Cryo-EM structure in situ reveals a molecular switch that safeguards virus against genome loss
Figures
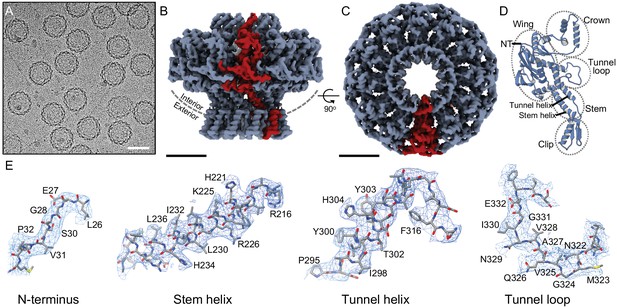
Structure of the portal protein in situ.
(A) Cryo-electron micrograph of P23- 45 procapsids, scale bar 50 nm. (B) Cryo-EM reconstruction map with one subunit coloured red, scale bar 50 Å, and same for (C) but rotated 90°, viewed along 12-fold axis. (D) Ribbon diagram of one portal protein subunit. (E) Regions of the map and corresponding atomic models with residue numbers.
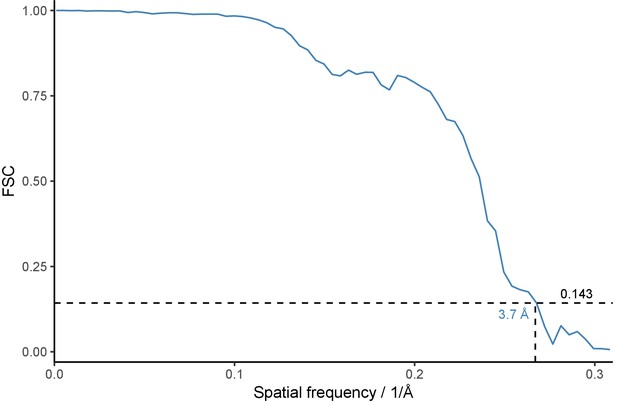
FSC curve for the portal protein reconstruction.
Fourier Shell Correlation is plotted as function of spatial frequency. Dotted lines denote resolution estimate at FSC = 0.143 according to the gold-standard method.

Mass spectrometry analysis of the portal protein from unexpanded capsids.
Blue bars beneath the sequence denote regions for which peptides were detected.
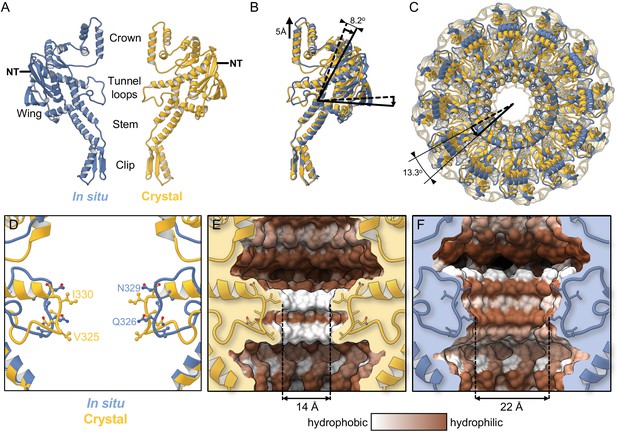
Comparison with the crystal structure.
(A) Single subunit of the in situ structure is in blue and an apposing chain from the crystal structure is in yellow. (B) Superposition of single subunits, exposing structural differences between the crystal structure and the in situ structure. The curved arrow indicates the pivoting of the Wing domain by ~8° in the in situ structure. (C) The two dodecamers overlaid, viewed from Crown (top domain in A), along the tunnel axis. Dodecamers are superposed based on residues 26–376 (Clip, Stem, and Wing), revealing a ~13° rotation of the Crown domain about the tunnel axis. (D) Overlay of in situ (blue) and crystal structure (yellow), ribbon diagram, with side-chains shown. (E) Van der Waals surface of the crystal structure (PDB 6IBG) showing tunnel loop-constricted region, with tunnel colouring by the hydrophobicity on the Kyte-Doolittle scale where white is hydrophobic and brown is hydrophilic, and same for (F) but for the in situ structure (PDB 6QJT). Diameters of most constricted part of tunnels measured from Van der Waals surfaces are shown.
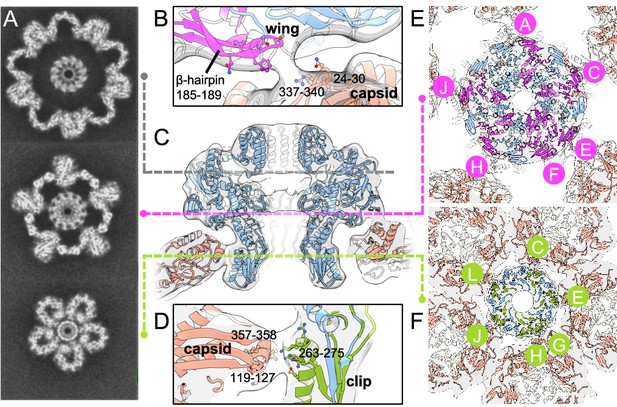
Portal–capsid interactions.
(A) Sections through the capsid reconstruction perpendicular to the portal tunnel axis, at three different heights as denoted on (C) by dashed lines. (B) Interactions between the portal Wing and capsid. Portal protein subunit making interactions with the capsid is in pink. Portal subunits not making interactions are in blue. (C) Ribbon diagram of the in situ portal protein fitted into the procapsid map. (D) Interactions between the portal Clip and capsid. Portal protein subunit making closest interactions with the capsid is in green. (E) Subunits of the portal protein interacting with the capsid by their Wing regions are in magenta, labelled clockwise. (F) Subunits of the portal protein interacting via their Clip are in green. View is from the center of the portal with chains labelled as in (E).
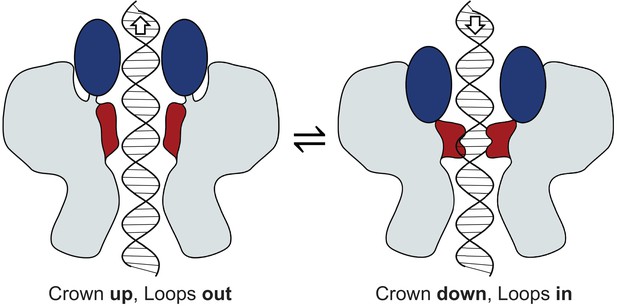
Mechanism of portal tunnel closure.
Left - the open state where the Crown (blue) is elevated, facilitating partial retreat of the tunnel loops (terracotta) toward the crown, widening the tunnel. Right – the closed state where the Crown is depressed into the body of the portal protein, facilitating closure of the tunnel where tunnel loops adopt a conformation that extends into the tunnel.
Videos
Reconstruction of the in situ portal.
Surface rendering, first viewed perpendicular to the tunnel axis, then viewed along the tunnel axis.
Morph between the in situ structure (first) and crystal structure (second).
Ribbon diagram, first viewed perpendicular to the tunnel axis, then viewed along the tunnel axis, then rotated back to initial view with two apposing chains displayed.
Portal–capsid registers.
One-degree step change in portal register (inner 12-fold circle) with respect to capsid vertex (outer 5-fold circle), beginning with ‘0°’. Portal register ‘6°’ is superposable on register ‘0°’ by 144° rotation of the whole capsid (i.e. rotating both inner and outer circles together).
Additional files
-
Supplementary file 1
Cryo-EM data collection and refinement statistics.
- https://cdn.elifesciences.org/articles/55517/elife-55517-supp1-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/55517/elife-55517-transrepform-v2.pdf





