Targeting mir128-3p alleviates myocardial insulin resistance and prevents ischemia-induced heart failure
Figures
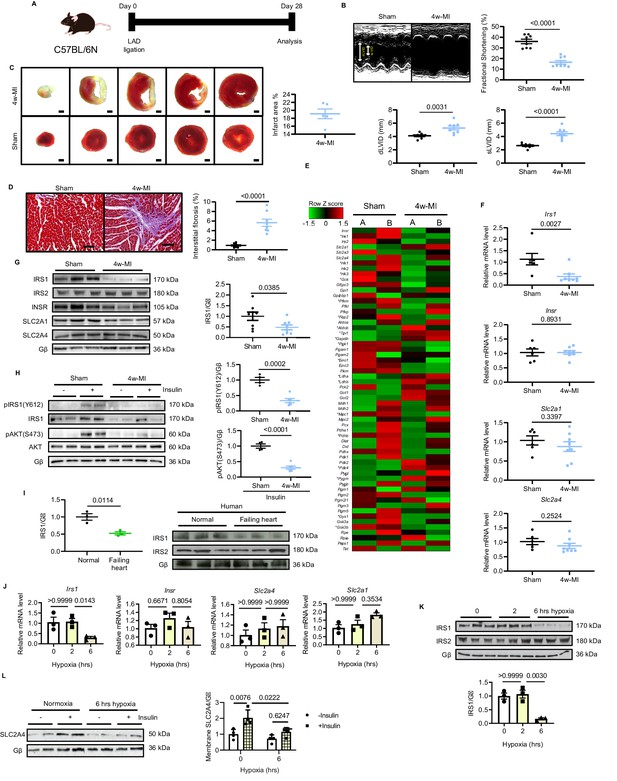
IRS1 is downregulated in failing hearts and prolonged hypoxia-stressed cardiomyocytes.
(A) Experimental design of male C57BL/6N mice subject to 4w-MI. (B) M-mode images captured by echocardiography determined cardiac dysfunction after 4w-MI, measured by fractional shortening (FS%) and end-diastolic and systolic left ventricular internal diameter (dLVID and sLVID) (N = 8 mice). (C) Infarct area (from one heart) detected by triphenyltetrazolium chloride (TTC) staining (N = 5 mice). Scale bar: 1 mm. (D) Interstitial fibrosis in the non-infarct region determined by Masson’s Trichrome staining (N = 8 mice). Scale bar: 100 μm. (E) RNA-sequencing (RNA-Seq) heat map depicts expression levels of the selected genes in the category of glycose pathways. * indicates genes identified as significantly different between sham and MI for four weeks (Log2 fold change according to adjusted P value < 0.05) (N = 2 mice). (F–G) qPCR validation and immunoblot analyses of IRS1, IRS2, INSR, SLC2A1 (also known as GLUT1), and SLC2A4 (also known as GLUT4) in the myocardium (N = 6–8 mice). (H) Insulin-dependent pathway evaluated by immunoblots after insulin injection (1 U/kg for 30 mins). (I) IRS1 and IRS2 protein level in failing human hearts (N = 3 mice). (J) qPCR detection of Insr, Irs1, Slc2a1, and Slc2a4 in ARCMs under hypoxia (N = 3 experiments). (K) IRS1, but not IRS2, was reduced in ARCMs under long-term hypoxia (N = 3 experiments). (L) Insulin stimulation-induced (0.1 µM, 30 mins) SLC2A4 membrane translocation in cardiomyocytes was diminished after 6 hr hypoxia (N = 4 experiments). Data presented as mean ± SEM. Comparisons between two groups were performed using two-tailed Student’s t-test. One-way or two-way ANOVA followed by Bonferroni post hoc tests were employed as appropriate.
-
Figure 1—source data 1
Insulin sensitivity and Irs1 downregulation in MI-induced heart failure and rat cardiomyocytes under prolonged hypoxic stress.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig1-data1-v2.xlsx
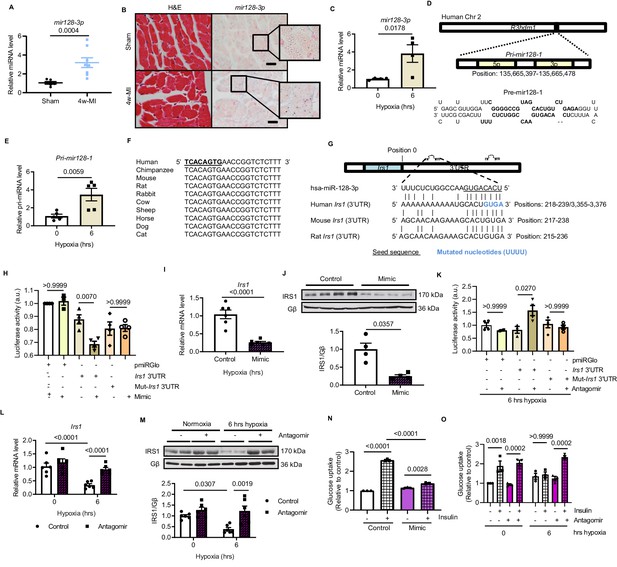
Hypoxia-upregulates mir128-3p and degrades Irs1.
(A) Mature mir128-3p was increased in 4w-MI myocardium (N = 6–8 mice). Data were normalized against mature mir191-5p expression. (B) In situ hybridization with digoxigenin-labeled mir128-3p probes (right) was accompanied with consecutive section stained by H and E (left) (N = 3–5 mice). Scale bar: 20 µm. (C) Upregulation of mir128-3p in ARCMs after 6 hr hypoxia (N = 4 experiments). (D) Location of Pri-mir128-1 hosted by human R3HDM1 gene and stem-loop structure of Pre-mir128-1. (E) Pri-mir128-1 level was determined by qPCR (N = 5 experiments). (F) Alignment of conserved mir128-3p sequence among mammals. (G) Sequences of mir128-3p conserved complementarity in human, mouse, and rat at Irs1 3’UTR. (H) Irs1 3’UTR luciferase reporter assay was performed in rat H9C2 cardiomyocytes transfected with pmiRGlo, pmiRGlo-Irs1 3’UTR, or mutant form (pmiRGlo-mut-Irs1 3’UTR). Luciferase activity was suppressed by mir128-3p mimic (N = 4 experiments). (I–J) qPCR and immunoblots illustrated miR-128–3 p mimic repression of IRS1 in ARCMs (N = 4–6 experiments). (K) Irs1 3’UTR luciferase reporter activity recovered by mir128-3p antagomiR (N = 4 experiments). (L–M) qPCR and immunoblots showed mir128-3pantagomiR maintained IRS1 expression in hypoxia-treated ARCMs (N = 4–6 experiments). Glucose uptake capacity by insulin stimulation (0.1 µM for 30 mins) was measured by the accumulation of 2-deoxy-D-glucose in ARCMs in the presence of (N–O) mir128-3p mimic or mir128-3p antagomiR (N = 3 experiments). Data presented as mean ± SEM. Comparisons between two groups were performed using two-tailed Student’s t-test. One-way or two-way ANOVA followed by Bonferroni post hoc tests were employed as appropriate.
-
Figure 2—source data 1
Study of mir128-3p expression under hypoxia and its function in regulating Irs1.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig2-data1-v2.xlsx
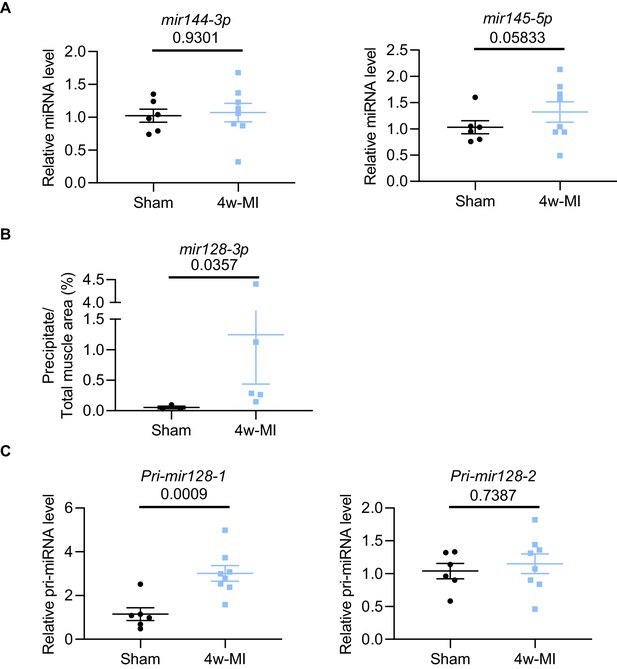
MiRNA analysis in 4w-MI myocardium.
(A) Mature mir144-3p and mir145-5p in 4w-MI myocardium were measured using TaqMan Advanced miRNA qPCR (N = 6–8 mice). Data were normalized against mature mir191-5p expression. (B) In situ hybridization with digoxigenin-labeled mir128-3p probes was quantified using ImageJ software (N = 3–5 mice). Groups were compared using the Mann-Whitney test. (C) Pri-mir128-1 and Pri-mir128-2 measurement in 4w-MI myocardium using TaqMan Gene expression (N = 6–8 mice). Data were normalised against 18S expression. Data presented as mean ± SEM. Groups were compared using two-tailed Student’s t-test.
-
Figure 2—figure supplement 1—source data 1
MiRNA analysis in 4w-MI myocardium.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig2-figsupp1-data1-v2.xlsx

Mir128-3p level in rat cardiomyocytes after hypoxia.
Mature mir128-3p was measured using TaqMan Advanced miRNA qPCR (N = 6 experiments). Data were normalized against mature mir191-5p expression. Data presented as mean ± SEM. Groups were compared using one-way ANOVA followed by Bonferroni post hoc tests.
-
Figure 2—figure supplement 2—source data 1
Mir128-3p level in rat cardiomyocytes after hypoxia.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig2-figsupp2-data1-v2.xlsx
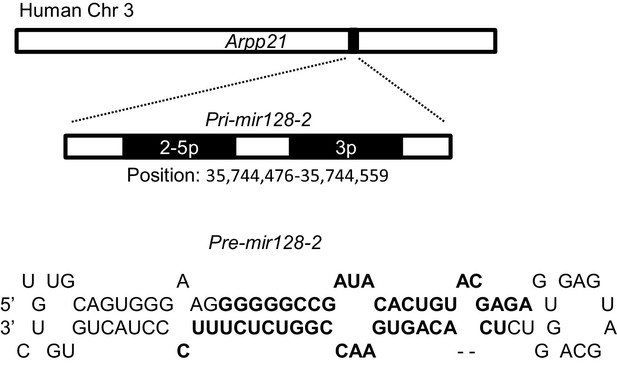
Pri-mir128-2 location and structure.
Location of Pri-miR-128-2 hosted by human ARPP21 gene (GRCh38:3:35638945:35794496:1, Ensembl ENSG00000172995) and the stem-loop structure of Pre-miR-128-2.
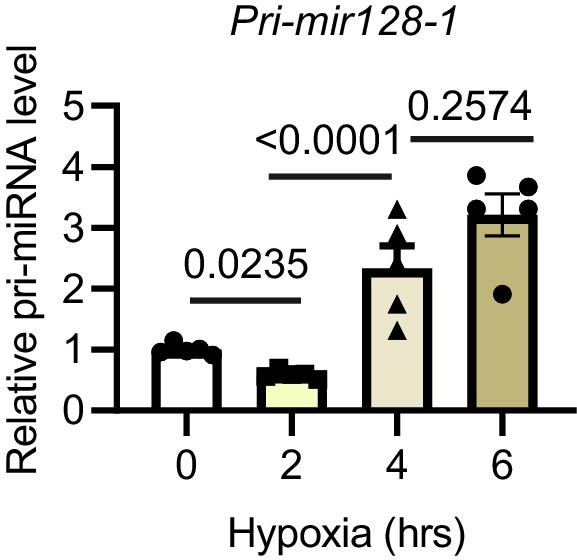
Pri-mir128-1 level in rat cardiomyocytes after hypoxia.
Pri-mir128-1 in cardiomyocytes was performed using TaqMan Gene expression (N = 5 experiments). Data were normalised against 18S expression. Data presented as mean ± SEM. Groups were compared using one-way ANOVA followed by Bonferroni post hoc tests.
-
Figure 2—figure supplement 4—source data 1
Pri-mir128-1 level in rat cardiomyocytes after hypoxia.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig2-figsupp4-data1-v2.xlsx
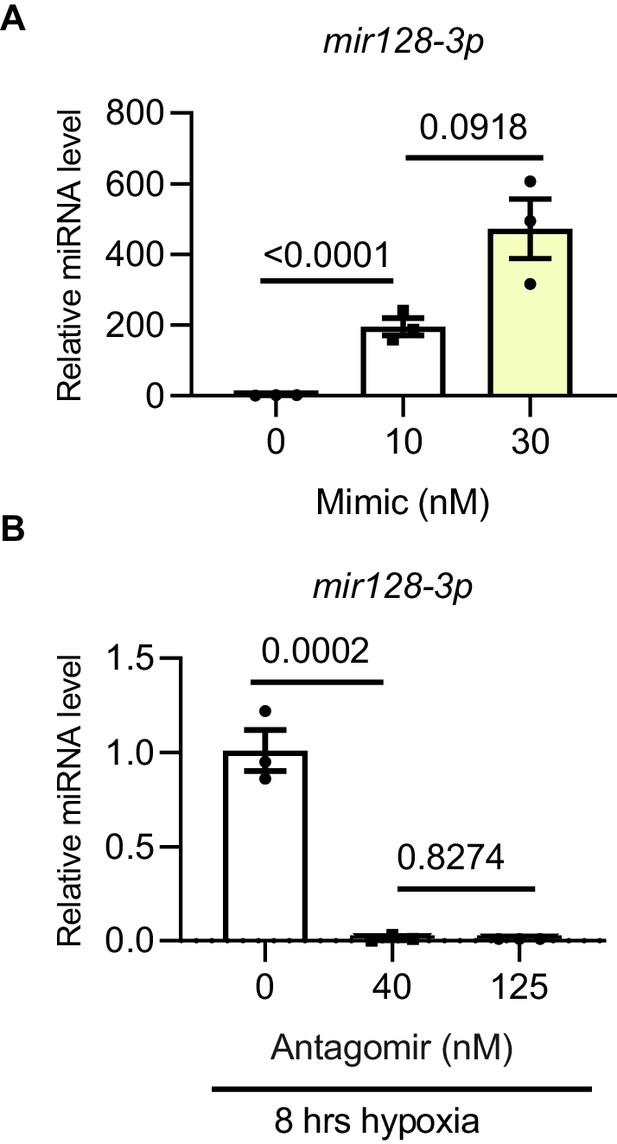
Mir128-3p overexpression and inhibition optimization.
Validation of mir128-3p mimic and antagomir. (A) mir128-3p mimic and (B) antagomir validation was carried out on H9C2 cells at different concentrations. Mature miRNA was measured by TaqMan Advanced miRNA qPCR. Data were normalized against mature mir191-5p expression (N = 3 experiments). Data presented as mean ± SEM. Groups were compared using one-way ANOVA followed by Bonferroni post hoc tests.
-
Figure 2—figure supplement 5—source data 1
Mir128-3p overexpression and inhibition optimization.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig2-figsupp5-data1-v2.xlsx
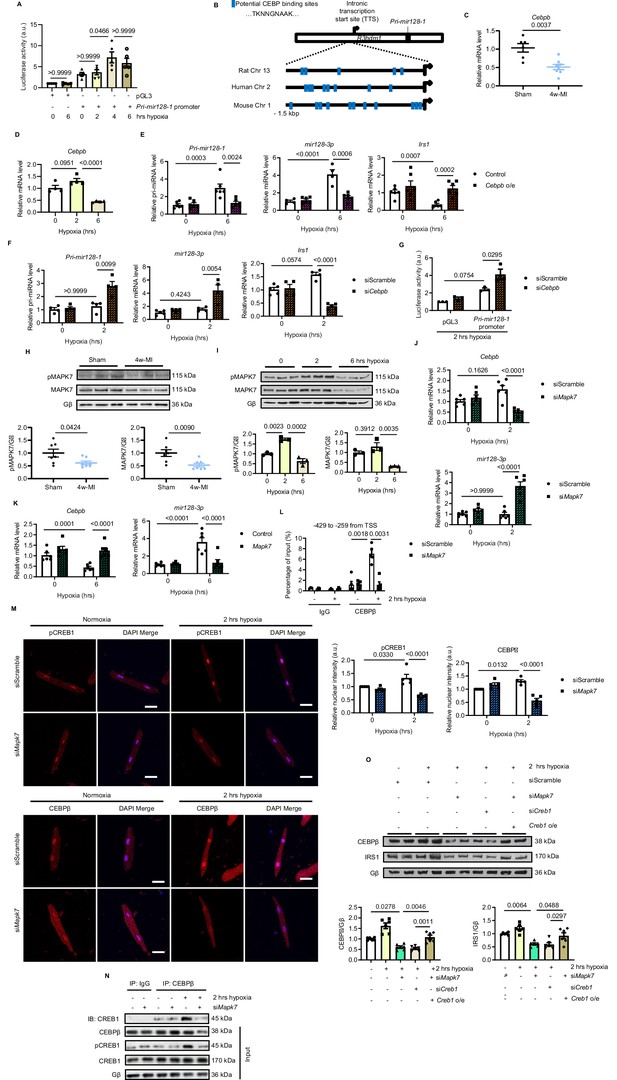
MAPK7 regulation of CEBPβ transcriptionally represses mir128-3p.
(A) Pri-mir128-1 promotor luciferase activity in H9C2 was increased by long-term hypoxia (N = 4 experiments). (B) Schematic figure showing CEBP consensus binding sites in the intronic promoter region between exon1 and exon2 of R3hdm1 gene conserved in human, rat, and mouse. (C–D) Cebpβ was downregulated in myocardium after 4w-MI (N = 6–8 mice) and ARCMs (N = 4 experiments) stressed by short and long-lasting hypoxia. (E–F) Measurements of Pri-mir128-1, mir128-3p and Irs1 were performed in ARCMs with CEBPβ overexpression (Cebpb o/e) under long-term hypoxia or Cebpb knockdown in response to short-term hypoxia (N = 4–6 experiments). (G) Pri-mir128-1 promotor luciferase activity was augmented by CEBPβ deficiency (N = 3 experiments). (H) MAPK7 (also known as ERK5) phosphorylation and expression were decreased in the myocardium four weeks after MI (N = 7 mice). (I) MAPK7 was activated by acute hypoxic stress, while it was declined by prolonged stimulation in ARCMs (N = 3 experiments). (J–K) qPCR analyses of Cebpb and mir128-3p in Mapk7-deficient or Mapk7-overexpressing (Mapk7 o/e) ARCMs (N = 6 experiments). (L) Chromatin Immunoprecipitation (ChIP) was performed on ARCMs DNA using anti-CEBPβ antibody (anti-IgG antibody as negative control) (N = 3–5 experiments). Hypoxia-induced CEBPβ binding to the proximal Pri-mir128-1 promoter was inhibited by MAPK7 deficiency. Data were normalized to the input chromatin. (M) Immunofluorescent staining of ARCMs demonstrated that hypoxia-triggered nuclear translocation of phosphorylated CREB1 or CEBPβ (red) was blocked by Mapk7 knockdown. DAPI was used for nuclear staining (blue) (N = 5 experiments). Scale bar: 30 µm. (N) Immunoprecipitation showed that hypoxia promoted association of phosphorylated CREB1 and CEBPβ in ARCMs; however, the affinity was reduced in the absence of MAPK7. IgG was used as negative control. (O) Decreased CEBPβ and reduced IRS1 were detected in the absence of MAPK7 or CREB1, while the reduction was rescued by CREB1 overexpression in Mapk7-knockdown ARCMs (N = 6 experiments). Data presented as mean ± SEM. Comparisons between two groups were performed using two-tailed Student’s t-test. One-way or two-way ANOVA followed by Bonferroni post hoc tests were employed as appropriate.
-
Figure 3—source data 1
Role of MAPK7 and CEBPβ in mir128-3p transcriptional regulation.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig3-data1-v2.xlsx
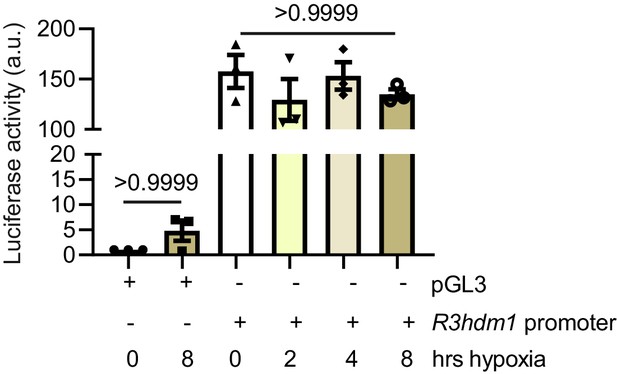
Hypoxia effect on R3hdm1 expression.
Luciferase reporter assay performed on H9C2 cells transfected with pGL3 or pGL3-R3hdm1 promoter (−1.5 kbp of transcription start site TSS downstream of R3hdm1 exon1) demonstrates that R3hdm1 promoter luciferase activity was not affected during different periods of hypoxia. Data were normalized to renilla expression and control (N = 3 experiments). Data presented as mean ± SEM. Groups were compared using one-way ANOVA followed by Bonferroni post hoc tests.
-
Figure 3—figure supplement 1—source data 1
Hypoxia effect on R3hdm1 expression.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig3-figsupp1-data1-v2.xlsx
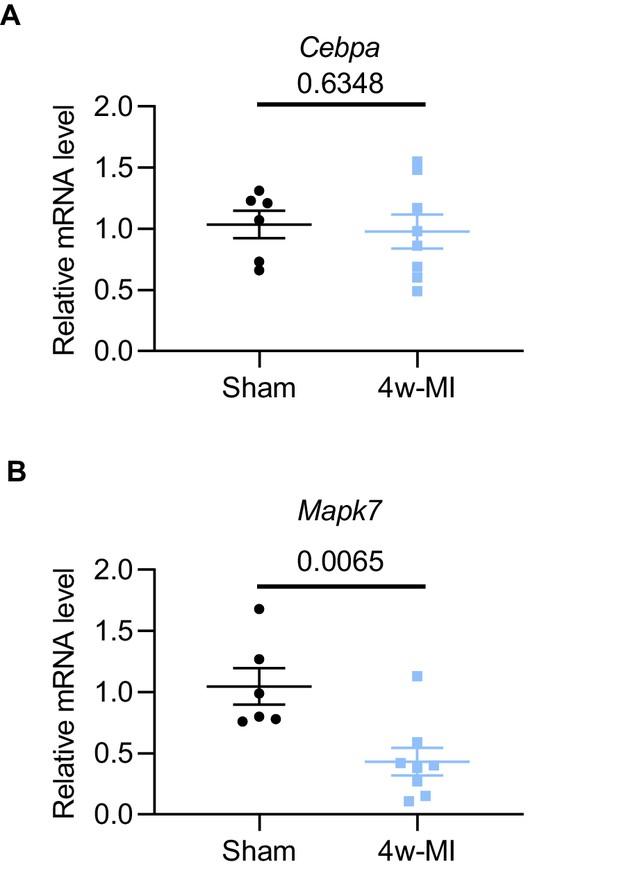
Mapk7 and Cebpa expression in 4w-MI myocardium.
(A) Cebpa transcript expression was not changed in 4w-MI myocardium, while (B) Mapk7 was decreased measured by qPCR (N = 6–8 mice). Data were normalized against 18S expression. Data presented as mean ± SEM. Groups were compared using two-tailed Student’s t-test.
-
Figure 3—figure supplement 2—source data 1
Mapk7 and Cebpa expression in 4w-MI myocardium.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig3-figsupp2-data1-v2.xlsx
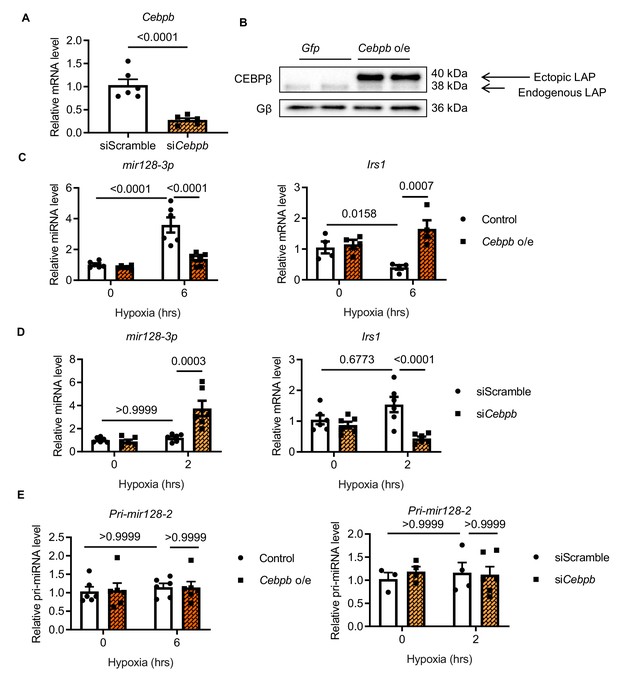
Role of CEBPβ in miR128-3p and Irs1 regulation in H9C2 cells.
CEBPβ repression of mir128-3p retains Irs1 in H9C2 cardiomyocytes. (A) Cebpb was knocked down more than 70% by siRNA transfection with Lipofectamine LTX (N = 6 experiments). (B) Human Cebpb overexpression (Cebpb o/e) in H9C2 cells showing LAP as the dominant isoform. qPCR detection of mir128-3p and Irs1 in H9C2 cardiomyocytes with (C) Cebpb overexpression under long-term hypoxia or (D) siRNA-mediated Cebpb knockdown under short-term stress (N = 5–6 experiments). (E) Pri-mir128-2 was not affected by Cebpb overexpression or knockdown (N = 4–6 experiments). Data were normalized against 18S expression. Data presented as mean ± SEM. Comparisons between two groups were performed using two-tailed Student’s t-test. Two-way ANOVA followed by Bonferroni post hoc tests was employed as appropriate.
-
Figure 3—figure supplement 3—source data 1
Role of CEBPβ in miR128-3p and Irs1 regulation in H9C2 cells.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig3-figsupp3-data1-v2.xlsx
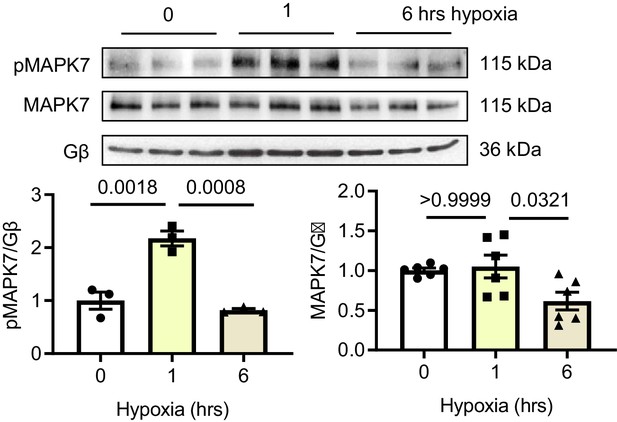
MAPK7 activation and expression in short and long-term hypoxia.
MAPK7 activation was observed in neonatal rat cardiomyocytes (NRCMs) in response to short-term hypoxia, whereas MAPK7 was decreased by prolonged stress. Quantification is provided (N = 3–6 experiments). Data presented as mean ± SEM. One-way ANOVA followed by Bonferroni post hoc tests was employed to compare groups.
-
Figure 3—figure supplement 4—source data 1
MAPK7 activation and expression in short and long-term hypoxia.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig3-figsupp4-data1-v2.xlsx
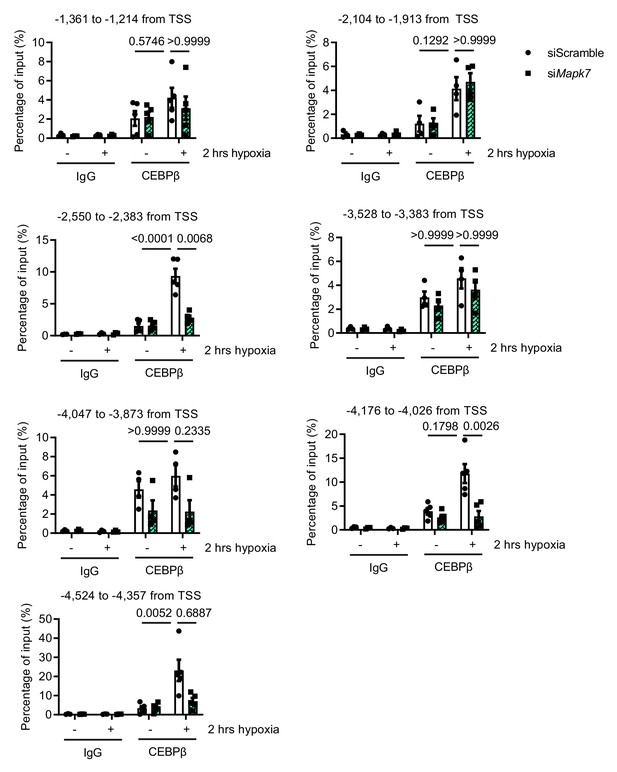
CEBPβ binding analysis to Pri-mir128-1 intronic promoter.
Chromatin immunoprecipitation (ChIP) assay was performed on ARCMs DNA using anti-CEBPβ antibody (anti-IgG antibody as the negative control). The enrichment of CEBPβ binding to different regions in Pri-mir128-1 intronic promoter localized between exon1 and exon2 of R3hdm1 was examined by qPCR in the presence and absence of Mapk7 in response to hypoxia. TSS: transcription start site. Data were normalized to input chromatin (N = 3–5 experiments). Data presented as mean ± SEM. Two-way ANOVA followed by Bonferroni post hoc tests was employed to compare groups.
-
Figure 3—figure supplement 5—source data 1
CEBPβ binding analysis to Pri-mir128-1 intronic promoter.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig3-figsupp5-data1-v2.xlsx
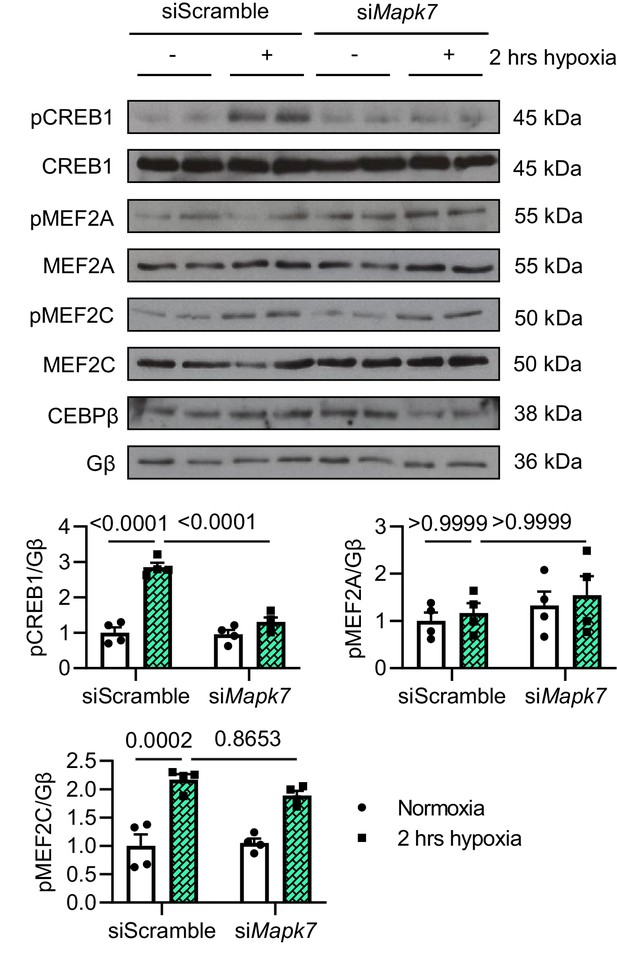
Analysis of MAPK7 downstream effectors after hypoxia.
Potential MAPK7 downstream effectors were analysed in ARCMs by immunoblots and quantification. CREB1 phosphorylation under short-term hypoxia was blocked by Erk5 deficiency (N = 4 experiments). Data presented as mean ± SEM. Two-way ANOVA followed by Bonferroni post hoc tests was employed to compare groups.
-
Figure 3—figure supplement 6—source data 1
Analysis of MAPK7 downstream effectors after hypoxia.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig3-figsupp6-data1-v2.xlsx
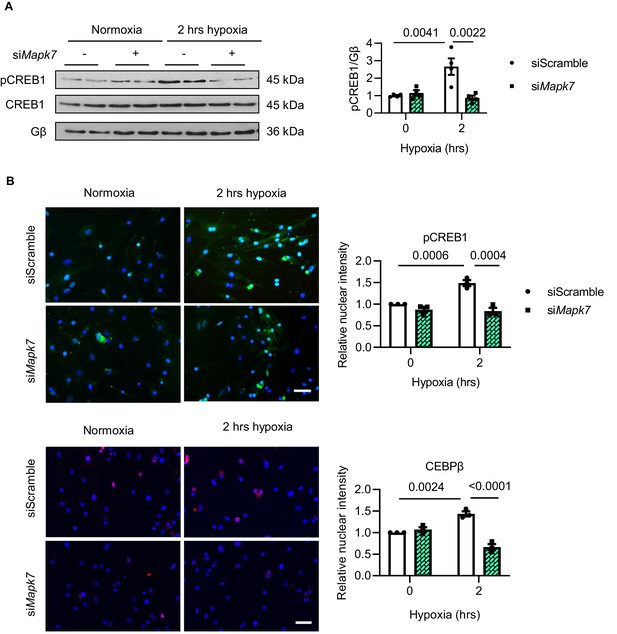
Hypoxia-induced CREB1 and CEBPβ nuclear translocation in NRCMs.
The impacts of Mapk7 knockdown on CREB1 activation and CEBPβ translocation to the nucleus in NRCMs in response to short-term hypoxia, which was determined by (A) immunoblots and (B) immunofluorescent detection of nuclear translocation of pCREB1 (green in upper panels) and CEBPβ (red in lower panels). DAPI (blue) stains nuclei (N = 3 experiments). Scale bar: 50 µm. Data presented as mean ± SEM. Two-way ANOVA followed by Bonferroni post hoc tests was employed to compare groups.
-
Figure 3—figure supplement 7—source data 1
Hypoxia-induced CREB1 and CEBPβ nuclear translocation in NRCMs.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig3-figsupp7-data1-v2.xlsx
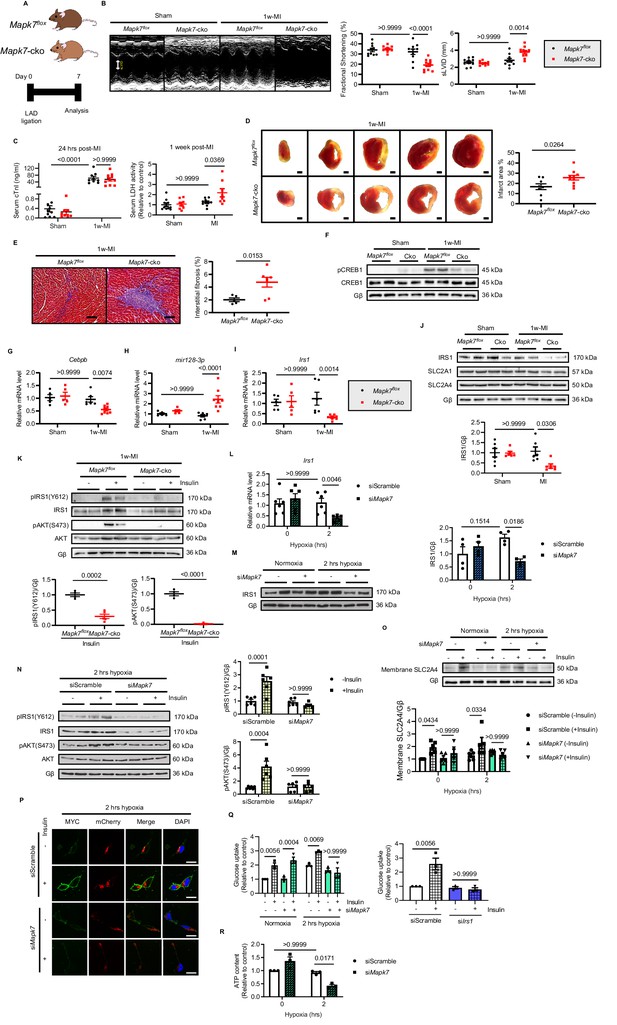
MAPK7 deficiency promotes cardiac insulin resistance and injury under hypoxic condition.
(A) Experimental design of male cardiac-specific Mapk7-deleted mice subject to 1w-MI. (B) Echocardiography determined detrimental cardiac function in Mapk7-cko mice post-MI, indicated by fractional shortening (%) and sLVID (N = 9–11 mice). (C–E) MAPK7 ablation led to increased serum cTnI (24 hr post-ligation) and LDH (one week post-ligation) concentration, TTC-detected infarct area (from one heart, scale bar: 1 mm), and interstitial fibrosis in the non-infarct region detected by Masson’s Trichrome (scale bar: 100 μm) (N = 8–10 mice). (F) Immunoblots analyses of phosphorylation and expression of CREB1 in the myocardium (N = 6 mice). (G–H) Real-time qPCR detected reduced Cebpb accompanied by upregulated mature mir128-3p due to Mapk7 deletion in response to MI (N = 5–8 mice). (I–J) Transcript and protein level of IRS1 was decreased by MAPK7 disruption (N = 5–8 mice). (K) Myocardial insulin sensitivity was assessed by phosphorylation of IRS1 and AKT after insulin injection (1 U/kg for 30 mins) (N = 5–6 mice). (L–N) In ARCMs, the effects of Mapk7 knockdown on Irs1 transcript, IRS1 protein level, and insulin-stimulated (0.1 µM for 30 mins) phosphorylation of IRS1 and AKT were observed upon 2 hr hypoxia (N = 4–6 experiments). (O–P) SLC2A4 plasma membrane translocation was detected by immunoblots of the membrane fraction and confocal images of MYC-Slc2a4-mCherry immunofluorescence in H9C2 cells (N = 5 experiments). Fused mCherry (red) allows for SLC2A4 detection, while membrane translocated SLC2A4 is visualized by MYC (green). DAPI stains nuclear (blue). Scale bar: 25 µm. (Q) Insulin-dependent (0.1 µM for 30 mins) glucose uptake capacity under hypoxia was deferred in Mapk7 or Irs1 deficient ARCMs (N = 3 experiments). (R) ATP production was reduced in Mapk7-knockdown ARCMs under hypoxic condition (N = 3 experiments). Data presented as mean ± SEM. Comparisons between two groups were performed using two-tailed Student’s t-test. Two-way ANOVA followed by Bonferroni post hoc tests was employed as appropriate.
-
Figure 4—source data 1
Effect of MAPK7 deficiency on cardiac insulin resistance and in vivo and in vitro.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig4-data1-v2.xlsx
-
Figure 4—source data 2
Echocardiographic parameters of Mapk7-cko and control mice one week post-MI.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig4-data2-v2.xlsx
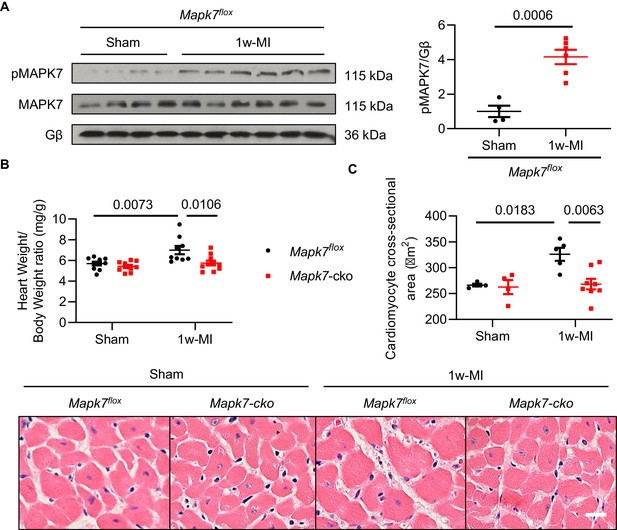
MAPK7 activation and hypertrophy in mouse myocardium one week post-MI.
(A) Contrary to the analysis after 4w-MI, MAPK7 activation was observed after 1w-MI in Mapk7flox mice (N = 4–6 mice). Cardiac hypertrophic growth in Mapk7-cko and Mapk7flox mice was evaluated by (B) heart weight vs body weight ratio (N = 9–10 mice), and (C) cardiomyocyte size measured from H and E stained heart sections (N = 4–8 mice). Scale bar: 50 µm. Data presented as mean ± SEM. Comparisons between two groups were performed using two-tailed Student’s t-test. Two-way ANOVA followed by Bonferroni post hoc tests was employed as appropriate.
-
Figure 4—figure supplement 1—source data 1
MAPK7 activation and hypertrophy in mouse myocardium one week post-MI.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig4-figsupp1-data1-v2.xlsx
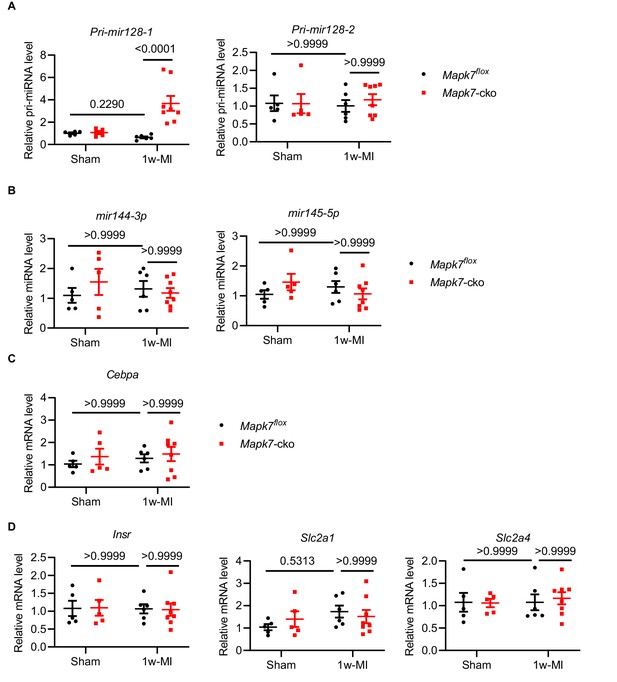
Gene expression comparison between Mapk7flox and Mapk7-cko mice one week post-MI.
Real-time qPCR detection of (A) Pri-mir128-1 and Pri-mir128-2, (B) mir144-3p and mir145-5p and (C) Cebpa in Mapk7flox and Mapk7-cko subject to 1 week MI (N = 5–8 mice). (D) qPCR determined that Insr, Slc2a1 and Slc2a4 transcripts levels were not altered in Mapk7-cko one week post-MI (N = 5–8 mice). Data werenormalized against 18S or mir191-5p. Data presented as mean ± SEM. Two-way ANOVA followed by Bonferroni post hoc tests was employed to compare groups.
-
Figure 4—figure supplement 2—source data 1
Gene expression comparison between Mapk7flox and Mapk7-cko mice one week post-MI.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig4-figsupp2-data1-v2.xlsx
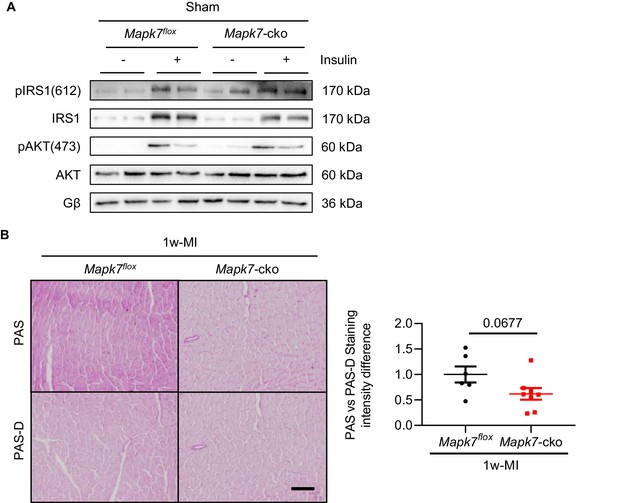
Activation of the insulin cascade in sham mice and cardiac glycogen detection one week post-MI.
(A) Immunoblots illustrated that insulin-stimulated (insulin IP injection, 1 U/kg for 30 mins) activation of IRS1 and AKT was at a comparable level in the myocardium from Mapk7flox and Mapk7-cko under basal condition (N = 5–6 mice). (B) Glycogen in the hearts from 1w-MI stressed Mapk7flox and Mapk7-cko mice was detected by Periodic Acid Schiff (PAS) staining. Diastase treatment was to discern unspecific staining (PAS-D) (N = 6–8 mice). Scale bar: 50 µm. Quantification of positive glycogen content is provided. Data presented as mean ± SEM. Groups were compared using two-tailed Student’s t-test.
-
Figure 4—figure supplement 3—source data 1
Activation of the insulin cascade in sham mice and cardiac glycogen detection one week post-MI.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig4-figsupp3-data1-v2.xlsx
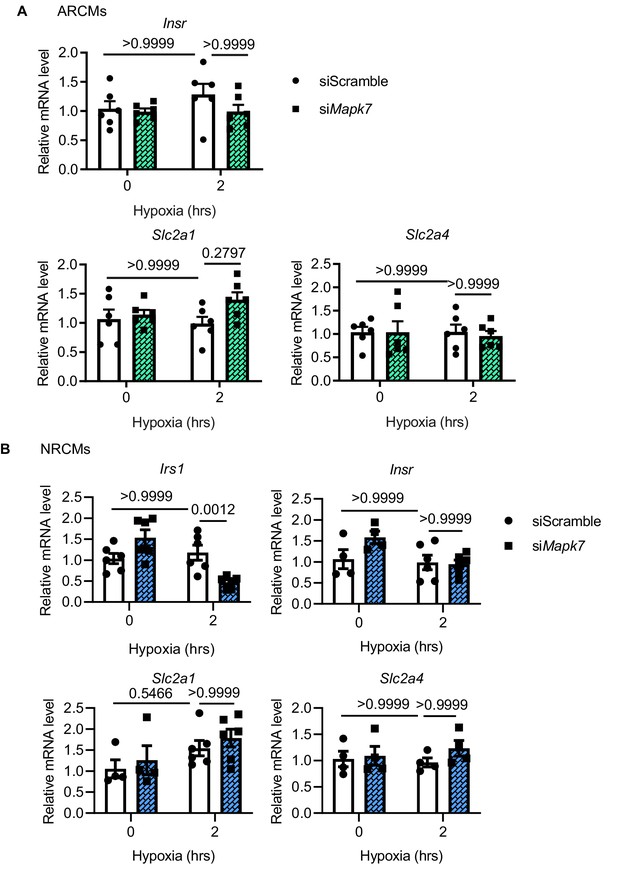
Insulin-related gene expression in MAPK7-deficient rat cardiomyocytes under hypoxic stress.
(A) Insr, Slc2a1 and Slc2a4 levels were comparable in adult rat cardiomyocytes (ARCMs) with MAPK7 deficiency despite hypoxic stress (N = 6 experiments). (B) Irs1 was reduced in neonatal cardiomyocytes (NRCMs) due to Mapk7 knockdown, while the others remained at a similar level in response to hypoxia (N = 4–6 experiments). Data presented as mean ± SEM. Two-way ANOVA followed by Bonferroni post hoc tests was employed to compare groups.
-
Figure 4—figure supplement 4—source data 1
Insulin-related gene expression in MAPK7-deficient rat cardiomyocytes under hypoxic stress.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig4-figsupp4-data1-v2.xlsx
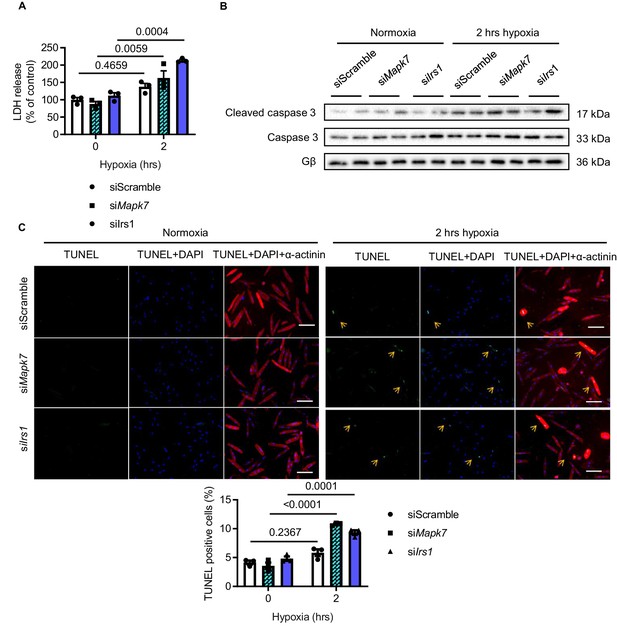
Effect of Mapk7 and Irs1 knockdown in ARCMs cell death after hypoxia.
ARCMs cell death under hypoxia was augmented by Mapk7 or Irs1 knockdown, as evidenced by increased (A) lactate dehydrogenase (LDH) released to the culture medium (N = 3 experiments), (B) cleaved caspase three level (N = 3 experiments), and (C) the number of TUNEL positive nuclei (green). DAPI (blue) detects nuclear and actinin (red) is cardiomyocyte indicator (N = 3 experiments). Scale bar: 100 µm. Data presented as mean ± SEM. Two-way ANOVA followed by Bonferroni post hoc tests was employed to compare groups.
-
Figure 4—figure supplement 5—source data 1
Effect of Mapk7 and Irs1 knockdown in ARCMs cell death after hypoxia.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig4-figsupp5-data1-v2.xlsx
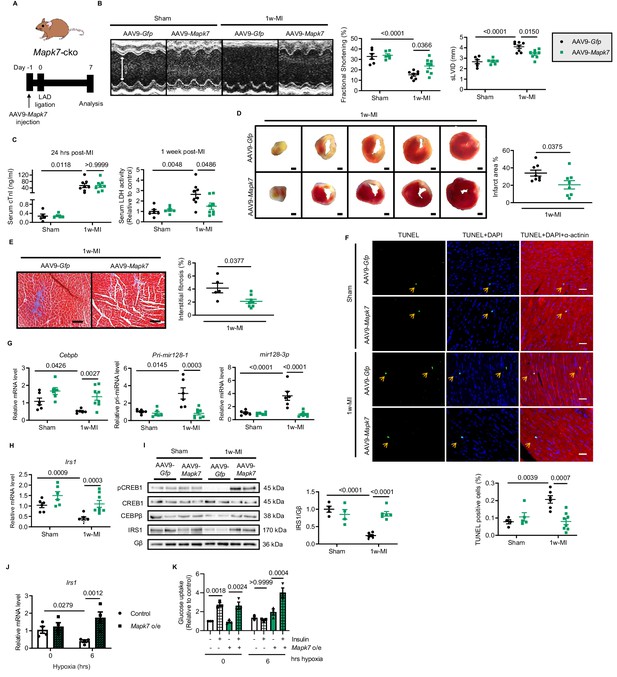
Cardiac MAPK7 restoration preserves IRS1 pathway in the myocardium and cardiac function in response to hypoxia.
(A) Experimental design of cardiac MAPK7 restoration in Mapk7-cko mice via tail vein administration of AAV9-Mapk7 (1 × 1011 viral particles), followed by LAD ligation for one week. (B) Echocardiography determined improved cardiac function by AAV9-Mapk7 injection, measured by fractional shortening (%) and sLVID (N = 5–8 mice). (C–F) MAPK7 restoration decreased serum cTnI (24 hr post-ligation) and LDH (one week post-ligation), infarct area (scale bar: 1 mm), interstitial fibrosis in non-infarct area (scale bar: 100 μm), and apoptotic cells detected by TUNEL assay by triple staining (scale bar: 30 µm) of TUNEL (green), DAPI (blue for nuclear) and α-actinin (red for marking cardiomyocytes). Arrows indicate TUNEL positive nuclei (N = 4–8 mice). (G) qPCR showed that Cebpb, Pri-mir128-1, and mature mir128-3p were retained at the normal range by AAV9-Mapk7 injection (N = 6–8 mice). (H–I) Transcript and protein level of IRS1 was maintained by MAPK7 restoration (N = 4–8 mice). (J) Irs1 transcript in ARCMs was preserved by MAPK7 overexpression (Mapk7 o/e) (N = 4 experiments). (K) Glucose uptake capacity of ARCMs was rescued by MAPK7 overexpression in response to insulin stimulation (0.1 µM for 30 mins) during long-term hypoxia (N = 3 experiments). Data presented as mean ± SEM. Comparisons between two groups were performed using two-tailed Student’s t-test. Two-way ANOVA followed by Bonferroni post hoc tests was employed as appropriate.
-
Figure 5—source data 1
Functional and transcriptional effects of MAPK7 restoration in Mapk7-cko mice after MI and rat cardiomyocytes under hypoxia.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig5-data1-v2.xlsx
-
Figure 5—source data 2
Echocardiographic parameters of Mapk7-cko mice injected with AAV9-Gfp or AAV9-Mapk7 one week post-MI.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig5-data2-v2.xlsx
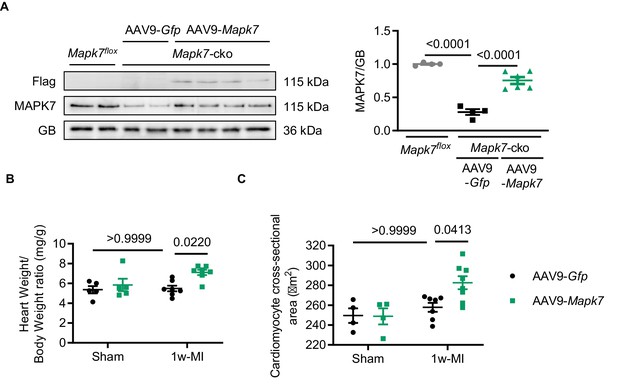
MAPK7 expression and cardiac hypertrophy in AAV9-Mapk7-injected mice one week post-MI.
(A) MAPK7 expression in myocardium was restored to physiological level in Mapk7-cko mice using AAV9-Mapk7 injection (AAV9-Gfp as control) (N = 4–6 mice). Cardiac hypertrophic response was observed in Mapk7-cko hearts with restored MAPK7 expression 1 week after MI, indicated by (B) heart weight vs. body weight ratio (N = 5–7 mice) and (C) cardiomyocyte size (N = 4–7 mice). Data presented as mean ± SEM. One or two-way ANOVA followed by Bonferroni post hoc tests were employed as appropriate.
-
Figure 5—figure supplement 1—source data 1
MAPK7 expression and cardiac hypertrophy in AAV9-Mapk7-injected mice one week post-MI.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig5-figsupp1-data1-v2.xlsx
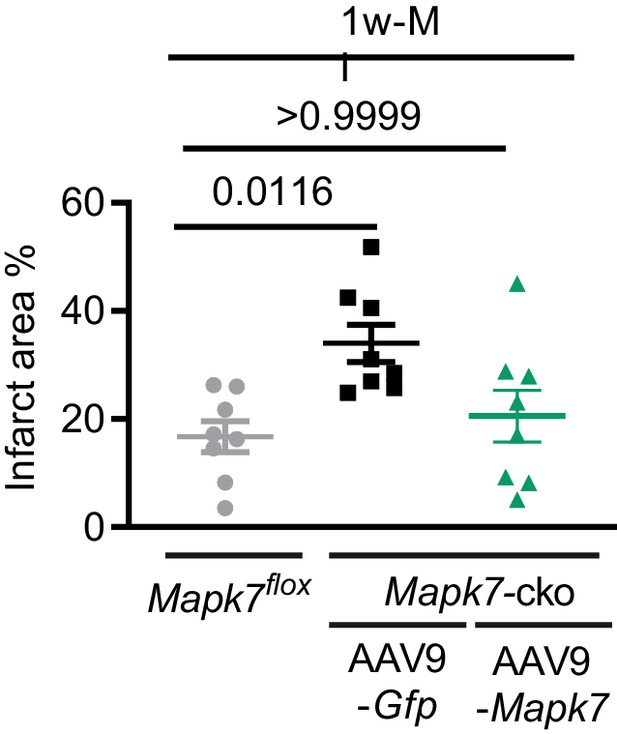
Infarct area comparison between Mapk7flox and Mapk7-cko injected with AAV9-Mapk7 seven days after MI.
Infarct area detected by triphenyltetrazolium chloride (TTC) staining in (N = 8 mice). Mapk7flox and Mapk7-cko mice with control injection and Mapk7-cko mice with AAV9-Mapk7 injection one week post-MI. Scale bar: 1 mm.
-
Figure 5—figure supplement 2—source data 1
Infarct area comparison between Mapk7flox and Mapk7-cko injected with AAV9-Mapk7 seven days after MI.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig5-figsupp2-data1-v2.xlsx
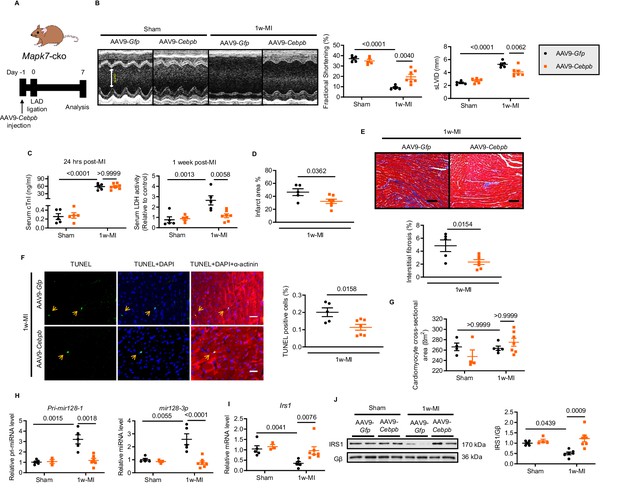
Cardiac-specific CEBPβ overexpression ameliorates cardiac injury in Mapk7-cko post-MI.
(A) Experimental design of CEBPβ overexpression in Mapk7-cko myocardium via tail vein injection of AAV9-Cebpb (1 × 1011 viral particles), followed by MI for one week. (B) Echocardiography showing preserved cardiac function by AAV9-Cebpb injection (N = 5–7 mice). (C–F) CEBPβ overexpression reduced serum cTnI (24 hr post-ligation) and LDH (one week post-ligation), infarct area, interstitial fibrosis in the non-infarct region (scale bar: 100 μm), and TUNEL positive apoptotic cells (green, scale bar: 30 µm), co-stained with DAPI (blue) and α-actinin (red). Arrows indicate TUNEL positive nuclei (N = 5–7 mice). (G) Cardiomyocyte size was measured from H and E stained sections (N = 4–7 mice). (H) Pri-mir128-1 and mature mir128-3p levels were examined in the myocardium by qPCR (N = 3–7 mice). (I–J) qPCR and immunoblotting analyses showed the maintenance of IRS1 expression by CEBPβ overexpression after MI despite MAPK7 depletion (N = 3–7 mice). Data presented as mean ± SEM. Comparisons between two groups were performed using two-tailed Student’s t-test. Two-way ANOVA followed by Bonferroni post hoc tests was employed as appropriate.
-
Figure 6—source data 1
CEBPβ overexpression effect on mir128-3p and Irs1 expression in Mapk7-cko post-MI.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig6-data1-v2.xlsx
-
Figure 6—source data 2
Echocardiographic parameters of Mapk7-cko mice injected with AAV9-Gfp or AAV9-Cebpb one week post-MI.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig6-data2-v2.xlsx
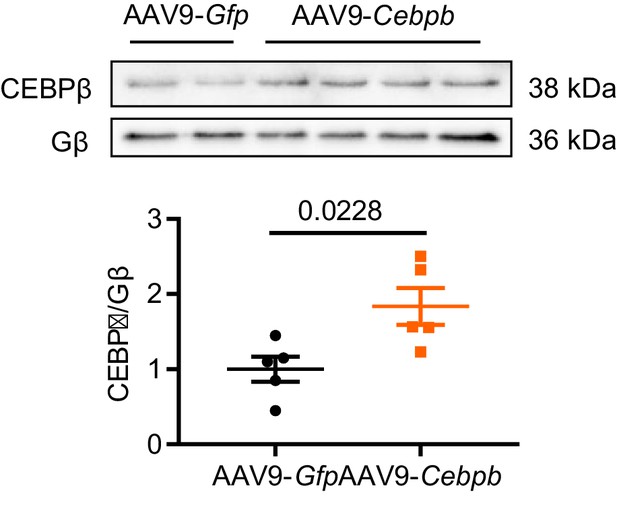
Myocardial CEBPβ expression level after AAV9-Cebpb tail vein injection.
Immunoblots and quantification determined that overexpression of ectopic CEBPβ at a physiological range was achieved by AAV9-Cebpb injection (N = 5 mice). Data presented as mean ± SEM. Groups were compared using two-tailed Student’s t-test.
-
Figure 6—figure supplement 1—source data 1
Myocardial CEBPβ expression level after AAV9-Cebpb tail vein injection.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig6-figsupp1-data1-v2.xlsx
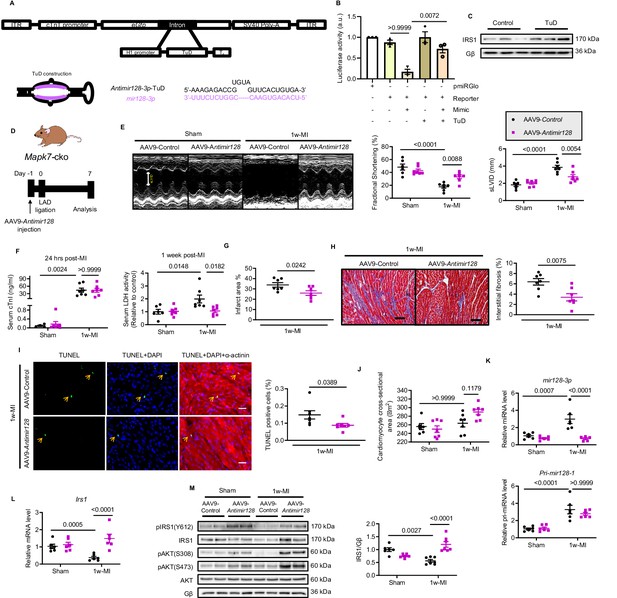
Silencing mir128-3p prevents Irs1 degradation and decelerates HF progression post-MI.
(A) Construction of antimir128-TuD built in the intronic region Gfp of AAV9-Gfp capsid. The sequence of antimir128-TuD is provided. (B) pmiRGlo-mir128-3p (four repeated complementary sequences of mir128-3p built in the 3’ region of pmiRGlo) luciferase reporter activity was inhibited by mir128-3p mimic in H9C2 cardiomyocytes, but recovered by antimir128-TuD (N = 3 experiments). (C) Antimir128-TuD increased IRS1 protein level in cardiomyocytes (N = 3 experiments). (D) Experimental design of AAV9-delivered antimir128-TuD into Mapk7-cko mice by tail vein administration of AAV9-antimir128 (1 × 1011 viral particles), followed by MI. (E) Cardiac function was evaluated by echocardiography (N = 6–7 mice). (F–I) AAV9-antimir128 inhibition of endogenous mir128-3p upregulated due to MAPK7 deficiency led to less serum cTnI (24 hr post-ligation) and LDH (one week post-ligation), infarct area, fibrosis (scale bar: 100 μm), and apoptosis is detected by TUNEL assay (green, scale bar: 30 µm), co-stained with DAPI (blue) and α-actinin (red). Arrows indicate TUNEL positive nuclei (N = 6–7 mice). (J) Cardiomyocyte size was measured according to H and E staining (N = 6–7 mice). (K) Mature mir128-3p was silenced by AAV9-antimir128 injection; however, Pri-mir128-1 upregulation was not affected (N = 6 mice). (L–M) qPCR and immunoblotting analyses illustrated that expression and activation of IRS1 and AKT were preserved by AAV9-antimir128 application in the absence of MAPK7 despite MI stress (N = 6–7 mice). Data presented as mean ± SEM. Comparisons between two groups were performed using two-tailed Student’s t-test. One-way or two-way ANOVA followed by Bonferroni post hoc tests were employed as appropriate.
-
Figure 7—source data 1
Functional effect of silencing mir128-3p post-MI.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig7-data1-v2.xlsx
-
Figure 7—source data 2
Echocardiographic parameters of Mapk7-cko mice injected with AAV9-Control or AAV9-Antimir128 one week post-MI.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig7-data2-v2.xlsx
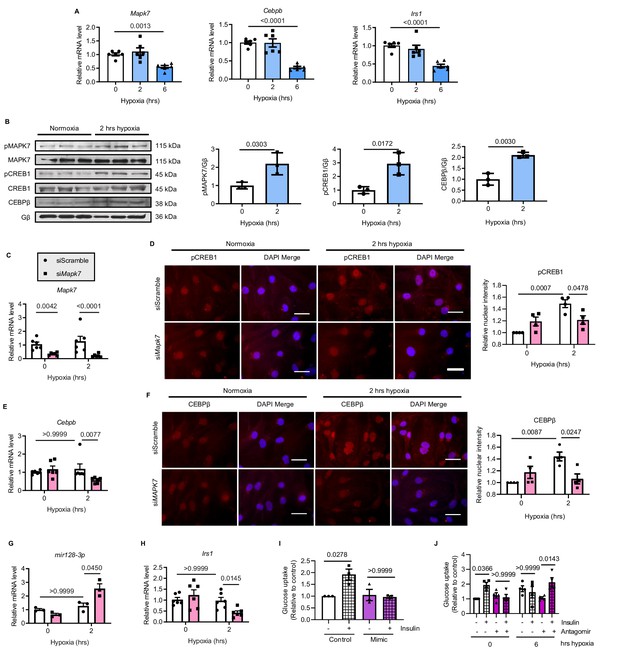
MAPK7-CEBPβ repression of mir128-3p maintains insulin sensitivity in human iPSC-CMs under hypoxia.
(A) Transcripts of Mapk7, Cebpb, and Irs1 were reduced in human iPSC-CMs in response to long-term hypoxia (N = 6 experiments). (B) Immunoblots and quantifications showed that phosphorylation of MAPK7 and CREB1 were accompanied by increased CEBPβ under short-term hypoxia (N = 3 experiments). (C) Mapk7 knockdown using its siRNA in iPSC-CMs was validated by qPCR (N = 6 experiments). (D) Phosphorylated CREB1 nuclear accumulation (red) induced by short-term hypoxia. However, Mapk7 knockdown blocked its nuclear deposition, counterstained with DAPI (blue) (N = 4 experiments, more than 50 nuclei measured per group per experiment). Scale bar: 20 µm. (E–F) qPCR and immunofluorescent staining (scale bar: 20 µm) determined that MAPK7 deficiency reduced expression and nuclear localization of CEBPβ (N = 4–6 experiments, more than 50 nuclei measured per group per experiment). (G–H) Augmented mature mir128-3p and decreased Irs1 were detected by qPCR in Mapk7-knockdown iPSC-CMs under hypoxia (N = 3–6 experiments). (I–J) mir128-3p mimic abrogated insulin stimulation (0.1 µM for 30 mins)-induced glucose uptake in iPSC-CMs. On the contrary, mir128-3p antagomiR rescued insulin-dependent glucose uptake capability, albeit prolonged hypoxia circumstance (N = 3 experiments). Data presented as mean ± SEM. Comparisons between two groups were performed using two-tailed Student’s t-test. One-way or two-way ANOVA followed by Bonferroni post hoc tests were employed as appropriate.
-
Figure 8—source data 1
Analysis of the MAPK7-CEBPβ-mir128-3p axis in human iPSC-CMs under hypoxia.
- https://cdn.elifesciences.org/articles/54298/elife-54298-fig8-data1-v2.xlsx
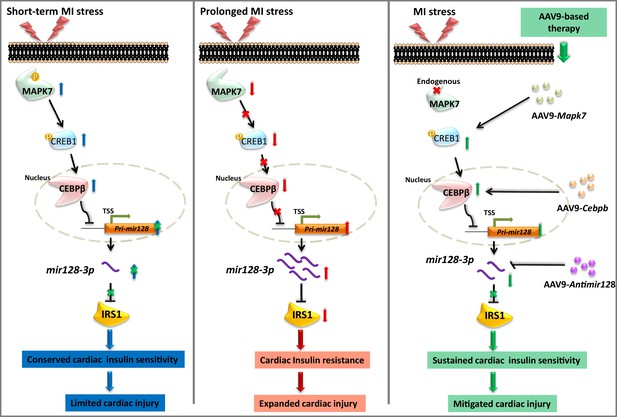
Schematic representation of MAPK7-CEBPβ signaling cascade negative regulation of mir128-3p under ischemia/hypoxic condition.
Short-term ischemia/hypoxia activates MAPK7-CREB1, which facilitates CEBPβ expression and its nucleus translocation to act as transcriptional repressor on Pri-mir128-1, consequently maintaining IRS1 in the myocardium. However, prolonged stress induces MAPK7 loss and defective CEBPβ repression on mir128-3p, which results in IRS1 loss and impaired insulin pathway. Targeting this signaling cascade manifests the beneficial effects on maintenance of cardiac insulin sensitivity in response to ischemic/hypoxic stress, which is evidenced by AAV9-delivered cardiac MAPK7 restoration, cardiac-specific Cebpb overexpression, or silencing endogenous mir128-3p. List of source data files.
Additional files
-
Supplementary file 1
Key resources table.
- https://cdn.elifesciences.org/articles/54298/elife-54298-supp1-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/54298/elife-54298-transrepform-v2.docx





