The functional organization of excitation and inhibition in the dendrites of mouse direction-selective ganglion cells
Figures
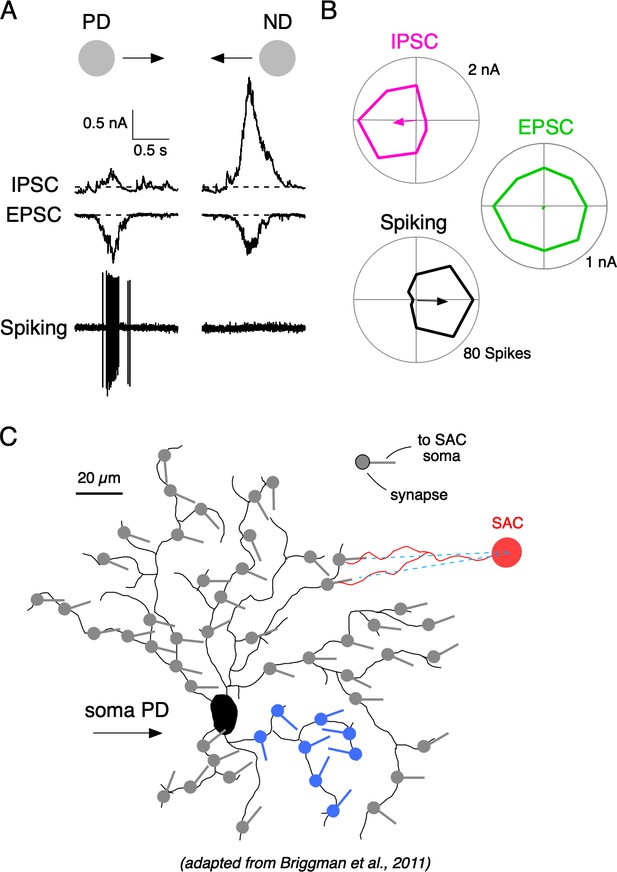
Synaptic mechanisms underlying direction selectivity in DSGCs.
(A) Spiking, IPSCs, and EPSCs recorded from the soma of an ON-OFF DSGC in response to a positive contrast spot (200 µm diameter; 500 µm/s) moving in the DSGC’s preferred (PD) or null direction (ND). (B) Polar plots of the peak amplitudes of the IPSC, EPSCs, and spiking responses evoked by spots moving in eight directions. Arrows indicate the vector sum angle, and its radius indicates the strength of the tuning. (C) Reconstructed DSGC overlaid with vectors representing the orientation of some of the presynaptic starburst amacrine cell (SAC) dendrites, which supply inhibition to it (adapted from SBEM reconstructions from Briggman et al., 2011; see their Figure 5). The orientations of SAC dendrites can predict the PD of DSGCs, because SACs release GABA maximally for stimuli moving from their soma towards their dendritic tips. The known anatomy predicts variable and sometimes disorganized local tuning in DSGCs (blue region).
-
Figure 1—source data 1
Currents and spiking responses in DSGCs.
- https://cdn.elifesciences.org/articles/52949/elife-52949-fig1-data1-v2.xlsx
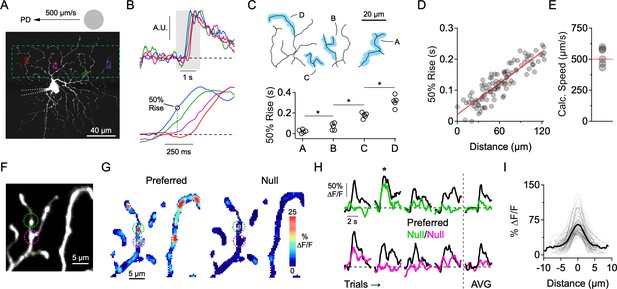
Independent activity in dendritic branches in the presence of NaV blockers.
(A) 2-photon image of the ON-stratifying dendritic arbor of an ON-OFF DSGC, which was filled with a Ca2+ indicator (Oregon Green Bapta-1, 200 µM), and a NaV channel blocker (QX-314, 2 mM) through a patch electrode. The dotted box indicates the imaging area. (B) Top, Normalized Ca2+ signals for the four regions of interest (ROIs) demarcated in A. Bottom, expanded view of the shaded region, showing clear differences in onset time at the different ROIs. (C) 50% rise time from each of the four dendritic branches outlined in the above reconstruction. Asterisks indicate p<0.05, t test. (D) In an another cell, 50% rise time of the Ca2+ signals versus lateral distance along the stimulus trajectory for many small ROIs that were placed throughout the dendritic tree. The red line is a linear fit to the data. (E) The slope of the linear fit predicts the speed of the stimulus (524 ± 28 µm/s), which was nominally set to be 500 µm/s. This data set included DSGCs that were recorded with (n = 3) and without (n = 3) the presence of the NMDA receptor antagonist D-AP5 (50 µM). No differences were found in terms of the onset timing of the Ca2+ signals, so the data sets were combined. (F) 2-photon image of a section of dendrites from an ON-OFF DSGC, with two ROIs demarcated. (G) Spatial map of the peak ∆F/F for preferred (left) and null (right) motion. (H) Left, Time-varying Ca2+ responses over multiple trials extracted from the 2 ROIs in F (green and magenta). Right, Average signals over four trials. The ∆F/F maps in G are from the second trial, in which a strong Ca2+ signal in the green ROI (top) was observed during null motion. (I) Line profiles of the ∆F/F across active dendritic sites during null motion. The solid line is the average line profile, and the thin lines are from individual ROIs. Gaussian fits to the line profiles had an average FWHM of 3.0 ± 1.2 µm (µ ± s.d., n = 61 ROIs from 7 cells).
-
Figure 2—source data 1
Calcium responses and rise time measurements.
- https://cdn.elifesciences.org/articles/52949/elife-52949-fig2-data1-v2.xlsx
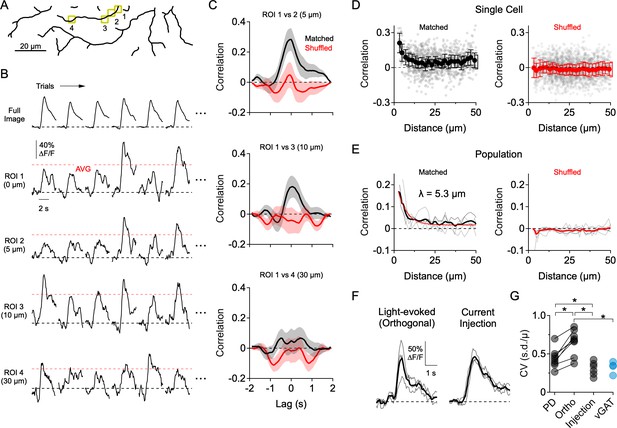
Dendritic Ca2+ signals form independently over small dendritic regions.
(A) Reconstruction of the imaging area in the ON arbor of an ON-OFF DSGC. (B) Ca2+ signals over 30 consecutive trials in response to motion in a direction orthogonal to the preferred-null axis (only six trials are shown). Ca2+ signals were extracted from the ROIs highlighted in A, or from the entire arbor within the imaging window (Full Image). Red dotted lines indicate the average peak Ca2+ signal over all trials. (C) Cross-correlation of the mean-subtracted residual Ca2+ signals between ROI one and the other highlighted ROIs, each of which is at an increased distance away. Correlations were performed using matched trials (black) or shuffled trials (red). (D) Peak correlation versus distance between the ROIs for matched (black) and shuffled trials (red) in a single DSGC. Faded data points are individual correlation values, solid points are the average (± s.e.m.) over 2 µm bins. (E) Peak correlation versus distance for a population of 7 ON-OFF DSGCs for matched and shuffled trials. Faded lines are the average correlation versus distance data for single cells, bold lines are the population average. For matched trials, the thin red line is an exponential fit, with a space constant of λ = 5.3 µm. (F) Light-evoked Ca2+ signals from an example ROI in response to orthogonal motion compared to amplitude-matched Ca2+ signals evoked by current injection at the soma (measured in synaptic blockers: 50 µM DL-AP4, 20 µM CNQX, 10 µM UBP-310). Thin lines are individual trials; thick lines are the average. (G) Coefficient of variance (standard deviation divided by the mean) for the peak Ca2+ signals evoked by preferred motion, orthogonal motion, somatic current injection, and in response to motion in a mouse that has the vesicular GABA transporter knocked out of SACs (Slc32a1fl/fl::ChatCre; vGAT KO). Asterisks indicate p<0.05, t-test.
-
Figure 3—source data 1
Calcium responses, cross-correlation, and variability measurements.
- https://cdn.elifesciences.org/articles/52949/elife-52949-fig3-data1-v2.xlsx
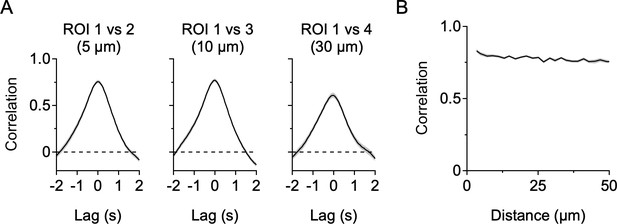
Stimulus-driven correlations between DSGC dendritic sites.
(A) Correlations between the raw Ca2+ signals from the ROIs shown in Figure 3 (without mean subtraction). (B) Correlation between ROIs as a function of cable distance. Here, since Ca2+ signals arise at all sites during the period of stimulus presentation, the spatial decay of the site-to-site correlations is minimal.
-
Figure 3—figure supplement 1—source data 1
Stimulus correlation measurements.
- https://cdn.elifesciences.org/articles/52949/elife-52949-fig3-figsupp1-data1-v2.xlsx
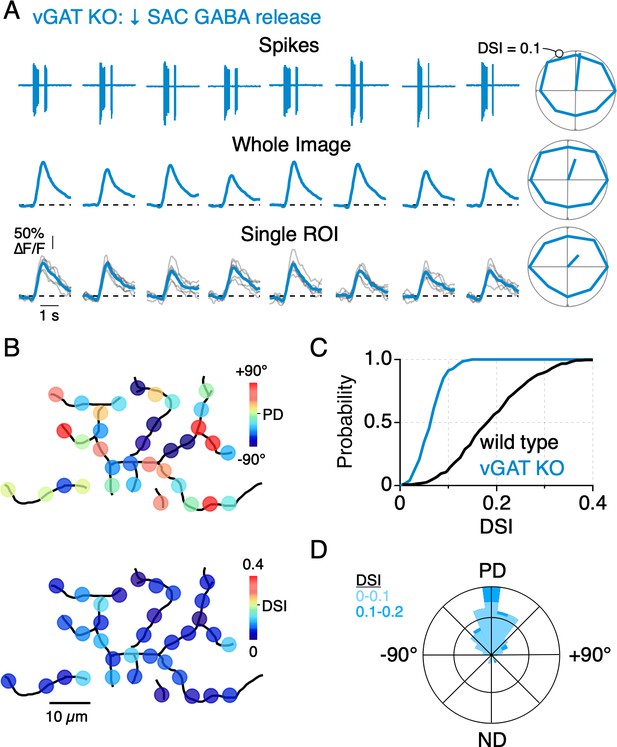
Dendritic Ca2+ signals are untuned when SAC inhibition is compromised.
(A) Spiking response and Ca2+ signals from a genetically labeled ON-OFF DSGC in vGAT KO mice, in which the vesicular GABA transporter is knocked out of SACs, preventing them from releasing GABA to DSGCs (Slc32a1fl/fl::ChatCre). Right, Polar plots of the spike rate and the peak Ca2+ responses over the whole image and over a small ROI. Lines extending from the origin indicate the preferred direction and the DSI (note, DSI scale is from 0 to 0.1). (B) Maps of the PD (top) and DSI (bottom) at dendritic ROIs in vGAT KO mice. (C) Cumulative distribution of the DSI from wild type and vGAT KO mice. (D) Rose plot of the preferred direction of dendritic ROIs in vGAT KO mice. Colors indicate the proportion of ROIs within different ranges of DSI. Note. subtle asymmetries present in the spiking responses, which would usually be considered ‘non-directional’ (Pei et al., 2015), were also present in the dendritic Ca2+ responses measured in the absence of NaV activity. Although the Ca2+ signals extracted from single ROIs in the dendritic tree were weakly tuned for direction, they encoded a similar direction to the spiking response, indicating that their weak tuning did not arise from technical noise sources.
-
Figure 3—figure supplement 2—source data 1
Spiking responses, calcium responses, and tuning in DSGCs in the vGAT KO mouse.
- https://cdn.elifesciences.org/articles/52949/elife-52949-fig3-figsupp2-data1-v2.xlsx
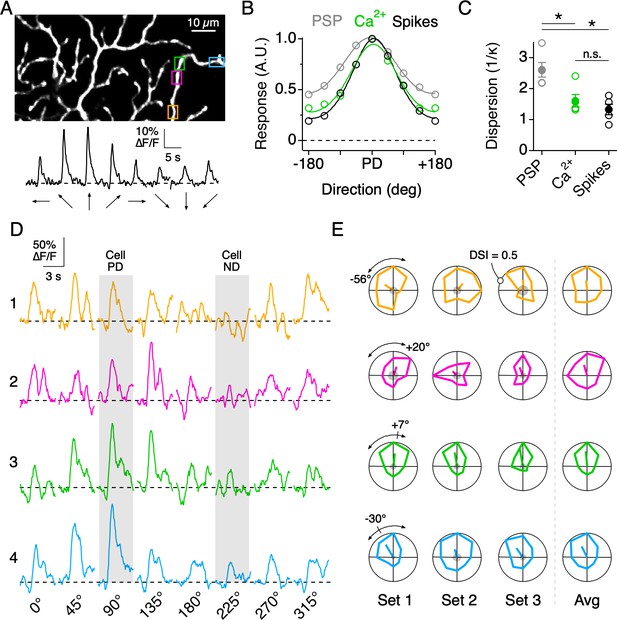
Directional tuning in DSGC dendrites in the absence of NaV activity.
(A) Top, Imaging area of the ON arbor of an ON-OFF DSGC. Bottom, Ca2+ signals over the entire imaging window in response to a positive contrast spot moving in eight different directions. (B) The average tuning curves of the spiking response, subthreshold somatic voltage (PSP), and dendritic Ca2+ signals (n = 7 cells). Solid lines indicate Von Mises fits to the data. The PD is normalized to that of the spiking response. (C) The value 1/κ, extracted from the Von Mises fits, is a measure of tuning width, or angular dispersion, for circular data. The tuning curve from the somatic voltage was significantly broader than both the Ca2+ signals and the spiking responses. Asterisks indicate p<0.05, t test. (D) Ca2+ signals over a single stimulus set (one set = 8 directions) at four nearby ROIs on the same dendritic branch, as demarcated by the colored boxes in A. The PD and ND, determined using the extracellular spiking response, are indicated by the shading. (E) Polar plots of the peak Ca2+ responses over consecutive stimulus sets, with the PD of each dendritic ROI indicated by the line extending from the origin, the length of which denoting the DSI, a measure of tuning strength (see Materials and methods). The arcs above the polar plot indicate the range in the preferred angle across these four sites on a single stimulus set. Right, the average polar plot for each ROI over the displayed stimulus sets.
-
Figure 4—source data 1
Calcium responses and tuning curves.
- https://cdn.elifesciences.org/articles/52949/elife-52949-fig4-data1-v2.xlsx
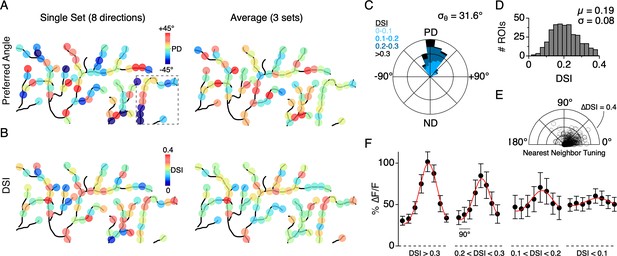
Distribution of directional tuning across the DSGC dendritic arbor.
(A) Maps of the preferred angle of each dendritic ROI, as measured over a single stimulus set (left), or averaged over multiple sets (right). The boxed area denotes the region used for the analysis presented in Figure 4. (B) Same as in A, but for the DSI. (C) Rose plot showing the distribution of the average PD for ROIs across the dendritic arbors of 7 ON-OFF DSGCs, relative to the PD of each DSGC’s spiking response (angular standard deviation, σθ = 31.6°). Colors indicate the proportion of ROIs within different ranges of DSI. (D) Distribution of DSIs for all dendritic ROIs (DSI = 0.19 ± 0.08, n = 353 ROIs from 7 cells). (E) Polar plot showing the difference in the DSI and PD between each ROI and its nearest neighboring ROI. (F) Average tuning curve for dendritic ROIs within different ranges of DSI. Note the amplitude of the null responses for weakly tuned sites (DSI <0.1) are on average larger than null responses measured for the strongly tuned sites (DSI >0.3).
-
Figure 5—source data 1
Vector angle and DSI measurements.
- https://cdn.elifesciences.org/articles/52949/elife-52949-fig5-data1-v2.xlsx
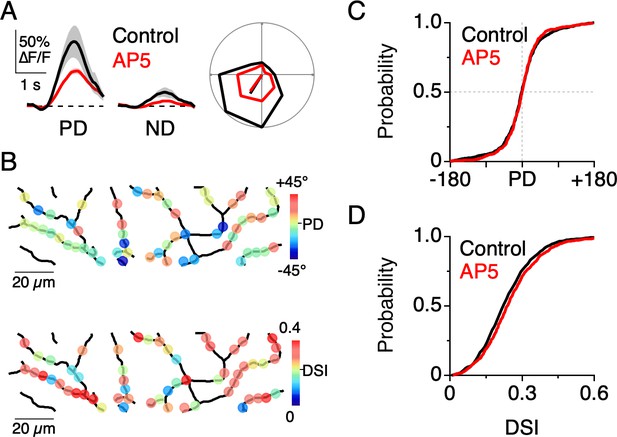
NMDA receptors amplify dendritic responses without altering direction selectivity.
(A) Example Ca2+ signals in response to preferred and null motion in control conditions (black) and in the presence of the NMDA receptor antagonist, D-AP5 (50 µM; red). Shading is ± s.e.m. Right, Example polar plot showing the directional tuning of dendritic Ca2+ signals before and after NMDA receptor blockade. (B) Map of the PD (top) and DSI (bottom) for dendritic ROIs during NMDA receptor blockade. (C) Cumulative distributions of the PD for dendritic ROIs. (D) Cumulative distributions of the DSI for dendritic ROIs.
-
Figure 6—source data 1
Calcium responses, and dendritic tuning distributions.
- https://cdn.elifesciences.org/articles/52949/elife-52949-fig6-data1-v2.xlsx
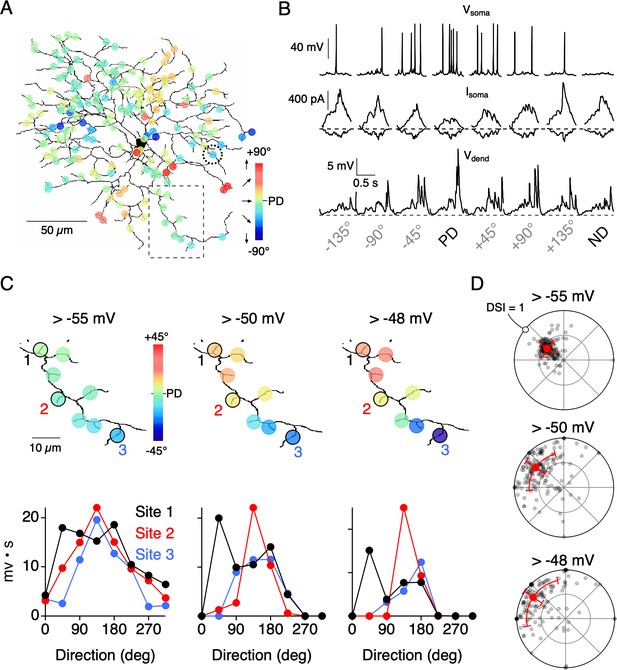
Threshold nonlinearities encourage dendritic compartmentalization.
(A) Reconstruction of an ON-OFF DSGC used in a multicompartmental in NEURON (Poleg-Polsky and Diamond, 2016; see Materials and methods for details). Simulated responses to motion in eight directions gave rise to directional tuning at 177 dendritic sites. These responses were measured in the absence of NaV activity. (B) Top, Somatic voltage (Vsoma) measured with NaV activity intact in response to 8 directions of motion. Middle and Bottom, somatic excitatory and inhibitory currents (Isoma), and voltage measured from a single dendritic site (Vdend; from site circled in A) in the absence of NaV activity. The PD of the integrated voltage responses (above a threshold of −55 mV) was used to compute the PDs shown in A. (C) Expanded view of the dendritic branch indicated by the boxed area in A, showing how the stringency of the voltage threshold (indicated above images) can affect the directional tuning in nearby dendritic sites. The integrated voltage at three sites is shown below. (D) Polar plots showing the distribution of PDs and DSIs measured over the dendritic tree, measured using different threshold values. Note, in the model all sites have the same average tuning, and the differences in tuning between sites only occur on single trial sets (eight directions) due to the stochastic nature of transmitter release.
-
Figure 7—source data 1
Dendritic voltage responses and directional tuning measurements.
- https://cdn.elifesciences.org/articles/52949/elife-52949-fig7-data1-v2.xlsx
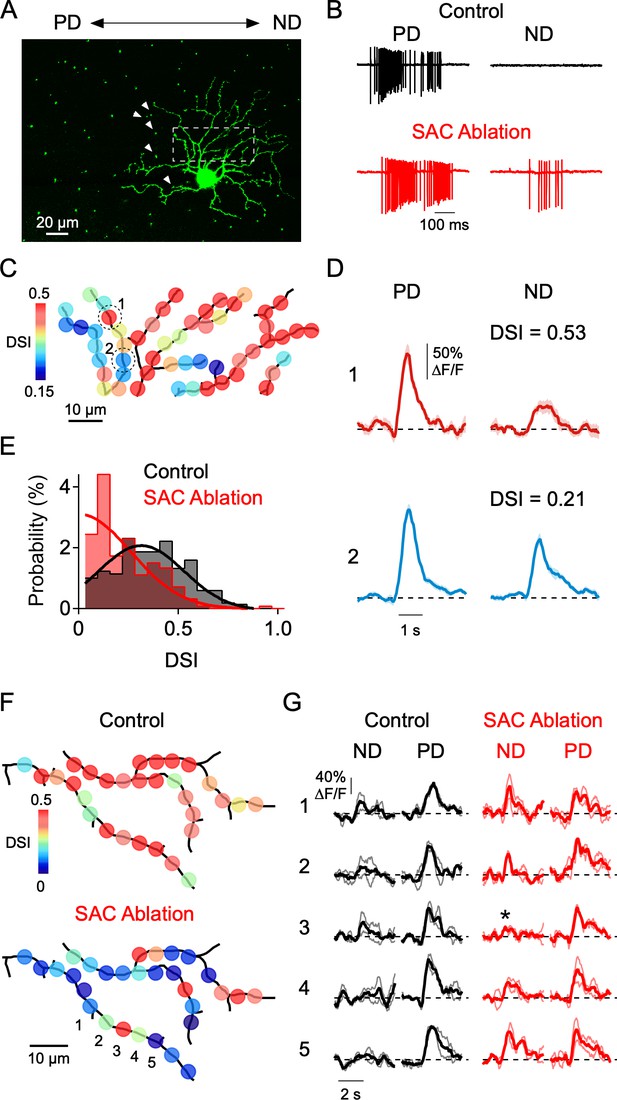
Local disruption of dendritic direction selectivity by SAC ablations.
(A) Image showing the mosaic of SAC somas labeled with a nuclear GFP marker surrounding a DSGC filled with OGB-1. Arrowheads indicate null-side SACs that were individually targeted for electrical ablation with an electrode (see Materials and methods). (B) Spiking responses of a DSGC before (black) and after SAC ablation (red). (C) DSI map of the imaged area (dotted square in A), after 5 SACs were ablated from the null side of its dendritic arbor. (D) Average Ca2+ signals (10 trials) in 2 ROIs (indicated in C) with strong and weak DSIs after SAC ablation. Shading is ± s.e.m. (E) Distribution of DSIs for a population of sites in 4 DSGCs after SAC ablation (red), compared with the distribution from DSGCs imaged under control conditions (black). Note, the DSI was calculated from only preferred and null stimulation, resulting in higher DSI values than those reported in Figures 4–5, where eight directions were used. Solid lines are Gaussian fits to the data. (F) Maps of the DSI before and after SAC ablation in the same DSGC. (G) Average Ca2+ signals from five ROIs (1-5) in F, before (black) and after (red) SAC ablation. Thin lines are individual trials, and thick lines are the average response. Asterisk denotes a strongly tuned dendritic site flanked by poorly tuned regions.
-
Figure 8—source data 1
Electrophysiology, calcium responses, and dendritic tuning measurements.
- https://cdn.elifesciences.org/articles/52949/elife-52949-fig8-data1-v2.xlsx
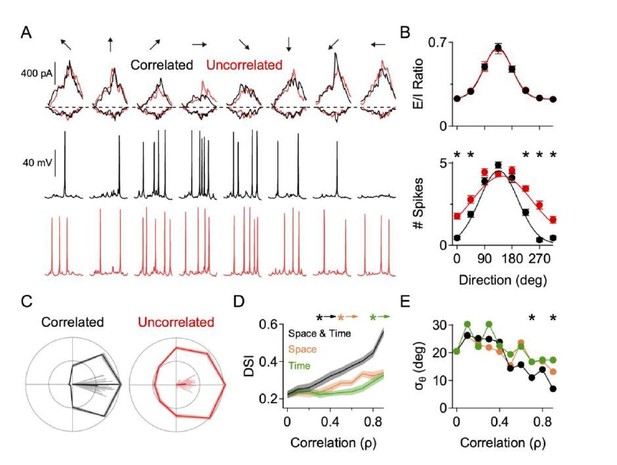
Correlated excitation and inhibition is critical for accurate and reliable direction selectivity.
These data are from a NEURON model (as described in the main manuscript) where the spatiotemporal correlation between the excitatory and inhibitory inputs was manipulated. Temporal correlation is the likelihood that, at a given dendritic site, that excitation and inhibition arrive at the same time. Spatial correlation is defined as the likelihood that, at a given site, if excitation arrives that inhibition will arrive at the same location. A, Top, EPSCs and IPSCs measured at the soma in correlated and uncorrelated states in response to 8 different simulated directions of motion. NaV activity is blocked for these recordings. Middle, Somatic spiking in response to correlated E/I inputs. Bottom, Somatic spiking in response to uncorrelated E/I inputs. Somatic and dendritic NaV were active for the spike recordings. Note, the somatic currents are roughly the same in correlated and uncorrelated conditions, but more null spikes arise in the uncorrelated case. B, Top, E/I ratio across direction (measured at the soma) is similar in the correlated and uncorrelated states. Bottom, Directional tuning of the spiking response in the correlated and uncorrelated states. Note the increase in null spikes during non-preferred motion for the uncorrelated state. C, Polar plot form and vector angles (lines) of the directional tuning in correlated and uncorrelated states. D, DSI as a function of spatial and temporal correlation strength. Here, the spatial and temporal correlation was modeled independently. E, Same as D, but for the angular standard deviation. * indicates statistical significance p < 0.05 (t-test)..
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (M. musculus) | B6.129S6-Chattm1(cre)lowl/J | PMID: 21284986 | RRID: IMSR_JAX:006410 | |
| Gene (M. musculus) | B6.129-Gt (ROSA)26Sortm1Joe/ | PMID: 18395835 | RRID: IMSR_JAX:008516 | |
| Gene (M. musculus) | Slc32a1tm1Lowl | PMID: 19160495 | RRID: IMSR_JAX:012897 | |
| Gene (M. musculus) | Trhr-EGFP | PMID: 14586460 | RRID: MMRRC_030036-UCD | |
| Gene (M. musculus) | Hb9-EGFP | PMID: 12176325 | RRID: IMSR_JAX:005029 |





