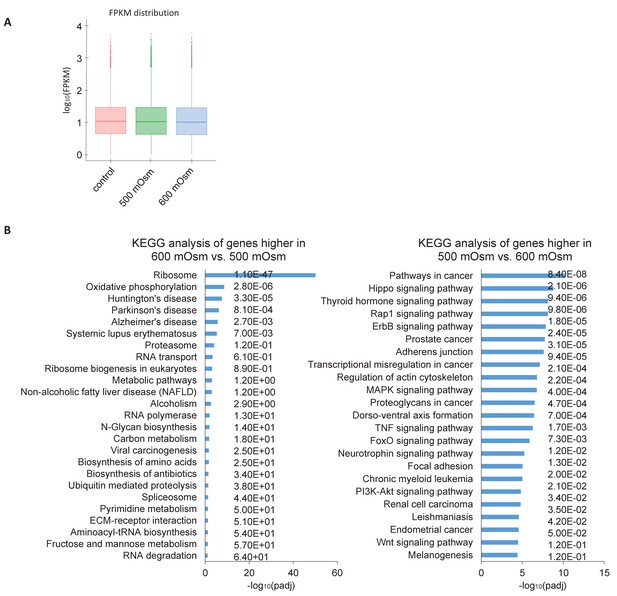
Hyperosmotic stress intensity alters expression of genes involved in ribosomal machinery and cancer signaling pathways.
(A) Similar numbers of reads were obtained between treatments with low- and high-intensity hyperosmotic stress. (B) KEGG analysis shows the most common pathways shared between differentially expressed genes under the indicated treatments. Q scores representing false discovery rate (FDR) are reported on the right.
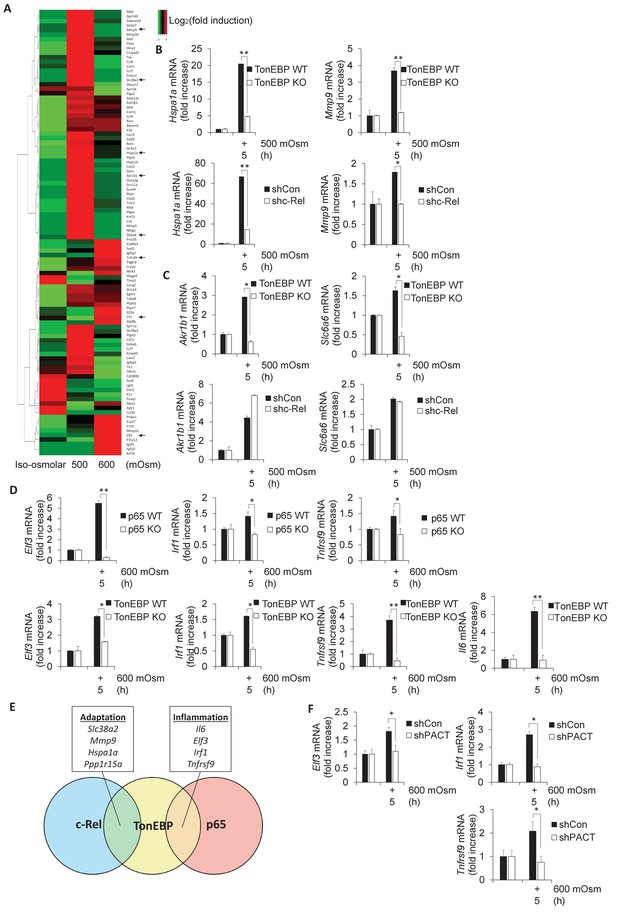
Subsets of genes induced by hyperosmotic stress are transcriptionally controlled by NF-κB c-Rel and TonEBP.
(A) TonEBP targets identified by literature demonstrate a general increase at mild intensity and decrease at high intensity hyperosmotic stress. (B–D) MEFs deficient in NF-κB c-Rel, NF-κB p65, and TonEBP were treated with the indicated stress intensity. RNA was isolated, and mRNA transcript levels analyzed via RT-qPCR. (E) Genes were categorized based on targeting of Rel-homology domain transcription factor complexes. (F) MEFs deficient in PACT were treated with the indicated stress intensity. RNA was isolated, and mRNA transcript levels analyzed via RT-qPCR.
-
Figure 5—source data 1
Graph values for RT-qPCR experiments in Figure 5.
- https://cdn.elifesciences.org/articles/52241/elife-52241-fig5-data1-v2.xlsx
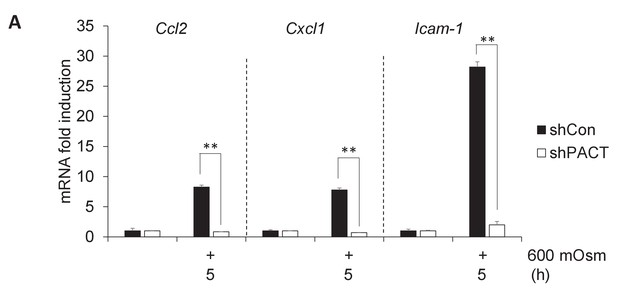
Additional proinflammatory gene expression programs are also dependent on PACT.
(A) MEFs deficient in PACT were treated with the indicated stress intensity. RNA was isolated, and mRNA transcript levels analyzed via RT-qPCR.
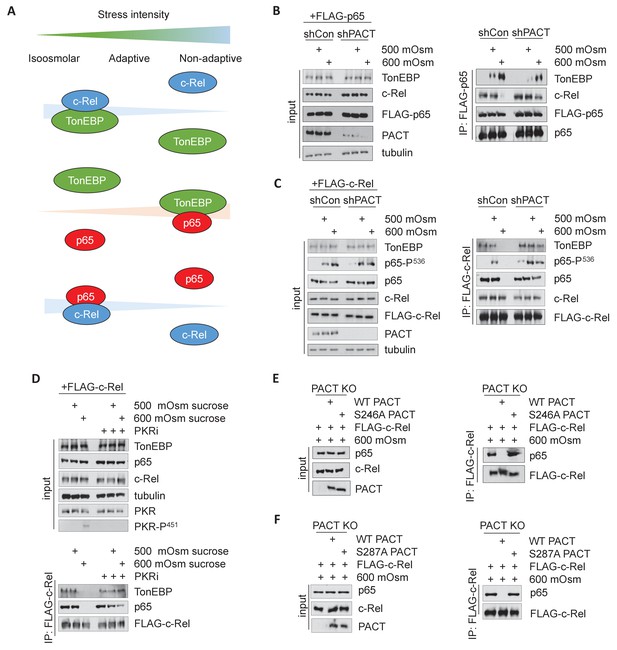
Activation of the PACT/PKR signaling axis inhibits the interaction of NF-κB c-Rel with NF-κB p65 or TonEBP, and increases the interaction of NF-κB p65 with TonEBP.
(A) As stress intensity increases, NF-κB c-Rel/TonEBP and NF-κB c-Rel/NF-κB p65 species dissociate, while NF-κB p65/TonEBP species accumulate. (B) Control and shPACT MEFs were transfected with a FLAG-NF-κB c-Rel construct, then treated with 500 or 600 mOsm sucrose for 3 hr. Total cell extracts and FLAG-co-immunoprecipitated proteins were analyzed via western blot. (C) Control and shPACT MEFs were transfected with a FLAG-NF-κB p65 construct, then treated with 500 or 600 mOsm sucrose for 3 hr. Total cell extracts and FLAG-co-immunoprecipitated proteins were analyzed via western blot. (D) WT MEFs were transfected with a FLAG-NF-κB c-Rel construct, then treated with the indicated stress for 3 hr in the presence or absence of a small molecule inhibitor of PKR. Total cell extracts and FLAG-co-immunoprecipitated proteins were analyzed via western blot. (E) PACT KO MEFs were reconstituted with WT- or S246A-PACT, and transfected with a FLAG-NF-κB c-Rel construct. After treatment with 600 mOsm sucrose for 3 hr, complexes were co-immunoprecipitated with the FLAG antibody and analyzed via western blot, in parallel with total cell extracts. (F) PACT KO MEFs were reconstituted with WT- or S287A-PACT, and transfected with a FLAG-NF-κB c-Rel construct. After treatment with 600 mOsm sucrose for 3 hr, complexes were co-immunoprecipitated with the FLAG antibody and analyzed via western blot in parallel with total extracts.
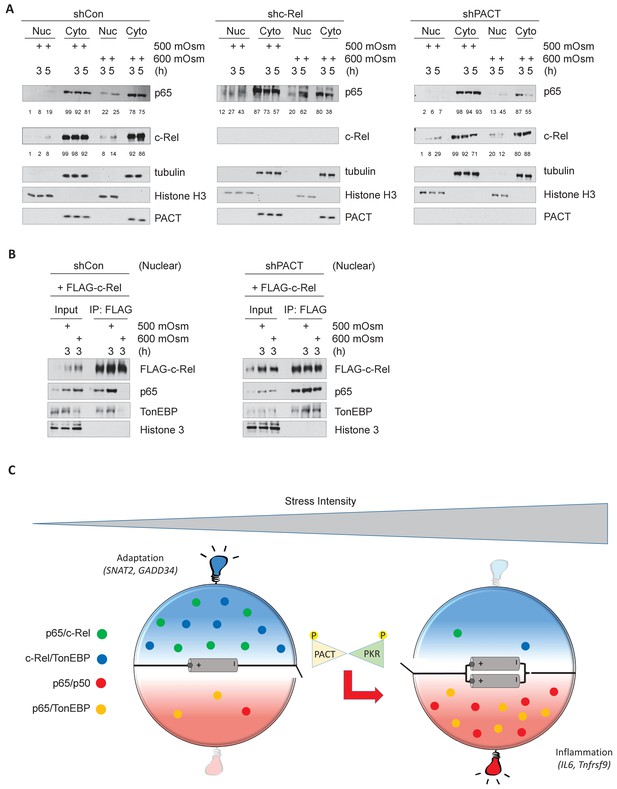
Hyperosmotic stress causes nuclear translocation of a small subset of NF-κB proteins.
(A) Control, shc-Rel, and shPACT MEFs were treated with the indicated intensities and durations of hyperosmotic stress, then cells were fractionated into cytoplasmic and nuclear lysates. Equal volumes of protein were analyzed via western blot. Protein localization was quantified as percentages in the corresponding nuclear or cytoplasmic fractions by densitometric analysis. (B) Control and shPACT MEFs were transfected with FLAG-NF-κB c-Rel. Cells were fractionated into nuclear and cytoplasmic lysates, and FLAG-NF-κB c-Rel was immunoprecipitated from nuclear lysates. Nuclear extracts and coimmunoprecipitated proteins were analyzed via western blot. (C) Adaptation to increased osmolarity of the extracellular environment involves Rel family transcription factors TonEBP and c-Rel. At high stress intensities, cells fail to adapt and proinflammatory signaling is induced via remodeling of Rel family transcription factor dimer formation. Loss of adaptation to hyperosmotic stress associates with increased TonEBP/p65 dimerization and loss of TonEBP/c-Rel and p65/c-Rel dimerization. PACT-mediated PKR activation marks the loss of adaptation and enhancement of proinflammatory gene expression.

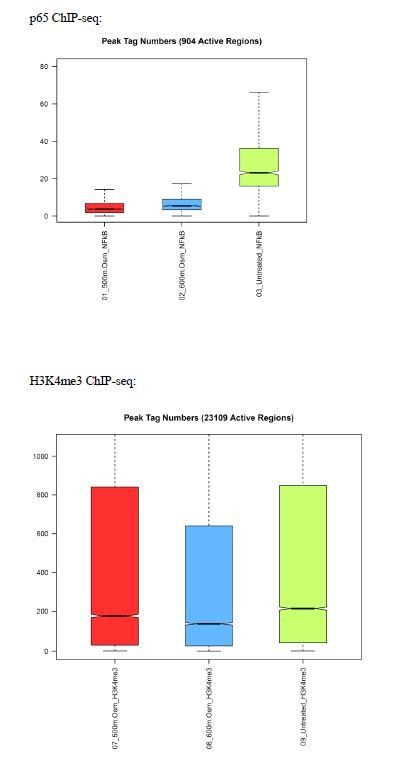
Number of ChIP-seq peaks for regions actively binding c-Rel were low, indicating failure of experiment.