Kainate receptors regulate development of glutamatergic synaptic circuitry in the rodent amygdala
Figures
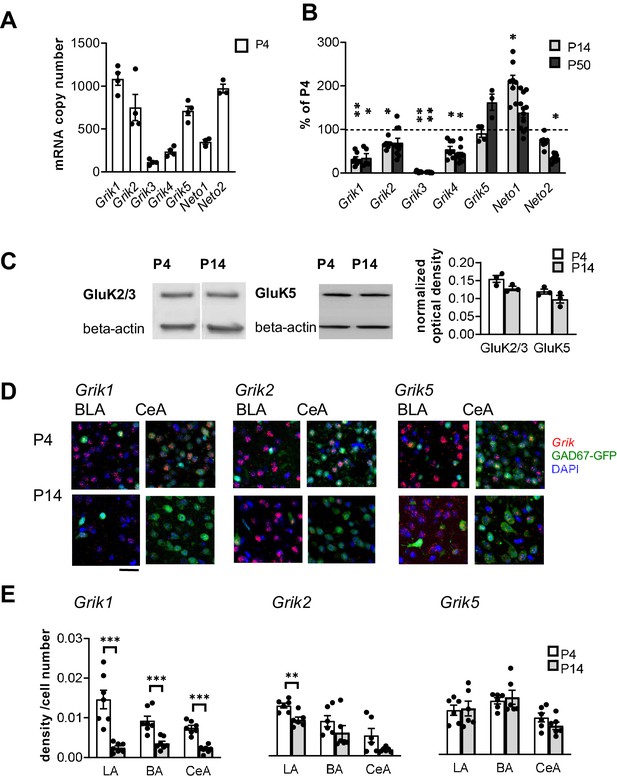
Kainate receptors are highly expressed in the amygdala during the first postnatal week.
(A) The predominant KAR subunits expressed in the newborn BLA are GluK1, GluK2 and GluK5 and the auxiliary subunit NETO2. RT-aqPCR data on Grik1-5, Neto1 and Neto2 mRNA expression in the BLA at P4 (n = 3–4 rats/group). (B) RT-qPCR analysis of Grik1-5, Neto1 and Neto2 mRNA expression in the BLA at different stages of development, expressed as a percentage of the level at P4 (n = 3–12 rats/group). Grik1 expression is strongly downregulated during early postnatal development. *p<0.05, **p<0.001 as compared to P4. (C) Western blot from neonatal (P4 and P14) BLA, using antibodies against GluK2/3 and GluK5. Beta-actin was used as a loading control. GluK2 and GluK5 protein was clearly detected in the BLA both at P4 and P14. Quantification of the signal indicates no significant developmental changes in the expression level. n = 3 rats/group. (D) Cell-type-specific expression pattern of KAR subunits in the amygdala during early postnatal development. Example images showing ISH staining (red) with antisense RNA probes against Grik1, Grik2 and Grik5 mRNA in the P4 mouse amygdala, in GAD67GFP positive (green) and negative (blue DAPI stain) cells. Scale bar, 100 µm. (E) Pooled data on the integrated density of the ISH signal, normalized to the number of cell bodies (identified by DAPI staining) within the analyzed region at P4 and P14. Grik1 mRNA expression is strongly downregulated in all amygdala subnuclei during first weeks of postnatal development. LA, Lateral amygdala; BA Basal Amygdala; CeA Central amygdala (data for each subunit is pooled from n = 3–4 mice and three rats). *p<0.05, **p<0.001.
-
Figure 1—source data 1
Raw data for results shown in the Figure 1.
- https://cdn.elifesciences.org/articles/52798/elife-52798-fig1-data1-v2.xlsx
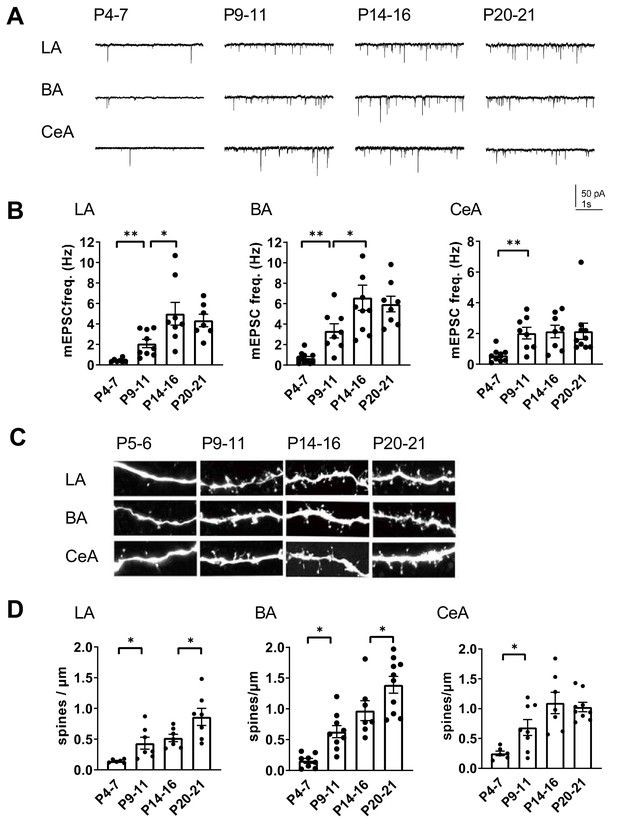
Development of glutamatergic synaptic connectivity in the amygdala.
(A) Example traces of action-potential independent glutamatergic events (mEPSCs), recorded from LA, BA and CeA neurons at different stages of development. (B) Pooled data of mEPSC frequency in amygdala subnuclei at different stages of development (n = 7–11 from 4 to 9 rats/group). The mEPSC frequency increases with a different developmental time course in CeA as compared to BA and LA. *p<0.05, **p<0.001. (C) Confocal images illustrating dendritic spines in biocytin filled neurons from LA, BA and CeA at P5, P10, P15 and P21 (D) Pooled data on spine density in LA, BA and CeA neurons at different stages of development. Data is obtained from 6 to 10 dendrites from three rats/group. *p<0.05; **p<0.001.
-
Figure 2—source data 1
Raw data for results shown in the Figure 2.
- https://cdn.elifesciences.org/articles/52798/elife-52798-fig2-data1-v2.xlsx
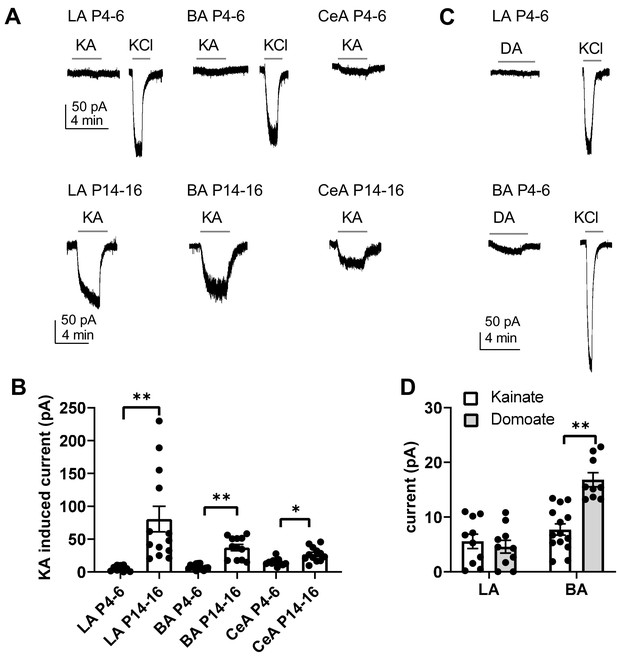
Functional map of ionotropic KAR activity in the amygdala during the first postnatal week.
(A) Example traces illustrating inward currents in response to application of 2 µM kainate (KA) in the presence of blockers for AMPA, NMDA and GABA-A receptors, in LA, BA and CeA neurons at P4-6 and P14-16. 33 mM KCl was applied as a positive control. In the neonatal BLA, no KA-induced currents are detected while a clear current is induced under similar conditions at P14-P16. (B) Pooled data on the effect of 2 µM kainate on holding current in LA, BA and CeA neurons at two different stages of development (P4-6 and P14-16; LA n = 10 [6] and 13[7]; BA: n = 14 [7] and 12 [6], CeA n = 12[10] and 12[8], respectively). (C) Example traces showing that application of 500 nM of domoate (DA) induced no current in neonate LA neurons, while a small inward current was detected in BA (P4-6). The recordings were made in the presence of blockers for AMPA, NMDA and GABA-A receptors. (D) Pooled data comparing the effect of 2 µM kainate (LA n = 10[4], BA n = 14[4]) and 500 nM domoate (LA: n = 10[4]; BA: n = 9[4]) on holding current in neonatal LA and BA neurons. The n numbers in this and following figure legends refer to the number of experiments, followed by the number of animals in brackets.
-
Figure 3—source data 1
Raw data for results shown in the Figure 3.
- https://cdn.elifesciences.org/articles/52798/elife-52798-fig3-data1-v2.xlsx
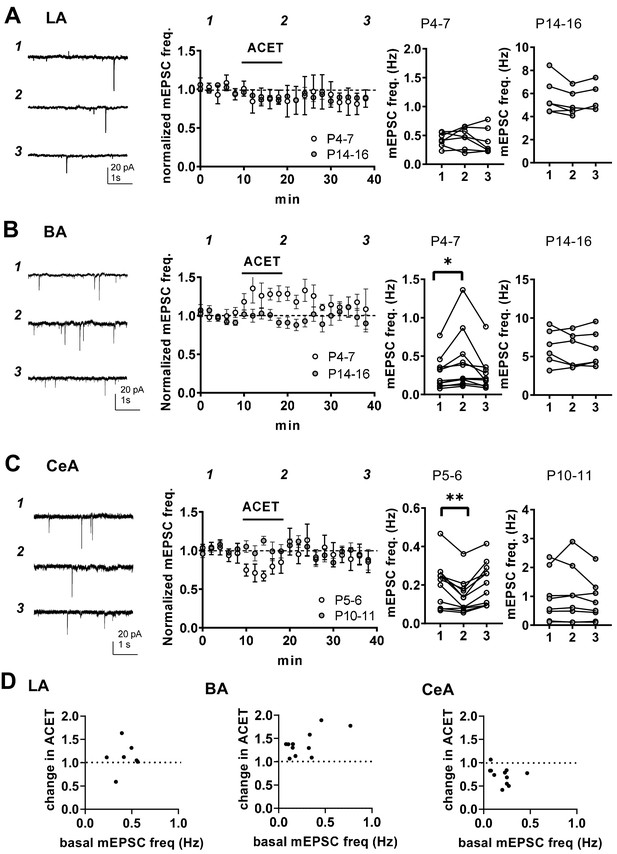
KARs regulate glutamatergic inputs to BA and CeA during the first postnatal week.
(A) KAR antagonism does not affect action-potential independent glutamatergic events (mEPSCs) to LA during the first postnatal week. Example traces (P5) from the time points indicated (1-3) and time course plot of averaged data (± s.e.m.) illustrating little or no effect of ACET (200 nM) on mEPSC frequency in LA at P4-7 and at P14-16. The graphs on the right show the mEPSC frequency under control conditions (1), in the presence of ACET (2), and after wash-out of the drug (3), in individual experiments at two different stages of development (P4-7 and P14-16; n = 7 [4] and 6 [5], respectively). (B) Similar data for recordings in the BA. In newborn BA, ACET application reversibly increases mEPSC frequency (n = 11[8]). This effect is no longer observed at P14-P16 (n = 6[4]). *p<0.05. (C) Similar data for recordings in the CeA. In the first week of life (P4-6; n = 9[6]) but not at P9-11 (n = 6[4]), ACET application leads to robust depression of mEPSC frequency. **p<0.01. (D) Scatter plots illustrating the effect of ACET on mEPSC frequency in individual cells in various amygdala subnuclei in the first postnatal week. The P4-7 data from A-C is plotted as a function of the basal mEPSC frequency.
-
Figure 4—source data 1
Raw data for results shown in the Figure 4.
- https://cdn.elifesciences.org/articles/52798/elife-52798-fig4-data1-v2.xlsx
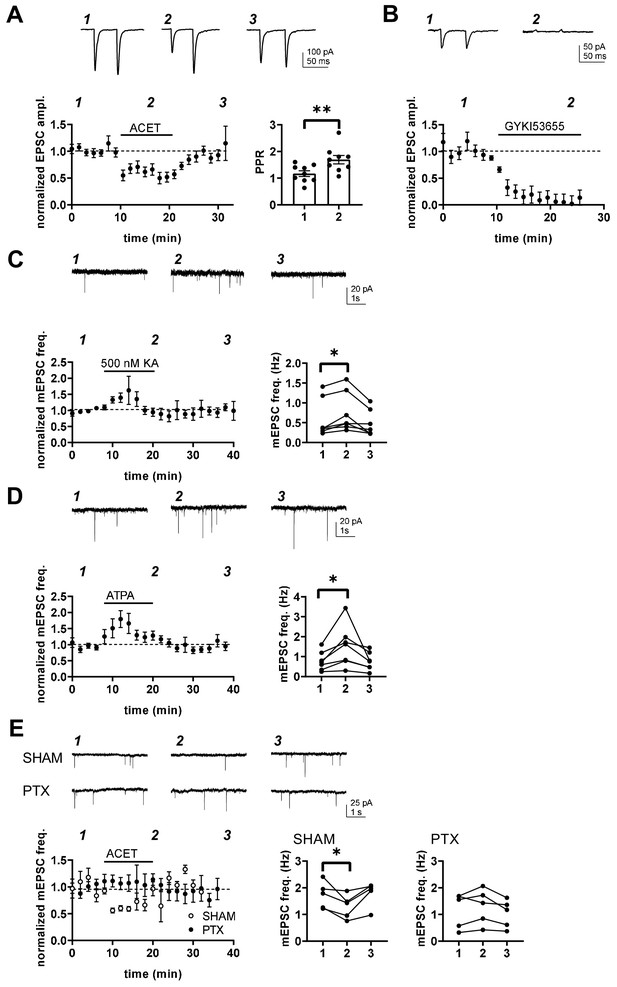
Transmission at immature BLA-CeA synapses is tonically facilitated by presynaptic G-protein-coupled KARs.
(A) KAR antagonism leads to reversible depression of glutamatergic transmission from to BLA-CeA during first postnatal week. Example traces of EPSCs, recorded from CeA neurons at P5 in response to paired-pulse stimulation of the LA, under control conditions and in the presence of 200 nM ACET. Pooled data (± s.e.m.) showing that ACET application is associated with a reduction in the EPSC amplitude and an increase in paired pulse ratio. n = 9[5]; **p<0.001. (B) Postsynaptic KARs do not contribute to transmission at immature BLA-CeA synapses. Similar data as in A, showing that the AMPA receptor antagonist GYKI53655 fully blocks the EPSC. n = 8 (4); p<0.001. (C) Example traces from the time points indicated (1-3) and time course plot (average ± s.e.m.) depicting a transient increase mEPSC frequency in response to application of KA (500 nM). The graphs show mEPSC frequency under control conditions (1), in the presence of KA (2), and after wash-out of the drug (3) in individual cells in CeA (P4-6). n = 7 (6); *p<0.05. (D) Application of the GluK1 selective agonist ATPA leads to an increase in mEPSCs frequency, similar to that observed in response to 500 nM KA. Example traces and pooled data as in C. n = 7 (4); *p<0.05. (E) The tonic KAR activity regulating glutamatergic transmission to CeA acts via a G-protein-dependent mechanism. The effect of ACET on mEPSC frequency is fully blocked in slices that are pre-incubated in the presence pertussis toxin (PTX; n = 5[3]), but not in SHAM incubated controls (n = 5[3]). *p<0.05.
-
Figure 5—source data 1
Raw data for results shown in the Figure 5.
- https://cdn.elifesciences.org/articles/52798/elife-52798-fig5-data1-v2.xlsx
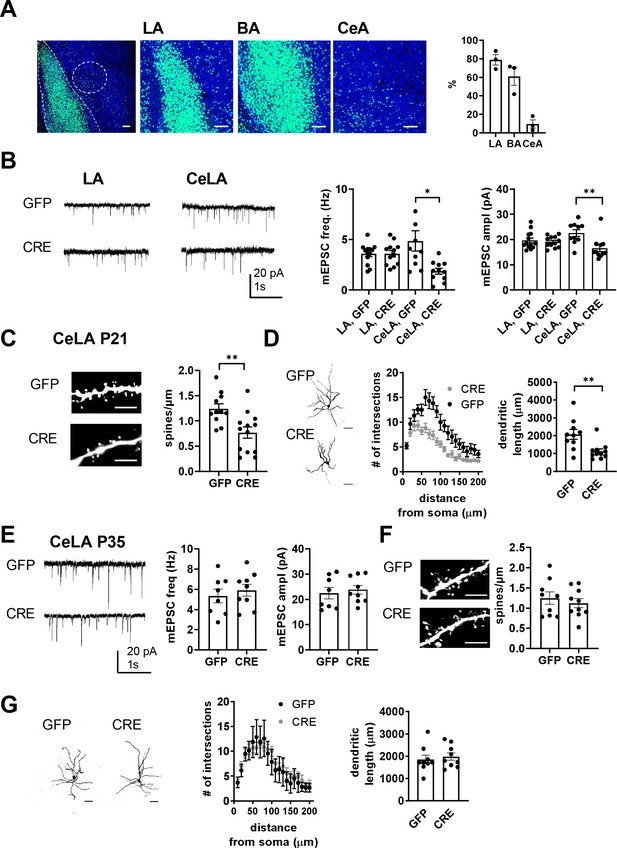
Loss of GluK1 in the amygdala in a cKO mouse model impairs glutamatergic innervation and maturation of CeLA neurons.
(A) Image illustrating the GFP fluorescence in the BLA and CeA of Grik1 cKO mice (P21), after in vivo injection of GFP-CRE AAV virus at P2. The viral transduction mainly targeted LA and BA (79 ± 5% and 61 ± 8% of the DAPI labeled cell bodies (blue), respectively), but also some cells in the CeA (10 ± 4% of cells). Scale bar, 100 µm. (B) Example traces and pooled data showing that loss of GluK1 in the BLA has no effect on the mEPSCs at LA neurons but results in lower mEPSC frequency and amplitude in the CeLA as compared to controls. AAV viruses encoding for GFP-CRE or GFP (control) were injected to the BLA at P2-4, and mEPSCs were recorded from acute slices at P21. LA: GFP n = 13(5), GFP-CRE n = 12(4); CeLA: GFP n = 10(3), GFP-CRE n = 11(3); *p<0.05. (C) Post hoc morphological characterization of the biocytin filled CeLA neurons (P21). Density of dendritic spines was significantly lower in GFP-CRE slices as compared to controls (GFP-CRE, n = 11(4); GFP n = 13(4)). **p<0.01. Scale bar, 5 µm (D) Sholl analysis of the same neurons. The number of dendritic intersections and the total dendritic length is significantly lower in GFP-CRE slices as compared to controls. Pooled data (GFP-CRE, n = 11(4); GFP n = 10(4)) and confocal images (z-stack) illustrating the morphology of the dendritic tree. **p<0.01. Scale bar, 50 µm E) Local inactivation of Grik1 at a later developmental stage has no effect on mEPSCs and dendritic morphology of CeLA neurons. AAV viruses encoding for GFP-CRE or GFP (control) were injected to the BLA of the Grik1 cKO mice at P14 and ex vivo analysis was done at P35. The image panels show mEPSCs (E), dendritic spines (F), intersections and total dendritic length (G) for 7-9(3) neurons from both GFP and GFP-CRE slices.
-
Figure 6—source data 1
Raw data for the results shown in the Figure 6.
- https://cdn.elifesciences.org/articles/52798/elife-52798-fig6-data1-v2.xlsx
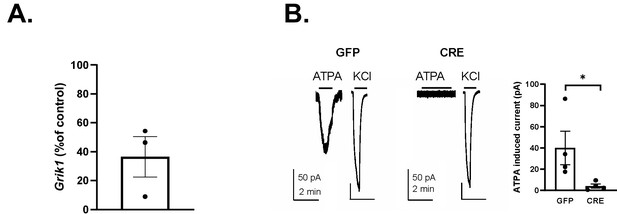
Validation of the GluK1 cKO mouse model.
(A) RT-qPCR results showing loss of Grik1 mRNA expression in the hippocampus (HC) of Grik1tm1c/tm1c mice 23 days after injection of AAV virus encoding for GFP-CRE. Injection was done in 9-week-old mice. The data represents the level of Grik1 mRNA expression in the HC of GFP-CRE group as a percentage of the GFP control (n = 3 mice for both groups). *p=0.01, ANOVA. (B) Loss of GluK1 function in the LA following injection of AAV virus encoding for GFP-CRE in the Grik1tm1c/tm1c mice. Example traces and pooled data illustrating ATPA (1 µM) induced currents in control (GFP expressing) but not in GFP-CRE expressing LA neurons in acute slices from Grik1tm1c/tm1c mice (GFP-CRE: n = 4, GFP: n = 4; ANOVA; p<0.05). GFP or GFP-CRE encoding AAV viruses were injected to the BLA of adult Grik1tm1c/tm1c mice, acute slices were cut for electrophysiological analysis 21 days after injection. Agonist induced currents were recorded as described in the main article.
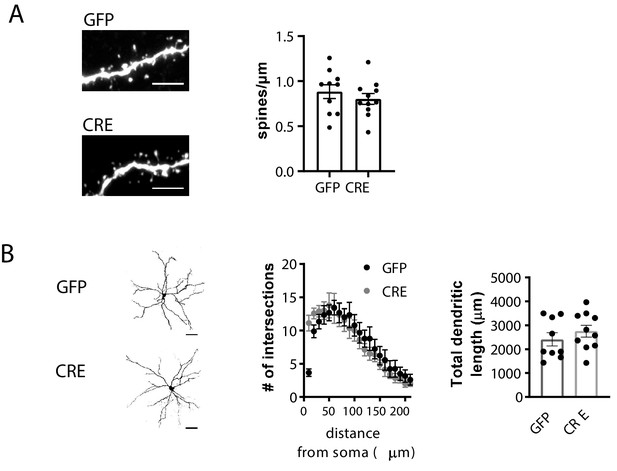
Morphological characterization of GFP and GFP-CRE expressing LA neurons in Grik1 cKO mice.
(A) GluK1 deficiency has no significant effect on the density of dendritic spines in LA neurons. AAV viruses encoding for GFP-CRE or GFP (control) were injected to the BLA of GluK1 cKO mice at P2-4 and morphological characterization was performed at P21. Example images and pooled data on the mean density of dendritic spines in GFP-CRE and GFP expressing neurons in the LA (GFP, n = 10(4); GFP-CRE, n = 11(4)). There is no significant difference between the groups (ANOVA, F(1,19) = 0.697, p=0.414). Scale bar, 5 µm. (B) GluK1 deficiency has no effect on mean dendritic length and branching in LA neurons. Sholl analysis of the biocytin filled LA neurons indicate that number of dendritic intersections and the total dendritic length is not different between GFP and GFP-CRE expressing neurons (GFP, n = 9 (4); GFP-CRE, n = 10(4); two-way ANOVA of dendritic intersections F(12,192)=1.262, p=0.24; dendritic length ANOVA F(1,17)=0.874, p=0.363). Confocal images (z-stack) illustrate the morphology of the dendritic tree. Scale bar, 50 µm. The n number refers to the number of experiments, followed by the number of animals in brackets.
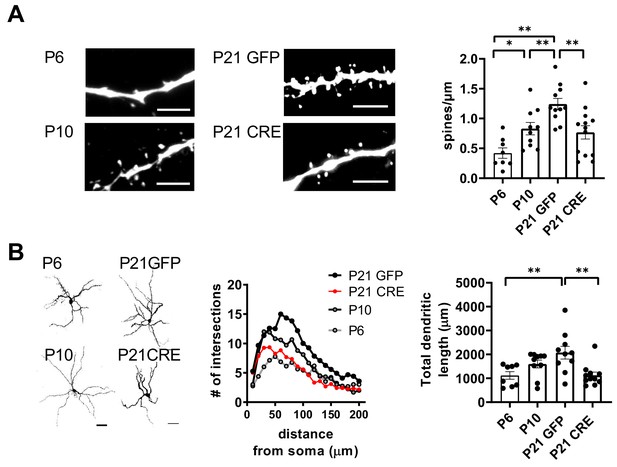
Development of CeA neuron morphology in control mice and in cKO mice lacking GluK1 in the BLA.
(A) Morphological characterization of biocytin filled CeA neurons from WT mice (P6 and P10) and P21 Grik1 cKO mice, injected with GFP-CRE or GFP (control) encoding AAV viruses to the BLA at P2-4. Example images and pooled data [P6 n = 8 (3); P10 n = 10(3); P21 GFP n = 11(4); P21 GFP-CRE n = 13(4)] illustrate developmental increase in spine density from P6 to P10 and from P10 to P21 in control mice (ANOVA F(3,38)=9.59, p<0.001; Holm-Sidak post-hoc comparison, P6 vs P10 t = 2.567, p=0.014; P10 vs P21 GFP t = 2.803, p=0.008). Following GluK1 inactivation, the spine density at P21 remains similar to P10 WT (Holm-Sidak, P6 vs P21 CRE t = 2.298, p=0.027; P10 vs P21 CRE: t = 0.441, p=0.662). **p<0.01. Scale bar, 5 µm. The n number refers to the number of experiments, followed by the number of animals in brackets. (B) Sholl analysis of the same neurons. The number of dendritic intersections and the total dendritic length is significantly increased during development from P6 to P21 in control mice (two-way ANOVA for dendritic intersections, F(19,418)=2.89, p<0.0001). Following GluK1 inactivation, the mean dendritic length and branching at P21 remains similar to P6–P10 WT’s (dendritic length, ANOVA F(3,35) = 5,724, p=0.003; Holm-Sidak post-hoc comparison P6 vs P10, t = 1.702, p=0.098; P10 vs P21 GFP, t = 1.771, p=0.085; P6 vs P21GFP t = 3.371, p=0.002; P6 vs P21CRE, t = 0.011, p=0.991; P10 vs P21CRE t = 1.835, p=0.075. Dendritic intersections, P10 vs P21CRE, two-way ANOVA F(11,240)=0.728,p=0.71). Confocal images (z-stack) illustrating the morphology of the dendritic tree. **p<0.01. Scale bar, 50 µm.
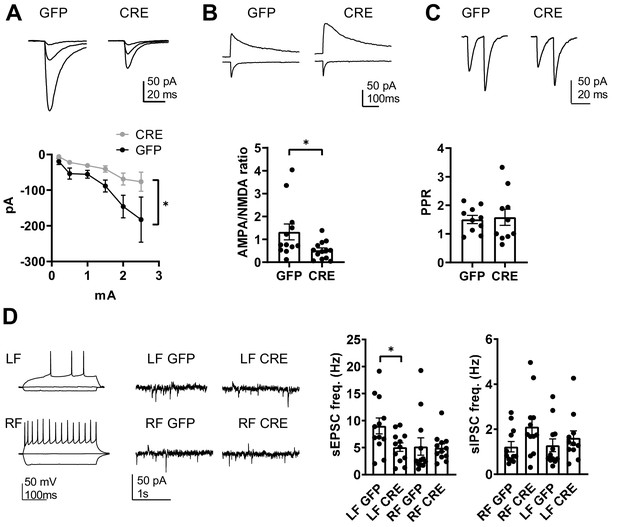
GluK1 deficiency in the amygdala perturbs glutamatergic transmission at LA-CeLA and associates with lower E/I ratio at CeLA late-firing neurons.
(A) Input – output characteristics of EPSCs, recorded from CeLA neurons in response to LA stimulation. Recordings for all the data presented in panels A-D were made in acute slices from P21 Grik1 cKO mice, injected with GFP or GFP-CRE encoding AAV viruses in BLA at P2-4. The EPSCs were significantly smaller in slices where GluK1 was locally inactivated as compared to controls. Example traces for responses evoked by three different stimulation intensities (0.2, 1 and 2.5 mA) for both groups and pooled data (GFP, n = 9[5], GFP-CRE n = 8[5], *p<0.05). (B) Example traces and pooled data illustrating the amplitude ratio of AMPAR and NMDAR–mediated EPSCs, evoked by LA stimulation in CeLA neurons. AMPA/NMDA ratio was significantly smaller in GFP-CRE group as compared to GFP controls (GFP n = 12[5], GFP-CRE n = 13[5], *p<0.05). (C) Paired –pulse ratio of LA-CeLA EPSCs was not different between GFP-CRE and GFP slices. Example traces and averaged data on paired-pulse ratio (PPR) of EPSCs (GFP, n = 10[4], GFP-CRE n = 10[4]). (D) Example traces of firing pattern of late-firing (LF) and regular-firing (RF) neurons in CeLA (P21) in response to depolarizing current pulses and recordings of spontaneous IPSCs and EPSCs under conditions where the sIPSCs are seen as outward currents and the sEPSCs are directed inwardly. sEPSC frequency was lower in LF neurons in CeLA in animals infected with GFP-CRE encoding AAV viruses at P2-4 as compared to GFP controls. Pooled data on frequencies of sEPSCs and sIPSCs in RF and LF neurons in CeLA (n = 12[4] for both groups, *p<0.05).
-
Figure 7—source data 1
Raw data for the results shown in the Figure 7.
- https://cdn.elifesciences.org/articles/52798/elife-52798-fig7-data1-v2.xlsx
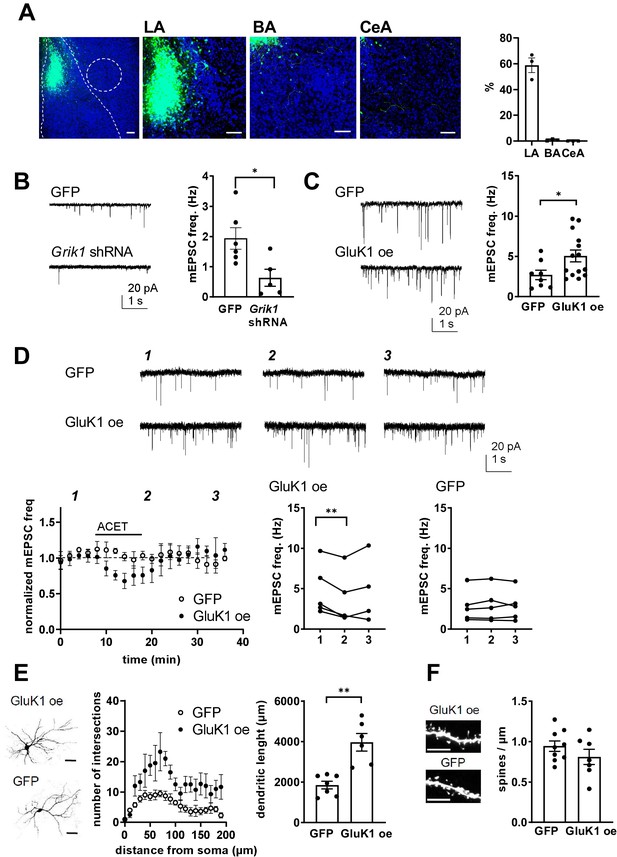
Presynaptic GluK1 KARs guide development of glutamatergic synaptic input to CeA.
(A) Image showing GFP expression specifically in the LA at P11, after in vivo injection of Grik1 shRNA/GFP encoding lentivirus at P5. The inset panel shows a magnified image illustrating GFP expression on average of 59 ± 5% of DAPI labeled (blue) cells in the LA, while practically no expression was detected in the BA (0.7 ± 0.4%) and CeA (0.1 ± 0.4%). Scale bar, 100 µm. (B) Example traces and pooled data showing that shRNA-mediated GluK1 knockdown in the LA results in lower mEPSC frequency in the CeA as compared to controls. Grik1 shRNA or mock lentivirus was injected to LA at P4-5 and the recordings were made from CeA neurons at P10-11 (GFP; n = 6 (5); Grik1 shRNA/GFP: n = 5 (3) *p<0.05). (C) Slices with GluK1 overexpression (GluK1 oe) in the LA show a significantly higher mEPSC frequency in the CeA as compared to controls (GFP). In vivo transduction was done at P6-9 and recordings were made at P14-15. GluK1 oe, n = 14(5), GFP n = 8 (4); *p<0.05. (D) GluK1 oe in the LA recapitulates the developmental phenotype of presynaptic KAR activity. Example traces from the time points indicated (1-3) and time course plots of pooled data illustrate that in GluK1 oe slices, ACET application causes a significant decrease in mEPSC frequency in the CeA neurons (n = 5 (3), p<0.05). At this developmental stage (P14-15), ACET has no effect on mEPSCs in control (GFP infected) slices (n = 5 (3)). Graphs on the right show mEPSC frequency under control conditions (1), in the presence of KA (2), and after wash-out of the drug (3) in individual cells. *p<0.05, paired t-test. (E) GluK1 oe results in an increase in dendritic length and branching. Sholl analysis of the biocytin filled CeA neurons indicate that number of dendritic intersections and the total dendritic length is significantly increased in slices with GluK1 oe in the LA. Pooled data (GluK1 oe, n = 6 (3); GFP n = 7 (3)) and confocal images (z-stack) illustrating the morphology of the dendritic tree. **p<0.01. Scale bar, 50 µm. (F) GluK1 oe does not influence the density of dendritic spines in CeA neurons. Pooled data of the mean density of dendritic spines (GluK1 oe n = 7 (4); GFP, n = 9 (4)). Example images shown on the right. Scale bar, 5 µm.
-
Figure 8—source data 1
Raw data for the results shown in the Figure 8.
- https://cdn.elifesciences.org/articles/52798/elife-52798-fig8-data1-v2.xlsx
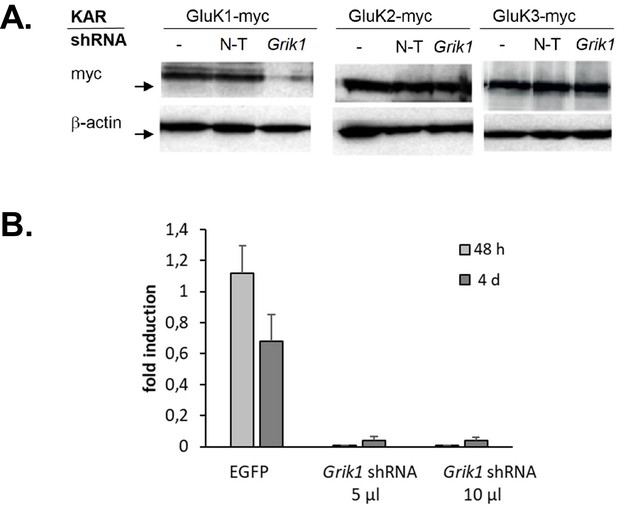
Validation of the Grik1 shRNA.
(A) A western blot illustrating that the Grik1 shRNA strongly inhibits expression of GluK1-myc but not GluK2-myc or GluK3-myc in HEK293T cells. (B) RT-qPCR results showing that the lentiviral Grik1 shRNA construct blocks expression of endogenous GluK1 in dorsal root gangion (DRG) neurons within 48 hr from infection.
Tables
Real-time PCR primers
| Target | Forward | Reverse | Size bp |
|---|---|---|---|
| Grik1 | ATGTGACGCAGAGGAACTGC | GCAGTTGAAGAATGGCAATCG | 126 |
| Grik2 | GTTTGTTACACAGCGGAACTG | CAGCTGAAGAATTGCTATGGTG | 127 |
| Grik3 | CATCGATTCCAAGGGCTACG | CGCCACCACTTCTCCTTCAT | 126 |
| Grik4 | GACACCAAGGGCTATGGGAT | ACCACTTCCGCTTCAGAATC | 118 |
| Grik5 | AGTACGGCACTATCCACGCT | CTCCTCTGTGCTCTTGACGA | 128 |
| Neto1 | TCATAGAAGCTGCCCCAAGG | AAGCCAAAGGGTCCATCTCG | 118 |
| Neto2 | TTTGGAAGCTGCTCCTCGTC | TCCAAGTGATCAAACCGGCA | 93 |
| Gadph | CAGTGCCAGCCTCGTCTCATA | TGGTAACCAGGCGTCCGATA | 79 |





