Structures of the ATP-fueled ClpXP proteolytic machine bound to protein substrate
Figures
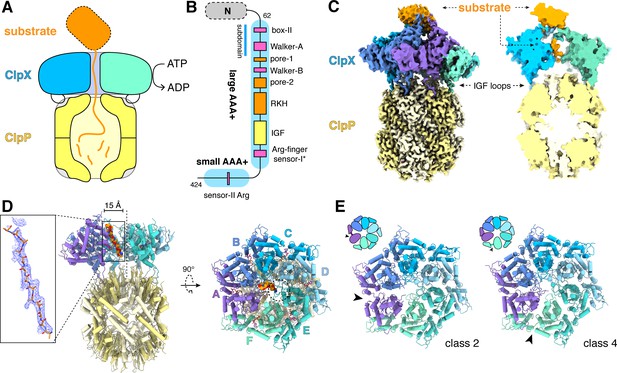
ClpXP protease.
(A) Schematic representation of ClpXP. (B) ClpX domain structure and positions of sequence motifs important for ATP binding and hydrolysis (magenta), substrate binding (orange), and ClpP binding (yellow). (C) Left. Composite cryo-EM density of ClpX∆N/ClpP complex. ClpP is yellow; ClpX∆N (class 3) is blue, green, or purple; and substrate is orange. Right. Slice through surface representation of the model showing substrate in the axial channel and the ClpP degradation channel. (D) Left. Cartoon representation of the class-4 ClpX∆N hexamer (subunit F removed) and its docking with a heptameric ClpP ring. Substrate in the ClpX channel is shown in space-filling representation as a poly-alanine chain; the inset shows substrate as a ball-and-stick model with associated density. The vertical dashed line shows the 7-fold symmetry axis of ClpP. Right. View rotated by 90°. The dashed circle shows the position of the ClpP pore. (E) Cartoon representation of class-2 and class-4 ClpX∆N hexamers with substrate removed for clarity. Arrows point to the different positions of the seam interface.
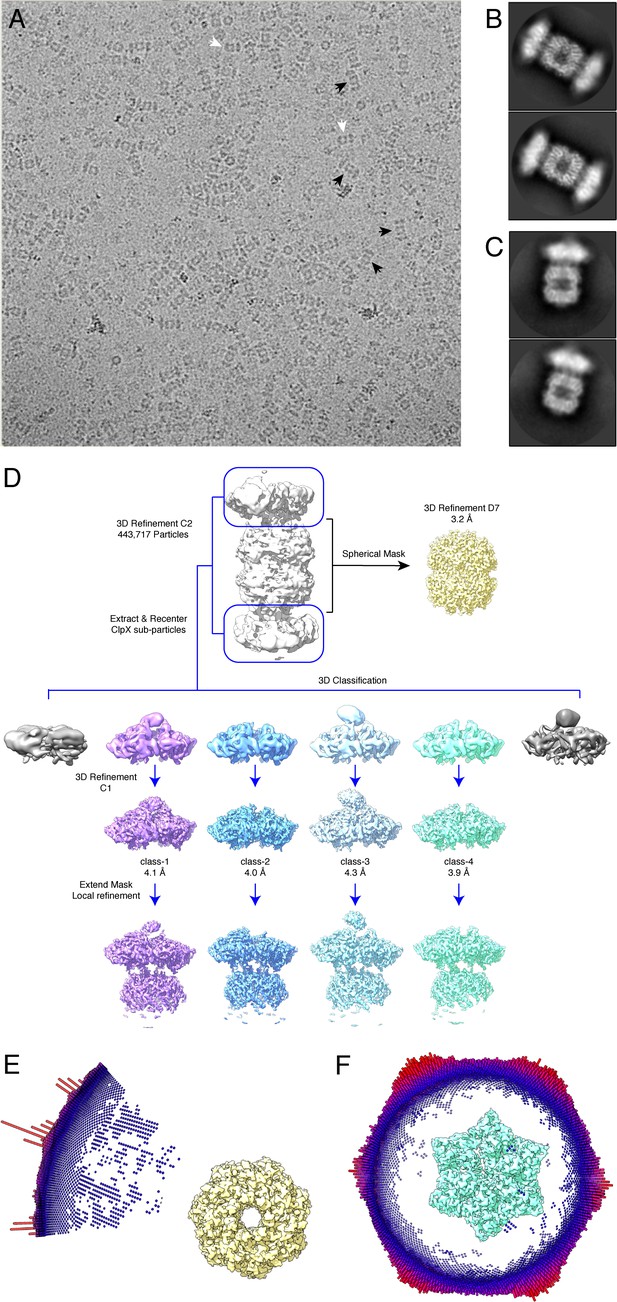
Cryo-EM data and strategy.
(A) Cryo-EM micrograph at 36000X magnification of doubly capped particles (black arrows) and singly capped particles (white arrows). (B) 2D-class averages of doubly capped complexes. (C) 2D-class averages of singly capped complexes. (D) Relion data processing scheme used to obtain 3D reconstructions. (E) Euler-angle distribution of particles used to reconstruct the D7 symmetric ClpP map. (F) Euler-angle distribution of particles used to reconstruct the non-symmetric class-4 ClpX∆N map.
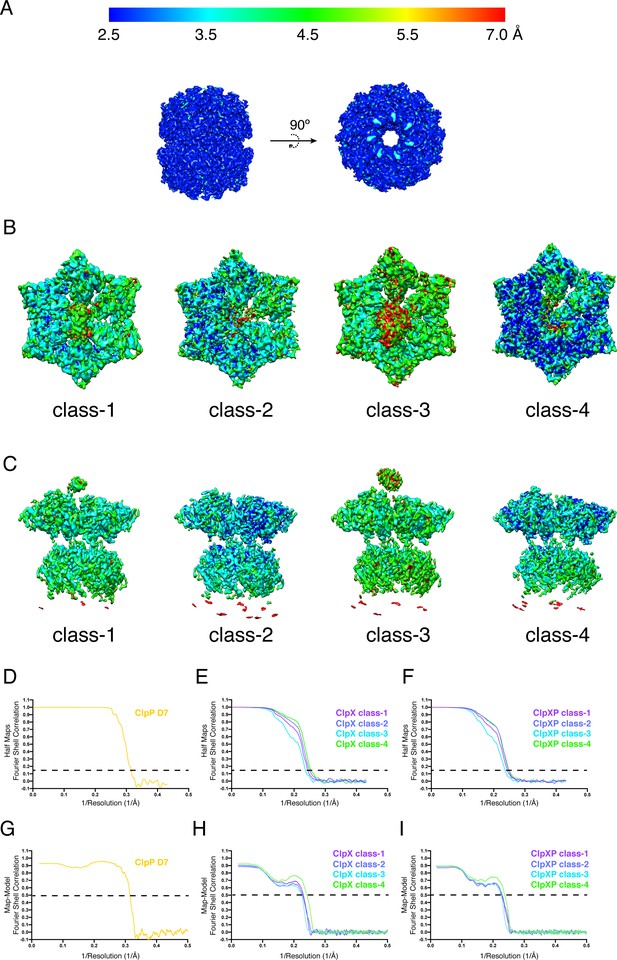
Local map resolution and FSC plots.
(A) Side views and top views of ClpP (D7). (B) Top views of the four ClpX∆N classes. (C) Side views of the four ClpXP classes. ResMap61 was used to estimate local resolutions. (D–F) FSC plots of all masked final 3D reconstructions. (G–I) FSC plots calculated using masked 3D reconstructions and models.
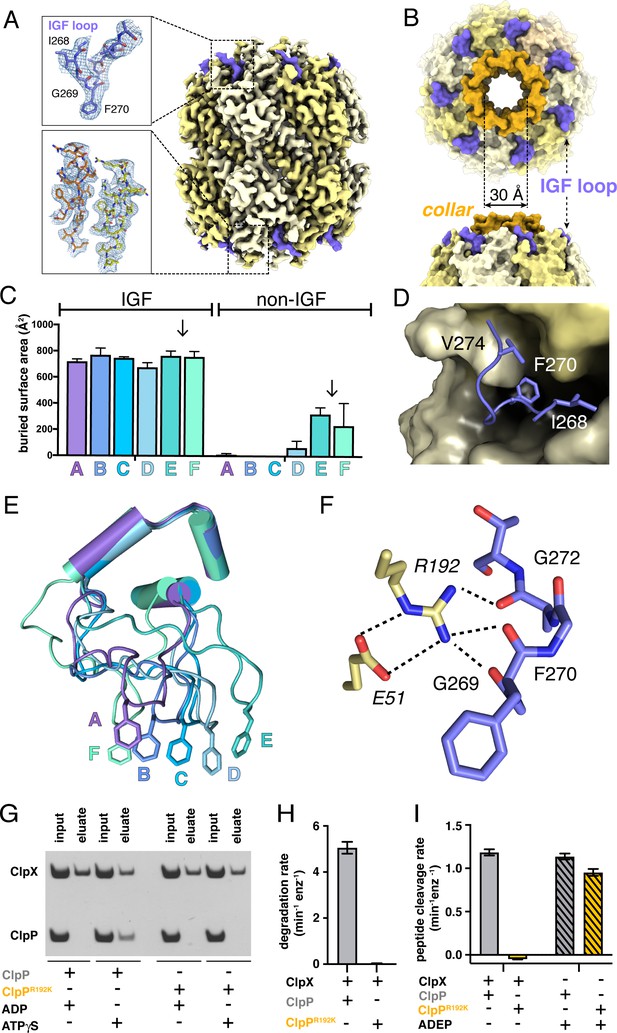
ClpP and symmetric IGF loop contacts.
(A) Density for ClpP14 in the D7 map is shown in yellow. Density for part of the IGF loops of ClpX is shown in blue. The upper inset shows the density as mesh and the fitted model for residues 267–275 of one IGF loop; the lower inset shows density and the fitted model for resides 2–18 of two β-hairpins in the ClpP collar. (B) Top and side views of ClpP showing the axial collar (darker yellow) and pore. (C) IGF loops of ClpX make the majority of contacts with ClpP as assessed by buried surface area. Each bar represents a different ClpX∆N subunit in the spiral and values are means ± 1 SD for the four classes. The arrows between subunits E and F mark the position of the unoccupied ClpP cleft. (D) The side chains of IGF residues Ile268 (I268) Phe270 (F270), and Val274 (V274) pack into hydrophobic ClpP clefts. (E) After alignment of the large AAA+ cores of the six subunits in the class-4 ClpX∆N hexamer, the IGF loops adopt a variety of conformations, helping mediate ClpP binding despite the symmetry mismatch. The side chains of Phe270 are shown in stick representation with the IGF loop (residues 263–283) and three flanking helices (residues 254–262 and 284–298) shown in cartoon representation. (F) The side chain of Arg192 (R192) in ClpP makes hydrogen bonds with carbonyl oxygens in the IGF loop of ClpX and also forms a salt bridge with Glu51 (E51) in a neighboring ClpP subunit. (G) ClpP but not R192KClpP binds ClpX∆N in pull-down assays performed in the presence of ATPγS. As a negative control, neither ClpP variant binds ClpX∆N in the presence of ADP (Joshi et al., 2004). (H) ClpX∆N supports degradation of cp7GFP-ssrA by ClpP but not by R192KClpP. (I) ClpX∆N supports degradation of a decapeptide by ClpP but not by R192KClpP. ADEP-2B activates decapeptide cleavage by both ClpP and R192KClpP. Error bars represent means (n = 3) ± 1 SD.

Structural features of ClpX∆N/ClpP interface.
(A) Cutaway showing axial collar of ClpP fitting into a bowl-shaped cavity at the bottom of the ClpX∆N hexamer. Black asterisks show contacts between the ClpX IGF loops and ClpP clefts. Red asterisks show contacts between ClpX and the ClpP collar. (B) Top views of ClpXP classes with substrate and most ClpX∆N except IGF loops (blue or purple) removed. Arrows show the position of the empty ClpP cleft without an IGF loop.
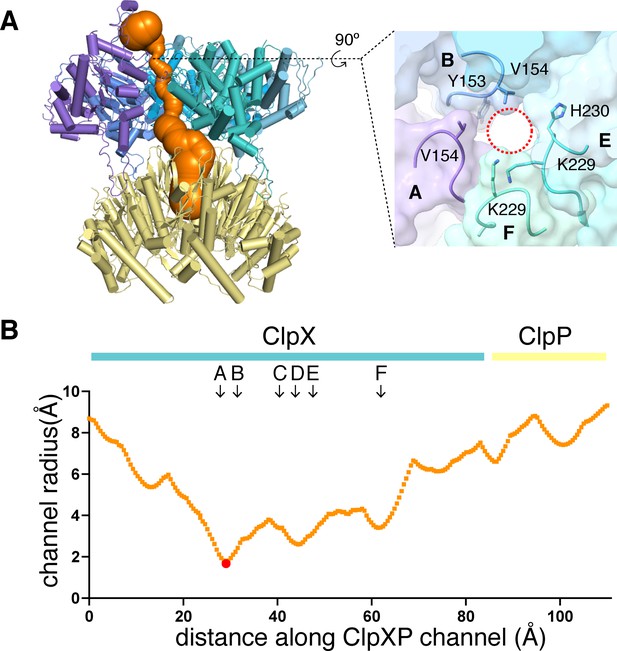
Dimensions of axial channel.
(A) The channel though ClpX and into ClpP (class-1) is most constricted near the top of the ClpX∆N hexamer. This panel was made using Caver (Pavelka et al., 2016). The inset shows that the constriction involves the pore-1 loops of subunits A and B and the RKH loops of subunits E and F. (B) Plot of channel radius as a function of channel length. Arrows mark positions of pore-1 loops. Red dot marks the bottleneck of this channel.
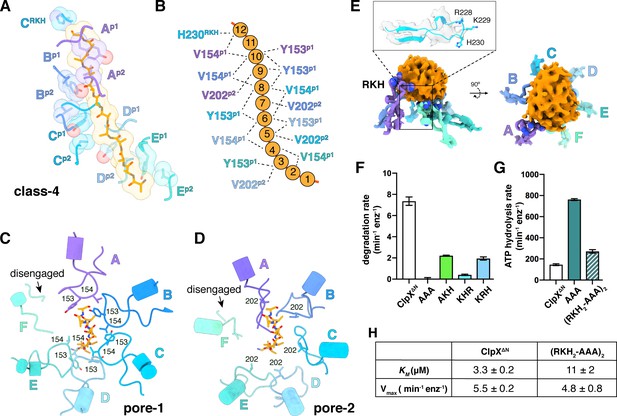
Substrate contacts.
(A) Interactions between substrate (orange, stick and surface representation) and ClpX∆N loops (blue and purple, stick and transparent space-filling representation) in the class-4 structure. Capital letters indicate subunits; superscripts indicate RKH loops, pore-1 (p1) loops, or pore-2 (p2) loops. (B) Scheme of interactions shown in panel (A). Dashed lines represent distances of 6.5 Å or less between the Cβ atoms of substrate alanines and the Cβ atoms of Y153/V154 (p1), or the Cγ atoms of V202 (p2), or the Cγ atom of H230 (RKH). (C) Top view of interactions of pore-1 loops with substrate (class 4). (D) Top view of interactions of pore-2 loops with substrate (class 4). (E) Interaction of RKH loops in the class-3 structure with the globular portion of the substrate above the channel. Inset – representative RKH-loop density (class 4, subunit C) and positions of R228, K229, and H230. (F) Mutation of RKH motifs in each subunit of a ClpX∆N hexamer inhibits degradation of Arc-st11-ssrA. (G) Effects of RKH mutations on ATP hydrolysis. (H) Mutating two RKH motifs in a single chain pseudohexamer to AAA increases KM for steady-state degradation of CP7GFP-ssrA.

Substrate interactions with ClpX.
(A) Substrate density within the ClpX channel is rendered as a transparent surface (grey) with atoms shown in ball and stick representation. Dashed lines shown approximate positions of the pore-1 loop from the ClpX subunit A (top) and subunit F (bottom). (B) Substrate density above the ClpX hexamer. Substrate density is colored in orange and ClpX subunits are colored in different shades from purple to aquamarine. (C) Interactions between substrate (orange, stick and surface representation) and ClpX pore loops (blue and purple, stick and transparent space-filling representation) in the class-2 structure. Capital letters indicate ClpX subunits; superscripts indicate RKH loops, pore-1 (p1) loops, or pore-2 (p2) loops. (D) Scheme of interactions shown in panel C. Dashed lines represent distances of 6.5 Å or less between the Cβ atoms of substrate alanines and the Cβ atoms of Y153/V154 (p1), or the Cε atom of R200 (p2), or the Cγ atom of H230 (RKH), or the Nζ atom of R228 (RKH).
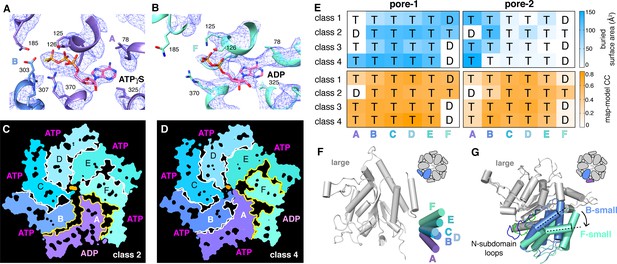
Nucleotide binding and subunit interactions in ClpX∆N hexamers.
(A) Density for ATPγS in subunit A of the class-4 hexamer. The positions of ATP-binding/hydrolysis motifs are shown in cartoon and stick representation. (B) Weaker nucleotide density built as ADP in subunit F of the class-4 hexamer. (C, D) Slices through the density maps of class-2 and class-4 hexamers show that the ADP-bound subunit (A or F, respectively) makes fewer neighbor contacts than ATP-bound subunits (yellow). (E) Contacts between substrate and pore-1 loops (left) or pore-2 loops (right) were characterized by buried surface area (top) or model-to-map correlation (bottom). T represents an ATPγS-bound subunit; D represents an ADP-bound subunit. (F) Superimposition of the large AAA+ domains of six subunits from a hexamer (class 4) showing variation in the angle between small and large domains of the same subunit. One of the large AAA+ domains is light grey, and the small AAA+ domains are in different shades of purple, blue and green. (G) Superimposition of two large AAA+ domains from a class-4 hexamer, illustrating different types of packing against the neighboring small AAA+ domain. The large domains of subunits A and C are light grey; the small domains of subunit F and B are cornflower blue and mint green. 65-111 subdomain loops are colored green (subunit F) and blue (subunit B). The dashed lines and arrow show an ~30° rotation at the interface. The F/A interface is the seam.

Nucleotide-binding pocket and mutations.
(A) Alignment of residues contacting bound nucleotide in the class-4 cryo-EM structure (A/B subunit interface; purple/blue) and crystal structure of ClpX∆N (pink/grey; PDB 3HWS; Glynn et al., 2009). (B) ATP-hydrolysis activity for ClpXΔN variants with mutations in nucleotide-binding pocket. Values are averages of three independent replicates ± SD. (C) Fitted Michealis-Menten-Hill parameters for ATP hydrolysis for variants with significant activity. Values are averages of three independent replicates ± SD. In panels B and C, CS (C169S) and YW (Y77W) indicate additional mutations in variants that do not, by themselves, alter ATP hydrolysis activity.
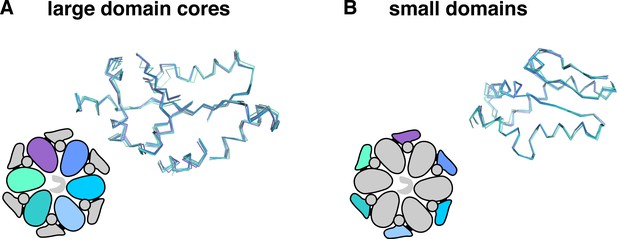
Invariant and flexible portions of ClpX∆N.
(A) Superimposition of the cores of the large AAA+ domains from each subunit of the class-4 ClpX∆N structure. (B) Superimposition of the small AAA+ domains from the class-4 ClpX∆N hexamer.
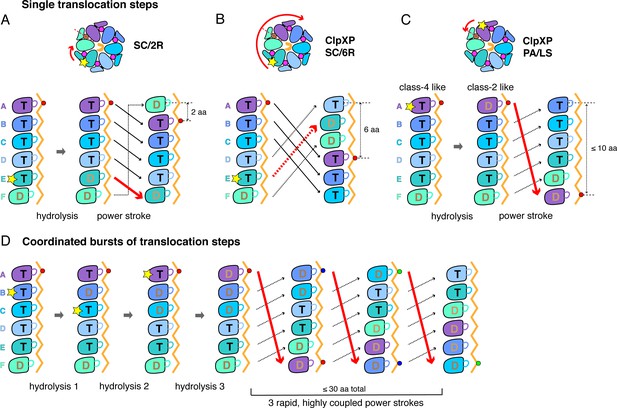
Single-step and burst translocation models.
In all panels, T represents ATP-bound subunits, and D represents subunits containing ADP and possibly inorganic phosphate. (A) SC/2R translocation model proposed for the Yme1 protease and 26S proteasome (Puchades et al., 2017; de la Peña et al., 2018; Dong et al., 2019). Only subunit E hydrolyzes ATP (depicted by a star). Hydrolysis and/or product release results in a two-residue translocation step, in which the top five subunits in the spiral move down one position in the clockwise direction and the bottom subunit moves up to the top position. (B) SC/6R translocation model. During a six-residue translocation step, subunits A, B, and C each move down in the spiral to positions D, E, and F, respectively, dragging substrate with them; at the same time, subunits D, E and F each move up to positions A, B, and C. Subunit displacement is in the clockwise direction. (C) PA/LS model in which ATP hydrolysis in subunit A results in a single translocation step of up to 2.5 nm in length as a consequence of anti-clockwise movement of this subunit to position F at the bottom of the spiral. At the same time, subunits BCDEF move up to positions ABCDE. (D) One variation of a PA/LS model resulting in a burst of three long translocation steps. Initial probabilistic ATP hydrolysis in subunits D and then B creates strain in the spiral, which is released in burst of fast steps upon ATP hydrolysis in subunit A. In the general PA/LS model, the initial ATP-hydrolysis event can occur with different probabilities in subunits A-E.
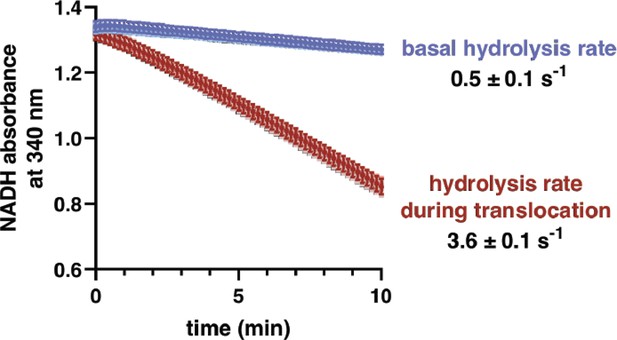
ATP-hydrolysis rates.
ATP-hydrolysis rates by ClpX∆N/ClpP were measured using a coupled assay (Martin et al., 2005) that results in loss of NADH absorbance at 23°C in PD buffer, conditions that approximate the conditions of optical-trapping experiments (Cordova et al., 2014). Shown are experiments in the absence (basal) or presence (translocation) of a near-saturating concentration (20 µM) of an unfolded titinI27-ssrA substrate. The figure shows an overlay of three independent replicates. The rates shown are averages of the three independent replicates ± SD in units corresponding ATP molecules hydrolyzed per second per enzyme.
Videos
Top and side views of a ClpX∆N hexamer and adjacent ClpP ring for classes 1, 2, 3, and 4.
ClpX∆N subunits (cartoon representation) are colored blue, green, or purple; carbons in substrate (space-filling representation) are colored orange; carbons in ATP or ADP (space-filling representation) are colored magenta; and ClpP subunits (cartoon representation) are colored yellow with the axial β-hairpin colored gold. Note that substrate above the axial channel of ClpX∆N is not shown because the corresponding density was not modeled.
Interaction of substrate with ClpX∆N.
Substrate in the axial channel of the class-4 ClpX structure adopts an extended conformation and is contacted by the pore-1, pore-2, and RKH loops of ClpX, which are arranged into a spiral and pack against each other.
Transition between class-2 and class-4 structures of ClpX∆N/ClpP.
A conformational morph between the class-2 and class-4 structures of ClpXP is shown. To generate this morph, ClpX subunits in the class-2 structure were renamed from ABCDEF to FABCDE and aligned to the class-4 ClpXP structure in two steps to maximize overlap between the ClpP portions of the two structures. The ADP-bound ClpX subunit (colored aquamarine) moves most dramatically, from the top to the bottom of the ClpX spiral. In the ADP-bound subunit, residues of the pore-1 loop move ~1.6–1.9 nm.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Escherichia coli) | clpX | E. coli (strain K12) EXPASY | UniProtKB- P0A6H1 | |
| Gene (Escherichia coli) | clpP | E. coli (strain K12) EXPASY | UniProtKB- P0A6G7 | |
| Recombinant DNA reagent | PACYC-ClpXΔN(E185Q)6-TEV-cHis6 (plasmid) | This paper, Material and methods | ClpX expression, can be obtained from the Sauer lab | |
| Recombinant DNA reagent | pT7ClpP-TEV-cHis6(plasmid) | (Stinson et al., 2013) | WTClpP expression, can be obtained from the Sauer lab | |
| Recombinant DNA reagent | pT7-ClpPR192K-TEV-cHis6 (plasmid) | This paper, Material and methods | ClpP mutant, can be obtained from the Sauer lab | |
| Recombinant DNA reagent | pT7-ClpXΔN-AAA (plasmid) | This paper, Material and methods | ClpX RKH loop mutant, can be obtained from the Sauer lab | |
| Recombinant DNA reagent | pT7-ClpXΔN-AKH (plasmid) | This paper, Material and methods | ClpX RKH loop mutant, can be obtained from the Sauer lab | |
| Recombinant DNA reagent | pT7-ClpXΔN-KHR (plasmid) | This paper, Material and methods | ClpX RKH loop mutant, can be obtained from the Sauer lab | |
| Recombinant DNA reagent | pT7-ClpXΔN-KRH (plasmid) | This paper, Material and methods | ClpX RKH loop mutant, can be obtained from the Sauer lab | |
| Recombinant DNA reagent | PACYC-ClpXΔN-(RKH2-AAA)2(plasmid) | This paper, Material and methods | ClpX RKH loop mutant, can be obtained from the Sauer lab | |
| Recombinant DNA reagent | pET4b-Arc-st11-ssrA (plasmid) | (Bell et al., 2018) | Expresses Arc substrate for degradation assay, can be obtained from the Sauer lab | |
| Recombinant DNA reagent | pT7-cp7GFP-ssrA (plasmid) | (Nager et al., 2011) | Expresses fluorescent substrate for degradation assay, can be obtained from the Sauer lab | |
| Strain, strain background (Escherichia coli) | ER2566 | NEB | 1312 | Chemical competent cells |
| Chemical compound, drug | ADEP-2B | (Schmitz et al., 2014) | Small molecule antibiotics binds ClpP. | |
| Software, algorithm | Relion | (Kimanius et al., 2016) | RRID:SCR_016274 | EM reconstruction software |
| Software, algorithm | UCSF Chimera | (Pettersen et al., 2004) | RRID:SCR_004097 | Molecular Visualization Software |
| Software, algorithm | UCSF ChimeraX | (Goddard et al., 2018) | RRID:SCR_015872 | Molecular Visualization Software |
| Software, algorithm | Phenix | (Adams et al., 2010) | RRID:SCR_014224 | Protein Model Building Software |
| Software, algorithm | Molprobity | (Williams et al., 2018) | RRID:SCR_014226 | Protein Model Evaluation Software |
| Software, algorithm | Caver | (Pavelka et al., 2016) | Molecular Visualization Software | |
| Software, algorithm | PyMOL | Schrödinger, LLC. | RRID:SCR_000305 | Molecular Visualization Software |
| Software, algorithm | Coot | (Emsley and Cowtan, 2004) | RRID:SCR_014222 | Protein Model Building Software |
| Software, algorithm | Ctffind | (Rohou and Grigorieff, 2015) | RRID:SCR_016732 | EM Image Analysis Software |
| Software, algorithm | PISA | 'Protein interfaces, surfaces and assemblies' service PISA at the European Bioinformatics Institute. (http://www.ebi.ac.uk/pdbe/prot_int/pistart.html) | RRID:SCR_015749 | Protein Model Analyzing Software |
Additional files
-
Supplementary file 1
contains four Tables with information about the structures.
- https://cdn.elifesciences.org/articles/52774/elife-52774-supp1-v2.doc
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/52774/elife-52774-transrepform-v2.docx