Loss of Ena/VASP interferes with lamellipodium architecture, motility and integrin-dependent adhesion
Figures
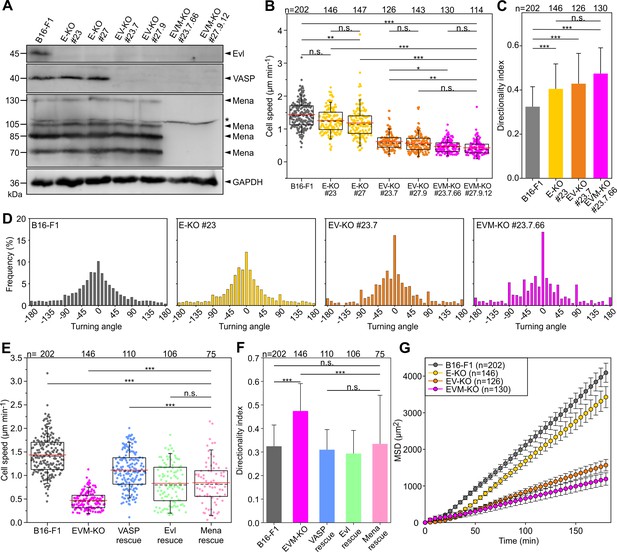
Loss of Ena/VASP-proteins impairs 2D cell migration in B16-F1 cells.
(A) To obtain independent triple-knockout mutant cell lines, the two independent Evl single knockout mutants (E-KO #23 and #27) derived from B16-F1 mouse melanoma cells, were separately used to the generate independent Evl/VASP double mutants (E/V-KO #23.7 and #27.9) followed by generation of two individual triple-KO mutants additionally lacking Mena (E/V/M-KO #23.7.66 and #27.9.12). Elimination of Evl, VASP and all Mena isoforms by CRISPR/Cas9 in B16-F1 cells was confirmed by immunoblotting using specific antibodies (asterisk indicates nonspecific band). GAPDH was used as loading control. (B) Consecutive gene disruption of the three Ena/VASP paralogues increasingly diminishes cell migration on laminin. At least three time-lapse movies from three independent experiments were analyzed for each cell line. (C) Directionality increased with consecutive inactivation of the three Ena/VASP paralogues. Bars represent arithmetic means ± SD. (D) Distribution of turning angles during migration of B16-F1 and derived mutant cells. (E–G) Cell speed and directionality are largely rescued by ectopic expression of EGFP-tagged VASP, Evl or Mena in EVM-KO #23 cells. (F) Bars represent arithmetic means ± SD. (G) Analyses of mean square displacement of wild-type versus mutant cells. Respective symbols and error bars represent means ± SEM. (B–E) The boxes in box plots indicate 50% (25–75%) and whiskers (5–95%) of all measurements, with dashed black lines depicting the medians, arithmetic means are highlighted in red. (B–C) and (E–F) Non-parametric, Kruskal-Wallis test and Dunn’s Multiple Comparison test were used to reveal statistically significant differences between datasets. *p≤0.05, **p≤0.01, ***p≤0.001; n.s.; not significant. n, number of cells analyzed from at least three independent experiments.
-
Figure 1—source data 1
Source data for details of cell migration including cell speeds, directionality indices and MSD values.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig1-data1-v2.xlsx
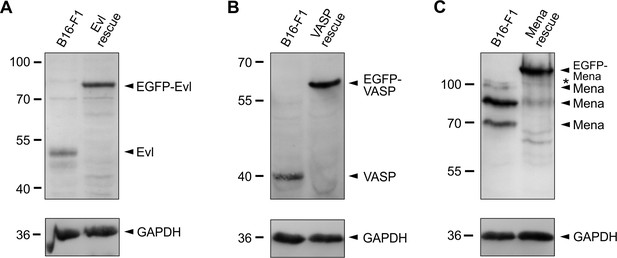
Expression levels of Ena/VASP proteins in reconstituted EVM-KO cells compared to endogenous levels assessed by immune blotting.
(A) Expression level of EGFP-Evl in EVM-KO Evl rescue cells was increased 2.5-fold compared to endogenous Evl levels in B16-F1 wild-type cells normalized to GAPDH. (B) Expression level of EGFP-VASP in EVM-KO VASP rescue cells was increased 1.5-fold as compared to endogenous VASP levels in B16-F1 cells. (C) Expression level of EGFP-Mena in EVM-KO Mena rescue cells was increased 1.1-fold compared to endogenous Mena levels in B16-F1 cells. Asterisk indicates nonspecific band. GAPDH was used as loading control. Relative changes of Ena/VASP expression levels normalized to GAPDH.
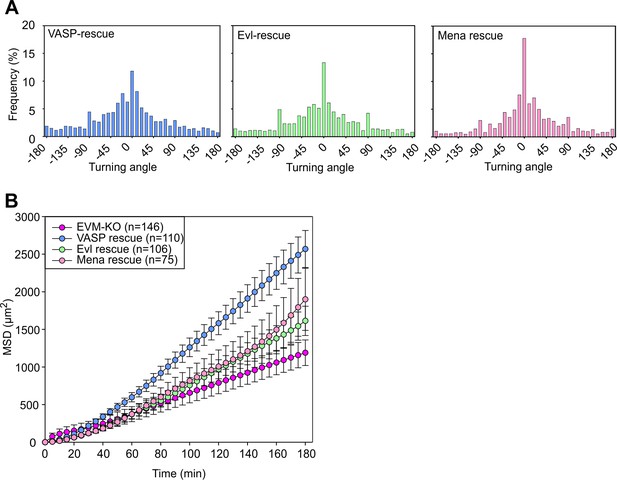
Loss of Ena/VASP proteins affects 2D migration and directionality.
(A) Distribution of turning angles during migration of EVM-KO cells reconstituted with VASP, Evl or Mena. As opposed to Mena, VASP and Evl both readily rescued turning angles. (B) Analyses of mean square displacement of mutant and reconstituted cells. All three Ena/VASP members rescued migration as assessed by MSD, but VASP was most efficient. Respective symbols and error bars represent means ± SEM.
-
Figure 1—figure supplement 2—source data 1
Source data for details of cell migration including turning angles and MSD values Figure 1—figure supplement 2.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig1-figsupp2-data1-v2.xlsx
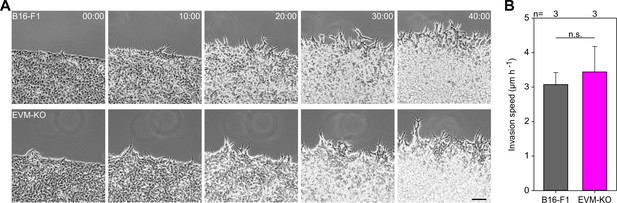

Loss of Ena/VASP-proteins does not affect invasion of B16-F1 cells.
(A) Representative invasion of B16-F1 and EVM-KO cells into 3D-matrigel over time. Bar, 100 µm. (B) Quantification of invasion speed of B16-F1 and EVM-KO cells. Bars represent arithmetic means ± SD. (B) Non-parametric, Kruskal-Wallis test was used to reveal statistically significant differences between datasets. n.s.; not significant. n, number of experiments.
-
Figure 1—figure supplement 3—source data 1
Source data for details of invasion including invasion speeds Figure 1—figure supplement 3.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig1-figsupp3-data1-v2.xlsx
Loss of Ena/VASP proteins in B16-F1 cells impairs 2D cell migration, related to Figure 1.
Random migration of B16-F1, and derived Evl-KO (E–KO), Evl/VASP-KO (EV-KO) and Evl/VASP/Mena-KO (EVM-KO) mutants on laminin. Cells were recorded by phase-contrast time-lapse imaging using a 4x objective and tracked by MTrackJ to illustrate representative cell trajectories. Note more directional movement of EVM-KO cells. Time is indicated in h:min. Bar, 300 µm.
Loss of Ena/VASP-proteins does not affect invasion of B16-F1 cells, related to Figure 1—figure supplement 3.
Representative time-lapse movies showing invasion of B16-F1 and EVM-KO cells confined beneath 3D-matrigel. Cells were recorded by phase-contrast time-lapse imaging using a 10x objective. Time is indicated in hr:min. Bar, 100 µm.
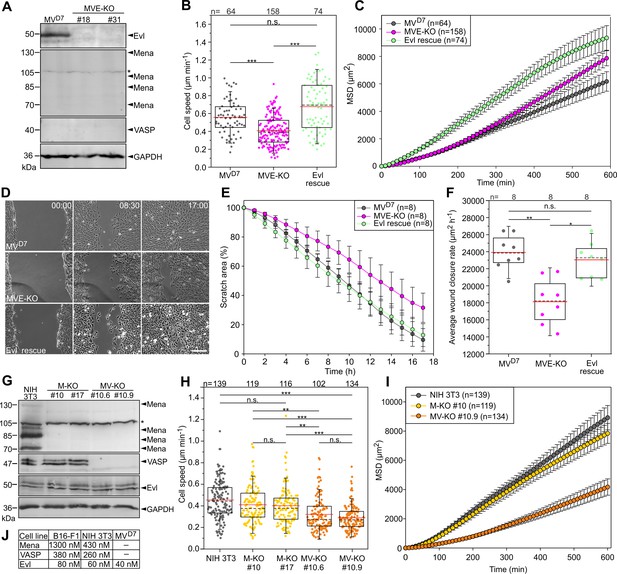
Inactivation of Ena/VASP proteins in various fibroblasts impairs 2D cell migration.
(A) MVD7 cells lack Mena and VASP, but still express Evl. Elimination of Evl by CRISPR/Cas9 in MVD7 fibroblasts was confirmed by immunoblotting in independent clonal cell lines (MVE-KO). GAPDH was used as loading control. Asterisk indicates nonspecific band in Mena blot. (B) Elimination of Evl in MVD7 cells decreased cell speed on fibronectin-coated glass and could be rescued by expression of Evl. (C) Analyses of mean square displacement of MVD7, MVE-KO and reconstituted cells as indicated. Data points represent arithmetic means ± SEM. At least three time-lapse movies from three independent experiments were analyzed for each cell line. (D) Representative frames from wound healing movies of MVD7, MVE-KO and reconstituted cells as indicated. MVE-KO cells were not able to close the wound after 17 hr. Bar, 200 µm. (E) Reduction of wound area over time. n, number of movies analyzed. Data are means ± SD. (F) Average wound closure rate. n, number of movies analyzed. (G) Immunoblot of independent single and double-KO mutants derived from NIH 3T3 fibroblasts lacking Mena (M–KO) or Mena and VASP (MV-KO) by CRISPR/Cas9 technology. GAPDH was used as loading control. (H) Consistent with findings in B16-F1 cells, consecutive gene disruption of these two Ena/VASP paralogues in NIH 3T3 fibroblasts again increasingly diminished cell migration on fibronectin. (I) Analyses of mean square displacement of NIH 3T3, M-KO and MV-KO cells as indicated. Data points represent arithmetic means ± SEM. n, number of cells tracked. At least three time-lapse movies from three independent experiments were analyzed for each cell line. (J) Total cytoplasmic concentrations of Ena/VASP proteins in investigated cell lines. (B, F and H) Boxes in box plots indicate 50% (25–75%) and whiskers (5–95%) of all measurements, with dashed black lines depicting the medians, arithmetic means are highlighted in red. Non-parametric, Kruskal-Wallis test and Dunn’s Multiple Comparison test were used to reveal statistically significant differences between datasets. *p≤0.05, **p≤0.01, ***p≤0.001; n.s.; not significant. n, number of cells analyzed from at least three independent experiments.
-
Figure 2—source data 1
Source data for details of cell migration including cell speeds, MSD values, wound healing scratch areas and average wound closure rates Figure 2.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig2-data1-v2.xlsx
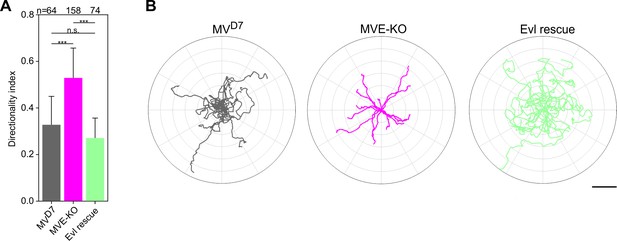
Elimination of Evl in MVD7 cells affects directionality.
(A) Quantification of directionality. Bars represent arithmetic means ± SD. ***p≤0.001 and n.s., not significant by Kruskal-Wallis test and Dunn’s Multiple Comparison test and n, number of cells analyzed. (B) Radar plots showing trajectories of 15 randomly migrating MVD7, MVE-KO and reconstituted cells as indicated. Note higher directional persistence of MVE-KO cells compared to MVD7 cells, which was diminished again by expression of Evl. Scale bar, 100 µm.
-
Figure 2—figure supplement 1—source data 1
Source data for details of cell migration including directionality indices Figure 2—figure supplement 1.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig2-figsupp1-data1-v2.xlsx
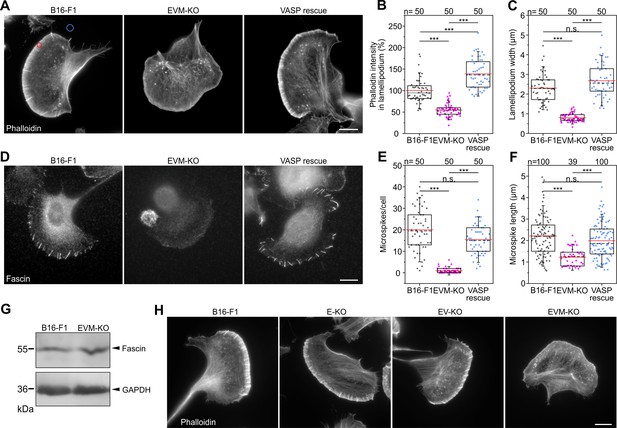
Loss of Ena/VASP perturbs lamellipodia and abrogates microspike formation in B16-F1 cells.
(A) Representative examples of lamellipodia from wild-type B16-F1, EVM-KO and reconstituted EVM-KO cells transiently expressing EGFP-VASP. Cells migrating on laminin and stained for the actin cytoskeleton with phalloidin. Bar, 10 µm. (B) Quantification of F-actin intensities in lamellipodia (red circle in A) of wild-type and mutant cells after subtraction of background (blue circle in A). (C) Quantification of lamellipodia width in wild-type and mutant cells. (D) Loss of Ena/VASP markedly diminishes fascin-decorated microspikes, which are rescued by EGFP-VASP. Immunostaining with fascin antibody. Bar, 10 µm. (E) Quantification of microspikes in wild-type and mutant cells. (F) Quantification of microspike length in wild-type and mutant cells. (G) Comparable expression of fascin in wild-type and EVM-KO cells was confirmed by immunoblotting using fascin-specific antibodies. GAPDH was used as loading control. (H) Representative examples of lamellipodia from wild-type B16-F1 cells, single E-KO, double EV-KO and triple EVM-KO mutants. Displayed cells migrating on laminin were fixed and stained with phalloidin to visualize their F-actin cytoskeleton. Bars, 10 µm. Note presence of microspikes in B16-F1 wild-type cells, single E-KO and double EV-KO mutants, but not in the triple EVM-KO mutant cell. (B–C and E–F) Boxes in box plots indicate 50% (25–75%) and whiskers (5–95%) of all measurements, with dashed black lines depicting the medians, arithmetic means are highlighted in red. Non-parametric, Kruskal-Wallis test and Dunn’s Multiple Comparison test were used to reveal statistically significant differences between datasets. ***p≤0.001; n.s.; not significant. n, number of cells analyzed (B, C, and E) or microspikes (F) from at least three independent experiments.
-
Figure 3—source data 1
Source data for details of lamellipodial actin including relative phalloidin intensities, lamellipodium widths, microspike numbers per cell and microspike length Figure 3.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig3-data1-v2.xlsx
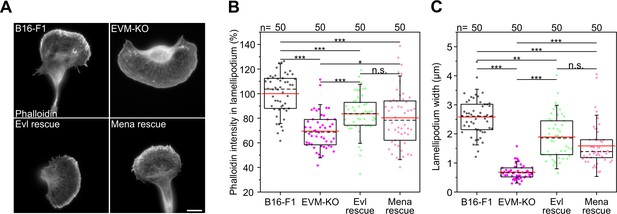
Expression of Evl and Mena is sufficient for rescuing lamellipodial parameters in B16-F1 cells.
(A) Representative examples of lamellipodia in wild-type B16-F1, EVM-KO and reconstituted EVM-KO cells transiently expressing EGFP-Evl or -Mena, as indicated. Cells migrating on laminin were fixed and stained with phalloidin to visualize the actin cytoskeleton. Bar, 10 µm. (B) Quantification of F-actin intensities in lamellipodia of wild-type, mutant and reconstituted cells after background subtraction. (C) Quantification of lamellipodia widths in cells indicated. (B–C) Boxes in box plots indicate 50% (25–75%) and whiskers (5–95%) of all measurements, with dashed black lines depicting the medians, arithmetic means are highlighted in red. Non-parametric, Kruskal-Wallis test and Dunn’s Multiple Comparison test were used to reveal statistically significant differences between datasets. *p≤0.05, **p≤0.01, ***p≤0.001; n.s.; not significant. n, number of cells analyzed from at least three independent experiments.
-
Figure 3—figure supplement 1—source data 1
Source data for details of lamellipodial actin including relative phalloidin intensities and lamellipodium widths Figure 3—figure supplement 1.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig3-figsupp1-data1-v2.xlsx
Elimination of all three Ena/VASP proteins impairs lamellipodium formation and abolishes microspikes, related to Figure 3.
Displayed cells were transfected with EGFP-LifeAct to monitor actin dynamics during migration on laminin by time-lapse imaging using a 40x objective. EVM-KO cells were additionally transfected with mCherry-VASP. Reconstitution of EVM-KO cells with mCherry-VASP restored lamellipodium and microspike formation. Time is indicated in min:s. Bar, 15 µm.
Elimination of all three Ena/VASP proteins abrogates microspikes, related to Figure 3.
The cells shown were transfected with EGFP-fascin to monitor microspikes in cells migrating on laminin by time-lapse imaging using a 40x objective. EVM-KO cells were additionally transfected with mCherry-VASP. Reconstitution of EVM-KO cells with mCherry-VASP fully restored microspike formation (right panel). Time is indicated in min:s. Bar, 15 µm.
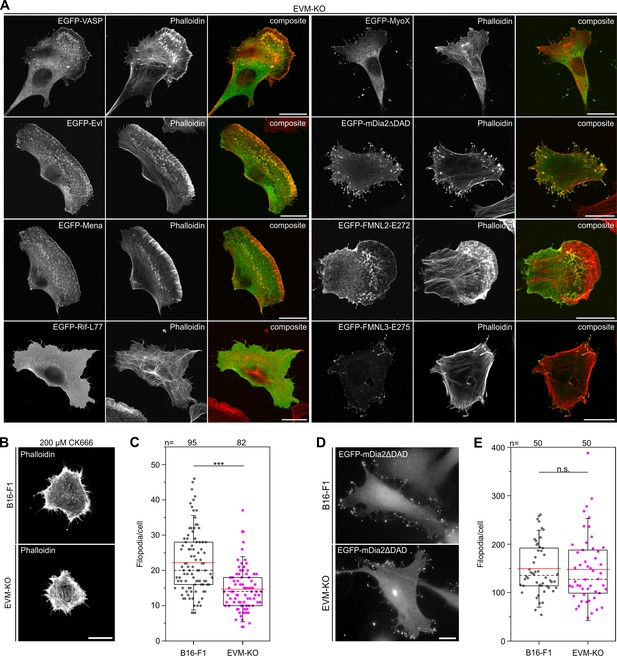
Microspike formation is exclusively rescued by all three Ena/VASP members, but not by active Rif-L77, myosin X, active mDia2ΔDAD or active FMNL2 and −3.
(A) Images display cells stained for the actin cytoskeleton with phalloidin and respective, expressed EGFP-tagged protein as indicated. Bars, 20 µm. (B) EVM-KO cells form filopodia in the absence of lamellipodia. Representative examples of phalloidin-stained wild-type B16-F1 and EVM-KO cells devoid of lamellipodia after treatment with 200 µM CK666 seeded at low laminin (1 µg/mL). Bar, 10 µm. (C) Quantification of filopodia in CK666-treated B16-F1 and EVM-KO cells. (D) Unchanged filopodium formation in B16-F1 and EVM-KO cells triggered by transient expression of active mDia2. Representative examples of B16-F1 and EVM-KO cells after labelling with GFP nanobodies are shown. Bar, 10 µm. (E) Quantification of filopodia revealed no difference between wild-type and mutant cells. (C and E) Boxes in box plots indicate 50% (25–75%) and whiskers (5–95%) of all measurements, with dashed black lines depicting the medians, arithmetic means are highlighted in red. Non-parametric, Mann-Whitney U rank sum test was used to reveal statistically significant differences between datasets. ***p≤0.001; n.s.; not significant. n, number of cells analyzed from at least three independent experiments.
-
Figure 4—source data 1
Source data for details of filopodia formation Figure 4.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig4-data1-v2.xlsx
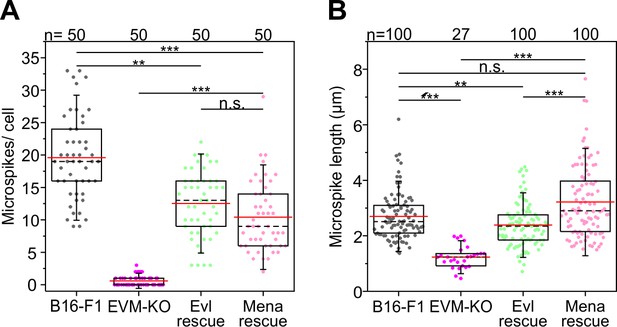
Quantification of microspikes after rescue of EVM-KO cells with EGFP-tagged Evl or Mena.
(A) Quantification of microspikes per cell. (B) Quantification of microspike lengths. Note longer microspikes in reconstituted EVM-KO cells expressing Mena. (A–B) **p≤0.01, ***p≤0.001; n.s. not significant by Kruskal-Wallis test and Dunn’s Multiple Comparison test. n, number of cells analyzed.
-
Figure 4—figure supplement 1—source data 1
Source data for details of microspike formation including microspike number per cell and microspike length Figure 4—figure supplement 1.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig4-figsupp1-data1-v2.xlsx
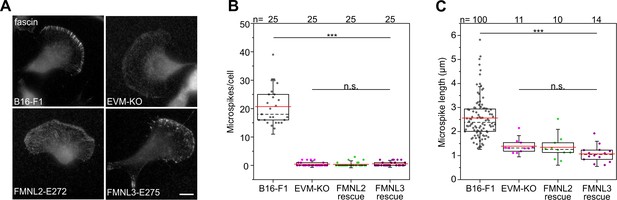
Active FMNL2 or −3 do not rescue microspikes in EVM-KO cell.
(A) Immunostaining with fascin antibody. Bar, 10 µm. (B) Quantification of microspikes per cell. (C) Quantification of microspike length. Note no difference in number and length of microspikes in reconstituted EVM-KO cells expressing EGFP-tagged FMNL2-E272 or FMNL3-E275 compared to EVM-KO cells. (B–C) ***p≤0.001; n.s. not significant by one-way ANOVA and Dunn’s Multiple Comparison test. n, number of cells analyzed.
-
Figure 4—figure supplement 2—source data 1
Source data for details of microspike formation including microspike number per cell and microspike length Figure 4—figure supplement 2.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig4-figsupp2-data1-v2.xlsx
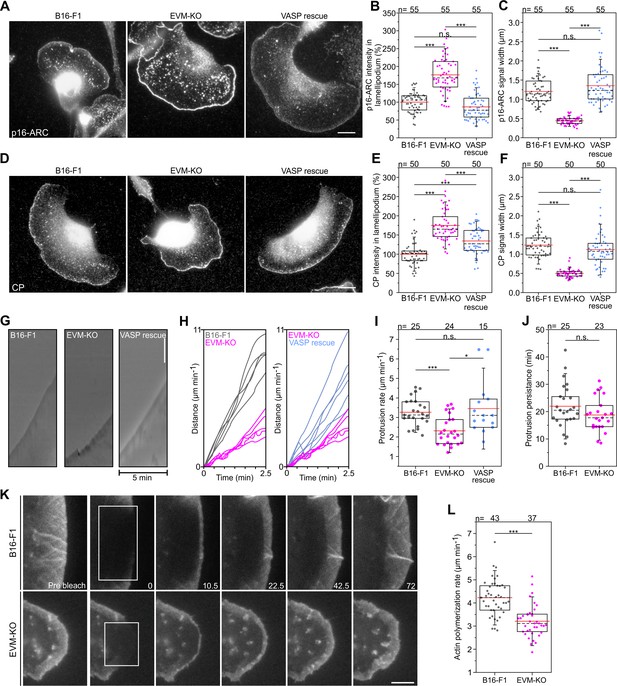
Loss of Ena/VASP affects lamellipodial parameters and protrusion dynamics.
(A) Immunolabelling of the Arp2/3 complex subunit p16-ARC (ArpC5A) in wild-type, mutant and reconstituted cells as indicated. Bar, 10 µm. (B) Quantification of Arp2/3 complex intensities in lamellipodia. (C) Quantification of Arp2/3 complex signal width. (D) Immunolabelling of CP in cell types as indicated. Bar, 10 µm. (E) Quantification of CP signal intensities in lamellipodia. (F) Quantification of CP signal width in lamellipodia. (G) Loss of Ena/VASP reduces the efficiency of lamellipodium protrusion. Kymographs of representative phase-contrast movies are shown. Bar, 10 µm. (H) Multiple examples of lamellipodium protrusion in B16-F1 versus EVM-KO versus EVM-KO cells reconstituted with VASP. (I) Quantification of protrusion rates. B-C, E-F and I, *p≤0.05, ***p≤0.001 and n.s., not significant by Kruskal-Wallis test and Dunn’s Multiple Comparison test. n, number of cells. (J) Quantification of protrusion persistence. (K) FRAP of EGFP-ß-actin in lamellipodia of B16-F1 and EVM-KO cells bleached as indicated by white rectangles. Numbers in post-bleach images correspond to seconds after bleach. Bar, 3 µm. (L) Average actin assembly rates of respective cell types expressing EGFP-ß-actin. Note reduced lamellipodial actin assembly in the mutant. J-L, ***p≤0.001, n.s., not significant by Mann-Whitney U rank sum test. n, number of cells analyzed.
-
Figure 5—source data 1
Source data for details of lamellipodial proteins including relative p16-ARC and CP intensities and signal widths, and of protrusions including protrusion rates and persistence, and actin polymerization rates Figure 5.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig5-data1-v2.xlsx
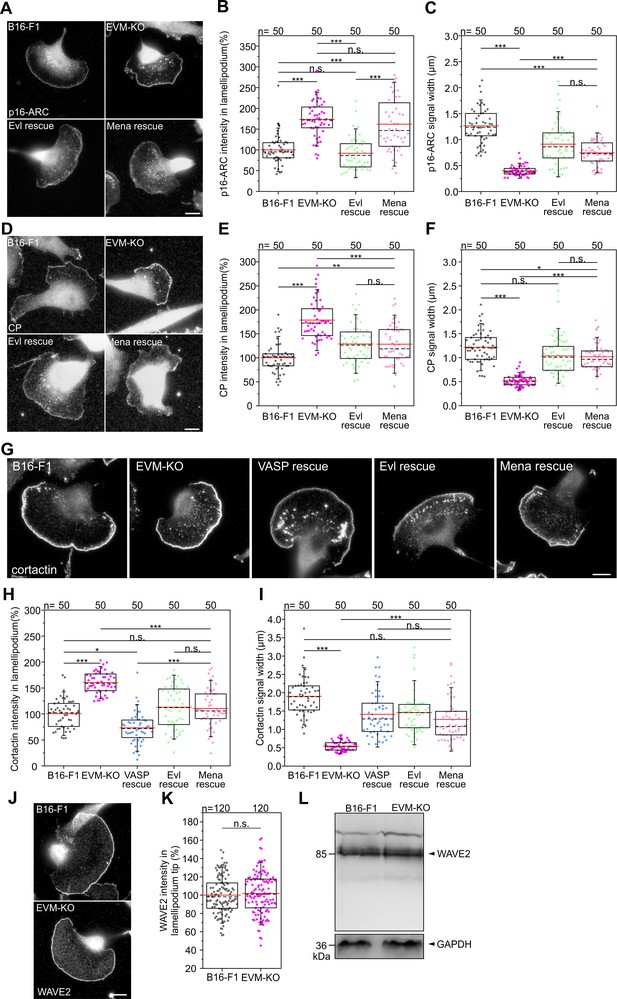
Loss of Ena/VASP affects additional lamellipodial parameters.
(A) Immunolabelling of endogenous Arp2/3 complex subunit p16-ARC (ArpC5A) in B16-F1 wild-type, EVM-KO and reconstituted EVM-KO cells expressing Evl or Mena. Bar, 10 µm. (B) Quantification of p16-ARC intensities in lamellipodia. (C) Quantification of lamellipodial p16-ARC width. (D) Immunolabeling of endogenous CP in wild-type, mutant and reconstituted cells. Bar, 10 µm. (E) Quantification of CP intensities in lamellipodia. (F) Quantification of CP width in lamellipodia. (G) Immunolabeling of endogenous cortactin in wild-type, mutant and reconstituted cells. Bar, 10 µm. (H) Quantification of cortactin intensities in lamellipodia. (I) Quantification of cortactin width in lamellipodia. B-C, E-F and H-I, *p≤0.05, **p≤0.01, ***p≤0.001, n.s., not significant by Kruskal-Wallis test and Dunn’s Multiple Comparison test. n, number of cells analyzed. (J–K) Immunolabeling of endogenous WAVE2 revealed no differences between B16-F1 wild-type and EVM-KO cells. Bar, 10 µm. n.s., not significant by Mann-Whitney U rank sum test. n, number of analyzed cells. (L) Comparable amounts of expression of WAVE2 in wild-type and EVM-KO cells were confirmed by immunoblotting using WAVE2-specific antibodies. GAPDH was used as loading control.
-
Figure 5—figure supplement 1—source data 1
Source data for details of lamellipodial proteins including relative p16-ARC, CP, cortactin and WAV2 intensities and signal widths Figure 5—figure supplement 1.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig5-figsupp1-data1-v2.xlsx
Impaired protrusion of EVM-KO cells migrating on laminin, related to Figure 5.
Reconstitution of the triple mutant cells with EGFP-VASP fully restored lamellipodial protrusion. Cells were recorded by time-lapse phase-contrast imaging using a 40x objective. Time is indicated in min:s. Bar, 15 µm.
Loss of Ena/VASP in B16-F1 cells reduces actin network polymerization, related to Figure 5.
Representative time-lapse movies showing FRAP experiments on lamellipodia of B16-F1 wild-type or EVM-KO cells, as indicated, expressing EGFP-β-actin on laminin. Time-lapse movies show periods before and after bleaching of lamellipodial regions with bleaching time points set to 0 s in each case. Bar, 3 µm.
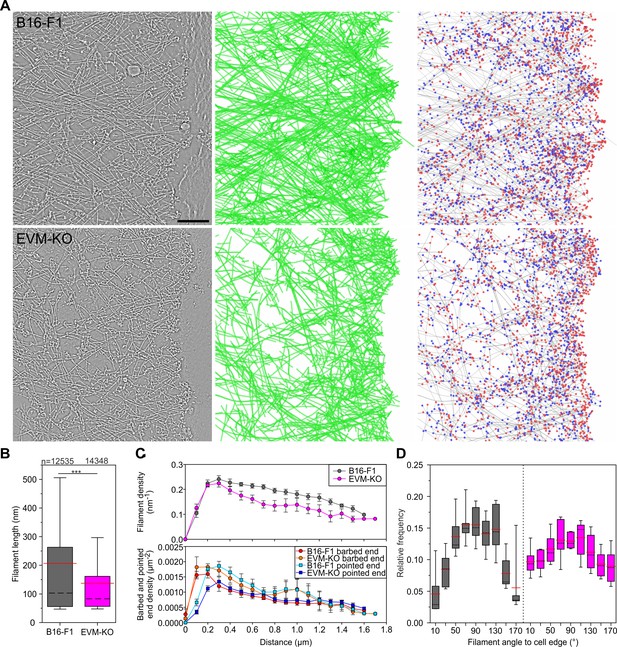
Electron tomography of ultrastructural changes in lamellipodial actin networks.
(A) Transmission electron micrographs of representative wild-type (B16–F1) and EVM-KO cells showing distinct actin filament networks at the leading edge (left), and 2D projections of digital 3D tomograms showing either actin filament trajectories in green (middle), or barbed ends on grey filaments in red, and pointed ends in blue (right). Scale bar, 100 nm. (B) Quantification of filament length in wild-type and EVM-KO cells. ***p≤0.001 by Mann-Whitney U rank sum test. Whiskers indicate 10% and 90% confidence intervals. n indicates the number of filaments analyzed. Dashed black lines show median, red lines arithmetic mean. (C) Densities of filaments (upper panel), barbed and pointed ends (lower panel) in 106 nm-wide spatial bins throughout the lamellipodium. Error bars indicate SEM. 6 tomograms for each cell line were analyzed. (D) Histogram showing frequencies of filament angles to the leading edge (90° corresponding to filaments perpendicular to the leading edge). Dashed black lines are medians, and red lines arithmetic means.
-
Figure 6—source data 1
Source data for details of lamellipodial actin networks including filament length, filament, barbed and pointed end densities and relative frequencies of filament angles Figure 6.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig6-data1-v2.xlsx
Loss of Ena/VASP deteriorates lamellipodium architecture in B16-F1 cells, related to Figure 6.
Animated visualization of the three-dimensional organization of lamellipodia of B16-F1 wild-type and EVM-KO cells resolved by electron tomography. A 3D model of the actin filament network corresponds to Figure 6A and was obtained by automatic tracking of filaments through the tomogram slices.
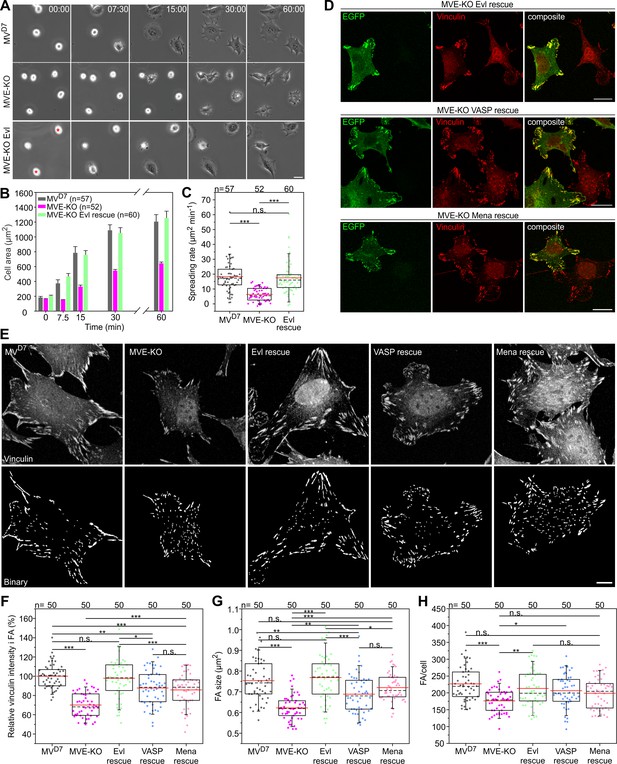
Inactivation of Evl in MVD7 cells impairs FA formation.
(A) Spreading of MVD7, EVM-KO and reconstituted cells on fibronectin. EGFP-Evl expressing cells are marked with red asterisks. Time is in minutes. Bar, 25 µm. (B) Quantification of cell area over time. Data are means ± SEM. n, number of analyzed cells from five independent experiments. (C) Quantification of spreading rate. (D) Expression of EGFP-tagged Evl, -VASP or -Mena in MVE-KO cells promotes the formation of FA. Immunolabeling of EGFP and vinculin in cells seeded on fibronectin. Bars, 20 µm. (E) Representative micrographs of MVD7, MVE-KO and reconstituted cells displaying vinculin staining before (upper panel) and after processing by Focal Adhesion Analysis Server (lower panel). Bar, 10 µm. (F) Quantification of vinculin intensities in FA. (G) Quantification of FA size. (H) Quantification of FA number per cell. (C and H) *p≤0.05, **p≤0.01, ***p≤0.001 and n.s., not significant by Kruskal-Wallis test and Dunn’s Multiple Comparison test or by one-way ANOVA and Tukey Multiple Comparison test (F–G), respectively. n, number of analyzed cells from three independent experiments. Boxes in box plots indicate 50% (25–75%) and whiskers (5–95%) of all measurements, with dashed black lines depicting the medians, arithmetic means are highlighted in red.
-
Figure 7—source data 1
Source data for details of cell spreading including cell areas and spreading rates, and of FA parameters including relative vinculin intensities, FA sizes and numbers per cell Figure 7.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig7-data1-v2.xlsx
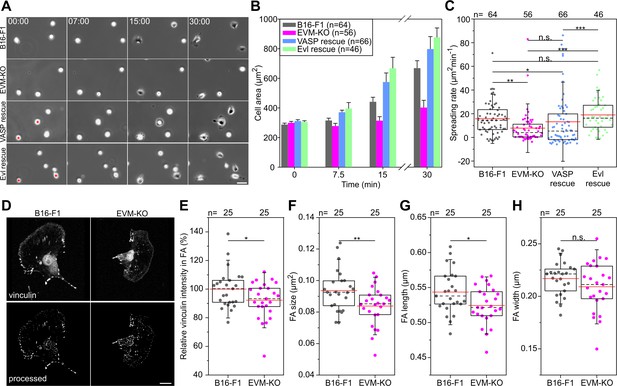
Loss of Ena/VASP proteins affects cell spreading and FA formation in B16-F1 cells.
(A) Spreading of B16-F1, EVM-KO and reconstituted cells on laminin. EGFP-VASP or EGFP-Evl expressing EVM-KO cells are marked by red asterisks. Time is in minutes. Scale bar, 25 µm. (B) Quantification of cell area over time. Bars and error bars represent arithmetic means and SEM, respectively. n, number of analyzed cells from five independent experiments. (C) Quantification of spreading rate. *p≤0.05, **p≤0.01, ***p≤0.001 and n.s., not significant by Kruskal-Wallis test and Dunn’s Multiple Comparison test. n, number of analyzed cells from at least three independent experiments. (D) Representative micrographs after vinculin staining of fixed B16-F1 and EVM-KO cells before (top) and after processing (bottom). Bar, 5 μm. (E). Quantification of vinculin intensities in FA. (F) Quantification of FA size. (G). Quantification of FA length. (H) Quantification of FA width. *p≤0.05, **p≤0.01 and n.s., not significant by student’s t-test. n, number of analyzed cells from at least three independent experiments.
-
Figure 7—figure supplement 1—source data 1
Source data for details of cell spreading including cell areas and spreading rates, and of FA parameters including relative vinculin intensities, FA sizes, length and widths Figure 7—figure supplement 1.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig7-figsupp1-data1-v2.xlsx
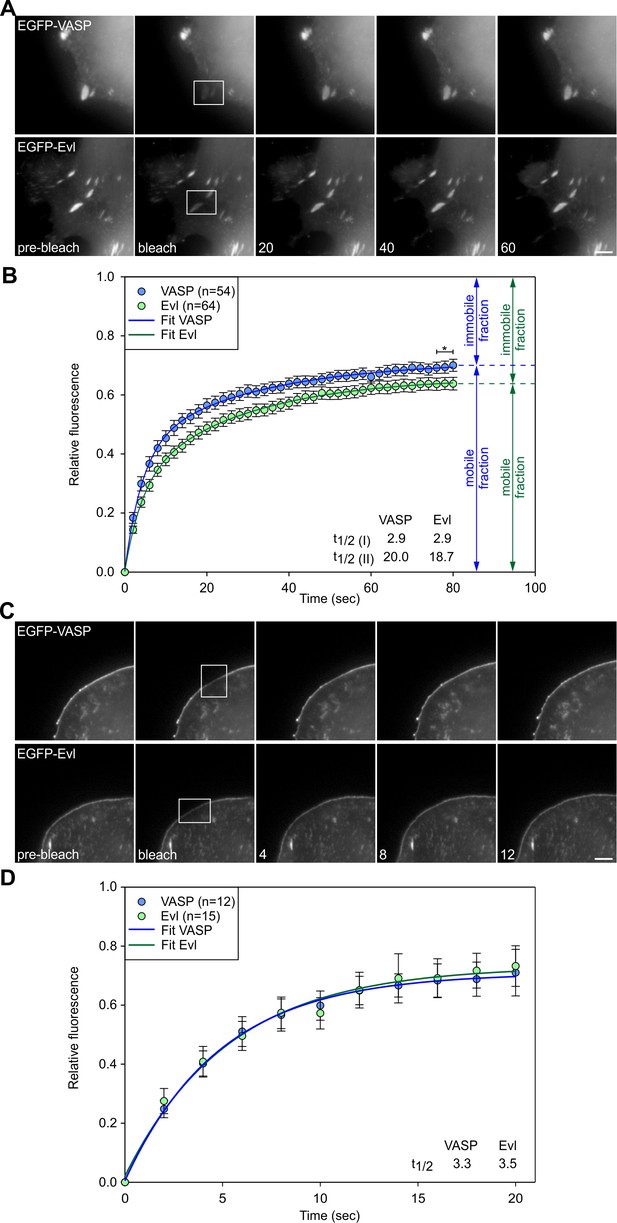
The fraction of Evl stably associated with focal adhesions is increased as compared to VASP.
(A) FRAP of EGFP-VASP or -Evl in focal adhesions of EVM-KO cells bleached as indicated by white rectangles. Numbers in post-bleach images correspond to seconds after bleach. Bar, 3 µm. (B) Relative fluorescence over time of bleached EGFP-VASP and -Evl in focal adhesions and respective curve fits using two phase association equation [Y(x)=a*(1-exp(-bx))+c*(1-exp(-dx))] assuming two independent dwell times for each component. Asterisk indicates statistically significant difference between immobile fractions assessed by averaging the final three data points (p≤0.05 by two-sided t-test); n, number of focal adhesions analyzed. (C) FRAP of EGFP-VASP or -Evl in lamellipodia of EVM-KO cells bleached as indicated by white rectangles. Numbers in post-bleach images correspond to seconds after bleach. Bar, 3 µm. (D) Relative fluorescence recovery over time of bleached EGFP-VASP or –Evl at the lamellipodium tip, and respective curve fits using one phase association equation [Y(x)=a∗(1-exp(-b∗x))] assuming one dwell time. n, number of lamellipodia analyzed.
-
Figure 7—figure supplement 2—source data 1
Source data for details of FRAP experiments including FA and lamellipodia Figure 7—figure supplement 2.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig7-figsupp2-data1-v2.xlsx
Loss of Evl in MVD7 cells diminishes spreading on fibronectin, related to Figure 7.
MVD7, MVE-KO and reconstituted cells expressing EGFP-Evl were seeded onto fibronectin and recorded by time-lapse phase-contrast imaging using a 20x objective. Reconstituted cells are marked with asterisks. Note enhanced spreading in MVE-KO cells expressing EGFP-Evl. Time, min:s. Scale bar, 25 µm.
FRAP of EGFP-tagged Evl versus -VASP in focal adhesions, related to Figure 7—figure supplement 2.
Representative time-lapse movies showing FRAP experiments of focal adhesions in EVM-KO cells expressing either EGFP-VASP or –Evl, as indicated, and plated on laminin. Time-lapse movies show periods before and after bleaching of focal adhesions (white rectangles) with bleaching time points set to 0 s in each case. Bar, 3 µm.
FRAP of EGFP-tagged VASP versus -Evl at the lamellipodium tip, related to Figure 7—figure supplement 2.
Representative time-lapse movies showing FRAP of EGFP-tagged VASP or -Evl residing at the tips of lamellipodia in EVM-KO cells migrating on laminin, as indicated. Time-lapse movies show periods before and after bleaching of indicated lamellipodial regions (white rectangles) with bleaching time points set to 0 s in each case. Bar, 3 µm.
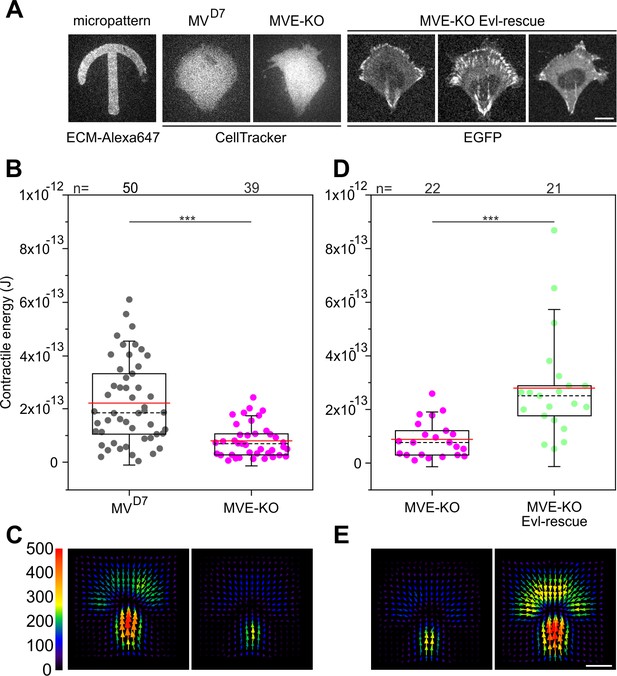
Confined MVE-KO cells exhibit diminished traction forces.
(A) Representative micrographs of cells plated on crossbow-shaped polyacrylamide micropatterns coated with fibronectin. MVD7 and MVE-KO were stained with CellTracker Green and rescued cells were stably expressing EGFP-Evl. Bar, 10 µm. (B and D) Quantifications of contractile energy of MVD7, MVE-KO and Evl expressing cells plated on crossbow-shaped polyacrylamide micropatterns in independent experiments. ***p≤0.001 by Mann-Whitney U rank sum test. n, number of cells analyzed. Boxes in box plots indicate 50% (25–75%) and whiskers (5–95%) of all measurements, with dashed black lines depicting the medians, arithmetic means are highlighted in red. (C and E) Images depicting averaged traction force field representations of cells indicated. Bar, 10 µm. Force scale bar is in Pascal and arrows represent the local force magnitude and orientation.
-
Figure 8—source data 1
Source data for details of contractile energies Figure 8.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig8-data1-v2.xlsx
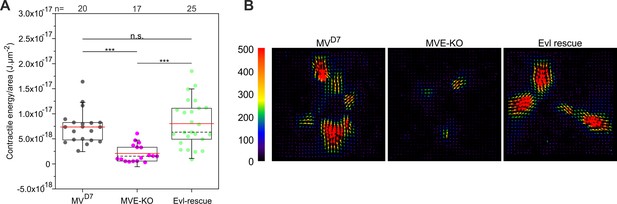
Inactivation of Evl in unconfined MVD7 cells diminishes cell contractile energy.
(A) Quantifications of contractile energy normalized by cellular area and measured for MVD7, MVE-KO and Evl-rescue of MVE-KO cells plated on plain polyacrylamide micropatterns coated with fibronectin. ***p≤0.001 or n.s., not significant by Kruskal-Wallis test and Dunn’s Multiple Comparison test. n, number of cells analyzed. Boxes in box plots indicate 50% (25–75%) and whiskers (5–95%) of all measurements, with dashed black lines depicting the medians, arithmetic means are highlighted in red. (B) Images depicting representative traction force field representations from cells of distinct genotypes, as indicated. Force scale bar is in Pascal and arrows represent local force magnitude and orientation.
-
Figure 8—figure supplement 1—source data 1
Source data for details of contractile energies Figure 8—figure supplement 1.
- https://cdn.elifesciences.org/articles/55351/elife-55351-fig8-figsupp1-data1-v2.xlsx
Additional files
-
Supplementary file 1
Key resources table.
- https://cdn.elifesciences.org/articles/55351/elife-55351-supp1-v2.docx
-
Supplementary file 2
Sequences of generated knock out clones.
- https://cdn.elifesciences.org/articles/55351/elife-55351-supp2-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/55351/elife-55351-transrepform-v2.docx





